Abstract
The endometrium contains a population of immune cells that undergo changes during implantation and pregnancy. The majority of these cells are uterine natural killer (uNK) cells; however, it is unclear how these cells interact with endometrial epithelial cells. Therefore, we investigated the paracrine effects of the uNK cell-secretion medium on the gene expression profile of endometrial epithelial cells in vitro through microarray analysis. Our results, which were verified by qRT-PCR and western blot, revealed that soluble factors from uNK cells alter the gene expression profiles of epithelial cells. The upregulated genes included interleukin-15 (IL-15) and interleukin-15 receptor alpha (IL-15RA), which result in a loop that stimulates uNK cell proliferation. In addition, vascular endothelial growth factor C (VEGF-C) and chemokine (C-X-C motif) ligand 10 (CXCL-10) were also determined to be upregulated in epithelial cells, which suggests that uNK cells work synergistically with epithelial cells to support implantation and pregnancy. In addition, oriental herbal medicines have been used to treat infertility since ancient times; however, we failed to find that Zi Dan Yin can regulate these endometrial paracrine effects.
1. Introduction
In the human endometrium, uterine leucocytes undergo cyclic changes during the menstrual cycle. Uterine adaptation to pregnancy includes highly regulated immune cell trafficking [1]. According to recent studies, uterine natural killer (uNK) cells are an in vivo factor that contributes to the pathological elongation of the window of endometrial receptivity [2]. The genetic deficiency of uNK cells results in a lack of remodeling of the uterine vasculature, leading to hypertrophied vascular media, swollen endothelial cells, and narrow vessel lumens [3]. Abnormal activation and numbers of uNK cells have been implicated in pregnancy complications [4].
uNK cells are the most abundant leucocytes during the implantation window and early pregnancy [5]. This sharp increase is likely due to the proliferation of existing uNK cells [6–8]. Hormonal effects may control this proliferation. However, to date, progesterone receptors have not been localized on these cells [9]; therefore, it has been proposed that progesterone may exert its effects on uNK cells indirectly via cytokines, such as interleukin-15 (IL-15), and other soluble factors [8, 10]. A previous study indicated that the paracrine communication between uterine leucocytes (>85% uNK) and uterine stromal cells can upregulate IL-15 and interleukin-15 receptor alpha (IL-15RA) to induce the proliferation of uNK cells and contributes to the support of trophoblast migration during implantation [11].
uNK cells are resident throughout the endometrium at the time of conception and throughout early pregnancy. Although uNK cells predominantly localize to the uterine stroma and the surrounding areas of the spiral arteries, Matrigel-supported cocultures of endothelial cells and uNK cells have shown that vascular endothelial growth factor C (VEGFC) producing uNK cells induce transporter 1 (TAP-1) expression in endothelial cells [12]. Furthermore, the endometrium undergoes precisely defined morphological and biochemical changes during implantation and early pregnancy; thus, there is no doubt that the preparation of epithelial cells is directed towards these process. We hypothesize that there similar paracrine communications occur between uNK cells and uterine epithelial cells. Our results revealed that soluble factors from uNK cells have substantial effects on endometrial epithelial gene expression. uNK cells were found to upregulate transcript levels in epithelial cells that are known to promote uNK cell proliferation, and these upregulated transcripts may contribute to the preparation for implantation and trophoblast migration and invasion.
In China, oriental herbal medicines have been used to treat infertility since ancient times [13]. Zi Dan Yin (ZDY), a compound of nine herbs (Table 1), was found to improve the morphology of the endometrium in mice [14]. Although the treatment effects of ZDY on mice have been confirmed, it is unclear whether ZDY contributes to the local immune responses. However, our results do not support the hypothesis that ZDY can improve the paracrine effects between uNK and endometrial epithelial cells.
Table 1.
Composition of Zi Dan Yin (ZDY).
| Components | Ratio |
|---|---|
| (1) Sheng Di (Rehmannia glutinosa (Gaertn.) Libosch., root) | 15 |
| (2) Dan Shen (Salviae Miltiorrhizae Bge., root) | 10 |
| (3) Dang gui (Angelica sinensis (Oliv.) Diels., root) | 12 |
| (4) Chuan Duan (Dipsacus asperoides C. Y. Cheng et T.M. Ai., root) | 15 |
| (5) Du Zhong (Eucommia ulmoides Oliv., cortex) | 12 |
| (6) Shan Yao (Dioscorea opposita Thunb., rhizome) | 15 |
| (7) Mei Gui-hua (Rosa rugosa Thunb., flower) | 6 |
| (8) Chuan Xiong (Ligusticum Chuanxiong Hort., rhizome) | 6 |
| (9) Yi Yi-ren (Coix lacryma-jobi L. var. ma-yuen (Roman.) Stapf., seed) | 12 |
2. Materials and Methods
2.1. Ethics Statement
All of the subjects understood and signed the informed consent form before participation. The experimental protocols were approved by the Ethics Committee of the Dongfang Hospital Human Ethics Committee (no. 2011090201).
2.2. Collection of Material
Decidual tissues were obtained from healthy women undergoing elective termination of a normal pregnancy between 7 and 8 weeks of gestation. The endometrial tissues were collected from endometrial biopsies taken during the proliferative phase of the cycle from women undergoing laparoscopy (Dongfang Hospital of Beijing University of Chinese Medicine, China) for benign disease. The average age was 30 ± 1.3 years, and the body mass index (BMI) was 22.1 ± 0.8 kg/m2. All of the nine women were Chinese of Han ethnicity. The exclusion criteria were hormonal stimulation, cancerous lesions, and irregular menstrual bleeding.
2.3. Isolation of the Uterine NK (uNK) Cells
The uNK cells were isolated as previously described [15, 16]. Briefly, the decidual tissues were extensively washed with Ca2+- and Mg2+-free Hank's balanced salt solution (HBSS) containing antibiotics, minced thoroughly between two scalpels into fragments of 1-2 mm3, and digested for 1 h at 37°C with gentle agitation in HBSS with 0.1% (w/v) collagenase I (Gibco, USA). The cell suspensions were layered over Ficoll-Hypaque medium (General Electric, USA) and centrifuged at 800 ×g and room temperature for 25 min. The cells at the interface were washed twice with Roswell Park Memorial Institute (RPMI)-1640 media with 10% fetal calf serum (FCS) and antibiotics. After incubation for 20 min at 4°C with anti-CD56 microbeads (Miltenyi Biotec, Ltd., Germany), the cells were washed with washing buffer (PBS, 2 mM EDTA, and 0.5%BSA (w/v)) and then loaded onto a MiniMACS Separator (MS) column in a MiniMACS magnet (Miltenyi Biotec, Ltd., Germany). After flushing the MS column three times, the CD56+ cells were flushed as indicated by the manufacturer. The purity of the uNK cells was >90% CD56+CD3− according to the flow cytometric analysis. The uNK cells were cultured in RPMI 1640 media with 1% FCS and IL-15 (10 ng/mL) (R&D systems, USA), in which to maintain the viability of the purified uNK cells [17].
2.4. Production of uNK Cell-Secretion Medium
The uNK cell-secretion medium was prepared using 200 μL of RPMI 1640 media with 1% FCS and IL-15 (10 ng/mL) containing 5 × 105 of the purified uNK cells. The cell suspension was placed in the upper chamber of a 0.4 μm pore hanging cell culture insert (Millipore, USA) in a 24-well tissue plate, and 1300 μL of the same media without cells was placed in the lower chamber. As a result, only the soluble molecules from the uNK cells can pass through the filter into the lower chamber. The control medium was 1500 μL of the RPMI 1640 media with 1% FCS and IL-15 (10 ng/mL), and this control medium was also used in the subsequent experiments. After incubation for 24 h at 37°C, the uNK cell-secretion medium (in the lower chamber) and the control medium were collected and frozen at −80°C. The cells in the upper chamber were collected, and the cell viability was measured using a live/dead viability kit (Invitrogen, USA). Only the conditional medium from uNK populations with less than 35% dead cells after overnight incubation was used in the subsequent experiments.
2.5. Human Endometrial Epithelial Cell Culture
Human endometrial tissue was cut using scissors and dissociated into single-cell suspensions using 0.1% (w/v) collagenase I for 50–60 min. Every 15 min, the digests were pipetted, and their dissociation was monitored by microscopy. The cell suspensions were filtered using a 40-mesh screen to separate the single cells from undigested tissue. To remove the erythrocytes, the cells were resuspended in 4 mL of Dulbecco's modified Eagle's medium and Ham's nutrient F12 (DMEM/F12) with 1% FCS, layered over Ficoll-Paque PLUS, and centrifuged for 25 min at 800 ×g. The leukocytes were removed with CD45-coated Dynabeads (Dynal Biotech, USA). The purified epithelial cell suspensions were then obtained through a further round of magnetic bead sorting using Collection Epithelial Dynabeads (Dynal Biotech, USA). The epithelial cell preparations were >95% pure.
The endometrial epithelial cells were cultured in serum-free bronchial epithelial growth medium (BEGM, Clonetics Crop) containing 2 mL of bovine pituitary extract (BPE), 0.5 mL of insulin, 0.5 mL of hydrocortisone (HC), 0.5 mL of gentamicin sulfate and amphotericin-B (GA-1000), 0.5 mL of retinoic acid, 0.5 mL of transferrin, 0.5 mL of triiodothyronine, 0.5 mL of epinephrine, and 0.5 mL of recombinant human epidermal growth factor (hEGF) (all supplied by Clonetics Corp., USA). The media was changed every five days. After two weeks, the epithelial cells were passed from six-well plates into 25 cm2 cell culture flasks.
2.6. ZDY Preparation
All of the crude drugs of ZDY were obtained from the Pharmacy Department of Dongfang Hospital of Beijing University of Chinese Medicine (Beijing Province, China). The quality of the raw herbs was controlled according to the requirement of the Pharmacopoeia of China. The aqueous extract of ZDY was prepared as follows. In brief, nine medicinal materials were mixed at the appropriate proportion, macerated for 1 h with eight volumes of distilled water and then decocted for 2 h. The cooled extract was then filtered. This extraction procedure was repeated twice. The extracts were then combined and concentrated by boiling to a final volume of 100 mL (10.3 g/mL). The ZDY extract concentration of 2 mg/mL was further diluted with the epithelial cell culture medium and passed through 0.2 μm sterilization filters for use in subsequent studies. The ZDY extract was stored at 2–8°C until further use. The concentration of ZDY was used according to previous analyses using a cell counting kit-8 (CCK-8).
2.7. Treatment of Epithelial Cells with uNK Cell-Secretion Medium
The endometrial epithelial cells were allocated into three groups: control, uNK, and ZDY groups. The cells in the control and uNK groups were treated with DMEM medium with 10% FBS for 24 h, and the cells in the ZDY group were treated with 2 mg/mL ZDY for 24 h. All of the cells were then washed twice with PBS and placed in serum-free DMEM for 24 h prior to the subsequent experiments. The uNK cell-secretion medium was pooled from different batches to decrease the variability. The DMEM in the ZDY and uNK groups was then replaced with 80% uNK cell-secretion medium and 20% serum-free DMEM. The control group was treated with 80% control medium and 20% serum-free DMEM. After 6 h of incubation, the cells from the three groups were collected.
2.8. Microarray Experiments
The total RNA from endometrial epithelial cells from the control, uNK, and ZDY groups was extracted using Trizol (Invitrogen, USA). The RNA was purified using the RNeasy Mini Kit (Qiagen, Germany) according to the manufacturer's instructions, quantified by spectrophotometry. The microarray analysis was performed using the GeneChip 3′ IVT Express Kit (Affymetrix). Briefly, the total RNA was subjected to reverse transcription, first-strand cDNA synthesis, double-strand DNA, in vitro transcription, and fragmentation. The samples were hybridized onto the GeneChip PrimeView Human Gene Expression Array (Affymetrix, USA), which covers more than 36,000 transcripts and variants. After 16 h of hybridization at 45°C, the arrays were washed on a Fluidics Station 450 (Affymetrix, USA) and then scanned with a Scanner 3000 (Affymetrix, USA) to obtain the quantitative gene expression levels. The endometrial epithelial cells from the control, uNK, and ZDY groups were processed simultaneously throughout the experimental process, and three chips were used for each group.
2.9. Quantitative Real-Time PCR (qRT-PCR) Analysis
To verify the results obtained from the microarray experiments, qRT-PCR verification was performed for five genes: chemokine (C-X-C motif) ligand 10 (CXCL-10), interleukin-15 (IL-15), intercellular adhesion molecule 1 (ICAM-1), NLR family CARD domain containing 5 (NLRC-5), and interferon regulatory factor 1 (IRF-1). The cells from the control, uNK, and ZDY groups were washed twice with PBS, and the total RNA was then extracted using Trizol. The RNA was quantified by spectrophotometry. Reverse transcription was performed with 8 μL of total RNA in a 20 µL reaction volume using a standard cDNA Synthesis Kit (Takara BIO, Japan). The qRT-PCR primer sequences for the target genes were self-designed, and the PCR primers were ordered from Invitrogen. The primer sequences for target genes are shown in Table 2.
Table 2.
Sequences of primers used in the quantitative real-time PCR analysis.
| Gene | Forward primer | Reverse primer | Amplicon |
|---|---|---|---|
| CXCL-10 | CTTTCTGACTCTAAGTGGCATTC | CACCCTTCTTTTTCATTGTAGCAA | 176 bp |
| IL-15 | TGGCTGCTGGAAACCC | CACAAGTAGCACTGGATGGAAAT | 123 bp |
| ICAM-1 | GAGGAAGGAGCAAGACTCAA | AGCATACCCAATAGGCAGCAAG | 141 bp |
| NLRC-5 | AAACTTGATGACTCCTCCCTTACTT | TTAGACCTGGCTTTGTCCCTTAC | 120 bp |
| IRF-1 | CCAGAAAAGCATAACACCAATCC | CCACTTTCCTTCACATTTCACTG | 144 bp |
| GAPDH | GAGCCAAAAGGGTCATCATCT | AGGGGCCATCCACAGTCTTC | 231 bp |
For each qRT-PCR reaction, the typical thermal cycling conditions included an initial activation step of 95°C for 5 min and 45 cycles of 95°C each for 30 sec, 65°C for 35 sec, and 72°C for 5 min. The PCR reactions were performed on an ABI Prism 7700 Sequence Detection System (Applied Biosystems, USA). For comparison of the target mRNA levels, the cDNA concentration was normalized with that of the glyceraldehyde-3-phosphate dehydrogenase (GAPDH) PCR product. The target mRNA expression was analyzed using the 2-ΔΔCt algorithm.
2.10. Western Blot Analysis of ICAM-1
The cells from the control, uNK, and ZDY groups were lysed with RIPA lysis buffer (Applygen, China) supplemented with a protease inhibitor cocktail (Applygen, China). The protein concentrations were quantified using bicinchoninic acid (BCA; Applygen, China). The proteins were separated using 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to nitrocellulose. The blots were blocked overnight in Tris-buffered-saline with Tween (TBST) containing 5% dried milk, washed three times for 5 min each in TBST, and then incubated overnight in a rocker at 4°C with a cocktail containing ICAM-1 (Abcam, UK; at a dilution of 1 : 1000) and 2% dried milk in TBST. The blots were then washed three times for 10 min each in TBST and incubated with the secondary antibody (Applygen, China; at a dilution of 1 : 10000) for 1 h at room temperature in a rocker. The blots were washed three times for 10 min each in TBST and then visualized with the Super ECL Plus Detection Reagent (Applygen, China). The ECL signals were detected with the Quantity One software (Bio-Rad, USA). GAPDH (Abcam, UK) was used as the internal control to validate the amount of protein.
2.11. Statistical Analysis
All of the variables were tested in three different culture experiments. The data were analyzed by ANOVA. In the microarray experiments, the significance level was determined by ANOVA with Benjamini-Hochberg correction (P < 0.001). We selected the up- and downregulated genes that showed a median fold change greater than 2 and a P < 0.001. The data from the qRT-PCR and western blot analyses are presented as means ± SEM after normalization with the average value of the housekeeping gene obtained for each group, and the significance level was P < 0.01. The graphs of the data were produced using the Microsoft Excel software.
3. Results
3.1. Microarray Experiments
Gene expression profiling using a microarray was used to compare the transcript expression levels in epithelial cells treated with control medium, uNK cell-secretion medium, and ZDY+ uNK cell-secretion medium.
This analysis identified 169 transcripts that exhibited a statistically significant change in their median expression level (either upregulated or downregulated) in response to uNK cell-secretion medium compared with their control level. These transcripts are listed in Table 3. A total of 40 genes were found upregulated, and 129 genes were downregulated. The largest group of upregulated genes included immunomodulatory gene regulators, cytokines, such as interleukin-15 (IL-15), chemokine (C-X-C motif) ligand 10 (CXCL-10), and vascular endothelial growth factor C (VEGF-C), and cytokine receptors, such as interleukin-15 receptor alpha (IL-15RA). Additionally, the upregulated transcripts encode proteins associated with transport, structuration, and transcription. The level of adhesion molecule intercellular adhesion molecule 1 (ICAM-1) was also altered. ICAM-1 binds to the leucocyte integrins LFA-1 and Mac-1 [18, 19], which are expressed on uNK cells. Thus, this change in the level of ICAM-1 should increase the localized interactions between uNK cells and epithelial cells. A number of transcripts regulating enzyme activity, such as the deltex 3-like (Drosophila) (DYX3L) and the proteasome (prosome, macropain) subunit beta type 9 (large multifunctional peptidase 2) (PSMB9), and several transcripts regulating nucleotide metabolism genes, such as guanylate binding protein 5 (GBP5) and guanylate binding protein 1 interferon-inducible (GBP1), were also upregulated. In addition, genes with various functions, such as the icon binding complement component 1S subcomponent (C1S), SP110 nuclear body protein (SP110), tryptophan metabolism indoleamine 2,3-dioxygenase 1 (IDO1), and tryptophanyl-tRNA synthetase (WARS), and several genes with unknown function were found to be upregulated. The largest group of downregulated genes encodes transcription factors, such as B double prime 1 subunit of RNA polymerase III transcription initiation factor IIIB (BDP1) and thyroid hormone receptor interactor 11 (TRIP11). The levels of immunomodulatory gene regulators, including interleukin-36 gamma (IL-36G) and interleukin-33 (IL-33), were also altered. The levels of transcripts encoding molecules responsible for signaling, transport, structuration, and cell-cell adhesion were also downregulated. In addition, genes with various functions, such as ion binding, kinase, enzyme activity, and nucleotide metabolism, were found to be downregulated.
Table 3.
Transcripts that were altered by more than twofold in endometrial epithelial cells by stimulation with the uNK cell-secretion medium (P < 0.001). The transcript levels were obtained using a GeneChip PrimeView Human Gene Expression Array.
| Gene symbol | Fold change |
Gene ID |
Description |
|---|---|---|---|
| Upregulated genes: 40 transcripts | |||
| Cytokines/membrane proteins/immunological factors |
|||
| NLRC5 | 14.1 | 84166 | NLR family, CARD domain containing 5 |
| IL15 | 5.8 | 3600 | Interleukin-15 |
| IFIT3 | 5.8 | 3437 | Interferon-induced protein with tetratricopeptide repeats 3 |
| CXCL10 | 3.1 | 3627 | Chemokine (C-X-C motif) ligand 10 |
| IFIH1 | 2.9 | 64135 | Interferon induced with helicase C domain 1 |
| IFIT5 | 2.8 | 24138 | Interferon-induced protein with tetratricopeptide repeats 5 |
| IL-15RA | 2.7 | 3601 | Interleukin-15 receptor, alpha |
| VEGFC | 2.5 | 7424 | Vascular endothelial growth factor C |
| PDCD1LG2 | 2.5 | 80380 | Programmed cell death 1 ligand 2 |
| Transporters | |||
| TAP1 | 3.4 | 6890 | Transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| TAP2 | 2.6 | 6891 | Transporter 2, ATP-binding cassette, sub-family B (MDR/TAP) |
| APOL3 | 2.1 | 80833 | Apolipoprotein L, 3 |
| VLDLR | 2.1 | 7436 | Very low density lipoprotein receptor |
| APOL2 | 2.1 | 23780 | Apolipoprotein L, 2 |
| Structural factors | |||
| LMNB1 | 4.3 | 4001 | Lamin B1 |
| Transcription | |||
| IRF1 | 11 | 3659 | Interferon regulatory factor 1 |
| IRF9 | 4.9 | 10379 | Interferon regulatory factor 9 |
| STAT1 | 4.5 | 6772 | Signal transducer and activator of transcription 1, 91 kda |
| TAF15 | 2.1 | 8148 | TAF15 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 68 kda |
| Cell-cell adhesion | |||
| ICAM1 | 5.1 | 3383 | Intercellular adhesion molecule 1 |
| Ion binding | |||
| C1S | 3.2 | 716 | Complement component 1, s subcomponent |
| SP110 | 2.9 | 3431 | SP110 nuclear body protein |
| PARP12 | 2.3 | 64761 | Poly (ADP-ribose) polymerase family, member 12 |
| ZC3HAV1 | 2.3 | 56829 | Zinc finger CCCH-type, antiviral 1 |
| Tryptophan metabolism | |||
| IDO1 | 6.2 | 3620 | Indoleamine 2,3-dioxygenase 1 |
| WARS | 4.2 | 7453 | Tryptophanyl-trna synthetase |
| Enzyme activity | |||
| DTX3L | 4.6 | 151636 | Deltex 3-like (Drosophila) |
| PSMB9 | 3.3 | 5698 | Proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) |
| MSRB1 | 2.5 | 51734 | Methionine sulfoxide reductase B1 |
| CTSZ | 2.2 | 1522 | Cathepsin Z |
| Nucleotide metabolism | |||
| GBP5 | 10.5 | 115362 | Guanylate binding protein 5 |
| GBP1 | 5.8 | 2633 | Guanylate binding protein 1, interferon-inducible |
| PARP14 | 5.2 | 54625 | Poly (ADP-ribose) polymerase family, member 14 |
| GBP2 | 3.2 | 2634 | Guanylate binding protein 2, interferon-inducible |
| MX1 | 2.5 | 4599 | Myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) |
| RAD23B | 2.3 | 5887 | RAD23 homolog B (S. Cerevisiae) |
| GBP7 | 2 | 388646 | Guanylate binding protein 7 |
| Others | |||
| FAM117B | 3.6 | 150864 | Family with sequence similarity 117, member B |
| C19orf66 | 2.6 | 55337 | Chromosome 19 open reading frame 66 |
| NUB1 | 2.5 | 51667 | Negative regulator of ubiquitin-like proteins 1 |
|
| |||
| Downregulated genes: 129 transcripts | |||
| Cytokines/membrane proteins/immunological factors | |||
| IL36G | −3.4 | 56300 | Interleukin-36, gamma |
| IL33 | −3.0 | 90865 | Interleukin-33 |
| IL1RN | −2.5 | 3557 | Interleukin-1 receptor antagonist |
| DNAJB14 | −2.5 | 79982 | Dnaj (Hsp40) homolog, subfamily B, member 14 |
| KITLG | −2.3 | 4254 | KIT ligand |
| LARP4 | −2.3 | 113251 | La ribonucleoprotein domain family, member 4 |
| CLDN4 | −2.2 | 1364 | Claudin 4 |
| CUL5 | −2.2 | 8065 | Cullin 5 |
| Intracellular/signalling factors | |||
| ITGB8 | −3.5 | 3696 | Integrin, beta 8 |
| RICTOR | −2.8 | 253260 | RPTOR independent companion of MTOR, complex 2 |
| IL6ST | −2.6 | 3572 | Interleukin-6 signal transducer (gp130, oncostatin M receptor) |
| SOCS4 | −2.5 | 122809 | Suppressor of cytokine signaling 4 |
| NRIP1 | −2.5 | 8204 | Nuclear receptor interacting protein 1 |
| SEMA3A | −2.5 | 10371 | Sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A |
| PLCB4 | −2.2 | 5332 | Phospholipase C, beta 4 |
| COPS2 | −2.2 | 9318 | COP9 constitutive photomorphogenic homolog subunit 2 (Arabidopsis) |
| BMPR2 | −2 | 659 | Bone morphogenetic protein receptor, type II (serine/threonine kinase) |
| Transporters | |||
| TPR | −3.4 | 7175 | Translocated promoter region, nuclear basket protein |
| KIAA1033 | −2.5 | 23325 | Kiaa1033 |
| BAZ2B | −2.5 | 29994 | Bromodomain adjacent to zinc finger domain, 2B |
| TMED5 | −2.2 | 50999 | Transmembrane emp24 protein transport domain containing 5 |
| SLC4A7 | −2 | 9497 | Solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| Structural factors | |||
| ROCK2 | −2.9 | 9475 | Rho-associated, coiled-coil containing protein kinase 2 |
| Transcription | |||
| BDP1 | −3.6 | 55814 | B double prime 1, subunit of RNA polymerase III transcription initiation factor IIIB |
| TRIP11 | −3.4 | 9321 | Thyroid hormone receptor interactor 11 |
| ZNF644 | −3.3 | 84146 | Zinc finger protein 644 |
| CHD9 | −3.2 | 80205 | Chromodomain helicase DNA binding protein 9 |
| ZNF480 | −3.1 | 147657 | Zinc finger protein 480 |
| ZNF292 | −3 | 23036 | Zinc finger protein 292 |
| ZNF148 | −2.8 | 7707 | Zinc finger protein 148 |
| ZNF267 | −2.8 | 10308 | Zinc finger protein 267 |
| ZBTB20 | −2.7 | 26137 | Zinc finger and BTB domain containing 20 |
| CREB1 | −2.7 | 1385 | Camp responsive element binding protein 1 |
| ZNF638 | −2.6 | 27332 | Zinc finger protein 638 |
| ZNF518A | −2.6 | 9849 | Zinc finger protein 518A |
| ESF1 | −2.6 | 51575 | ESF1, nucleolar pre-rrna processing protein, homolog (S. Cerevisiae) |
| FAR1 | −2.6 | 84188 | Fatty acyl coa reductase 1 |
| ZNF91 | −2.5 | 7644 | Zinc finger protein 91 |
| CENPF | −2.5 | 1063 | Centromere protein F, 350/400 kda (mitosin) |
| FOXN2 | −2.4 | 3344 | Forkhead box N2 |
| DNTTIP2 | −2.2 | 30836 | Deoxynucleotidyl transferase, terminal, interacting protein 2 |
| TMF1 | −2.2 | 7110 | TATA element modulatory factor 1 |
| BCLAF1 | −2.2 | 9774 | BCL2-associated transcription factor 1 |
| CEBPZ | −2.2 | 10153 | CCAAT/enhancer binding protein (C/EBP), zeta |
| ZNF146 | −2.1 | 7705 | Zinc finger protein 146 |
| HIF1A | −2 | 3091 | Hypoxia inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) |
| Cell-cell adhesion | |||
| DST | −2.9 | 667 | Dystonin |
| Ion binding | |||
| ATRX | −4.6 | 546 | Alpha thalassemia/mental retardation syndrome X-linked |
| EEA1 | −3.4 | 8411 | Early endosome antigen 1 |
| RSF1 | −2.6 | 51773 | Remodeling and spacing factor 1 |
| NIN | −2.6 | 51199 | Ninein (GSK3B interacting protein) |
| SCAF11 | −2.5 | 9169 | SR-related CTD-associated factor 11 |
| MIB1 | −2 | 57534 | Mind bomb E3 ubiquitin protein ligase 1 |
| TRMT13 | −2 | 54482 | Trna methyltransferase 13 homolog (S. Cerevisiae) |
| Kinase | |||
| CCDC88A | −3.1 | 55704 | Coiled-coil domain containing 88A |
| FER | −3.1 | 2241 | Fer (fps/fes related) tyrosine kinase |
| KTN1 | −3 | 3895 | Kinectin 1 (kinesin receptor) |
| SCYL2 | −2.6 | 55681 | SCY1-like 2 (S. Cerevisiae) |
| AKAP11 | −2.5 | 11215 | A kinase (PRKA) anchor protein 11 |
| PIK3C2A | −2.5 | 5286 | Phosphatidylinositol-4-phosphate 3-kinase, catalytic subunit type 2 alpha |
| CDK6 | −2.3 | 1021 | Cyclin-dependent kinase 6 |
| NEK1 | −2.3 | 4750 | NIMA (never in mitosis gene a)-related kinase 1 |
| SLK | −2.2 | 9748 | STE20-like kinase |
| UTRN | −2.2 | 7402 | Utrophin |
| RB1CC1 | −2 | 9821 | RB1-inducible coiled-coil 1 |
| YES1 | −2 | 7525 | V-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 |
| Enzyme activity | |||
| RANBP2 | −2.9 | 5903 | RAN binding protein 2 |
| JMJD1C | −2.7 | 221037 | Jumonji domain containing 1C |
| RNF6 | −2.7 | 6049 | Ring finger protein (C3H2C3 type) 6 |
| LTN1 | −2.6 | 26046 | Listerin E3 ubiquitin protein ligase 1 |
| RBBP6 | −2.5 | 5930 | Retinoblastoma binding protein 6 |
| PPIG | −2.5 | 9360 | Peptidylprolyl isomerase G (cyclophilin G) |
| PTAR1 | −2.4 | 375743 | Protein prenyltransferase alpha subunit repeat containing 1 |
| PYROXD1 | −2.3 | 79912 | Pyridine nucleotide-disulphide oxidoreductase domain 1 |
| LIG4 | −2.3 | 3981 | Ligase IV, DNA, ATP-dependent |
| RAD50 | −2.1 | 10111 | RAD50 homolog (S. Cerevisiae) |
| DBF4 | −2 | 10926 | DBF4 homolog (S. Cerevisiae) |
| Nucleotide metabolism | |||
| CHD1 | −2.6 | 1105 | Chromodomain helicase DNA binding protein 1 |
| CHML | −2.5 | 1122 | Choroideremia-like (Rab escort protein 2) |
| ELMOD2 | −2.1 | 255520 | ELMO/CED-12 domain containing 2 |
| NAA15 | −2 | 80155 | N(alpha)-acetyltransferase 15, nata auxiliary subunit |
| KIF20B | −2 | 9585 | Kinesin family member 20B |
| Others | |||
| GOLGA4 | −3.9 | 2803 | Golgin A4 |
| ASPM | −3.4 | 259266 | Asp (abnormal spindle) homolog, microcephaly associated (Drosophila) |
| CEP350 | −3.3 | 9857 | Centrosomal protein 350 kda |
| C10orf118 | −3.2 | 55088 | Chromosome 10 open reading frame 118 |
| GALNT5 | −3.2 | 11227 | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 5 (galnac-T5) |
| BOD1L1 | −3.1 | 259282 | Biorientation of chromosomes in cell division 1-like 1 |
| ANKRD12 | −3.1 | 23253 | Ankyrin repeat domain 12 |
| RIF1 | −3 | 55183 | RAP1 interacting factor homolog (yeast) |
| KIAA2026 | −3 | 158358 | Kiaa2026 |
| FAM111B | −2.9 | 374393 | Family with sequence similarity 111, member B |
| EIF5B | −2.8 | 9669 | Eukaryotic translation initiation factor 5B |
| OSBPL8 | −2.8 | 114882 | Oxysterol binding protein-like 8 |
| GOLGB1 | −2.8 | 2804 | Golgin B1 |
| MALAT1 | −2.7 | 378938 | Metastasis associated lung adenocarcinoma transcript 1 (non-protein coding) |
| PRPF40A | −2.7 | 55660 | PRP40 pre-mrna processing factor 40 homolog A (S. Cerevisiae) |
| QSER1 | −2.6 | 79832 | Glutamine and serine rich 1 |
| THOC2 | −2.6 | 57187 | THO complex 2 |
| RASSF6 | −2.6 | 166824 | Ras association (ralgds/AF-6) domain family member 6 |
| STAG2 | −2.6 | 10735 | Stromal antigen 2 |
| NUFIP2 | −2.5 | 57532 | Nuclear fragile X mental retardation protein interacting protein 2 |
| SMC2 | −2.5 | 10592 | Structural maintenance of chromosomes 2 |
| SLC30A1 | −2.5 | 7779 | Solute carrier family 30 (zinc transporter), member 1 |
| ZNF268 | −2.4 | 10795 | Zinc finger protein 268 |
| KIAA1109 | −2.4 | 84162 | Kiaa1109 |
| NIPBL | −2.4 | 25836 | Nipped-B homolog (Drosophila) |
| ANKRD30BP2 | −2.4 | 149992 | Ankyrin repeat domain 30B pseudogene 2 |
| KIF5B | −2.4 | 3799 | Kinesin family member 5B |
| SMC4 | −2.3 | 10051 | Structural maintenance of chromosomes 4 |
| SMC5 | −2.3 | 23137 | Structural maintenance of chromosomes 5 |
| NEMF | −2.3 | 9147 | Nuclear export mediator factor |
| GCC2 | −2.3 | 9648 | GRIP and coiled-coil domain containing 2 |
| EXOC5 | −2.3 | 10640 | Exocyst complex component 5 |
| FCHO2 | −2.3 | 115548 | FCH domain only 2 |
| SMC6 | −2.3 | 79677 | Structural maintenance of chromosomes 6 |
| HOOK3 | −2.2 | 84376 | Hook homolog 3 (Drosophila) |
| LGALSL | −2.1 | 29094 | Lectin, galactoside-binding-like |
| SEMA3C | −2.1 | 10512 | Sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3C |
| TMEM106B | −2.1 | 54664 | Transmembrane protein 106B |
| PCM1 | −2.1 | 5108 | Pericentriolar material 1 |
| KRTAP2-3 | −2.1 | 730755 | Keratin associated protein 2-3 |
| MACC1 | −2.1 | 346389 | Metastasis associated in colon cancer 1 |
| CCDC82 | −2.1 | 79780 | Coiled-coil domain containing 82 |
| TTC37 | −2.1 | 9652 | Tetratricopeptide repeat domain 37 |
| UACA | −2.1 | 55075 | Uveal autoantigen with coiled-coil domains and ankyrin repeats |
| CEP290 | −2 | 80184 | Centrosomal protein 290 kda |
| USP53 | −2 | 54532 | Ubiquitin specific peptidase 53 |
| METRNL | −2 | 284207 | Meteorin, glial cell differentiation regulator-like |
The microarray analysis identified 51 transcripts in epithelial cells that exhibited a statistically significant change in their median expression level (either upregulated or downregulated) in response to pretreatment with ZDY and treatment with the uNK cell-secretion medium compared with their level in response to treatment with uNK cell-secretion medium (Table 4). Of these genes, 33 were downregulated, and 18 were upregulated. The largest group of upregulated genes includes the following: enzyme asparagine synthetase (glutamine-hydrolyzing) (ASNS) and aldo-keto reductase family 1 member C1 (dihydrodiol dehydrogenase 1; 20-alpha (3-alpha)-hydroxysteroid dehydrogenase) (AKR1C1). The transcripts encoding molecules responsible for signal transduction, namely, aldo-keto reductase family 1 member C2 (dihydrodiol dehydrogenase 2; bile acid binding protein; 3-alpha hydroxysteroid dehydrogenase, type III) (AKR1C2) and structuration, such as talin 1 (TLN 1) and keratin 75 (KRT75), were also upregulated. The levels of transcripts encoding proteins that regulate transcription and ion binding, such as nuclear protein transcriptional regulator 1 (NUPR1) and S100 calcium binding protein A8 (S100A8), were also altered, and several genes with unknown function were found to be upregulated. The downregulated transcripts included only two genes of known function that were downregulated by more than 4fold: lamin B1 (LMNB1, a structural factor) and tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase 2 (TNKS2, an enzyme activator). The genes that were downregulated by more than 2fold included the genes encoding the cytokines chemokine (C-X-C motif) ligand 3 (CXCL3) and caprin family member 2 (CAPRIN2), the transporter component of the oligomeric golgi complex 5 (COG5), and various genes associated with transcription, kinase, ion binding, and nucleotide metabolism.
Table 4.
Transcripts that were altered by more than twofold in endometrial epithelial cells by stimulation with ZDY and uNK cell-secretion medium (P < 0.001). The transcript levels were obtained using a GeneChip PrimeView Human Gene Expression Array.
| Gene symbol | Fold change | Gene ID | Description |
|---|---|---|---|
| Upregulated genes: 18 transcripts | |||
| Intracellular/signalling factors | |||
| AKR1C2 | 2.7 | 1646 | Aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding protein; 3-alpha hydroxysteroid dehydrogenase, type III) |
| Structural factors | |||
| TLN1 | 2.7 | 7094 | Talin 1 |
| KRT75 | 2.0 | 9119 | Keratin 75 |
| Transcription | |||
| NUPR1 | 2.6 | 26471 | Nuclear protein, transcriptional regulator, 1 |
| Ion binding | |||
| S100A8 | 2.1 | 6279 | S100 calcium binding protein A8 |
| Enzyme activity | |||
| ASNS | 5.2 | 440 | Asparagine synthetase (glutamine-hydrolyzing) |
| AKR1C1 | 3.3 | 1645 | Aldo-keto reductase family 1, member C1 (dihydrodiol dehydrogenase 1; 20-alpha (3-alpha)-hydroxysteroid dehydrogenase) |
| NQO1 | 2.9 | 1728 | NAD(P)H dehydrogenase, quinone 1 |
| CBS | 2.5 | 875 | Cystathionine-beta-synthase |
| AKR1C3 | 2.3 | 8644 | Aldo-keto reductase family 1, member C3 (3-alpha hydroxysteroid dehydrogenase, type II) |
| PSAT1 | 2.2 | 29968 | Phosphoserine aminotransferase 1 |
| HMOX1 | 2.2 | 3162 | Heme oxygenase (decycling) 1 |
| GPX2 | 2.2 | 2877 | Glutathione peroxidase 2 (gastrointestinal) |
| DHRS3 | 2.1 | 9249 | Dehydrogenase/reductase (SDR family) member 3 |
| H1FX | 2.1 | 8971 | H1 histone family, member X |
| Others | |||
| SLC7A11 | 3.1 | 23657 | Solute carrier family 7 (anionic amino acid transporter light chain, xc-system), member 11 |
| CTH | 2.4 | 1491 | Cystathionase (cystathionine gamma-lyase) |
| HSPB8 | 2.2 | 26353 | Heat shock 22 kda protein 8 |
|
| |||
| Downregulated genes: 33 transcripts | |||
| Cytokines/membrane proteins/signalling factors | |||
| CXCL3 | −2.1 | 2921 | Chemokine (C-X-C motif) ligand 3 |
| CAPRIN2 | −2.0 | 65981 | Caprin family member 2 |
| Transporters | |||
| COG5 | −2.0 | 10466 | Component of oligomeric golgi complex 5 |
| Structural factors | |||
| LMNB1 | −5.5 | 4001 | Lamin B1 |
| PNN | −2.9 | 5411 | Pinin, desmosome associated protein |
| MFAP5 | −2.2 | 8076 | Microfibrillar associated protein 5 |
| RPL37A | −2.2 | 6168 | Ribosomal protein l37a |
| Transcription | |||
| HIRA | −3.6 | 7290 | HIR histone cell cycle regulation defective homolog A (S. Cerevisiae) |
| YAP1 | −2.4 | 10413 | Yes-associated protein 1 |
| ZNF226 | −2.4 | 7769 | Zinc finger protein 226 |
| TAF15 | −2.1 | 8148 | TAF15 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 68 kda |
| NFKBIZ | −2.1 | 64332 | Nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta |
| Ion binding | |||
| ZDHHC11 | −2.2 | 79844 | Zinc finger, DHHC-type containing 11 |
| RNF145 | −2.1 | 153830 | Ring finger protein 145 |
| Kinase | |||
| AHSA2 | −2.0 | 130872 | AHA1, activator of heat shock 90 kda protein atpase homolog 2 (yeast) |
| Enzyme activity | |||
| TNKS2 | −7.7 | 80351 | Tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase 2 |
| Nucleotide metabolism | |||
| RAD23B | −2.0 | 5887 | RAD23 homolog B (S. Cerevisiae) |
| RCC1 | −2.0 | 1104 | Regulator of chromosome condensation 1 |
| Others | |||
| GOLGA8A | −5.4 | 23015 | Golgin A8 family, member A |
| LUC7L3 | −3.8 | 51747 | LUC7-like 3 (S. Cerevisiae) |
| HNRNPA2B1 | −2.9 | 3181 | Heterogeneous nuclear ribonucleoprotein A2/B1 |
| XIST | −2.8 | 7503 | X (inactive)-specific transcript (non-protein coding) |
| SNORA33 | −2.6 | 594839 | Small nucleolar RNA, H/ACA box 33 |
| SNHG12 | −2.5 | 85028 | Small nucleolar RNA host gene 12 (non-protein coding) |
| C20orf197 | −2.4 | 284756 | Chromosome 20 open reading frame 197 |
| EML2 | −2.4 | 24139 | Echinoderm microtubule associated protein like 2 |
| PNISR | −2.3 | 25957 | PNN-interacting serine/arginine-rich protein |
| KIAA0020 | −2.3 | 9933 | Kiaa0020 |
| HMMR | −2.2 | 3161 | Hyaluronan-mediated motility receptor (RHAMM) |
| NOP58 | −2.2 | 51602 | NOP58 ribonucleoprotein homolog (yeast) |
| PILRB | −2 | 29990 | Paired immunoglobin-like type 2 receptor beta |
| NCAPG | −2 | 64151 | Non-SMC condensin I complex, subunit G |
| PICALM | −2 | 8301 | Phosphatidylinositol binding clathrin assembly protein |
3.2. Quantitative Real-Time RT-PCR (qRT-PCR)
To verify these changes, the transcript levels of the following genes were measured by qRT-PCR: CXCL-10, IL-15, ICAM-1, NLRC-5, and IRF-1 (Figure 1). The qRT-PCR results confirmed the substantial and significant changes in expression observed in the microarray analysis. Epithelial cells treated with the uNK cell-secretion medium showed increased expression levels of CXCL-10 (18.4-fold), IL-15 (11.7-fold), ICAM-1 (12.5-fold), NLRC-5 (31.5-fold), and IRF-1 (15.6-fold) (all P < 0.001) compared with the control group.
Figure 1.
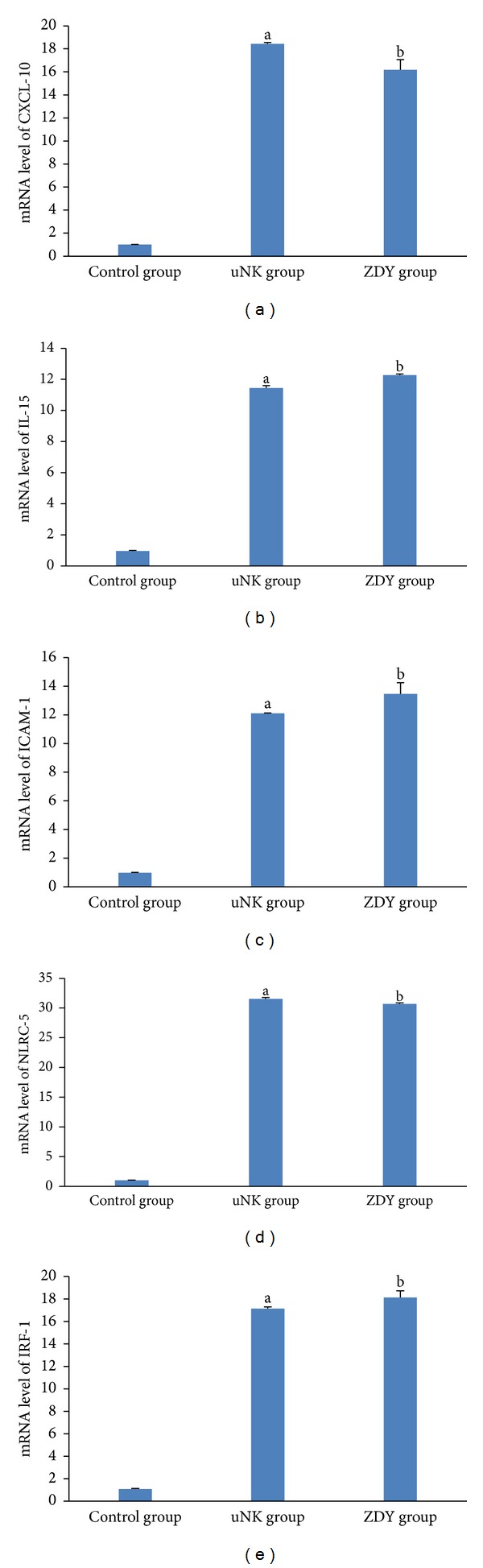
Transcription levels of (a) CXCL-10, (b) IL-15, (c) ICAM-1, (d) NLRC-5, and (e) IRF-1 determined by qRT-PCR. The relative fold changes are shown in Section 3. The qRT-PCR results were consistent with the microarray analysis. The data are expressed as means ± SEM. a P < 0.001 for the comparison between the uNK and control groups; b P < 0.001 for the comparison between the ZDY and control groups.
We also confirmed that epithelial cells pretreated with ZDY and then treated with the uNK cell-secretion medium exhibited downregulated expression levels of CXCL-10 (0.9-fold; 0.01 < P < 0.05) and NLRC-5 (1.0-fold;0.01 < P < 0.05) and upregulated levels of IL-15 (1.1-fold; 0.01 < P < 0.05), ICAM-1(1.1-fold; P > 0.05), and IRF-1 (1.1-fold; P > 0.05) compared with epithelial cells treated with the uNK cell-secretion medium. However, none of these changes were significant.
3.3. Western Blot
To confirm these effects at the protein level, the amount of ICAM-1 was analyzed by western blot (Figure 2). Epithelial cells treated with the uNK cell-secretion medium showed a 2.1-fold (P < 0.01) increase in the expression of ICAM-1. However, the epithelial cells treated with ZDY and the uNK cell-secretion medium showed a 1.4-fold increase (0.01 < P < 0.05) in the expression of ICAM-1 compared with the epithelial cells treated with the uNK cell-secretion medium. Thus, there was no significant difference between the uNK and ZDY groups.
Figure 2.
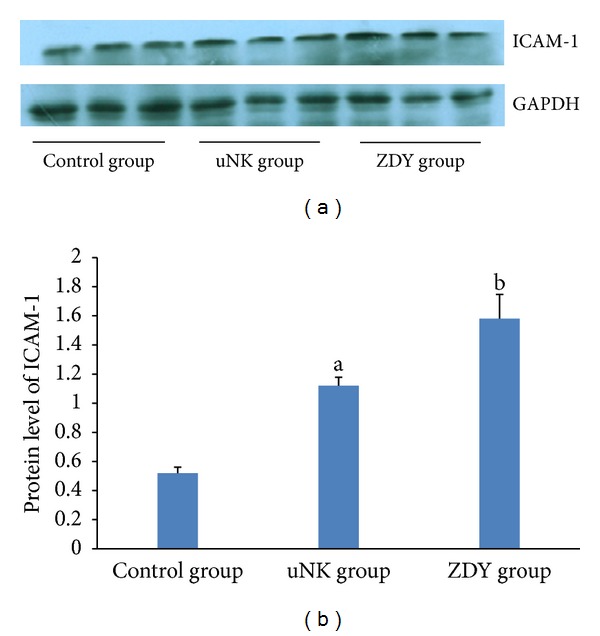
(a) Expression of endometrial ICAM-1 protein in endometrial epithelial cells. The GAPDH band was used as the internal loading control in each lane. (b) Comparison of the normalized expression intensity of endometrial ICAM-1 protein in the control, uNK, and ZDY groups. The data are expressed as means ± SEM. a P < 0.01 for the comparison between the uNK and control groups; b P < 0.01 for the comparison between the ZDY and control groups.
4. Discussion
Progesterone plays a central role in reproduction and is involved in implantation and pregnancy. When pregnancy occurs, high progesterone levels are critical not only for facilitating implantation but also for maintaining pregnancy by stimulating uterine growth [20, 21]. Several observations suggest that uNK cells also play an important role in reproduction: (a) these cells are hormonally regulated because they increase in number during the luteal phase when implantation occurs [7], (b) these cells are present in early gestation at the time placental cells invade the maternal arteries [22], and (c) these cells are particularly abundant around trophoblast cells [23]. The proliferation of uNK cells is the foundation of their reproductive function. One of the mechanisms underlying the increase in uNK cell numbers is the stimulation of the proliferation of existing uNK cells by progesterone. In mouse uterine tissues, the significant upregulation in the decidual NK population is regulated by progesterone [24]. Because there are no progesterone receptors on uNK cells, it is hypothesized that the paracrine communication between uNK and epithelial cells participates in this process. In this study, we used the uNK cell-secretion medium to stimulate epithelial cells in an attempt to find the interaction and/or paracrine effects between these cells. The data obtained in the present study demonstrate that uNK cells have a significant effect on the gene expression in epithelial cells in the nonpregnant state, and these paracrine effects also have a profound effect on uNK proliferation.
The largest group of upregulated genes induced in epithelial cells by the soluble factors derived from uNK cells induced cytokines and immunological factors, including IL-15 and IL-15RA. IL-15 is critical for NK cell differentiation in mice and human lymphoid tissue [25, 26]. IL-15 acts on precursor and immature NK cells. In humans, IL-15 transcription is more abundant during the secretory compared with the proliferative cycle phase, is maintained in early pregnancy, and localizes to the endothelium and decidual spiral arteries [27]. The correlative time strongly suggests that IL-15 participates in uNK cell proliferation and that IL-15 may also contribute to the localization of uNK cells. Our results show that uNK cells exhibit upregulated levels of both IL-15 and IL-15RA. This may activate a powerful proliferation loop because IL-15 is presented by IL-15RA on epithelial cells to promote uNK proliferation.
There are temporal and quantitative features associated with the activities of uNK cells and trophoblast cells in uterine spiral artery maturation. There is evidence that IL-15 upregulates the level of VEGF-C mRNA expression, which suggests that uNK cells may play an important role in endometrial angiogenesis and regeneration [28]. In addition, CXCL-10 is involved in uNK cell production, directs the migration and invasion of CXCR1+, CXCR3+, CXCR4+, and CCR3+ trophoblast [12], and promotes angiogenesis in the placental bed [29]. Furthermore, in the absence of uNK cells, spiral artery extension is delayed, and the tunica media remain intact [30], and these studies are consistent with an angiogenic/vascular remodeling role for uNK cells [4, 12, 31]. We found some evidence that uNK cells promote changes in endometrial epithelial cells in the nonpregnant state. The level of VEGF-C, which is upregulated in epithelial cells by the uNK cell-secretion medium, increases markedly during implantation and early pregnancy [32]. Our data suggest that uNK may prime some of the decidualization changes that occur in epithelial cells prior to the onset of full decidualization.
The exact function of uNK cells has not yet been unequivocally determined, but it is known that these cells express a different cytokine profile compared with peripheral NK cells. For example, these cells express colony-stimulating factor (CSF), macrophage colony-stimulating factor (M-CSF), granulocyte-macrophage colony-stimulating factor (GM-CSF), tumor necrosis factor alpha (TNF-α), interferon-γ (IFN-γ), and leukemia inhibitory (LIF) [33, 34]. During human pregnancy, these cells are found in close proximity to the implantation site and in close contact with the infiltrating extravillous trophoblasts. Furthermore, uNK cells increase their numbers in early pregnancy due to hormonal dependence and exhibit close proximity to trophoblasts. Taken together, these data support the conclusion that uNK cells play an important role in the regulation for the maternal immune response for the control of implantation and trophoblast invasion during human pregnancy. The summary of the paracrine effects of uNK cells on the epithelial cells is shown in Figure 3.
Figure 3.
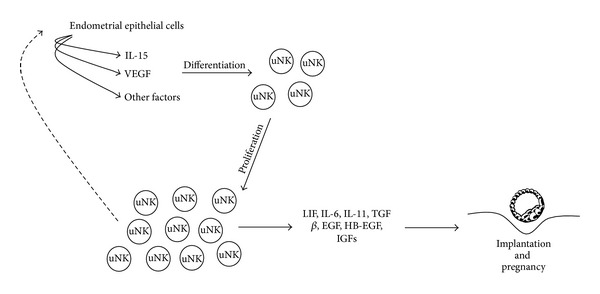
Model of the proliferation and function of uNK cells. During implantation and early pregnancy, the uNK paracrine signals stimulate endometrial epithelial cells to produce IL-15, VEGF, and other unidentified factors that may regulate uNK cell proliferation. The immune paracrine communication between uNK and endometrial epithelial cells may support implantation and trophoblast development.
In addition, this study provides the first demonstration of the paracrine communication between uNK and endometrial epithelial cells. Because our previous study showed that ZDY can improve the morphology of the endometrium in mice, we attempted to identify a relationship between the paracrine communication (uNK/epithelial cells) and the mechanism of action of ZDY. However, the majority of the up- and downregulated genes induced in the ZDY pretreated epithelial cells were of unknown function. To the best of our knowledge, there are no published studies on the effects of traditional Chinese medicine on the human endometrial paracrine communication. Our results suggest that ZDY may not regulate this process in vitro. ZDY may improve the morphology of the uterus through another mechanism.
5. Conclusions
Overall, this study provides the first detailed demonstration of the paracrine interaction between immune cells and epithelial cells in the endometrial environment. The findings indicate that these paracrine signals may contribute to uNK cell proliferation during implantation and early pregnancy. In addition, we failed to find evidence indicating that ZDY regulates the paracrine effects between uNK cells and epithelial cells in vitro.
Supplementary Material
Hierarchical Clustering of microarray experiments. SP-ZY represented the ZDY group; SP-uNK represented the uNK group and SP-K represented the control group. Three chips were used for each group.
Acknowledgments
This project was supported by the Natural Science Foundation of China. The project number is 81173292.
Conflict of Interests
The authors declare that they have no conflict of interests.
References
- 1.Zhang J, Chen Z, Smith GN, Croy BA. Natural killer cell-triggered vascular transformation: maternal care before birth? Cellular & Molecular Immunology. 2010;8(1):1–11. doi: 10.1038/cmi.2010.38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Quenby S, Nik H, Innes B, et al. Uterine natural killer cells and angiogenesis in recurrent reproductive failure. Human Reproduction. 2009;24(1):45–54. doi: 10.1093/humrep/den348. [DOI] [PubMed] [Google Scholar]
- 3.Guimond M-J, Luross JA, Wang B, Terhorst C, Danial S, Croy BA. Absence of natural killer cells during murine pregnancy is associated with reproductive compromise in TgE26 mice. Biology of Reproduction. 1997;56(1):169–179. doi: 10.1095/biolreprod56.1.169. [DOI] [PubMed] [Google Scholar]
- 4.Lash GE, Robson SC, Bulmer JN. Review: functional role of uterine natural killer (uNK) cells in human early pregnancy decidua. Placenta. 2010;31:S87–S92. doi: 10.1016/j.placenta.2009.12.022. [DOI] [PubMed] [Google Scholar]
- 5.Bulmer JN, Morrison L, Longfellow M, Ritson A, Pace D. Granulated lymphocytes in human endometrium: histochemical and immunohistochemical studies. Human Reproduction. 1991;6(6):791–798. doi: 10.1093/oxfordjournals.humrep.a137430. [DOI] [PubMed] [Google Scholar]
- 6.Okada H, Nakajima T, Sanezumi M, Ikuta A, Yasuda K, Kanzaki H. Progesterone enhances interleukin-15 production in human endometrial stromal cells in vitro. Journal of Clinical Endocrinology & Metabolism. 2000;85(12):4765–4770. doi: 10.1210/jcem.85.12.7023. [DOI] [PubMed] [Google Scholar]
- 7.King A, Wellings V, Gardner L, Loke YW. Immunocytochemical characterization of the unusual large granular lymphocytes in human endometrium throughout the menstrual cycle. Human Immunology. 1989;24(3):195–205. doi: 10.1016/0198-8859(89)90060-8. [DOI] [PubMed] [Google Scholar]
- 8.Verma S, Hiby SE, Loke YW, King A. Human decidual natural killer cells express the receptor for and respond to the cytokine interleukin 15. Biology of Reproduction. 2000;62(4):959–968. doi: 10.1095/biolreprod62.4.959. [DOI] [PubMed] [Google Scholar]
- 9.Henderson TA, Saunders PTK, Moffett-King A, Groome NP, Critchley HOD. Steroid receptor expression in uterine natural killer cells. Journal of Clinical Endocrinology and Metabolism. 2003;88(1):440–449. doi: 10.1210/jc.2002-021174. [DOI] [PubMed] [Google Scholar]
- 10.Gubbay O, Critchley HOD, Bowen JM, King A, Jabbour HN. Prolactin induces ERK phosphorylation in epithelial and CD56+ natural killer cells of the human endometrium. Journal of Clinical Endocrinology & Metabolism. 2002;87(5):2329–2335. doi: 10.1210/jcem.87.5.8515. [DOI] [PubMed] [Google Scholar]
- 11.Germeyer A, Sharkey AM, Prasadajudio M, et al. Paracrine effects of uterine leucocytes on gene expression of human uterine stromal fibroblasts. Molecular Human Reproduction. 2009;15(1):39–48. doi: 10.1093/molehr/gan075. [DOI] [PubMed] [Google Scholar]
- 12.Hanna J, Goldman-Wohl D, Hamani Y, et al. Decidual NK cells regulate key developmental processes at the human fetal-maternal interface. Nature Medicine. 2006;12(9):1065–1074. doi: 10.1038/nm1452. [DOI] [PubMed] [Google Scholar]
- 13.Crimmel AS, Conner CS, Monga M. Withered Yang: a review of traditional Chinese medical treatment of male infertility and erectile dysfunction. Journal of Andrology. 2001;22(2):173–182. [PubMed] [Google Scholar]
- 14.Qian C, Zhe J, Qiu-dan L, Ying C, Nan L. Affection of Zidan decoction containing serum on endometrial receptivity in uterine immunity environment. Guo Ji Zhong Yi Zhong Yao Za Zhi. 2012;2012(3):220–223. [Google Scholar]
- 15.Carlone DL, Rider V. Embryonic modulation of basic fibroblast growth factor in the rat uterus. Biology of Reproduction. 1993;49(4):653–665. doi: 10.1095/biolreprod49.4.653. [DOI] [PubMed] [Google Scholar]
- 16.Viganò P, Gaffuri B, Somigliana E, Infantino M, Vignali M, Di Blasio AM. Interleukin-10 is produced by human uterine natural killer cells but does not affect their production of interferon-γ . Molecular Human Reproduction. 2001;7(10):971–977. doi: 10.1093/molehr/7.10.971. [DOI] [PubMed] [Google Scholar]
- 17.Verma S, Hiby SE, Loke YW, King A. Human decidual natural killer cells express the receptor for and respond to the cytokine interleukin 15. Biology of Reproduction. 2000;62(4):959–968. doi: 10.1095/biolreprod62.4.959. [DOI] [PubMed] [Google Scholar]
- 18.Diamond MS, Staunton DE, De Fougerolles AR, et al. ICAM-1 (CD54): a counter-receptor for Mac-1 (CD11b/CD18) Journal of Cell Biology. 1990;111(6):3129–3139. doi: 10.1083/jcb.111.6.3129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Marlin SD, Springer TA. Purified intercellular adhesion molecule-1 (ICAM-1) is a ligand for lymphocyte function-associated antigen 1 (LFA-1) Cell. 1987;51(5):813–819. doi: 10.1016/0092-8674(87)90104-8. [DOI] [PubMed] [Google Scholar]
- 20.Clarke CL, Sutherland RL. Progestin regulation of cellular proliferation. Endocrine Reviews. 1990;11(2):266–301. doi: 10.1210/edrv-11-2-266. [DOI] [PubMed] [Google Scholar]
- 21.Dinny Graham J, Clarke CL. Physiological action of progesterone in target tissues. Endocrine Reviews. 1997;18(4):502–519. doi: 10.1210/edrv.18.4.0308. [DOI] [PubMed] [Google Scholar]
- 22.Bulmer J, Johnson P, Bulmer D. Leukocyte populations in human decidua and endometrium. In: Gill TC, Wegmann T, editors. Immunoregulation and Fetal Survival. New York, NY, USA: Oxford University Press; 1987. pp. 111–134. [Google Scholar]
- 23.Loke Y. Human Implantation: Cell Biology and Immunology. Cambridge, UK: Cambridge University Press; 1995. [Google Scholar]
- 24.Choe YS, Shim C, Choi D, Lee CS, Lee KK, Kim K. Expression of galectin-1 mRNA in the mouse uterus is under the control of ovarian steroids during blastocyst implantation. Molecular Reproduction and Development. 1997;48(2):261–266. doi: 10.1002/(SICI)1098-2795(199710)48:2<261::AID-MRD14>3.0.CO;2-0. [DOI] [PubMed] [Google Scholar]
- 25.Kennedy MK, Glaccum M, Brown SN, et al. Reversible defects in natural killer and memory CD8 T cell lineages in interleukin 15-deficient mice. Journal of Experimental Medicine. 2000;191(5):771–780. doi: 10.1084/jem.191.5.771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Carson WE, Giri JG, Lindemann MJ, et al. Interleukin (IL) 15 is a novel cytokine that activates human natural killer cells via components of the IL-2 receptor. Journal of Experimental Medicine. 1994;180(4):1395–1403. doi: 10.1084/jem.180.4.1395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kitaya K, Yasuda J, Yagi I, Tada Y, Fushiki S, Honjo H. IL-15 expression at human endometrium and decidua. Biology of Reproduction. 2000;63(3):683–687. doi: 10.1095/biolreprod63.3.683. [DOI] [PubMed] [Google Scholar]
- 28.Li XF, Charnock-Jones DS, Zhang E, et al. Angiogenic growth factor messenger ribonucleic acids in uterine natural killer cells. Journal of Clinical Endocrinology & Metabolism. 2001;86(4):1823–1834. doi: 10.1210/jcem.86.4.7418. [DOI] [PubMed] [Google Scholar]
- 29.Moffett-King A. Natural killer cells and pregnancy. Nature Reviews Immunology. 2002;2(9):656–663. doi: 10.1038/nri886. [DOI] [PubMed] [Google Scholar]
- 30.Chakraborty D, Karim Rumi MA, Konno T, Soares MJ. Natural killer cells direct hemochorial placentation by regulating hypoxia-inducible factor dependent trophoblast lineage decisions. Proceedings of the National Academy of Sciences of the United States of America. 2011;108(39):16295–16300. doi: 10.1073/pnas.1109478108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Smith SD, Dunk CE, Aplin JD, Harris LK, Jones RL. Evidence for immune cell involvement in decidual spiral arteriole remodeling in early human pregnancy. The American Journal of Pathology. 2009;174(5):1959–1971. doi: 10.2353/ajpath.2009.080995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lash GE, Schiessl B, Kirkley M, et al. Expression of angiogenic growth factors by uterine natural killer cells during early pregnancy. Journal of Leukocyte Biology. 2006;80(3):572–580. doi: 10.1189/jlb.0406250. [DOI] [PubMed] [Google Scholar]
- 33.Jokhi PP, King A, Sharkey AM, Smith SK, Loke YW. Screening for cytokine messenger ribonucleic acids in purified human decidual lymphocyte populations by the reverse-transcriptase polymerase chain reaction. Journal of Immunology. 1994;153(10):4427–4435. [PubMed] [Google Scholar]
- 34.Saito S, Nishikawa K, Morii T, et al. Cytokine production by CD16-CD56(bright) natural killer cells in the human early pregnancy decidua. International Immunology. 1993;5(5):559–563. doi: 10.1093/intimm/5.5.559. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Hierarchical Clustering of microarray experiments. SP-ZY represented the ZDY group; SP-uNK represented the uNK group and SP-K represented the control group. Three chips were used for each group.


