Abstract
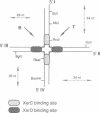
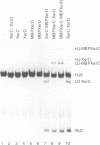
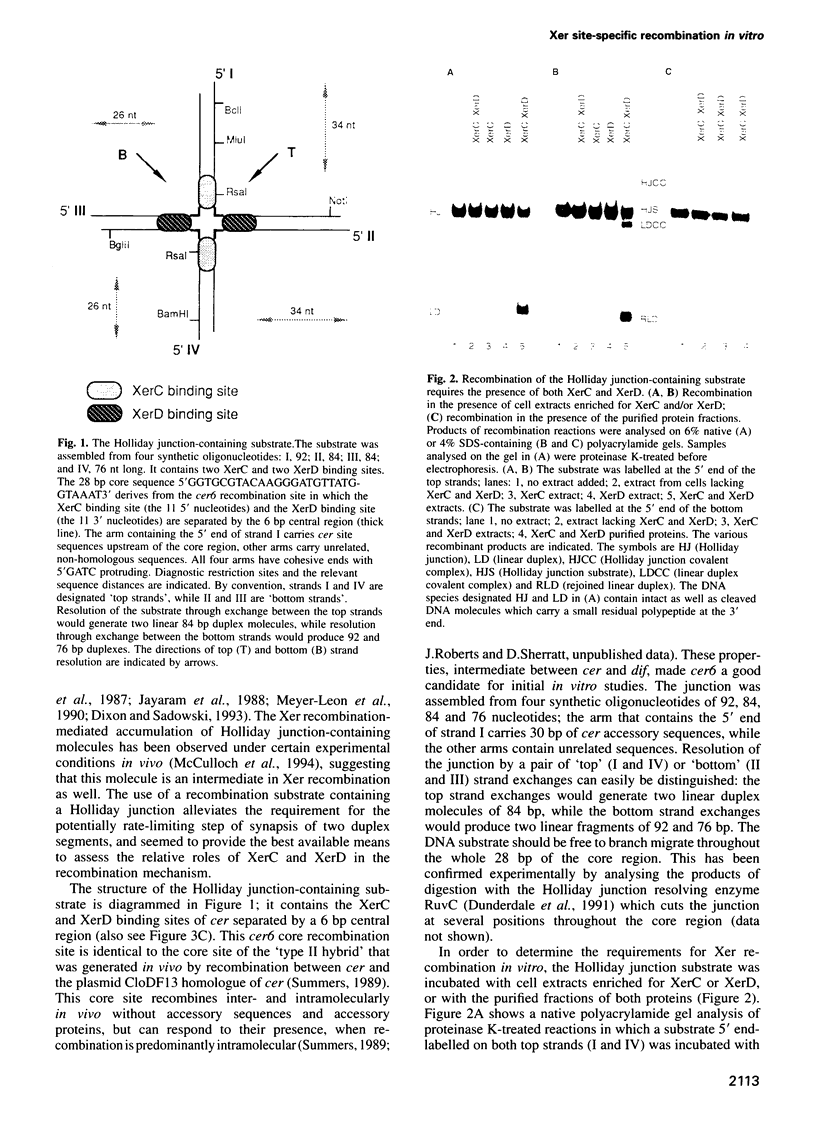
Two related recombinases, XerC and XerD, belonging to the lambda integrase family of enzymes, are required for Xer site-specific recombination in vivo. In order to understand the roles of these proteins in the overall reaction mechanism, an in vitro recombination system using a synthetic Holliday junction-containing substrate has been developed. Recombination of this substrate is efficient and requires both XerC and XerD. However, only exchange of one pair of strands, the one corresponding to the conversion of the Holliday junction intermediate back to the substrate, has been observed. Recombination reactions using XerC and XerD derivatives that are mutant in their presumptive catalytic residues, or are maltose-binding fusion recombinase derivatives, have demonstrated that this pair of strand exchanges is catalysed by XerC. The site of XerC-mediated cleavage has been located to between the last nucleotide of the XerC binding site and the first nucleotide of the central region. Cleavage at this site generates a free 5'-OH and a covalent complex between XerC and the 3' end of the DNA.
Full text
PDF


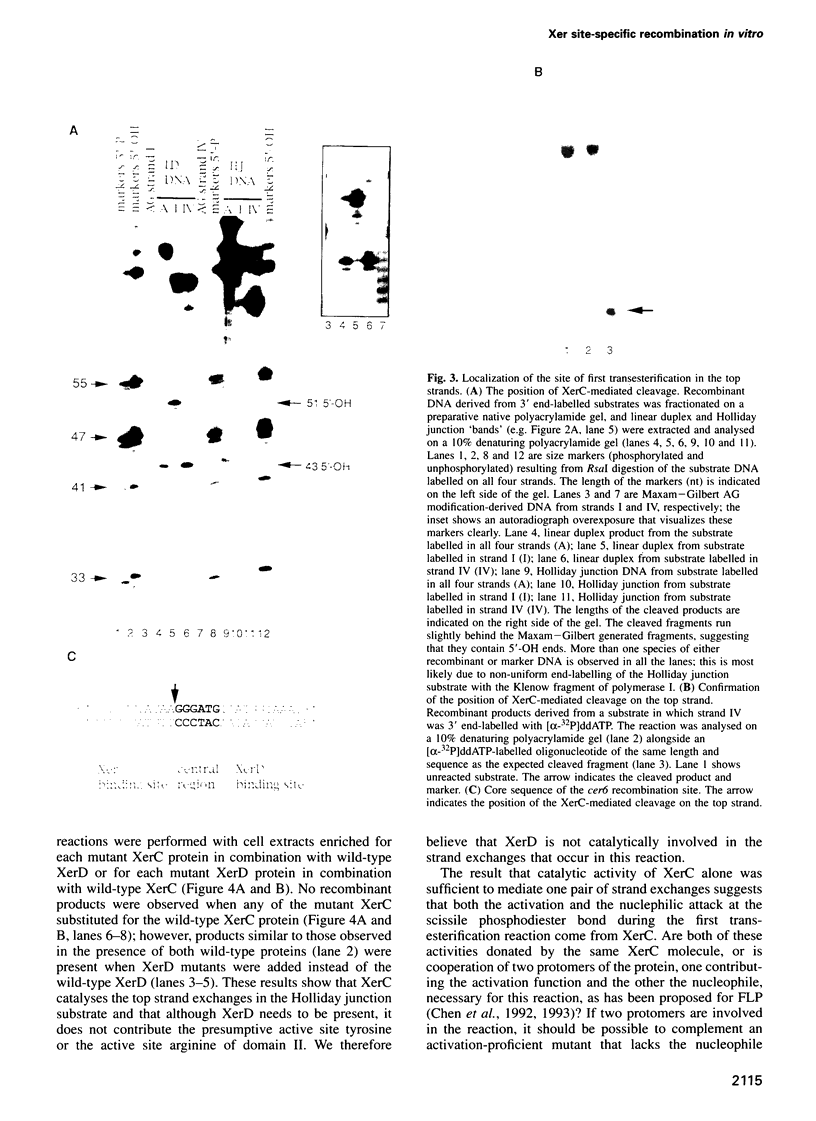





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abremski K. E., Hoess R. H. Evidence for a second conserved arginine residue in the integrase family of recombination proteins. Protein Eng. 1992 Jan;5(1):87–91. doi: 10.1093/protein/5.1.87. [DOI] [PubMed] [Google Scholar]
- Abremski K., Hoess R., Sternberg N. Studies on the properties of P1 site-specific recombination: evidence for topologically unlinked products following recombination. Cell. 1983 Apr;32(4):1301–1311. doi: 10.1016/0092-8674(83)90311-2. [DOI] [PubMed] [Google Scholar]
- Andrews B. J., Proteau G. A., Beatty L. G., Sadowski P. D. The FLP recombinase of the 2 micron circle DNA of yeast: interaction with its target sequences. Cell. 1985 Apr;40(4):795–803. doi: 10.1016/0092-8674(85)90339-3. [DOI] [PubMed] [Google Scholar]
- Argos P., Landy A., Abremski K., Egan J. B., Haggard-Ljungquist E., Hoess R. H., Kahn M. L., Kalionis B., Narayana S. V., Pierson L. S., 3rd The integrase family of site-specific recombinases: regional similarities and global diversity. EMBO J. 1986 Feb;5(2):433–440. doi: 10.1002/j.1460-2075.1986.tb04229.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blakely G., Colloms S., May G., Burke M., Sherratt D. Escherichia coli XerC recombinase is required for chromosomal segregation at cell division. New Biol. 1991 Aug;3(8):789–798. [PubMed] [Google Scholar]
- Blakely G., May G., McCulloch R., Arciszewska L. K., Burke M., Lovett S. T., Sherratt D. J. Two related recombinases are required for site-specific recombination at dif and cer in E. coli K12. Cell. 1993 Oct 22;75(2):351–361. doi: 10.1016/0092-8674(93)80076-q. [DOI] [PubMed] [Google Scholar]
- Burgin A. B., Jr, Nash H. A. Symmetry in the mechanism of bacteriophage lambda integrative recombination. Proc Natl Acad Sci U S A. 1992 Oct 15;89(20):9642–9646. doi: 10.1073/pnas.89.20.9642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J. W., Lee J., Jayaram M. DNA cleavage in trans by the active site tyrosine during Flp recombination: switching protein partners before exchanging strands. Cell. 1992 May 15;69(4):647–658. doi: 10.1016/0092-8674(92)90228-5. [DOI] [PubMed] [Google Scholar]
- Chen J. W., Yang S. H., Jayaram M. Tests for the fractional active-site model in Flp site-specific recombination. Assembly of a functional recombination complex in half-site and full-site strand transfer. J Biol Chem. 1993 Jul 5;268(19):14417–14425. [PubMed] [Google Scholar]
- Colloms S. D., Sykora P., Szatmari G., Sherratt D. J. Recombination at ColE1 cer requires the Escherichia coli xerC gene product, a member of the lambda integrase family of site-specific recombinases. J Bacteriol. 1990 Dec;172(12):6973–6980. doi: 10.1128/jb.172.12.6973-6980.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Craig N. L., Nash H. A. The mechanism of phage lambda site-specific recombination: site-specific breakage of DNA by Int topoisomerase. Cell. 1983 Dec;35(3 Pt 2):795–803. doi: 10.1016/0092-8674(83)90112-5. [DOI] [PubMed] [Google Scholar]
- Craig N. L. The mechanism of conservative site-specific recombination. Annu Rev Genet. 1988;22:77–105. doi: 10.1146/annurev.ge.22.120188.000453. [DOI] [PubMed] [Google Scholar]
- Dixon J. E., Sadowski P. D. Resolution of synthetic chi structures by the FLP site-specific recombinase. J Mol Biol. 1993 Dec 5;234(3):522–533. doi: 10.1006/jmbi.1993.1608. [DOI] [PubMed] [Google Scholar]
- Dunderdale H. J., Benson F. E., Parsons C. A., Sharples G. J., Lloyd R. G., West S. C. Formation and resolution of recombination intermediates by E. coli RecA and RuvC proteins. Nature. 1991 Dec 19;354(6354):506–510. doi: 10.1038/354506a0. [DOI] [PubMed] [Google Scholar]
- Friesen H., Sadowski P. D. Mutagenesis of a conserved region of the gene encoding the FLP recombinase of Saccharomyces cerevisiae. A role for arginine 191 in binding and ligation. J Mol Biol. 1992 May 20;225(2):313–326. doi: 10.1016/0022-2836(92)90924-9. [DOI] [PubMed] [Google Scholar]
- Han Y. W., Gumport R. I., Gardner J. F. Complementation of bacteriophage lambda integrase mutants: evidence for an intersubunit active site. EMBO J. 1993 Dec;12(12):4577–4584. doi: 10.1002/j.1460-2075.1993.tb06146.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoess R. H., Abremski K. Mechanism of strand cleavage and exchange in the Cre-lox site-specific recombination system. J Mol Biol. 1985 Feb 5;181(3):351–362. doi: 10.1016/0022-2836(85)90224-4. [DOI] [PubMed] [Google Scholar]
- Hoess R., Wierzbicki A., Abremski K. Isolation and characterization of intermediates in site-specific recombination. Proc Natl Acad Sci U S A. 1987 Oct;84(19):6840–6844. doi: 10.1073/pnas.84.19.6840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu P. L., Landy A. Resolution of synthetic att-site Holliday structures by the integrase protein of bacteriophage lambda. Nature. 1984 Oct 25;311(5988):721–726. doi: 10.1038/311721a0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jayaram M., Crain K. L., Parsons R. L., Harshey R. M. Holliday junctions in FLP recombination: resolution by step-arrest mutants of FLP protein. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7902–7906. doi: 10.1073/pnas.85.21.7902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jayaram M. Phosphoryl transfer in Flp recombination: a template for strand transfer mechanisms. Trends Biochem Sci. 1994 Feb;19(2):78–82. doi: 10.1016/0968-0004(94)90039-6. [DOI] [PubMed] [Google Scholar]
- Kitts P. A., Nash H. A. An intermediate in the phage lambda site-specific recombination reaction is revealed by phosphorothioate substitution in DNA. Nucleic Acids Res. 1988 Jul 25;16(14B):6839–6856. doi: 10.1093/nar/16.14.6839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitts P. A., Nash H. A. Homology-dependent interactions in phage lambda site-specific recombination. Nature. 1987 Sep 24;329(6137):346–348. doi: 10.1038/329346a0. [DOI] [PubMed] [Google Scholar]
- Kuempel P. L., Henson J. M., Dircks L., Tecklenburg M., Lim D. F. dif, a recA-independent recombination site in the terminus region of the chromosome of Escherichia coli. New Biol. 1991 Aug;3(8):799–811. [PubMed] [Google Scholar]
- Landy A. Dynamic, structural, and regulatory aspects of lambda site-specific recombination. Annu Rev Biochem. 1989;58:913–949. doi: 10.1146/annurev.bi.58.070189.004405. [DOI] [PubMed] [Google Scholar]
- Landy A. Mechanistic and structural complexity in the site-specific recombination pathways of Int and FLP. Curr Opin Genet Dev. 1993 Oct;3(5):699–707. doi: 10.1016/s0959-437x(05)80086-3. [DOI] [PubMed] [Google Scholar]
- Lee J., Jayaram M. Mechanism of site-specific recombination. Logic of assembling recombinase catalytic site from fractional active sites. J Biol Chem. 1993 Aug 15;268(23):17564–17570. [PubMed] [Google Scholar]
- Lee J., Serre M. C., Yang S. H., Whang I., Araki H., Oshima Y., Jayaram M. Functional analysis of Box II mutations in yeast site-specific recombinases Flp and R. Significance of amino acid conservation within the Int family and the yeast sub-family. J Mol Biol. 1992 Dec 20;228(4):1091–1103. doi: 10.1016/0022-2836(92)90317-d. [DOI] [PubMed] [Google Scholar]
- Lilley D. M., Clegg R. M. The structure of branched DNA species. Q Rev Biophys. 1993 May;26(2):131–175. doi: 10.1017/s0033583500004054. [DOI] [PubMed] [Google Scholar]
- McCulloch R., Coggins L. W., Colloms S. D., Sherratt D. J. Xer-mediated site-specific recombination at cer generates Holliday junctions in vivo. EMBO J. 1994 Apr 15;13(8):1844–1855. doi: 10.1002/j.1460-2075.1994.tb06453.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer-Leon L., Huang L. C., Umlauf S. W., Cox M. M., Inman R. B. Holliday intermediates and reaction by-products in FLP protein-promoted site-specific recombination. Mol Cell Biol. 1988 Sep;8(9):3784–3796. doi: 10.1128/mcb.8.9.3784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer-Leon L., Inman R. B., Cox M. M. Characterization of Holliday structures in FLP protein-promoted site-specific recombination. Mol Cell Biol. 1990 Jan;10(1):235–242. doi: 10.1128/mcb.10.1.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer-Leon L., Senecoff J. F., Bruckner R. C., Cox M. M. Site-specific genetic recombination promoted by the FLP protein of the yeast 2-micron plasmid in vitro. Cold Spring Harb Symp Quant Biol. 1984;49:797–804. doi: 10.1101/sqb.1984.049.01.090. [DOI] [PubMed] [Google Scholar]
- Mizuuchi K., Weisberg R., Enquist L., Mizuuchi M., Buraczynska M., Foeller C., Hsu P. L., Ross W., Landy A. Structure and function of the phage lambda att site: size, int-binding sites, and location of the crossover point. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):429–437. doi: 10.1101/sqb.1981.045.01.057. [DOI] [PubMed] [Google Scholar]
- Nunes-Düby S. E., Matsumoto L., Landy A. Site-specific recombination intermediates trapped with suicide substrates. Cell. 1987 Aug 28;50(5):779–788. doi: 10.1016/0092-8674(87)90336-9. [DOI] [PubMed] [Google Scholar]
- Nunes-Düby S. E., Tirumalai R. S., Dorgai L., Yagil E., Weisberg R. A., Landy A. Lambda integrase cleaves DNA in cis. EMBO J. 1994 Sep 15;13(18):4421–4430. doi: 10.1002/j.1460-2075.1994.tb06762.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan G., Sadowski P. D. Ligation activity of FLP recombinase. The strand ligation activity of a site-specific recombinase using an activated DNA substrate. J Biol Chem. 1992 Jun 25;267(18):12397–12399. [PubMed] [Google Scholar]
- Pargellis C. A., Nunes-Düby S. E., de Vargas L. M., Landy A. Suicide recombination substrates yield covalent lambda integrase-DNA complexes and lead to identification of the active site tyrosine. J Biol Chem. 1988 Jun 5;263(16):7678–7685. [PubMed] [Google Scholar]
- Parsons R. L., Prasad P. V., Harshey R. M., Jayaram M. Step-arrest mutants of FLP recombinase: implications for the catalytic mechanism of DNA recombination. Mol Cell Biol. 1988 Aug;8(8):3303–3310. doi: 10.1128/mcb.8.8.3303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rice P. A., Steitz T. A. Model for a DNA-mediated synaptic complex suggested by crystal packing of gamma delta resolvase subunits. EMBO J. 1994 Apr 1;13(7):1514–1524. doi: 10.2210/pdb1gdr/pdb. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadowski P. D., Lee D. D., Andrews B. J., Babineau D., Beatty L., Morse M. J., Proteau G., Vetter D. In vitro systems for genetic recombination of the DNAs of bacteriophage T7 and yeast 2-micron circle. Cold Spring Harb Symp Quant Biol. 1984;49:789–796. doi: 10.1101/sqb.1984.049.01.089. [DOI] [PubMed] [Google Scholar]
- Sadowski P. Site-specific recombinases: changing partners and doing the twist. J Bacteriol. 1986 Feb;165(2):341–347. doi: 10.1128/jb.165.2.341-347.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Senecoff J. F., Bruckner R. C., Cox M. M. The FLP recombinase of the yeast 2-micron plasmid: characterization of its recombination site. Proc Natl Acad Sci U S A. 1985 Nov;82(21):7270–7274. doi: 10.1073/pnas.82.21.7270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stark W. M., Boocock M. R., Sherratt D. J. Catalysis by site-specific recombinases. Trends Genet. 1992 Dec;8(12):432–439. [PubMed] [Google Scholar]
- Stirling C. J., Colloms S. D., Collins J. F., Szatmari G., Sherratt D. J. xerB, an Escherichia coli gene required for plasmid ColE1 site-specific recombination, is identical to pepA, encoding aminopeptidase A, a protein with substantial similarity to bovine lens leucine aminopeptidase. EMBO J. 1989 May;8(5):1623–1627. doi: 10.1002/j.1460-2075.1989.tb03547.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stirling C. J., Szatmari G., Stewart G., Smith M. C., Sherratt D. J. The arginine repressor is essential for plasmid-stabilizing site-specific recombination at the ColE1 cer locus. EMBO J. 1988 Dec 20;7(13):4389–4395. doi: 10.1002/j.1460-2075.1988.tb03338.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Summers D. K. Derivatives of ColE1 cer show altered topological specificity in site-specific recombination. EMBO J. 1989 Jan;8(1):309–315. doi: 10.1002/j.1460-2075.1989.tb03378.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Summers D. K., Sherratt D. J. Multimerization of high copy number plasmids causes instability: CoIE1 encodes a determinant essential for plasmid monomerization and stability. Cell. 1984 Apr;36(4):1097–1103. doi: 10.1016/0092-8674(84)90060-6. [DOI] [PubMed] [Google Scholar]
- Summers D. K., Sherratt D. J. Resolution of ColE1 dimers requires a DNA sequence implicated in the three-dimensional organization of the cer site. EMBO J. 1988 Mar;7(3):851–858. doi: 10.1002/j.1460-2075.1988.tb02884.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vetter D., Andrews B. J., Roberts-Beatty L., Sadowski P. D. Site-specific recombination of yeast 2-micron DNA in vitro. Proc Natl Acad Sci U S A. 1983 Dec;80(23):7284–7288. doi: 10.1073/pnas.80.23.7284. [DOI] [PMC free article] [PubMed] [Google Scholar]