1 Introduction
Gene expression microarrays have become a standard technology in the biologist’s toolbox. For two-color microarrays, an immediate question in the early stages of planning a microarray study is how to choose the arrangement of samples onto the microarrays. Multiple papers have taken up this question, including Kerr and Churchill (2001); Dobbin and Simon (2002); Yang and Speed (2002); Kerr (2003a); Wit et al. (2005); Kerr (2006); Bailey (2007); Sanchez and Glonek (2009). Some of these papers have utilized the so-called ”alphabet” design criteria such as A- and D-optimality to evaluate microarray designs. Unfortunately, these design criteria are defined differently in the regression and block design settings, which seems to have caused some confusion in the literature.
This paper has two purposes. First, we explicate the definitions of A-, D-, and E-optimality to address the confusion in the literature. Second, we give our assessment of the relative value of these criteria for choosing microarray designs. From years of collaborating with biologists using microarrays, a guiding principle of our design recommendations will stem from our experience that a study that is most likely to be successful is one where particular comparisons are of interest a priori. It follows from this principle that design criteria should, as much as possible, focus on the comparisons of interest, which is a point of broad agreement.
2 Optimality Criteria Defined
Design optimality criteria are defined differently in the regression and block design settings. For each setting we provide the necessary background and then define the criteria.
2.1 Optimality Criteria in the Regression Setting
In linear regression, there is a model
where y is the data vector, X is the design matrix, β is the (unknown) vector of regression parameters, and ε is the random error vector standardly assumed to have expectation 0 and cov(ε)=σ2I. In the regression setting it is reasonable to assume that X has full column rank; the model would not be worth fitting otherwise. This assumption implies that (X′X)−1 exists, and that there is a unique least-squares estimate of β given by β̂ = (X′X)−1X′y. Under the assumption of independent errors that have a common variance σ2, β̂ has covariance σ2(X′X)−1.
Some classical design optimality criteria are A-, D-, and E-optimality (Atkinson and Donev, 1992). The A-optimal design minimizes trace((X′X)−1). This has the interpretation of minimizing the average variance among parameter estimates. The D-optimal design minimizes |(X′X)−1|, or equivalently, maximizes |X′X|. This has the interpretation of minimizing the volume of the confidence ellipsoid for the regression parameters under the assumption that ε is multivariate normal. The E-optimal design minimizes the variance of the least-well estimated linear combination: a′β is minimized subject to a′a = 1. In the field of optimal design, ‘A’ stands for average, ‘D’ stands for determinant, and ‘E’ stands for extreme.
2.2 Optimality Criteria in the Block Design Setting
In block design there is also a linear model. Following the notation of John and Williams (1995), suppose there are υ treatments in b blocks of size k. In the microarray application, we consider each two-color microarray as the “block,” so k = 2. The different varieties of biological samples of RNA are the “treatments” and υ ≥ 2. For simplicity, suppose each treatment is replicated r times – some adjustment is needed when there are unequal replications. We write the model as
or, in matrix notation,
μ is the model intercept, β is the vector of parameters βj, 1 ≤ j ≤ b, representing block effects, and τ is the vector of parameters τi, 1 ≤ i ≤ υ, representing treatment effects. There are two important points. First, the overall design matrix
does not have full column rank and design criteria from general regression do not apply directly. Second, μ and β are nuisance parameters, so it is most appropriate to focus design questions on the treatment effects. More specifically, we translate the scientific question into a question about contrasts among treatment effects. Continuing to follow John and Williams (1995), the normal equations for the treatment effects are
| (2.1) |
where
| (2.2) |
and
| (2.3) |
In (2.2) and (2.3), N is the incidence matrix of the design with υ rows and b columns: nij=1 if treatment i appears in block j and otherwise nij=0. T is the data vector of treatment totals and B is the data vector of block totals.
A is called the information matrix of the block design. The information matrix A always has a set of orthogonal eigenvectors pi with corresponding eigenvalues θi. A is singular and exactly one eigenvalue is 0. We leave out the 0 eigenvalue and its eigenvector in forming
which is a generalized inverse of A. Since the eigenvectors are orthogonal and pυ ∝ 1, this tells us that p1, …, pυ−1 are contrasts and therefore are estimable for i = 1, …, υ − 1. The are the basic contrasts of the design (Pearce et al., 1974). The variance of is . Further, any contrast vector d can be written Σaipi, and .
Define the efficiency factor of a contrast as
where varCBD(d′τ̂) is the variance of the contrast in an orthogonal design using the same number of experimental units1. The efficiency factor of a basic contrast is . For a generic contrast we can write , and since ei = 1 for a complete block design,
Let θmin be the smallest eigenvalue and θmax be the largest. These correspond to the smallest and largest efficiency factors among the basic contrasts, emin and emax. Clearly
and thus
Therefore, the canonical efficiency factors bound the efficiency of any contrast, and they can be considered a summary of the properties of a design. A design that maximizes the smallest canonical efficiency factor is said to be E-optimal. One can also choose a design to maximize the harmonic mean of the efficiency factors. Since it can be shown that the average variance over all pairwise treatment contrasts is proportional to (Kempthorne, 1956), this design is called A-optimal. This is the classical definition of A-optimality for incomplete block designs – minimizing the average variance over pairwise treatment contrasts. This was the definition used by Kerr and Churchill (2001) to study microarray designs. A design that maximizes the geometric mean of the efficiency factors is D-optimal. The D-optimality criterion is the product of eigenvalues just like in the regression setting. However, in the regression setting we can interpret the D-criterion as a summary of the generalized variance of the regression parameters. We do not have the analogous interpretation in the block design setting – the D-criterion does not directly relate to the generalized variance of the model parameters (John and Williams, 1995, p. 32).
3 Regression-style D-Optimality Criteria for Microarrays
Linear models for the analysis of microarray data were first introduced as ANOVA models (Kerr et al., 2000; Wolfinger et al., 2001). Smyth (2004) and others introduced regression-style models. Briefly, by collapsing the data from a single microarray into logratios, an ANOVA-style model can be transformed into a regression-style model. The regression-style model uses parameters δii′ ≡ τi − τi′, where the τi are the “treatment effects” in the ANOVA model. Kerr (2003b) describes in detail the relationship between the regression and ANOVA approaches in the analysis of microarray data. For analytic purposes, choice of a regression-style or ANOVA-style model is largely a matter of personal taste. However, the fact remains that there are different definitions of some alphabetic optimality criteria that are not equivalent.
Wit et al. (2005) employed regression-style models to evaluate microarray designs. They use υ − 1 pairwise treatment contrasts in a no-intercept model and consider A-, D-, and L-optimality as defined in the regression setting. It is important to note that Wit et al’s A- and D- criteria are not as traditionally defined in block design settings, generating some confusion. It is such confusion that this paper aims to ameliorate. The L-optimality criterion in Wit et al. (2005) is the criterion that this paper will advocate for.
Wit et al. (2005) note that D-optimality does not depend on the particular choice of treatment contrasts that parametrize the regression model. They imply that this invariance to parametrization is an advantage of D-optimality. We do not agree for three reasons. First, this invites investigators to naively seek a D-optimal design without considering the scientific question. As noted in the Introduction, we believe the scientific question should always be a driving force in the selection of a design. Second, the scientific question may not correspond to υ − 1 treatment contrasts, so invariance to choice of υ − 1 treatment contrasts is beside the point.
Our third reason for being less favorably inclined toward D-optimality is because its invariance leads to some indifference among designs. Consider a situation where there is a “control” treatment and three “experimental” treatments. Number the control treatment 1 and the experimental treatments 2, 3, 4. In the notation of Wit et al. (2005), scientific interest is in the parameters δ12, δ13, δ14, i.e. the comparisons between the control treatment 1 and the experimental treatment 2, 3, and 4. This seems to be a case that is well-suited to the approach in Wit et al. (2005), since the contrasts of interest correspond to three parameters that can parametrize the regression model.
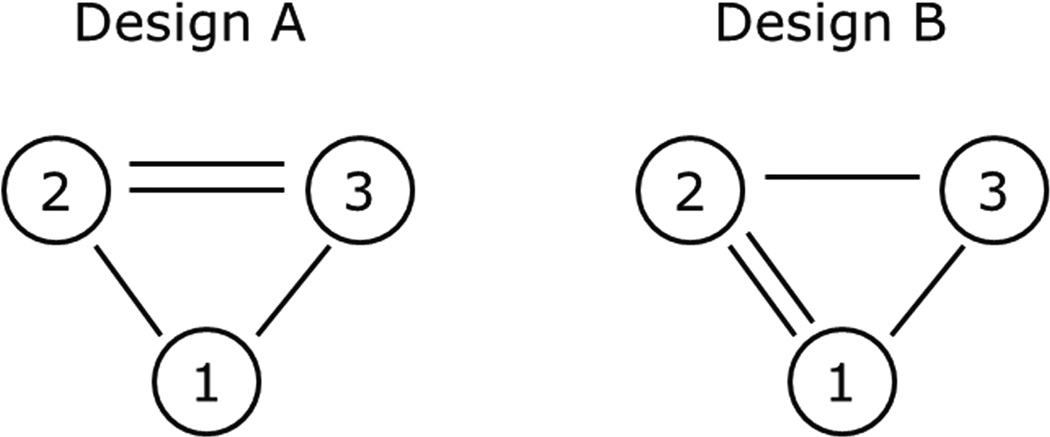
Consider the designs in Figure 1. Intuitively, Design A matches the scientific objective. In design A each of the experimental treatments is compared directly to the control treatment. Direct comparisons are most efficient, and it seems ideal to allot the three direct comparisons to the three pairwise contrasts that are of interest. Designs B and C are intuitively unappealing. However, D-Optimality says that all three designs are equally good. This can be viewed as a consequence of invariance to parametrization: if we parametrized the model instead with δ14, δ24, δ34 then Design B becomes the more natural choice; if we parametrized the model with δ12, δ23, δ34 then Design C becomes the natural choice. However, D-optimality is invariant to parametrization and so has no preference among these three designs.
Figure 1.
Designs for four varieties and three arrays/blocks. Designs are represented as undirected graphs. Each number represents a variety and each edge represents a microarray (a block of size 2).
In contrast to D-optimality and in accordance with intuition, if we consider the average variance among δ12, δ13, δ14, Design A is strongly preferred. The average variance is 66% larger with Design B and twice as large with Design C. This is not to say that A-optimality is the ideal criterion, but in many block design problems we believe it is preferable to D-optimality.
As a second example, suppose there are two experimental treatments and one control treatment. The control treatment is included as way of checking that things are working as expected, but primary interest is in comparing the two experimental treatments. As before, label the control treatment 1 and the experimental treatments 2 and 3. Two possible designs are shown in Figure 2. Design A clearly matches the scientific objective of the study. The parameter of interest is δ23; the variance of its estimate is 50% larger with Design B. However, D-optimality is indifferent between Designs A and B.
Figure 2.
Designs for three varieties and four arrays/blocks.
4 Block Design Optimality Criteria
There is broad agreement that a design criterion should match, as closely as possible, the scientific objectives of a study (Sanchez and Glonek, 2009; Wit et al., 2005). In a microarray study there are scientific questions of interest that correspond to contrasts of interest. In the previous section we argued against applying regression-style D-optimality. What about D- and E-optimality as defined in block design?
The E-optimal design maximizes the efficiency of the worst-case-scenario contrast. However, the worst-case-scenario contrast may have not relevance to the scientific question at hand. Similarly, the D-optimal design minimizes the generalized variance of the basic contrasts of the design (defined in Section 2.2). Again, the basic contrasts are not necessarily of scientific interest. In fact, the basic contrasts might be different from design to design. In total, we believe that block design D- and E-optimality fail to satisfy the basic requirement of reflecting the scientific objectives of a study.
5 Concluding Remarks
Definitions of the “alphabet” design criteria differ in the regression and block design settings, which has created some confusion in the literature. Wit et al. (2005) suggest that Kerr and Churchill (2001) use a non-standard definition of A-optimality. To the contrary, Kerr and Churchill (2001) use a definition consistent with the literature on block design. The approach in Wit et al. (2005) is unconventional in that it considers incomplete block designs in a regression setting. On the other hand, the approach in Sacan et al. (2009) is unclear since the design matrices in their formulation do not have full rank yet their stated design criteria are only defined for models with full-rank design matrices. Tan and Passos (2011) criticize the dependence of the A-optimality criterion on model parametrization but do not make it clear that their criticism applies to the definition of A-optimality in the regression setting, which is not the definition traditionally used in microarray design.
E- and D-optimal designs are advantageous in terms of the basic contrasts, but these are not the contrasts of interest. These criteria may be associated with good “all-purpose” designs. There are occasions when such designs may be appropriate. However, in our collaborations we prefer to choose designs that best-suit the scientific objectives of particular studies.
We have essentially argued for A-style optimality criteria when evaluating microarray designs. The A-optimality criteria can be applied to all pairwise contrasts, as in Kerr and Churchill (2001), or modified to apply to a different set of contrasts of interest. Wit et al. (2005), following Pukelsheim (1993), use the term L-optimality. A valid concern about A-optimality is that it does not account for the correlation among estimates. It is not clear how to address this, and we consider this an open question. However, for the reasons described, we do not consider D- optimality to be the solution.
Bailey (2007) points out that, unlike for larger block sizes, for block size two there can be severe conflict between the A- and D-criteria. She shows, for example, that the loop design is D-optimal for b = t ≥ 10, whereas the loop design is extremely A-inefficient. Similarly, the A-optimal design is severely D-inefficient. Bailey (2007, p. 376) proposes that robustness considerations can be used to choose between the A-optimal and D-optimal designs. We wholeheartedly endorse the consideration of robustness. However, given our general disfavor for D-optimality and preference for A-style optimality criteria directed at the contrasts of interest, we propose a slightly modified version of this strategy. We do not advocate automatic acceptance of the optimal design on the preferred criterion, but rather considering all efficient designs in light of robustness considerations. A slightly sub-optimal design might easily be preferable if it is more robust.
Footnotes
CBD stands for “complete block design.” Such a design may not exist, but this serves as a reasonable basis for comparison.
References
- Atkinson AC, Donev AN. Optimum Experimental Designs. Oxford, UK: Oxford University Press; 1992. [Google Scholar]
- Bailey RA. Designs for two-colour microarray experiments. Applied Statistics. 2007;56:365–394. [Google Scholar]
- Dobbin K, Simon R. Comparison of microarray designs for class comparison and class discovery. Bioinformatics. 2002;18:1438–1445. doi: 10.1093/bioinformatics/18.11.1438. [DOI] [PubMed] [Google Scholar]
- John JA, Williams ER. Cyclic and Computer Generated Designs. second edition. London: Chapman and Hall; 1995. [Google Scholar]
- Kempthorne O. The efficiency factor of an incomplete block design. Anals of Mathematical Statistics. 1956;27:846–849. [Google Scholar]
- Kerr KF. 2k factorials in blocks of size 2, with application to two-color microarray experiments. Journal of Quality Technology. 2006;38:349–364. [Google Scholar]
- Kerr MK. Design considerations for efficient and effective microarray studies. Biometrics. 2003a;59:822–828. doi: 10.1111/j.0006-341x.2003.00096.x. [DOI] [PubMed] [Google Scholar]
- Kerr MK. Linear models for microarray data analysis: Hidden similarities and differences. Journal of Computational Biology. 2003b;10:891–901. doi: 10.1089/106652703322756131. [DOI] [PubMed] [Google Scholar]
- Kerr MK, Churchill GA. Experimental design for gene expression microarrays. Biostatistics. 2001;2:183–201. doi: 10.1093/biostatistics/2.2.183. [DOI] [PubMed] [Google Scholar]
- Kerr MK, Martin M, Churchill GA. Analysis of variance for gene expression microarrays. Journal of Computational Biology. 2000;7:819–837. doi: 10.1089/10665270050514954. [DOI] [PubMed] [Google Scholar]
- Pearce SC, Calinski T, de C, Marshall TF. The basic contrasts of an experimental design with special reference to the analysis of data. Biometrika. 1974;61:449–460. [Google Scholar]
- Pukelsheim F. Optimal Design of Experiments. New York: Wiley; 1993. [Google Scholar]
- Sacan A, Ferhatosmanoglu N, Ferhatosmanoglu H. Microarraydesigner: an online search tool and repository for near-optimal microarray experimental designs. BMC Bioinformatics. 2009;10:304. doi: 10.1186/1471-2105-10-304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanchez P, Glonek G. Optimal designs for 2-color microarray experiments. Biostatistics. 2009;10:561–574. doi: 10.1093/biostatistics/kxp012. [DOI] [PubMed] [Google Scholar]
- Smyth GK. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Statistical Applications in Genetics and Molecular Biology. 2004;3 doi: 10.2202/1544-6115.1027. Article 3. [DOI] [PubMed] [Google Scholar]
- Tan FE, Passos VL. On the A-optimality criterion for finding two-color microarray optimal designs. Journal of Computational Biology. 2011;18:835–841. doi: 10.1089/cmb.2010.0009. [DOI] [PubMed] [Google Scholar]
- Wit E, Nobile A, Khanin R. Near-optimal designs for dual channel microarray studies. Applied Statistics. 2005;54:817–830. [Google Scholar]
- Wolfinger RD, Gibson G, Wolfinger E, Bennett L, Hamadeh H, Bushel P, Afshari C, Paules RS. Assessing gene significance from cdna microarray expression data via mixed models. Journal of Computational Biology. 2001;8:625–637. doi: 10.1089/106652701753307520. [DOI] [PubMed] [Google Scholar]
- Yang YH, Speed T. Design issues for cDNA microarray experiments. Nature Reviews. 2002;3:579–588. doi: 10.1038/nrg863. [DOI] [PubMed] [Google Scholar]