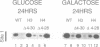Abstract
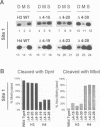
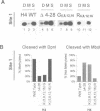
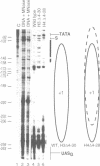
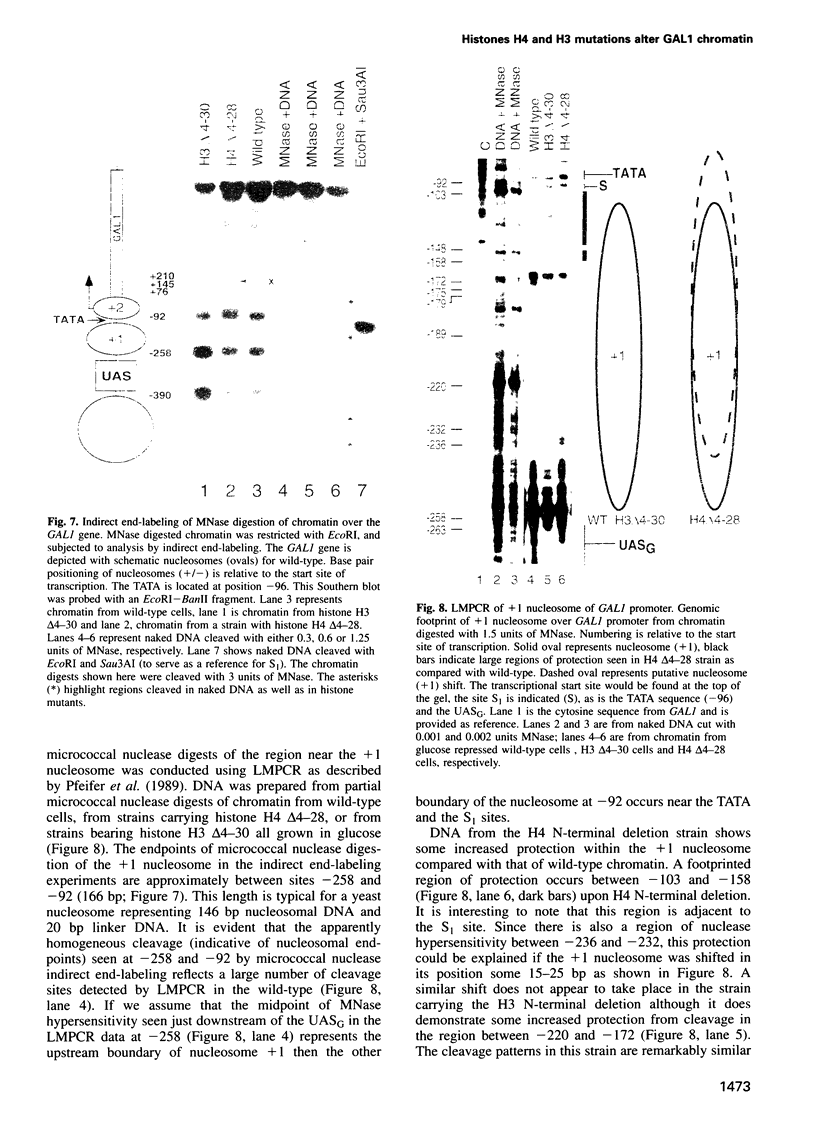
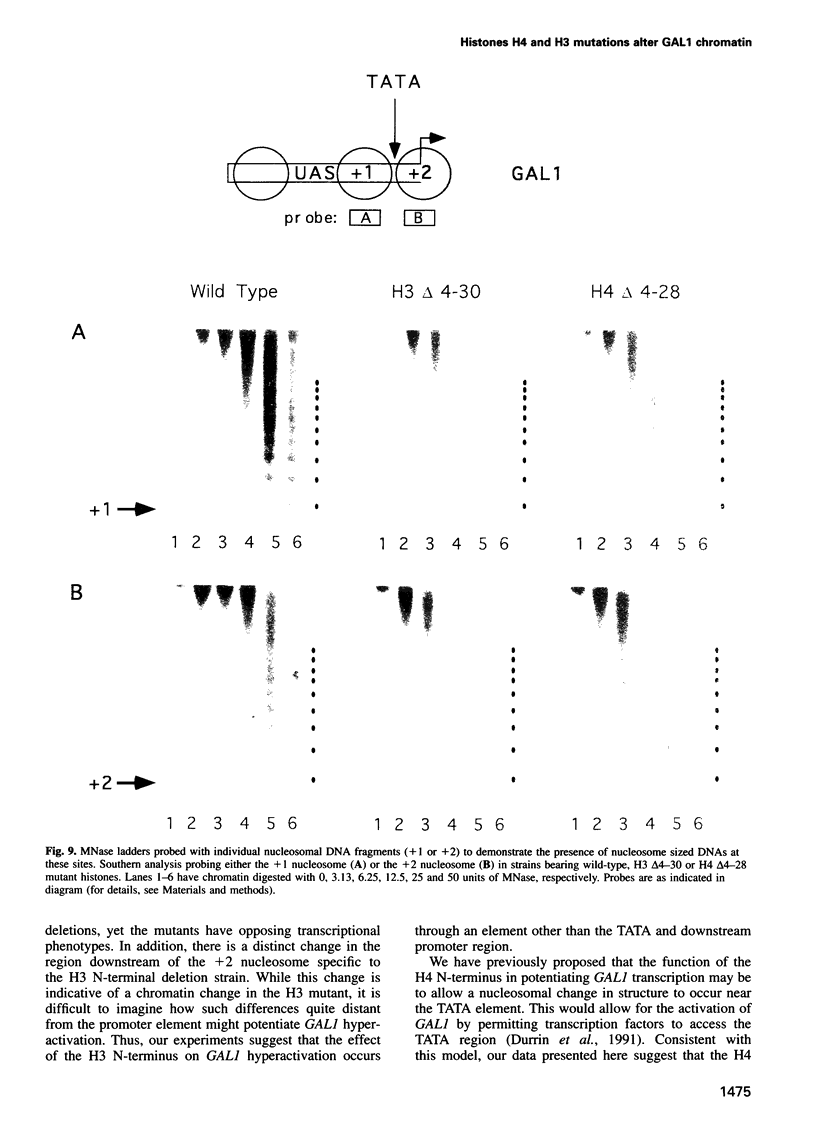
Deletion of the histone H4 N-terminal residues 4-23 decreases activation of the GAL1 promoter as much as 20-fold, while deletion of histone H3 N-terminal residues 4-15 hyperactivates GAL1 approximately 3-fold. In an attempt to understand the mechanisms by which these two different events take place, we have examined the effects of the H4 and H3 lesions on GAL1 chromatin structure. The bacterial dam methylase, which methylates adenine residues of GATC sequences, was used as an in vivo probe for chromatin structure and both indirect end-labeling and ligation mediated PCR (LMPCR) analysis of micrococcal nuclease digestions were used to analyze chromatin in isolated nuclei. We show that while deletions of the H4 and H3 N-termini have similar effects on dam methylase access in the GAL1 coding region, the H4 N-terminal deletion uniquely alters dam access at a region near the TATA element. This change is independent of the transcriptional state of GAL1. In addition, LMPCR analysis of micrococcal nuclease digests of yeast nuclei demonstrate that H4 N-terminal deletion has unique effects on nuclease accessibility in the nucleosomal region upstream of the TATA element. Our results are consistent with the H4 N-terminus mediating activation of GAL1 through its effect on the proximal promoter region near the TATA box. These data also suggest that the H3 N-terminus affects GAL1 hyperactivation through a different promoter element than that affected by H4.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aparicio O. M., Billington B. L., Gottschling D. E. Modifiers of position effect are shared between telomeric and silent mating-type loci in S. cerevisiae. Cell. 1991 Sep 20;66(6):1279–1287. doi: 10.1016/0092-8674(91)90049-5. [DOI] [PubMed] [Google Scholar]
- Axelrod J. D., Reagan M. S., Majors J. GAL4 disrupts a repressing nucleosome during activation of GAL1 transcription in vivo. Genes Dev. 1993 May;7(5):857–869. doi: 10.1101/gad.7.5.857. [DOI] [PubMed] [Google Scholar]
- Durrin L. K., Mann R. K., Grunstein M. Nucleosome loss activates CUP1 and HIS3 promoters to fully induced levels in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1992 Apr;12(4):1621–1629. doi: 10.1128/mcb.12.4.1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Durrin L. K., Mann R. K., Kayne P. S., Grunstein M. Yeast histone H4 N-terminal sequence is required for promoter activation in vivo. Cell. 1991 Jun 14;65(6):1023–1031. doi: 10.1016/0092-8674(91)90554-c. [DOI] [PubMed] [Google Scholar]
- Fedor M. J., Kornberg R. D. Upstream activation sequence-dependent alteration of chromatin structure and transcription activation of the yeast GAL1-GAL10 genes. Mol Cell Biol. 1989 Apr;9(4):1721–1732. doi: 10.1128/mcb.9.4.1721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Gietz D., St Jean A., Woods R. A., Schiestl R. H. Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res. 1992 Mar 25;20(6):1425–1425. doi: 10.1093/nar/20.6.1425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottschling D. E. Telomere-proximal DNA in Saccharomyces cerevisiae is refractory to methyltransferase activity in vivo. Proc Natl Acad Sci U S A. 1992 May 1;89(9):4062–4065. doi: 10.1073/pnas.89.9.4062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han M., Grunstein M. Nucleosome loss activates yeast downstream promoters in vivo. Cell. 1988 Dec 23;55(6):1137–1145. doi: 10.1016/0092-8674(88)90258-9. [DOI] [PubMed] [Google Scholar]
- Hebbes T. R., Clayton A. L., Thorne A. W., Crane-Robinson C. Core histone hyperacetylation co-maps with generalized DNase I sensitivity in the chicken beta-globin chromosomal domain. EMBO J. 1994 Apr 15;13(8):1823–1830. doi: 10.1002/j.1460-2075.1994.tb06451.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hebbes T. R., Thorne A. W., Crane-Robinson C. A direct link between core histone acetylation and transcriptionally active chromatin. EMBO J. 1988 May;7(5):1395–1402. doi: 10.1002/j.1460-2075.1988.tb02956.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoekstra M. F., Malone R. E. Expression of the Escherichia coli dam methylase in Saccharomyces cerevisiae: effect of in vivo adenine methylation on genetic recombination and mutation. Mol Cell Biol. 1985 Apr;5(4):610–618. doi: 10.1128/mcb.5.4.610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffman C. S., Winston F. A transcriptionally regulated expression vector for the fission yeast Schizosaccharomyces pombe. Gene. 1989 Dec 14;84(2):473–479. doi: 10.1016/0378-1119(89)90523-4. [DOI] [PubMed] [Google Scholar]
- Johnson L. M., Fisher-Adams G., Grunstein M. Identification of a non-basic domain in the histone H4 N-terminus required for repression of the yeast silent mating loci. EMBO J. 1992 Jun;11(6):2201–2209. doi: 10.1002/j.1460-2075.1992.tb05279.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson L. M., Kayne P. S., Kahn E. S., Grunstein M. Genetic evidence for an interaction between SIR3 and histone H4 in the repression of the silent mating loci in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1990 Aug;87(16):6286–6290. doi: 10.1073/pnas.87.16.6286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston M. A model fungal gene regulatory mechanism: the GAL genes of Saccharomyces cerevisiae. Microbiol Rev. 1987 Dec;51(4):458–476. doi: 10.1128/mr.51.4.458-476.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kayne P. S., Kim U. J., Han M., Mullen J. R., Yoshizaki F., Grunstein M. Extremely conserved histone H4 N terminus is dispensable for growth but essential for repressing the silent mating loci in yeast. Cell. 1988 Oct 7;55(1):27–39. doi: 10.1016/0092-8674(88)90006-2. [DOI] [PubMed] [Google Scholar]
- Kladde M. P., Simpson R. T. Positioned nucleosomes inhibit Dam methylation in vivo. Proc Natl Acad Sci U S A. 1994 Feb 15;91(4):1361–1365. doi: 10.1073/pnas.91.4.1361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knezetic J. A., Luse D. S. The presence of nucleosomes on a DNA template prevents initiation by RNA polymerase II in vitro. Cell. 1986 Apr 11;45(1):95–104. doi: 10.1016/0092-8674(86)90541-6. [DOI] [PubMed] [Google Scholar]
- Lohr D. Organization of the GAL1-GAL10 intergenic control region chromatin. Nucleic Acids Res. 1984 Nov 26;12(22):8457–8474. doi: 10.1093/nar/12.22.8457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorch Y., LaPointe J. W., Kornberg R. D. Nucleosomes inhibit the initiation of transcription but allow chain elongation with the displacement of histones. Cell. 1987 Apr 24;49(2):203–210. doi: 10.1016/0092-8674(87)90561-7. [DOI] [PubMed] [Google Scholar]
- Mann R. K., Grunstein M. Histone H3 N-terminal mutations allow hyperactivation of the yeast GAL1 gene in vivo. EMBO J. 1992 Sep;11(9):3297–3306. doi: 10.1002/j.1460-2075.1992.tb05408.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Megee P. C., Morgan B. A., Mittman B. A., Smith M. M. Genetic analysis of histone H4: essential role of lysines subject to reversible acetylation. Science. 1990 Feb 16;247(4944):841–845. doi: 10.1126/science.2106160. [DOI] [PubMed] [Google Scholar]
- Park E. C., Szostak J. W. Point mutations in the yeast histone H4 gene prevent silencing of the silent mating type locus HML. Mol Cell Biol. 1990 Sep;10(9):4932–4934. doi: 10.1128/mcb.10.9.4932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfeifer G. P., Steigerwald S. D., Mueller P. R., Wold B., Riggs A. D. Genomic sequencing and methylation analysis by ligation mediated PCR. Science. 1989 Nov 10;246(4931):810–813. doi: 10.1126/science.2814502. [DOI] [PubMed] [Google Scholar]
- Roth S. Y., Shimizu M., Johnson L., Grunstein M., Simpson R. T. Stable nucleosome positioning and complete repression by the yeast alpha 2 repressor are disrupted by amino-terminal mutations in histone H4. Genes Dev. 1992 Mar;6(3):411–425. doi: 10.1101/gad.6.3.411. [DOI] [PubMed] [Google Scholar]
- Selleck S. B., Majors J. In vivo "photofootprint" changes at sequences between the yeast GAL1 upstream activating sequence and "TATA" element require activated GAL4 protein but not a functional TATA element. Proc Natl Acad Sci U S A. 1988 Aug;85(15):5399–5403. doi: 10.1073/pnas.85.15.5399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh J., Klar A. J. Active genes in budding yeast display enhanced in vivo accessibility to foreign DNA methylases: a novel in vivo probe for chromatin structure of yeast. Genes Dev. 1992 Feb;6(2):186–196. doi: 10.1101/gad.6.2.186. [DOI] [PubMed] [Google Scholar]
- St John T. P., Davis R. W. The organization and transcription of the galactose gene cluster of Saccharomyces. J Mol Biol. 1981 Oct 25;152(2):285–315. doi: 10.1016/0022-2836(81)90244-8. [DOI] [PubMed] [Google Scholar]
- Stewart A. F., Reik A., Schütz G. A simpler and better method to cleave chromatin with DNase 1 for hypersensitive site analyses. Nucleic Acids Res. 1991 Jun 11;19(11):3157–3157. doi: 10.1093/nar/19.11.3157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Straka C., Hörz W. A functional role for nucleosomes in the repression of a yeast promoter. EMBO J. 1991 Feb;10(2):361–368. doi: 10.1002/j.1460-2075.1991.tb07957.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson J. S., Ling X., Grunstein M. Histone H3 amino terminus is required for telomeric and silent mating locus repression in yeast. Nature. 1994 May 19;369(6477):245–247. doi: 10.1038/369245a0. [DOI] [PubMed] [Google Scholar]
- Turner B. M. Histone acetylation and control of gene expression. J Cell Sci. 1991 May;99(Pt 1):13–20. doi: 10.1242/jcs.99.1.13. [DOI] [PubMed] [Google Scholar]
- Workman J. L., Roeder R. G. Binding of transcription factor TFIID to the major late promoter during in vitro nucleosome assembly potentiates subsequent initiation by RNA polymerase II. Cell. 1987 Nov 20;51(4):613–622. doi: 10.1016/0092-8674(87)90130-9. [DOI] [PubMed] [Google Scholar]
- Wright J. H., Gottschling D. E., Zakian V. A. Saccharomyces telomeres assume a non-nucleosomal chromatin structure. Genes Dev. 1992 Feb;6(2):197–210. doi: 10.1101/gad.6.2.197. [DOI] [PubMed] [Google Scholar]