Abstract
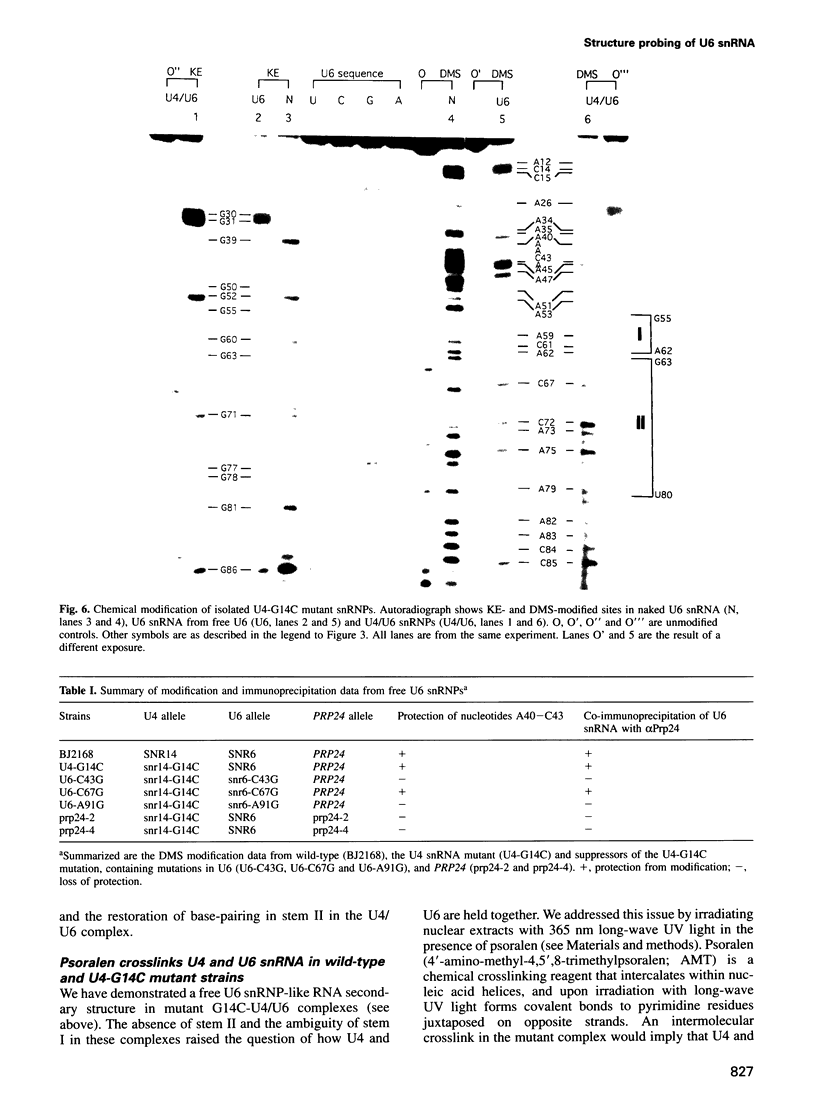
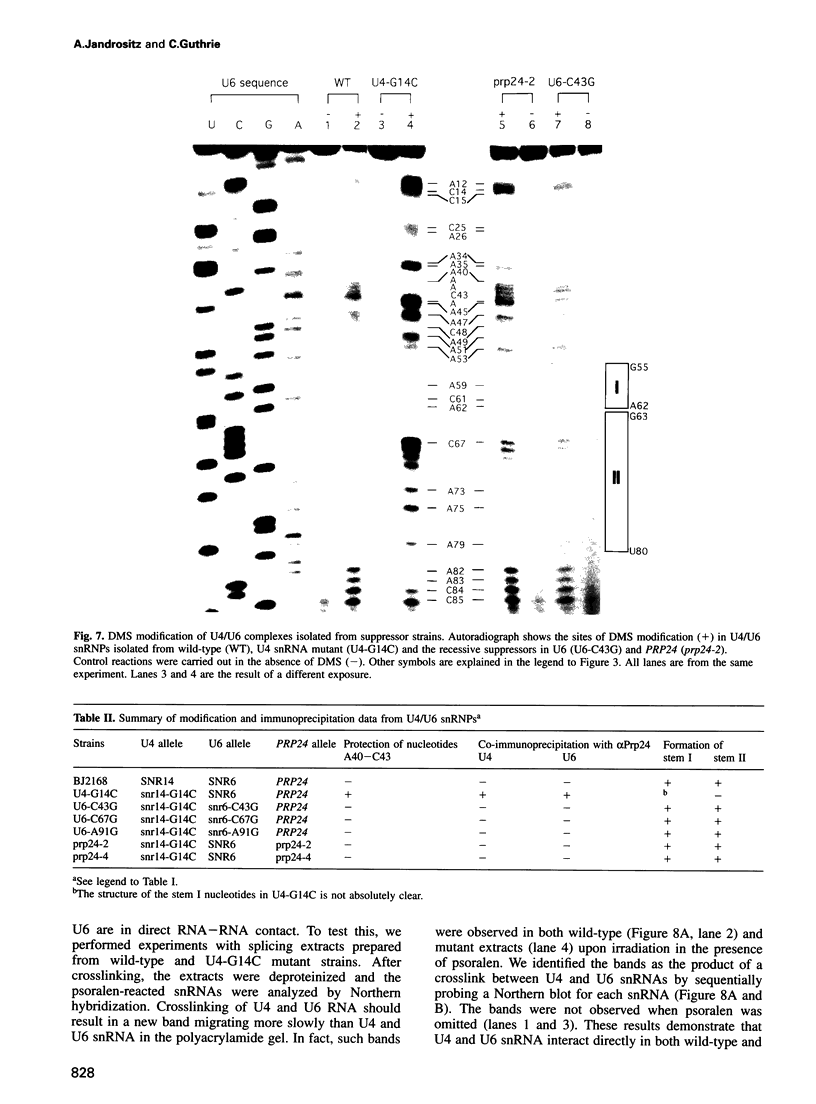
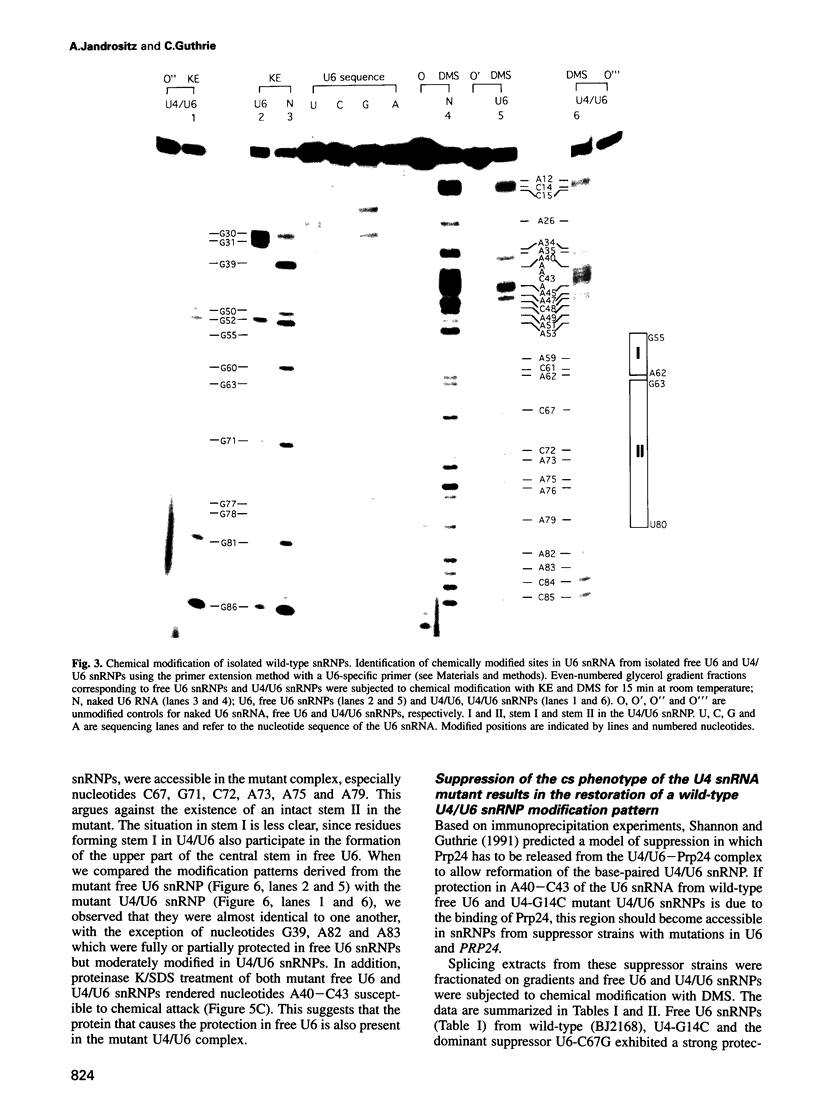
A mutation (U4-G14C) that destabilizes the base-pairing interaction between U4 and U6 snRNAs causes the accumulation of a novel complex containing U4, U6 and Prp24, a protein with RNA binding motifs. An analysis of suppressors of this cold-sensitive mutant led to the hypothesis that this complex is normally a transient intermediate in the annealing of U4 and U6. It was proposed that Prp24 must be released to form a fully base-paired U4/U6 snRNP. By using a chemical probing method we have tested the prediction that nucleotides A40-C43 in U6 mediate the binding of Prp24. Consistent with the location of recessive suppressors in U6, we find that residues A40-C43 are protected from chemical modification in U4/U6 complexes from the U4-G14C mutant but not from the wild-type or suppressor strains carrying mutations in U6 or PRP24. Furthermore, we find that base-pairing is substantially disrupted in the mutant complexes. Notably, the base-paired structure is restored in recessive suppressors despite the presence of a mismatched base-pair at the U4-G14C site. Our results support the model that Prp24 binds to U6 to promote its association with U4, but must dissociate to allow complete annealing.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Banroques J., Abelson J. N. PRP4: a protein of the yeast U4/U6 small nuclear ribonucleoprotein particle. Mol Cell Biol. 1989 Sep;9(9):3710–3719. doi: 10.1128/mcb.9.9.3710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bindereif A., Wolff T., Green M. R. Discrete domains of human U6 snRNA required for the assembly of U4/U6 snRNP and splicing complexes. EMBO J. 1990 Jan;9(1):251–255. doi: 10.1002/j.1460-2075.1990.tb08102.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Black D. L., Pinto A. L. U5 small nuclear ribonucleoprotein: RNA structure analysis and ATP-dependent interaction with U4/U6. Mol Cell Biol. 1989 Aug;9(8):3350–3359. doi: 10.1128/mcb.9.8.3350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blencowe B. J., Sproat B. S., Ryder U., Barabino S., Lamond A. I. Antisense probing of the human U4/U6 snRNP with biotinylated 2'-OMe RNA oligonucleotides. Cell. 1989 Nov 3;59(3):531–539. doi: 10.1016/0092-8674(89)90036-6. [DOI] [PubMed] [Google Scholar]
- Bordonné R., Banroques J., Abelson J., Guthrie C. Domains of yeast U4 spliceosomal RNA required for PRP4 protein binding, snRNP-snRNP interactions, and pre-mRNA splicing in vivo. Genes Dev. 1990 Jul;4(7):1185–1196. doi: 10.1101/gad.4.7.1185. [DOI] [PubMed] [Google Scholar]
- Bringmann P., Appel B., Rinke J., Reuter R., Theissen H., Lührmann R. Evidence for the existence of snRNAs U4 and U6 in a single ribonucleoprotein complex and for their association by intermolecular base pairing. EMBO J. 1984 Jun;3(6):1357–1363. doi: 10.1002/j.1460-2075.1984.tb01977.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brow D. A., Guthrie C. Spliceosomal RNA U6 is remarkably conserved from yeast to mammals. Nature. 1988 Jul 21;334(6179):213–218. doi: 10.1038/334213a0. [DOI] [PubMed] [Google Scholar]
- Brown J. D., Beggs J. D. Roles of PRP8 protein in the assembly of splicing complexes. EMBO J. 1992 Oct;11(10):3721–3729. doi: 10.1002/j.1460-2075.1992.tb05457.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng S. C., Abelson J. Spliceosome assembly in yeast. Genes Dev. 1987 Nov;1(9):1014–1027. doi: 10.1101/gad.1.9.1014. [DOI] [PubMed] [Google Scholar]
- Datta B., Weiner A. M. Genetic evidence for base pairing between U2 and U6 snRNA in mammalian mRNA splicing. Nature. 1991 Aug 29;352(6338):821–824. doi: 10.1038/352821a0. [DOI] [PubMed] [Google Scholar]
- Fabrizio P., McPheeters D. S., Abelson J. In vitro assembly of yeast U6 snRNP: a functional assay. Genes Dev. 1989 Dec;3(12B):2137–2150. doi: 10.1101/gad.3.12b.2137. [DOI] [PubMed] [Google Scholar]
- Fortner D. M., Troy R. G., Brow D. A. A stem/loop in U6 RNA defines a conformational switch required for pre-mRNA splicing. Genes Dev. 1994 Jan;8(2):221–233. doi: 10.1101/gad.8.2.221. [DOI] [PubMed] [Google Scholar]
- Frank D., Guthrie C. An essential splicing factor, SLU7, mediates 3' splice site choice in yeast. Genes Dev. 1992 Nov;6(11):2112–2124. doi: 10.1101/gad.6.11.2112. [DOI] [PubMed] [Google Scholar]
- Green M. R. Biochemical mechanisms of constitutive and regulated pre-mRNA splicing. Annu Rev Cell Biol. 1991;7:559–599. doi: 10.1146/annurev.cb.07.110191.003015. [DOI] [PubMed] [Google Scholar]
- Guthrie C., Patterson B. Spliceosomal snRNAs. Annu Rev Genet. 1988;22:387–419. doi: 10.1146/annurev.ge.22.120188.002131. [DOI] [PubMed] [Google Scholar]
- Hamm J., Mattaj I. W. An abundant U6 snRNP found in germ cells and embryos of Xenopus laevis. EMBO J. 1989 Dec 20;8(13):4179–4187. doi: 10.1002/j.1460-2075.1989.tb08603.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hashimoto C., Steitz J. A. U4 and U6 RNAs coexist in a single small nuclear ribonucleoprotein particle. Nucleic Acids Res. 1984 Apr 11;12(7):3283–3293. doi: 10.1093/nar/12.7.3283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hausner T. P., Giglio L. M., Weiner A. M. Evidence for base-pairing between mammalian U2 and U6 small nuclear ribonucleoprotein particles. Genes Dev. 1990 Dec;4(12A):2146–2156. doi: 10.1101/gad.4.12a.2146. [DOI] [PubMed] [Google Scholar]
- Inoue T., Cech T. R. Secondary structure of the circular form of the Tetrahymena rRNA intervening sequence: a technique for RNA structure analysis using chemical probes and reverse transcriptase. Proc Natl Acad Sci U S A. 1985 Feb;82(3):648–652. doi: 10.1073/pnas.82.3.648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kandels-Lewis S., Séraphin B. Involvement of U6 snRNA in 5' splice site selection. Science. 1993 Dec 24;262(5142):2035–2039. doi: 10.1126/science.8266100. [DOI] [PubMed] [Google Scholar]
- Lamond A. I., Konarska M. M., Grabowski P. J., Sharp P. A. Spliceosome assembly involves the binding and release of U4 small nuclear ribonucleoprotein. Proc Natl Acad Sci U S A. 1988 Jan;85(2):411–415. doi: 10.1073/pnas.85.2.411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lesser C. F., Guthrie C. Mutations in U6 snRNA that alter splice site specificity: implications for the active site. Science. 1993 Dec 24;262(5142):1982–1988. doi: 10.1126/science.8266093. [DOI] [PubMed] [Google Scholar]
- Li Z., Brow D. A. A rapid assay for quantitative detection of specific RNAs. Nucleic Acids Res. 1993 Sep 25;21(19):4645–4646. doi: 10.1093/nar/21.19.4645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin R. J., Newman A. J., Cheng S. C., Abelson J. Yeast mRNA splicing in vitro. J Biol Chem. 1985 Nov 25;260(27):14780–14792. [PubMed] [Google Scholar]
- Lossky M., Anderson G. J., Jackson S. P., Beggs J. Identification of a yeast snRNP protein and detection of snRNP-snRNP interactions. Cell. 1987 Dec 24;51(6):1019–1026. doi: 10.1016/0092-8674(87)90588-5. [DOI] [PubMed] [Google Scholar]
- Madhani H. D., Guthrie C. A novel base-pairing interaction between U2 and U6 snRNAs suggests a mechanism for the catalytic activation of the spliceosome. Cell. 1992 Nov 27;71(5):803–817. doi: 10.1016/0092-8674(92)90556-r. [DOI] [PubMed] [Google Scholar]
- Madhani H. D., Guthrie C. Dynamic RNA-RNA interactions in the spliceosome. Annu Rev Genet. 1994;28:1–26. doi: 10.1146/annurev.ge.28.120194.000245. [DOI] [PubMed] [Google Scholar]
- Moazed D., Noller H. F. Intermediate states in the movement of transfer RNA in the ribosome. Nature. 1989 Nov 9;342(6246):142–148. doi: 10.1038/342142a0. [DOI] [PubMed] [Google Scholar]
- Moazed D., Stern S., Noller H. F. Rapid chemical probing of conformation in 16 S ribosomal RNA and 30 S ribosomal subunits using primer extension. J Mol Biol. 1986 Feb 5;187(3):399–416. doi: 10.1016/0022-2836(86)90441-9. [DOI] [PubMed] [Google Scholar]
- Nilsen T. W. RNA-RNA interactions in the spliceosome: unraveling the ties that bind. Cell. 1994 Jul 15;78(1):1–4. doi: 10.1016/0092-8674(94)90563-0. [DOI] [PubMed] [Google Scholar]
- Parker R., Siliciano P. G., Guthrie C. Recognition of the TACTAAC box during mRNA splicing in yeast involves base pairing to the U2-like snRNA. Cell. 1987 Apr 24;49(2):229–239. doi: 10.1016/0092-8674(87)90564-2. [DOI] [PubMed] [Google Scholar]
- Patterson B., Guthrie C. An essential yeast snRNA with a U5-like domain is required for splicing in vivo. Cell. 1987 Jun 5;49(5):613–624. doi: 10.1016/0092-8674(87)90537-x. [DOI] [PubMed] [Google Scholar]
- Pikielny C. W., Rymond B. C., Rosbash M. Electrophoresis of ribonucleoproteins reveals an ordered assembly pathway of yeast splicing complexes. 1986 Nov 27-Dec 3Nature. 324(6095):341–345. doi: 10.1038/324341a0. [DOI] [PubMed] [Google Scholar]
- Rinke J., Appel B., Digweed M., Lührmann R. Localization of a base-paired interaction between small nuclear RNAs U4 and U6 in intact U4/U6 ribonucleoprotein particles by psoralen cross-linking. J Mol Biol. 1985 Oct 20;185(4):721–731. doi: 10.1016/0022-2836(85)90057-9. [DOI] [PubMed] [Google Scholar]
- Sawa H., Abelson J. Evidence for a base-pairing interaction between U6 small nuclear RNA and 5' splice site during the splicing reaction in yeast. Proc Natl Acad Sci U S A. 1992 Dec 1;89(23):11269–11273. doi: 10.1073/pnas.89.23.11269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawa H., Shimura Y. Association of U6 snRNA with the 5'-splice site region of pre-mRNA in the spliceosome. Genes Dev. 1992 Feb;6(2):244–254. doi: 10.1101/gad.6.2.244. [DOI] [PubMed] [Google Scholar]
- Shannon K. W., Guthrie C. Suppressors of a U4 snRNA mutation define a novel U6 snRNP protein with RNA-binding motifs. Genes Dev. 1991 May;5(5):773–785. doi: 10.1101/gad.5.5.773. [DOI] [PubMed] [Google Scholar]
- Sharp P. A. Split genes and RNA splicing. Cell. 1994 Jun 17;77(6):805–815. doi: 10.1016/0092-8674(94)90130-9. [DOI] [PubMed] [Google Scholar]
- Siliciano P. G., Brow D. A., Roiha H., Guthrie C. An essential snRNA from S. cerevisiae has properties predicted for U4, including interaction with a U6-like snRNA. Cell. 1987 Aug 14;50(4):585–592. doi: 10.1016/0092-8674(87)90031-6. [DOI] [PubMed] [Google Scholar]
- Sontheimer E. J., Steitz J. A. The U5 and U6 small nuclear RNAs as active site components of the spliceosome. Science. 1993 Dec 24;262(5142):1989–1996. doi: 10.1126/science.8266094. [DOI] [PubMed] [Google Scholar]
- Steitz J. A. Splicing takes a holliday. Science. 1992 Aug 14;257(5072):888–889. doi: 10.1126/science.1386941. [DOI] [PubMed] [Google Scholar]
- Vankan P., McGuigan C., Mattaj I. W. Domains of U4 and U6 snRNAs required for snRNP assembly and splicing complementation in Xenopus oocytes. EMBO J. 1990 Oct;9(10):3397–3404. doi: 10.1002/j.1460-2075.1990.tb07541.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wassarman D. A. Psoralen crosslinking of small RNAs in vitro. Mol Biol Rep. 1993 Feb;17(2):143–151. doi: 10.1007/BF00996222. [DOI] [PubMed] [Google Scholar]
- Wassarman D. A., Steitz J. A. Interactions of small nuclear RNA's with precursor messenger RNA during in vitro splicing. Science. 1992 Sep 25;257(5078):1918–1925. doi: 10.1126/science.1411506. [DOI] [PubMed] [Google Scholar]
- Watkins K. P., Agabian N. In vivo UV cross-linking of U snRNAs that participate in trypanosome trans-splicing. Genes Dev. 1991 Oct;5(10):1859–1869. doi: 10.1101/gad.5.10.1859. [DOI] [PubMed] [Google Scholar]
- Wise J. A. Guides to the heart of the spliceosome. Science. 1993 Dec 24;262(5142):1978–1979. doi: 10.1126/science.8266091. [DOI] [PubMed] [Google Scholar]
- Wise J. A., Tollervey D., Maloney D., Swerdlow H., Dunn E. J., Guthrie C. Yeast contains small nuclear RNAs encoded by single copy genes. Cell. 1983 Dec;35(3 Pt 2):743–751. doi: 10.1016/0092-8674(83)90107-1. [DOI] [PubMed] [Google Scholar]
- Wolff T., Bindereif A. Conformational changes of U6 RNA during the spliceosome cycle: an intramolecular helix is essential both for initiating the U4-U6 interaction and for the first step of slicing. Genes Dev. 1993 Jul;7(7B):1377–1389. doi: 10.1101/gad.7.7b.1377. [DOI] [PubMed] [Google Scholar]
- Wolff T., Bindereif A. Reconstituted mammalian U4/U6 snRNP complements splicing: a mutational analysis. EMBO J. 1992 Jan;11(1):345–359. doi: 10.1002/j.1460-2075.1992.tb05057.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu J. A., Manley J. L. Base pairing between U2 and U6 snRNAs is necessary for splicing of a mammalian pre-mRNA. Nature. 1991 Aug 29;352(6338):818–821. doi: 10.1038/352818a0. [DOI] [PubMed] [Google Scholar]
- Yean S. L., Lin R. J. U4 small nuclear RNA dissociates from a yeast spliceosome and does not participate in the subsequent splicing reaction. Mol Cell Biol. 1991 Nov;11(11):5571–5577. doi: 10.1128/mcb.11.11.5571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Youvan D. C., Hearst J. E. Reverse transcriptase pauses at N2-methylguanine during in vitro transcription of Escherichia coli 16S ribosomal RNA. Proc Natl Acad Sci U S A. 1979 Aug;76(8):3751–3754. doi: 10.1073/pnas.76.8.3751. [DOI] [PMC free article] [PubMed] [Google Scholar]