Abstract
A mutant of Mycobacterium smegmatis is a potential class I model substitute for Mycobacterium tuberculosis. Because not all of the rifamycins have been tested in this organism, we determined bactericidal profiles for the 6 major rifamycin derivatives. The profiles closely mirrored that established for Mycobacterium tuberculosis. Rifalazil was confirmed to be the most potent rifamycin.
Because the tuberculous granuloma presents a harshly oxidizing environment we explored the effects of oxidation on rifamycins. Mass spectrometry confirmed that three of the six major rifamycins showed autoxidation in the presence of trace metals. Oxidation could be monitored by distinctive changes including isosbestic points in the ultraviolet-visible spectrum. Oxidation of rifamycins abrogated antimycobacterial activity in Mycobacterium smegmatis. Protection from autoxidation was conferred by binding susceptible rifamycins to tear lipocalin, a promiscuous lipophilic protein.
Rifalazil was not susceptible to autoxidation but was insoluble in aqueous. Solubility was enhanced when complexed to tear lipocalin and was accompanied by a spectral red shift. The positive solvatochromism was consistent with robust molecular interaction and binding. Other rifamycins also formed a complex with lipocalin, albeit to a lesser extent. Protection from oxidation and enhancement of solubility with protein binding may have implications for delivery of select rifamycin derivatives.
Keywords: rifamycin, lipocalin-1, tear lipocalin, oxidation, minimal inhibitory concentration, rifalazil, Mycobacteria smegmatis
1. Introduction
Mycobacterium tuberculosis infects one-third of the human population, 9 million people per year, and accounts for 1.7 million deaths per year [1]. The bactericidal rifamycins are considered a mainstay of therapy. However, treatment requires continued low dosage for 6 to 9 months. Prolonged therapy often leads to emergence of resistant strains and accompanying toxicity to liver and bone marrow. Very few rifamycins, namely rifabutin and rifapentine, have been marketed for tuberculosis during the 40 years after the release of rifampin. Rifalazil is potent but presumed toxicity in clinical trials has stirred a debate [2, 3].
Because one in ten patients infected with tuberculosis will transmit the disease to ten others, a small improvement in treatment could have dramatic effects [1]. A number of approaches are needed. One approach is the development of new drugs but the process is expensive, slow and requires specialized facilities for handling Mycobacterium tuberculosis. Utilization of rapid growing non-pathogenic mycobacterial species could facilitate testing. Efficient plasmid transformation has potentiated mutants of Mycobacterium smegmatis [4]. The transposon insertion in RHS 234 disrupts the arr gene, which encodes rifampin ADP ribosyltransferase, the enzyme that inactivates rifampin. RHS 234 is a rifampin-hypersensitive mutant that in initial studies may mimic the drug sensitivity of Mycobacterium tuberculosis [5]. However, carefully controlled studies are needed for all major rifamycins to compare effectiveness.
A second approach is to enhance existing drugs for a shorter, more potent and less toxic drug regimen to effectively eradicate dormant organisms residing in macrophages of tuberculous granulomata. Delivery of rifamycins on complexes of micro [6,7] or nanoparticles [8-15] has been used to vitiate systemic toxicity. However, inhalable foreign microparticles may have their own toxicity such as altering cytokine profiles in the lung [9]. We investigated the feasibility of binding rifamycins to a human protein that binds lipophilic ligands in a calyx. Tear lipocalin is produced in large quantities in the lacrimal gland (57-165 μM in tears) [16] but only in small amounts in the tracheobronchial tree. Tear lipocalin has been shown to bind rifampin and block its autoxidation to the napthoquinone form [17-19]. Rifampin is released from tear lipocalin under acidic conditions and is displaced by long chain fatty acids such as those encountered in granulomata of tuberculosis. Here, we compare six rifamycins for potency, binding to tear lipocalin as well as the efficacy and resultant protection from oxidation in vitro.
2. Materials and methods
All chemicals and solvents were purchased from Thermo Fisher Scientific (Fairlawn, NJ, USA) if not mentioned otherwise and were at least spectrophotometric grade. Rifampin, rifabutin, rifaximin and rifamycin SV were purchased from Sigma-Aldrich (St. Louis, MO, USA), rifapentine was kindly provided from Sanofi Aventis (Gentilly Cedex, France) and rifalazil from ActivBiotics Pharma, LLC (Tucker, GA, USA). Stock solutions of the drugs were prepared in DMSO or ethanol (Sigma-Aldrich, St. Louis, MO, USA) and stored at −80°C.
2.1 Bacterial strains, media and growth conditions
Wt Mycobacterium smegmatis strain mc2155 [4] and φMycoMar transposon insertion mutant Mycobacterium smegmatis strain RHS 234, kindly provided by Jun Liu (University of Toronto) [5], were used for drug susceptibility testing. All mycobacterial strains were grown in Middlebrook 7H9 liquid medium supplemented with 0.2% glycerol and 0.05% Tween 80 or on Middlebrook 7H10 plates (Difco, Voigt Global Distribution Inc, Lawrence, KS, USA) at 35°C.
2.2 Expression and purification of recombinant tear lipocalin
Tear lipocalin cDNA in PCR II (Invitrogen, San Diego, CA, USA) [20], was used as a template to amplify and clone the gene spanning bases 115-592 of the previously published sequence [21] into pET-20b(+) vector (Novagen, Milwaukee, WI, USA). Flanking restriction sites for NdeI and BamHI were added to produce the major isoform of the native protein sequence as found in tears with the addition of an initiating methionine [22].
The plasmid was transformed into E. coli BL21 (DE3) and cells were cultured and protein was expressed according to the manufacturer’s protocol (Novagen, Milwaukee, WI, USA). Following cell lysis the supernatant was treated with methanol (40% final concentration) at 4°C for 2.5 h [23]. The suspension was centrifuged in a Sorvall RC-5B (Thermo Fisher Scientific, Fairlawn, NJ, USA) with a SLA-1500 rotor at 3000 × g for 30 min. The supernatant was dialyzed against 50 mM Tris-HCl, pH 8.4. The dialysate was treated stepwise with ammonium sulfate from 50% to 70% saturation. The resulting precipitate was dissolved in 50 mM Tris-HCL, pH 8.4, dialyzed against 50 mM Tris-HCL, pH 8.4 and applied to a DEAE Sephadex A-25 column (GE Healthcare, Uppsala, Sweden). The fraction containing the protein was eluted with a stepwise 0/ 50/ 300 mM NaCl gradient and applied to a 2.5 × 100 Sephadex G-100 column (GE Healthcare, Uppsala, Sweden) equilibrated with 50 mM Tris-HCL, 100 mM NaCl, pH 8.4. Eluted fractions containing tear lipocalin were centrifugally concentrated (Amicon Ultra-4 10K, Millipore, Billericia, MA, USA), transferred to 50 mM Tris-HCL buffer and stored at −20°C. Purity of the proteins was confirmed by SDS-tricine gel electrophoresis [24]. The protein concentration was determined by the biuret method [25] and was confirmed by the calculated extinction coefficient of 13,760 M−1cm−1 for tear lipocalin [26].
2.3 Absorption spectroscopy
UV absorption spectra were recorded at room temperature (RT) using a Shimadzu UV-2401PC spectrophotometer (Columbia, MD, USA). All spectra were repeated at least three times. Molar extinction coefficients of gravimetrically measured drug concentrations were calculated using Beer’s law, A= ε | c, where A is the absorbance, ε is the molar extinction coefficient (M−1cm−1), l is the path length (cm), and c is the concentration of the solution (M).
Sodium phosphate buffer, 100 mM, pH 7.3, was used from the same reagent stock in all autoxidation experiments. The sodium phosphate is known to contain 0.5 parts per billion of iron. ETDA (Sigma-Aldrich, St. Louis, MO, USA) was used to remove divalent metal ions from the samples. K3Fe(CN)6 was also used in oxidation experiments (Sigma-Aldrich, St. Louis, MO, USA).
2.4 Mass spectrometry
Mass spectrometry is an effective technique to confirm the loss of two hydrogen atoms responsible for the oxidation of the napthohydroquinone structure to the napthoquinone form of rifampin [17,18,27]. Therefore, mass spectrometry was performed on the other rifamycins with an Agilent 6460 LC/MS/MS triple quadrupole and an electrospray ionization source. After attempted oxidation (Section 2.7), samples were diluted to a concentration of 20 μM in methanol/formic acid .1%), directly infused via the Agilent 1290 at 100 μL/min and scanned in the positive ion mode in the m/z range of 200-1200. Background ion counts from extracted chromatographs of the reagents without the rifamycins were subtracted from the samples using the Masshunter software. The instrument parameters for the identification of rifamycins included (Ultra High Purity Nitrogen) gas at 300 °C, gas flow 5 l/min, 45 psi nebulizer, fragmentation energy of 135 V, Capillary 3500 positive V, 3500 negative, 9 nA Nozzle Voltage.
2.5 Drug susceptibility testing
Drug susceptibility of Mycobacterium smegmatis wt and mutant (2.1 above), was tested for six rifamycin drugs by the broth microdilution method after the NCCLS standard [28, 29] with some minor modifications. Sterile 96-well polystyrene microtiter plates with U-shaped wells (Greiner Bio-One, Monroe, NC, USA) [29] were prepared with 2-fold serial dilutions of antibiotics in Middlebrook 7H9 broth. Antibiotics ranged in concentration from 1.28 × 102 to 1.2 × 10−4 μg/ml for Mycobacterium smegmatis mc2155 and 3.2 × 10 to 1.2 × 10−4 μg/ml for Mycobacterium smegmatis RHS 234. Inoculum suspensions of bacteria were prepared from swabs of 7H10 agar plates placed in 4.5 ml of sterile water, vortexed for 15 s, and allowed to stand at RT for 1 minute to remove coarse grained particles. The turbidity was adjusted to an OD625 of 0.08 to 0.10 to obtain a density of 1.5 × 108 CFU/ml. The final bacterial density of 1.5 × 106 CFU/ml was achieved by transferring 0.2 ml of the suspension to 20 ml sterile water with 0.05% Tween 80. Each well was inoculated with 10 μl of the bacterial suspensions. Complete inhibition of bacterial growth after 5 days of incubation at 35°C was interpreted as the minimal inhibitory concentration (MIC) in the presence of three paired control wells that showed visible colonies. In tandem, a 50 μl sample from each inoculum (1.5 × 106 CFU/ml) was transferred to a 5 ml N-saline solution (1.5 × 104 CFU/ml), vortexed and 10 μl streaked on 7H10 plates as a control for the colony forming units and to exclude contamination. All experiments were done in triplicate.
2.6 Rifamycin binding to tear lipocalin
A precise binding curve was determined for rifalazil based on its profound insolubility and strong binding to tear lipocalin. Progressive additional increments of concentrated rifalazil in ethanol were added to tear lipocalin in PBS, maintaining the final ethanol concentration at less than 1%, followed by centrifugation in a Sorvall Discovery M150 (Thermo Fisher Scientific, Fairlawn, NJ, USA), with an S150AT rotor at ~196,000 × g for 1 h at 20°C. The concentrations of bound and free drug were both experimentally determined from the absorbance at 632 nm of the supernatant and pellet, respectively, in ethanol, using the experimentally determined extinction coefficient of 37,900 M−1cm−1. The concentrations were mathematically verified by subtraction from the total drug used in the experiment. The experiment was repeated 6 times and the means were fit to the Hill equation (Microcal Origin, Northhampton, MA, USA).
For the other more aqueous soluble rifamycins estimates of binding were calculated from comparison of the relative binding to the known dissociation constant (Kd) for rifampin of 128 μM [18]. Relative binding was determined by gel filtration column chromatography with 0.5 × 10 cm glass columns (Bio-Rad, Richmond, CA, USA) containing Sephadex G-25 medium (GE Healthcare, Uppsala, Sweden) equilibrated and eluted with 100 mM Na2HPO4, pH 7.3. After incubation of tear lipocalin and rifamycin, 260 μM each, for 1 h at 25°C, 200 μl were applied to the column at a flow rate of 0.2 ml/min. 0.2 ml fractions were collected and monitored for protein at 280 nm and at selected wavelengths of each drug based on their individual spectral characteristics.
High molecular peak fractions containing tear lipocalin, both bound and free were pooled. Low molecular peak fractions containing unbound rifamycin were pooled and remeasured spectrophotometrically. The bound drug concentrations in the high molecular weight fractions were calculated from the absorbance of the rifamycins at their respective peaks at wavelengths >320 nm where tear lipocalin has a small contribution. Although exiguous, the fractional absorbance contributed by tear lipocalin was predetermined at the given wavelength from the spectrum of tear lipocalin alone and then after incubation with the drug. The absorbance attributed only to rifamycin at a given wavelength was determined in the bound fractions.
2.7 Effect of oxidation of rifamycins on anti-mycobacterial activity
So called autoxidation of rifampin and rifamycin SV occurs in the presence of only trace divalent metal cations acting as a catalyst, and oxygen. In this study we investigated the oxidative profiles of six different rifamycin drugs (4 previously untested) and assessed the effect of oxidation on antimicrobial activity. Each susceptible rifamycin, 10 μg/ml, was incubated in PBS, pH 7.4 at 24 h at RT and oxidation was verified spectrophotometrically. Gravimetric correction was made for evaporation. For rifamycins that showed oxidation, 50 μl of oxidized and non-oxidized drug, as wells as a control without drugs, were transferred to a 96 well-plate. An inoculum of 5 × 104 CFU/ml Mycobacterium smegmatis mc2155 was added to each well. The plate was incubated for 30 min at 37°C. Immediately thereafter, 150 μl 7H9-G-TW were added to give a final drug concentration of 2.5 μg/ml, time-point 0 h. Luminescence as an indicator of microbial survival (see below) was measured after 0 h and 48 h. Each experiment was repeated at least 9 times. P-values were calculated using non-parametric methods (paired Wilcoxon Signed Rank, Sign Test), (Microcal Origin, Northhampton, MA, USA).
2.8 Luminescence assay
Bacterial luciferase (Vibrio harveyi) previously introduced into Mycobacterium smegmatis mc2155 with the vector pSMT3LxEGFP, was provided by Brian D. Robertson (Imperial College London) [30]. Luminescence measurements were obtained using a Berthold Sirius single tube luminometer (Berthold Technologies, Bad Wildbad, Germany). Ten μl samples were diluted with 190 μl PBS, pH 7.4. Bacterial luminescence was measured immediately after addition of 20 μl substrate, 1% n-decyl aldehyde (Sigma-Aldrich, St. Louis, MO, USA) in ethanol, for 10 s using an integration time of 1 s, and the results were expressed in relative light units (RLU). Sterile PBS without bacteria was used to measure background and subtracted from sample RLU measurements. The RLU:CFU ratio was determined to be 10:1. The exponential growth rate constant (k) was determined under control conditions using k=ln(N/ N0)/t-to where N0= initial number of bacteria at initial time to, and N= number at time t. The percent survival of cells was determined in the experimental groups from the ratio of the number of bacteria observed from that expected at 48 hours given k above and N0 verified from the RLU at time 0 for each experiment.
3. Results
3.1 Spectral characteristics of the six rifamycin derivatives
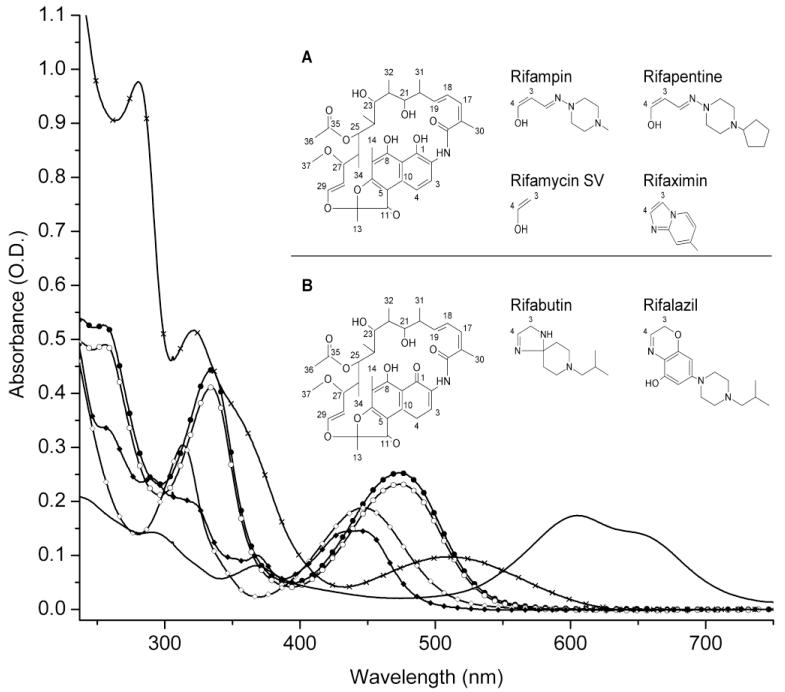
Comparison of antimicrobial activity and oxidation is predicated on accurate and consistent measurements of drug concentrations. Accurate extinction coefficients are important to make dilute antibiotic concentrations needed to determine the MIC. The absorbance spectra and chemical structures for the six major rifamycin derivatives are shown in Fig. 1. The spectra of rifampin and rifapentine are nearly superimposable, the latter chemically differing only by a cyclopentyl group. The spectra for the other drugs are unique. For two rifamycin derivatives, extinction coefficients (ε) were not previously published and limited for two others (Table 1). The ε used for the rifamycin derivatives in two solvents are shown in Table 1. As DMSO showed intrinsic absorbance in the spectra at shorter wavelengths, only wavelengths ≥320 nm were used for calculations of ε in this solvent. Comparison of values of ε with those previously published reveals close agreement [18, 31-33]. For drugs for which values of ε have not been published, the averages of three experimental values are provided. Rifalazil shows the highest ε of any of the rifamycins with intense red spectral absorption. This can be expected from π to π* transitions for a molecule with extended conjugation as well as substitutions on the additional aromatic rings by groups carrying nonbonding or π electrons in benzoxazinorifamycin.
Fig. 1.
Absorbance spectra of six rifamycin drugs at pH 7.3. Rifampin (15 μM) (
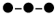 ), rifapentine (15 μM) (
), rifapentine (15 μM) (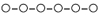 ), rifamycin SV (15 μM) (
), rifamycin SV (15 μM) ( ), rifalazil (15 μM) (————), rifaximin (15 μM) (
), rifalazil (15 μM) (————), rifaximin (15 μM) ( ), and rifabutin (30 μM) (
), and rifabutin (30 μM) (
 ). Inset (A, B): Base chemical structures of all six rifamycin drugs (left) and unique groups (right). The hydroxyls at positions 1 and 4 may be oxidized to form the hydroquinone. The sole hydroxyl of the benzoxanino moiety is a candidate for oxidation on rifalazil. Hydroxyls are not available at position 1 and 4 for autoxidation for rifabutin or rifalazil.
). Inset (A, B): Base chemical structures of all six rifamycin drugs (left) and unique groups (right). The hydroxyls at positions 1 and 4 may be oxidized to form the hydroquinone. The sole hydroxyl of the benzoxanino moiety is a candidate for oxidation on rifalazil. Hydroxyls are not available at position 1 and 4 for autoxidation for rifabutin or rifalazil.
Table 1. Extinction coefficients of rifamycin drugs.
| Drug | Wavelength | Current study | Literature | ||||
|---|---|---|---|---|---|---|---|
|
| |||||||
| (nm) | εDMSO (mM−1cm−1) |
SD | εMeOH (mM−1cm−1) |
SD | εLiterature (mM−1cm−1) |
Reference | |
| Rifampin | 237 | N.A.* | N.A.* | 34.53 | 1.90 | 33.2; 33.4 | [26]; [29] |
| 255 | N.A.* | N.A.* | 32.49 | 1.77 | 32.1 | [29] | |
| 334 | 26.83 | 0.24 | 27.16 | 1.49 | 27.0; 26.4 | [26]; [29] | |
| 470 | 15.49 | 0.16 | 15.11 | 0.84 | 15.3 | [26] | |
| 475 | 15.98 | 0.16 | 15.49 | 0.86 | 15.4 | [29] | |
|
| |||||||
| Rifapentine | 237 | N.A.* | N.A.* | 33.54 | 0.51 | - | - |
| 255 | N.A.* | N.A.* | 32.31 | 0.26 | - | - | |
| 334 | 26.39 | 0.48 | 26.46 | 0.14 | 26.7 | [29] | |
| 470 | 15.28 | 0.27 | 14.68 | 0.05 | - | - | |
| 475 | 15.74 | 0.27 | 15.04 | 0.05 | 15.2 | [29] | |
|
| |||||||
| Rifamycin SV | 237 | N.A.* | N.A.* | 37.97 | 0.22 | - | - |
| 255 | N.A.* | N.A.* | 21.43 | 0.14 | - | - | |
| 334 | 10.89 | 0.94 | 10.67 | 0.09 | - | - | |
| 445 | 14.29 | 1.24 | 14.61 | 0.13 | 14.2 | [30] | |
| 470 | 11.24 | 1.95 | 11.69 | 0.11 | - | - | |
| 475 | 11.08 | 0.19 | 10.32 | 0.08 | - | - | |
|
| |||||||
| Rifaximin | 232 | N.A.* | N.A.* | 38.14 | 0.60 | 38.4 | [29] |
| 260 | N.A.* | N.A.* | 26.57 | 0.45 | 26.6 | [29] | |
| 292 | N.A.* | N.A.* | 22.88 | 0.35 | 23.2 | [29] | |
| 320 | 16.97 | 0.13 | 16.74 | 0.22 | 17 | [29] | |
| 370 | 9.90 | 0.09 | 9.22 | 0.14 | 9.4 | [29] | |
| 450 | 12.89 | 0.14 | 12.54 | 0.29 | 12.5 | [29] | |
|
| |||||||
| Rifabutin | 320 | 15.73 | 0.72 | - | - | - | - |
| 510 | 3.13 | 0.21 | - | - | - | - | |
|
| |||||||
| Rifalazil | 368 | 9.93 | 0.99 | - | - | - | - |
| 654 | 30.85 | 2.98 | - | - | - | - | |
N.A. = not available, the extinction coefficient could not be determined accurately at the indicated wavelengths, because DMSO dominated absorbance.
3.2 Autoxidation of rifamycins
Autoxidation of rifampin can be readily monitored spectrophotometrically at 475 nm [18, 27]. Spectral changes of autoxidation of rifampin are nearly complete after 24 h [18]. Three rifamycin derivatives were found to show spectral changes under autoxidizing conditions (Fig. 2). Isosbestic points were noted for rifampin, rifamycin SV, and rifapentine indicative of 2 distinct chemical forms for each, the oxidized and reduced forms. The isosbestic points (Fig. 2) indicate that the oxidation reaction is limited to two molecular forms of the drugs that are isoabsorbant at specific wavelengths. For practical purposes isosbestic spectral points statistically exclude the possibility of nonspecific degradation involving multiple molecular species. As expected from the structure, rifampin and rifapentine share the same oxidation spectral characteristics with abrogation of the major absorbance peaks at 475 nm. Oxidation of rifamycin SV resulted in diminishing peaks at 448 nm.
Fig. 2.
Autoxidation of rifamycin drugs at pH 7.3 after various times in phosphate buffer. (a) 1 min, (b) 20 min, (c) 40 min, (d) 80 min, (e) 120 min, (f) 180 min, (g) 24 h. Absorbance spectra of (A) rifampin (15 μM), (B) rifapentine (15 μM), (C) rifamycin SV (15 μM). (D) Mass spectrum of oxidized rifapentine shows predicted monoisotopic m/z of 825.5, 897.2 and 913.1 for the M+H, M+Na and M+K ions.
Rifaximin, rifabutin and rifalazil varied in solubility with buffer composition but did not show spectral changes under autoxidizing conditions when properly solvated. Following oxidation experiments mass spectrometry confirmed the loss of two Daltons from the predicted M+H, M+Na, and/or M+K ions for samples of rifampin, rifamycin SV and rifapentine (Fig. 2D ). No oxidized species were identified for the other drugs; major monoisotopic peaks were observed for rifalazil, rifabutin, and rifaximin at m/z of 863.4, 869.4 and 808.4, respectively all representing the (M+Na)+ adducts of the reduced species. An additional peaks was noted for rifalazil representing the (M+K)+ adduct.( Figure S2).
3.3 Protection from oxidation of rifamycins by tear lipocalin
We also explored the protective effect of tear lipocalin on the oxidation for the susceptible rifamycin drugs as previously published for rifampin [18]. Tear lipocalin confers protection from autoxidation for at least 3 h for rifampin, rifapentine, and rifamycin SV (Fig. 3A-D). To obtain adequate spectral peaks, tear lipocalin was used in molar excess, but this concentration falls within the range in human tears, 57-165 μM [16]. Ethylenediaminetetraacetic acid (EDTA) blocked the oxidation for all susceptible drugs (Fig. 3) [18, 27].
Fig. 3.
A-D. Oxidation of rifamycin drugs in the presence of tear lipocalin. For comparison, autoxidation of rifamycin drugs alone (-●-), protected with tear lipocalin (100 μM) (-■-), and in the presence of the divalent ion chelator EDTA (5 mM) (-△-). (A) rifampin (15 μM), (B) rifapentine (15 μM), (C) rifamycin SV (15 μM). D-F. Addition of K3Fe(CN)6 to rifamycin drugs protected with tear lipocalin after 180 min at pH 7.3 in phosphate buffer. Absorbance spectra of rifamycin drugs and tear lipocalin (100 μM) after 1 min (————), 180 min (-- -- --), and 180 min with K3Fe(CN)6 (90 μM) (…). (E) rifampin (15 μM), (F) rifapentine (15 μM), (G) rifamycin SV (15 μM), (H) rifalazil (15 μM).
To determine if the binding of tear lipocalin provides protection from a strong oxidizing agent, K3Fe(CN)6 was added to rifamycin drugs after initial protection with tear lipocalin [17]. As illustrated in Fig. 3E and 3F, rifampin and rifapentine were both rapidly oxidized in the presence of K3Fe(CN)6 despite initial protection from autoxidation for 3 h. Rifamycin SV appeared better protected by tear lipocalin (Fig. 3F).
3.4 Influence of drug binding to tear lipocalin
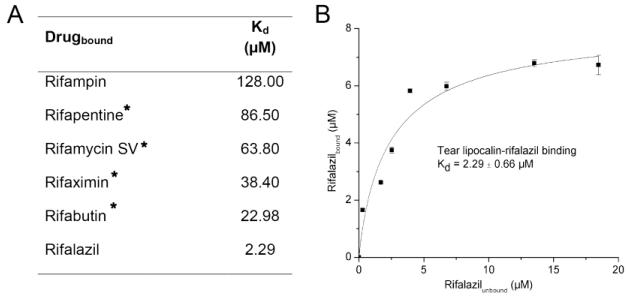
Because the complex of tear lipocalin and rifampin inhibits drug oxidation [18], relative binding was determined for the rifamycins. Additionally, tear lipocalin is known to solubilize lipids. Augmenting the solubility of rifalazil (completely insoluble in buffer) with tear lipocalin was explored. The binding curve for the complex of rifalazil and tear lipocalin is shown in Fig. 4. Analysis of size exclusion chromatograms provides a comparison of relative binding for the other rifamycins (Fig. S1, Fig. 4A). In the early eluted fractions the corrected absorbance profiles verify the co-elution of rifamycin drugs and tear lipocalin. For most rifamycins the protein-ligand complex consistently elutes in fraction 2. However, rifalazil binds more strongly to tear lipocalin by almost two orders of magnitude than rifampin (rifalazil Kd = 2.3 + 0.66 μM, rifampin Kd = 128 + 18 μM) and more than one order of magnitude than the other rifamycins (Fig. 4A).
Fig. 4.
A. Dissociation constants for rifamycins. *Estimated by gel filtration experiments (profiles shown in Figure S1). B. Ligand binding curve for rifalazil and tear lipocalin determined by spectrophotometric measurement of the soluble complex. The experiment was performed at pH 7.4 in PBS-buffer. The fitting curve is indicated (————).
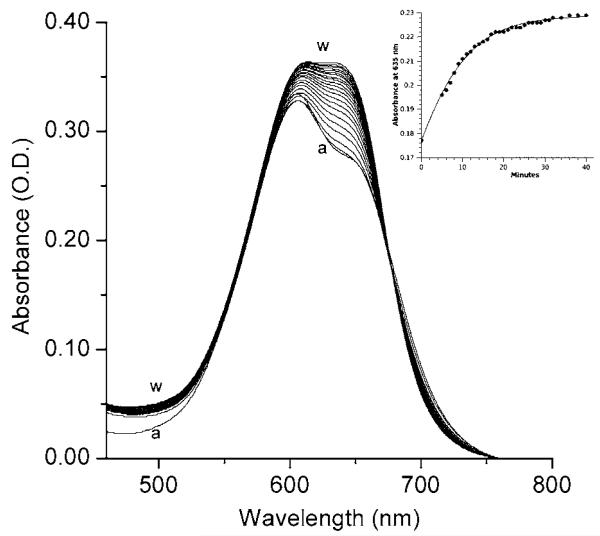
The spectra of the tear lipocalin-rifalazil complex shows a distinctive broadening of the peak at 605 nm comprised of expansion to the red side of the peak (Fig. 5). Successive scans reveal the reaction kinetics and show that the red shift increases over time up to 44 min and can be easily fit to a sigmoidal plot (Fig. 5 inset).
Fig. 5.
Red shift (solvatochromism) in absorbance spectra taken at successive 2 minute intervals from A to W of rifalazil (30 μM) added to tear lipocalin (100 μM), in 10 mM sodium phosphate, pH 7.3 Inset: Kinetics of solvatochromism, baseline corrected absorbance at 635 nm plotted (black circles) over 40 minutes fits a sigmoidal function (solid line).
3.5 MICs of rifamycins for Mycobacterium smegmatis
The MICs of rifamycins for Mycobacterium smegmatis wt and Mycobacterium smegmatis mutant were easily read after 5 days of incubation and the plaque count did not change after 7 to 10 days. The MICs for wt and mutant were all within the tested concentration range and results are depicted in Table 2. The inocula sizes for each round of drug susceptibility testing are shown in Table S2.
Table 2. Activity of antibiotics against M. smegmatis mc2155 and RHS 234.
| Drug | MIC (μM) | |
|---|---|---|
|
| ||
| mc2155 | RHS 234 | |
| Rifampin | 2.43 | 0.30 |
| Rifapentine | <2.28 | 0.14 |
| Rifamycin SV | <1.38 | 0.69 |
| Rifaximin | <1.27 | 0.16 |
| Rifabutin | 0.30 | 0.07 |
| Rifalazil | 0.13 | <0.06 |
Among the tested rifamycin derivatives, rifalazil was the most bactericidal with lower MICs for both Mycobacterium smegmatis wt and the mutant than the MICs of rifabutin, rifaximin and rifamycin SV (Table 2). The highest MICs were measured for rifampin and rifapentine. The mutant showed more sensitivity than the wt for all rifamycins. A relative comparison of drug sensitivity of the wt versus the mutant is provided by the resistance factor R, which is given as MICwt/MICmutant. Interestingly, rifalazil was closer in drug sensitivity between wt and mutant, R≈2. The Mycobacterium smegmatis mutant was much more sensitive to rifampin and rifapentine than was the wt; R = 8 for both drugs (Table 2).
3.6 Effect of rifamycin oxidation on bacterial growth
The three oxidized rifamycin drugs showed less antibacterial activity than non-oxidized forms (Fig. 6) (Table 3).
Fig. 6.
Influence of antibiotics. Reduced versus oxidized rifamycin drugs. M. smegmatis mc2155 was incubated at pH 7.4 in PBS-buffer with rifamycin drugs (10 μg/ml) for 30 min. Thereafter 150 μl 7H9-G-TW was added and incubated for 48 h. (A) rifampin, (B) rifapentine, (C) rifamycin SV, ( Error bars indicate SD, n = 6.)
Table 3. P-values for comparison of oxidized versus reduced rifamycins.
| DrugReduced | DrugOxidized | Sign-Test | Wilcoxon Signed Rank Test |
|---|---|---|---|
| Rifampin | Rifampin oxide | 0.002 | 0.002 |
| Rifapentine | Rifapentine oxide |
0.002 |
0.002 |
| Rifamycin SV | Rifamycin S | 0.02 | 0.006 |
| Rifabutin | Negative control | NS* | NS* |
NS= not significant, p-value >0.05
4. Discussion
The key findings of this paper are the elucidation of extinction coefficients for rifamycins not previously described, a head to head comparison of all major rifamycins for antimicrobial activity in Mycobacterium smegmatis, absorption spectroscopy of each major rifamycin in oxidized and/or reduced form, reduction in mycobacterial activity from oxidation including autoxidation of some rifamycins, protection from oxidation based on the protein binding affinity to tear lipocalin, and enhanced solubility for rifalazil by binding tear lipocalin.
4.1 Comparison of anti-mycobacterial activity of rifamycins
Prior studies of antimicrobial activity have been limited to historical comparisons of few experimentally obtained MICs for only a small number of drugs. Inter-laboratory variability is evident from different methodologies including different solvents to dissolve the drugs, varying concentrations of drugs due to a lack of accurate extinction coefficients, non-uniform inocula sizes and use of Tween affecting cell permeability [5, 29, 34-38]. In our study these factors were rigorously controlled. Oxidation was unlikely to affect our drug susceptibility assay because less than 10% of even the most susceptible rifamycin, rifamycin SV, is oxidized within 30 minutes. Rifamycins have been shown to penetrate cells to an intracellular concentration that is five-fold greater than the extracellular concentration within 30 minutes in vitro [39].
Rifalazil stands out as the most potent rifamycin tested in Mycobacterium smegmatis, with 16 times the mycobactericidal activity as that of rifampin and a resistance factor of only 2. Similar results have been published for rifalazil against Mycobacterium tuberculosis H37Rv (MICs ranging from 0.004 to 0.035 μg/ml) [40, 41]. Mycobacterium smegmatis appears to be a valid model for comparison.
Rifabutin was second in activity against Mycobacterium smegmatis (Table 2). Rifabutin is also a relatively strong anti-mycobacterial agent that has affinity for tear lipocalin.
4.2 Oxidation of rifamycins
Mycobacterium tuberculosis lives in a profoundly oxidizing environment. Orally administered rifamycins confront body fluids with abundant divalent cations. Oxidation of rifamycins may have implications for toxicity and efficacy. Three of the six major rifamycin derivatives, including rifampin are susceptible to auto-oxidation. Oxidation is readily monitored spectroscopically as a diminution in the major visible absorption peak after 400 nm. The isosbestic points occur only in rifamycins that have accessible hydroxyls in the hydroquinone and exclude non-specific degradation. The spectral changes were shown to be specific for the oxidation of rifampin and verified by mass spectrometry (MS/MS) [18]. Rifamycin SV is known to oxidize to form rifamycin S in the presence of divalent cations, a reaction used for the synthesis of rifamycin analogs [42, 43]. Rifampin and rifapentine are similar in structure and spectral characteristics. Rifabutin, rifaximin and rifalazil lack a hydroquinone susceptible to oxidation. These structural features fit our UV-Vis and mass spectra. Rifabutin served as a negative control for oxidation experiments (Table 3).
4.3 Antimycobacterial activity of oxidized rifamycins
While the relative potency of the rifamycins in Mycobacterium smegmatis is similar to Mycobacterium tuberculosis, the results with oxidized rifamycins may be dependent on the organism. For example rifamycin S has been reported to have a similar MIC as rifamycin SV for Mycobacterium tuberculosis (.05 μg/ml) [44]. The same study also reported that rifamycin S has a higher ED50, than rifamycin SV against Staphylococcus aureus, in mice. Further, the MIC’s of these drugs in Mycobacterium smegmatis are about 10 fold greater than that reported for Mycobacterium tuberculosis.
Another concern is that the rifampin-quinone has been shown to have greater binding to a subunit of RNA polymerase than the hydroquinone in a rolling transcription assay [45]. Discrepancies of the rolling transcription assay versus classic drug susceptibility testing have been attributed to off-target effects including efflux pumps [46]. The rifampin-quinone has also been published to bind irreversibly to proteins, RNA and poly-Lysine in a “non-enzymic reaction” [47] so that other intracellular events may account for differences.
4.4 Protection from oxidation by protein binding
Protection from oxidation conferred by tear lipocalin may be related to the Kd (Fig. 4A) for susceptible rifamycins. For rifamycin SV some protection is afforded even in the presence of an extremely strong oxidizing reagent K3Fe(CN)6 (Fig. 2C and 3C).
4.5 Mechanistic considerations of rifamycin binding to tear lipocalin
The binding of ligands to tear lipocalin has been well studied in solution and crystal structure [48-52]. The lipocalin calyx can accommodate a single ligand oriented with the hydrophobic moiety inside the cavity; the binding curves for both rifampin and rifalazil are consistent with a single ligand. Comparison of rifamycin structures depicted in Fig. 1 reveals that the three with the lowest relative dissociation constants for tear lipocalin share the presence of alkyl groups at position 3. Rifalazil and rifabutin have isopropyl and rifaximin has a methyl group in this position. The relationship between the length of the alkyl group on ligands and binding free energy has been studied for tear lipocalin. Each CH2 corresponds to a binding free energy difference of 600 cal/mole up to an alkyl chain length of C18 [24]. Estimates available from crystallography of tear lipocalin measure the cavity mouth as 10 Å in diameter with a cavity depth of 15 Å [53]. In solution, the functional cavity dimensions are such that tear lipocalin accommodates ligands that have simple ring structures as well as alkyl chains to about 18 Å in depth. However, molecules of 9.1 Å are rejected [50] presumably because of the overhanging loops at the calyx mouth [54]. The leading isopropyl chain in rifalazil and rifabutin as well as the additional six member ring in rifalazil fit within the calyx opening and cavity depth of tear lipocalin The increase in binding affinity of rifalazil over rifampin corresponds to an approximate free energy change of about 2.4 kcal/mole, which is reasonable given the extended structure from R1 (Fig. 1A, B). The increased binding fits the accommodation of the hydroxyl on the benzoxanino ring of rifalazil (Fig. 1), but the the large hydroquinone of other rifamycins would be restricted from entering the cavity mouth of tear lipocalin explaining the lower binding affinity.
The spectral features of rifalazil are also relevant to tear lipocalin binding. The peak of the spectrum for rifalazil appears at about 605 nm with a low shoulder on the red side of the spectrum at about 648 nm (Fig. 1). However, binding with tear lipocalin is accompanied by elevation of the shoulder at 648 nm (Fig. 4). This constitutes a bathochromic (red) shift and slight hyperchromism. Such a spectral change generally connotes a change in the environment surrounding the chromophore including a photoinduced charge transfer and an increase in the molecular dipole [55]. The Kd for rifalazil of 2.3 μM concurs with reduced solvent accessibility for spin labeled [56] and fluorescent labeled ligands buried in the lipocalin cavity [48].
4.6 Clinical implications for oxidation of rifamycins and toxicity
The clinical implications of oxidation of rifamycins have not been well studied. Hepatoxicity from anti-tuberculous therapy has been linked to polymorphisms in genes that express repressors to antioxidant pathways [57], but appears unrelated to deacetylated metabolites of rifamycins [58]. The toxicity of rifamycins has been better related to the amount of the unbound drug which can freely diffuse into cells of the bone marrow, lung, and liver [59]. One would expect to find the rifampin-quinone in the blood of patients taking rifampin. However the rifampin-quinone has not been identified as a plasma metabolite in patients despite the presence of the active 25-deacetylated and 3-formyl variants [60]. In the aforementioned study not all HPLC peaks were characterized. The rifampin-quinone has been proffered to “irreversibly” bind albumin [47], which might complicate extraction and detection. In addition, oxidation of rifampin may be limited to tissues with an oxidizing or inflammatory environment containing endogenous peroxidases [61].
Although rifalazil is a very attractive anti-mycobacterial agent, the side effects from once per week dosing have been reported and manifest as transient flue-symptoms, neutropenia, and thrombocytopenia [62]. Notably lacking with rifalazil is either evidence of hepatotoxicity or induction of CYP3A4 [63]. In the circulation of animals rifalazil is 99% bound to serum albumin as opposed to rifampin (80% to 85%) [64, 65]. The lack of oxidation of rifalazil and the lack of free drug in serum due to enhanced protein binding are just two of many possible explanations for a lack of hepatotoxicity.
4.7 Drug delivery implications
Rifampin binding and protection from oxidation by tear lipocalin is greater than that achieved by serum albumin [18]. These findings invite speculation regarding potential selective targeting in tuberculosis. Rifalazil could be delivered complexed to tear lipocalin or another binding protein via an inhaler to avoid the liver. Tear lipocalin is nominally produced in the lung so exogenous protein would not induce antibody formation. Lack of immunogenicity is a feature of one newly engineered anticalin, which is based on the calyx structure and promiscuous ligand binding of tear lipocalin [66]. In our case release of the drug at the target could be enhanced by a number of local conditions. An acidic environment is created in granulomata and may result in release of rifamycin compounds from tear lipocalin [18]. In addition mycobacteria produce abundant mycolic acids with an alkyl chain length in excess of 20 carbons [67, 68]. Tear lipocalin can accommodate an alkyl chain length of 18 with a Kd of 1.3 μM [69]. Based on the equilibrium constants, mycolic acids would be expected to displace rifamycins (Fig. 4) from tear lipocalin at the site of infection. Although rifalazil has extremely high intracellular penetration [65], specific and enhanced delivery to mycobacteria that reside dormant in macrophages may be possible with a lipocalin. A delivery mechanism through a lipocalin receptor is plausible since at least one receptor, that for neutrophil gelatinase-associated lipocalin (NGAL) is known to be expressed and upregulated in macrophages in the presence of mycobacterial infection [70].
Supplementary Material
Chemical compounds studied in this article
Rifampin (PubChem CID:24871024); Rifalazil (PubChem CID:6540558); Rifabutin (PubChem CID:6323490); Rifamycin SV (PubChem CID:6324616); Rifapentine (PubChem CID:6323497); Rifaximin (PubChem CID:6436173); Rifamycin S (PubChem CID:6436726)
Highlights.
A mutant of Mycobacterium smegmatis is validated as a model for M. tuberculosis.
Some rifamycins oxidize with apparent reduction of antimicrobial activity.
Tear lipocalin protects susceptible rifamycins from oxidation.
Tear lipocalin binds and solubilizes rifalazil with resulting solvatochromism.
Tear lipocalin is a potential drug delivery vehicle for rifamycins in tuberculosis.
Acknowledgements
This work was supported by Public Health Service grant EY 11224 from the National Eye Institute (BG), the Edith and Lew Wasserman Endowed Professorship (BG)
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- [1].WHO global tuberculosis control report 2010 Summary. Cent Eur J Public Health. 2010;18:237. [PubMed] [Google Scholar]
- [2].Aristoff PA, Garcia GA, Kirchhoff PD, Hollis Showalter HD. Rifamycins--obstacles and opportunities. Tuberculosis (Edinb) 2010;90:94–118. doi: 10.1016/j.tube.2010.02.001. [DOI] [PubMed] [Google Scholar]
- [3].Sayada C. Re: Review article titled, “Rifamycins - Obstacles and opportunities” by Paul A. Aristoff, George A. Garcia, Paul D. Kirchhoff, H.D. Hollis Showalter. Tuberculosis. 2010;90(2):94–118. doi: 10.1016/j.tube.2010.02.001. Tuberculosis (Edinb) 2010;90:326; author reply -7. [DOI] [PubMed] [Google Scholar]
- [4].Snapper SB, Melton RE, Mustafa S, Kieser T, Jacobs WR., Jr. Isolation and characterization of efficient plasmid transformation mutants of Mycobacterium smegmatis. Mol Microbiol. 1990;4:1911–9. doi: 10.1111/j.1365-2958.1990.tb02040.x. [DOI] [PubMed] [Google Scholar]
- [5].Alexander DC, Jones JR, Liu J. A rifampin-hypersensitive mutant reveals differences between strains of Mycobacterium smegmatis and presence of a novel transposon, IS1623. Antimicrob Agents Chemother. 2003;47:3208–13. doi: 10.1128/AAC.47.10.3208-3213.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Pandey R, Khuller GK. Solid lipid particle-based inhalable sustained drug delivery system against experimental tuberculosis. Tuberculosis (Edinb) 2005;85:227–34. doi: 10.1016/j.tube.2004.11.003. [DOI] [PubMed] [Google Scholar]
- [7].Takenaga M, Ohta Y, Tokura Y, Hamaguchi A, Igarashi R, Disratthakit A, et al. Lipid microsphere formulation containing rifampicin targets alveolar macrophages. Drug Deliv. 2008;15:169–75. doi: 10.1080/10717540801952530. [DOI] [PubMed] [Google Scholar]
- [8].Johnson CM, Pandey R, Sharma S, Khuller GK, Basaraba RJ, Orme IM, et al. Oral therapy using nanoparticle-encapsulated antituberculosis drugs in guinea pigs infected with Mycobacterium tuberculosis. Antimicrob Agents Chemother. 2005;49:4335–8. doi: 10.1128/AAC.49.10.4335-4338.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [9].Sharma R, Muttil P, Yadav AB, Rath SK, Bajpai VK, Mani U, et al. Uptake of inhalable microparticles affects defence responses of macrophages infected with Mycobacterium tuberculosis H37Ra. J Antimicrob Chemother. 2007;59:499–506. doi: 10.1093/jac/dkl533. [DOI] [PubMed] [Google Scholar]
- [10].Qurratul A, Sharma S, Khuller GK, Garg SK. Alginate-based oral drug delivery system for tuberculosis: pharmacokinetics and therapeutic effects. J Antimicrob Chemother. 2003;51:931–8. doi: 10.1093/jac/dkg165. [DOI] [PubMed] [Google Scholar]
- [11].Sharma A, Sharma S, Khuller GK. Lectin-functionalized poly (lactide-co-glycolide) nanoparticles as oral/aerosolized antitubercular drug carriers for treatment of tuberculosis. J Antimicrob Chemother. 2004;54:761–6. doi: 10.1093/jac/dkh411. [DOI] [PubMed] [Google Scholar]
- [12].Kisich KO, Gelperina S, Higgins MP, Wilson S, Shipulo E, Oganesyan E, et al. Encapsulation of moxifloxacin within poly(butyl cyanoacrylate) nanoparticles enhances efficacy against intracellular Mycobacterium tuberculosis. Int J Pharm. 2007;345:154–62. doi: 10.1016/j.ijpharm.2007.05.062. [DOI] [PubMed] [Google Scholar]
- [13].Saraogi GK, Gupta P, Gupta UD, Jain NK, Agrawal GP. Gelatin nanocarriers as potential vectors for effective management of tuberculosis. Int J Pharm. 2010;385:143–9. doi: 10.1016/j.ijpharm.2009.10.004. [DOI] [PubMed] [Google Scholar]
- [14].Saraogi GK, Sharma B, Joshi B, Gupta P, Gupta UD, Jain NK, et al. Mannosylated gelatin nanoparticles bearing isoniazid for effective management of tuberculosis. J Drug Target. 2011;19:219–27. doi: 10.3109/1061186X.2010.492522. [DOI] [PubMed] [Google Scholar]
- [15].Clemens DL, Lee BY, Xue M, Thomas CR, Meng H, Ferris D, et al. Targeted intracellular delivery of antituberculosis drugs to Mycobacterium tuberculosis-infected macrophages via functionalized mesoporous silica nanoparticles. Antimicrob Agents Chemother. 2012;56:2535–45. doi: 10.1128/AAC.06049-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [16].Yeh PT, Casey R, Glasgow BJ. A novel fluorescent lipid probe for dry eye: retrieval by tear lipocalin in humans. Invest Ophthalmol Vis Sci. 2013;54:1398–410. doi: 10.1167/iovs.12-10817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [17].Buss WC, Reyes E, Barela TD. Metal ion catalyzed oxidation of the antibiotic rifampicin. Res Commun Chem Pathol Pharmacol. 1977;17:547–50. [PubMed] [Google Scholar]
- [18].Gasymov OK, Abduragimov AR, Gasimov EO, Yusifov TN, Dooley AN, Glasgow BJ. Tear lipocalin: potential for selective delivery of rifampin. Biochim Biophys Acta. 2004;1688:102–11. doi: 10.1016/j.bbadis.2003.11.006. [DOI] [PubMed] [Google Scholar]
- [19].Taube H. Mechanisms of oxidation with oxygen. J Gen Physiol. 1965;49(Suppl):29–52. doi: 10.1085/jgp.49.1.29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [20].Glasgow BJ, Heinzmann C, Kojis T, Sparkes RS, Mohandas T, Bateman JB. Assignment of tear lipocalin gene to human chromosome 9q34-9qter. Curr Eye Res. 1993;12:1019–23. doi: 10.3109/02713689309029229. [DOI] [PubMed] [Google Scholar]
- [21].Redl B, Holzfeind P, Lottspeich F. cDNA cloning and sequencing reveals human tear prealbumin to be a member of the lipophilic-ligand carrier protein superfamily. J Biol Chem. 1992;267:20282–7. [PubMed] [Google Scholar]
- [22].Glasgow BJ. Tissue expression of lipocalins in human lacrimal and von Ebner’s glands: colocalization with lysozyme. Graefes Arch Clin Exp Ophthalmol. 1995;233:513–22. doi: 10.1007/BF00183433. [DOI] [PubMed] [Google Scholar]
- [23].Marston FAO. DNA Cloning. IRL Press; Oxford, England: 1987. A Practical Approach. [Google Scholar]
- [24].Glasgow BJ, Abduragimov AR, Farahbakhsh ZT, Faull KF, Hubbell WL. Tear lipocalins bind a broad array of lipid ligands. Curr Eye Res. 1995;14:363–72. doi: 10.3109/02713689508999934. [DOI] [PubMed] [Google Scholar]
- [25].Bozimowski D, Artiss JD, Zak B. The variable reagent blank: protein determination as a model. J Clin Chem Clin Biochem. 1985;23:683–9. doi: 10.1515/cclm.1985.23.10.683. [DOI] [PubMed] [Google Scholar]
- [26].Gasymov OK, Abduragimov AR, Glasgow BJ. The conserved disulfide bond of human tear lipocalin modulates conformation and lipid binding in a ligand selective manner. Biochim Biophys Acta. 2011;1814:671–83. doi: 10.1016/j.bbapap.2011.03.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [27].Li J, Zhu M, Rajamani S, Uversky VN, Fink AL. Rifampicin inhibits alpha-synuclein fibrillation and disaggregates fibrils. Chem Biol. 2004;11:1513–21. doi: 10.1016/j.chembiol.2004.08.025. [DOI] [PubMed] [Google Scholar]
- [28].Woods GL. Susceptibility testing for mycobacteria. Clin Infect Dis. 2000;31:1209–15. doi: 10.1086/317441. [DOI] [PubMed] [Google Scholar]
- [29].Woods GL. Susceptibility Testing of Mycobacteria, Nocardiae, and other aerobic Actinomycetes; Approved Standard. NCCLS. 2003;23:18. [PubMed] [Google Scholar]
- [30].Humphreys IR, Stewart GR, Turner DJ, Patel J, Karamanou D, Snelgrove RJ, et al. A role for dendritic cells in the dissemination of mycobacterial infection. Microbes Infect. 2006;8:1339–46. doi: 10.1016/j.micinf.2005.12.023. [DOI] [PubMed] [Google Scholar]
- [31].O’Neil MJ, editor. The Merck Index, An encyclopedia of Chemicals, Drugs and Biologicals. Merck Research Laboratories; Whitehouse Station, NJ: 2001. [Google Scholar]
- [32].Reisbig RR, Woody AM, Woody RW. Rifampicin as a spectroscopic probe of the mechanism of RNA polymerase from Escherichia coli. Biochemistry. 1982;21:196–200. [Google Scholar]
- [33].Steffek M, Newton GL, Av-Gay Y, Fahey RC. Characterization of Mycobacterium tuberculosis mycothiol S-conjugate amidase. Biochemistry. 2003;42:12067–76. doi: 10.1021/bi030080u. [DOI] [PubMed] [Google Scholar]
- [34].Chakravorty S, Aladegbami B, Motiwala AS, Dai Y, Safi H, Brimacombe M, et al. Rifampin resistance, Beijing-W clade-single nucleotide polymorphism cluster group 2 phylogeny, and the Rv2629 191-C allele in Mycobacterium tuberculosis strains. J Clin Microbiol. 2008;46:2555–60. doi: 10.1128/JCM.00666-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [35].Piddock LJ, Williams KJ, Ricci V. Accumulation of rifampicin by Mycobacterium aurum, Mycobacterium smegmatis and Mycobacterium tuberculosis. J Antimicrob Chemother. 2000;45:159–65. doi: 10.1093/jac/45.2.159. [DOI] [PubMed] [Google Scholar]
- [36].Ren H, Liu J. AsnB is involved in natural resistance of Mycobacterium smegmatis to multiple drugs. Antimicrob Agents Chemother. 2006;50:250–5. doi: 10.1128/AAC.50.1.250-255.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [37].Danilchanka O, Pavlenok M, Niederweis M. Role of porins for uptake of antibiotics by Mycobacterium smegmatis. Antimicrob Agents Chemother. 2008;52:3127–34. doi: 10.1128/AAC.00239-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [38].Liu J, Nikaido H. A mutant of Mycobacterium smegmatis defective in the biosynthesis of mycolic acids accumulates meromycolates. Proc Natl Acad Sci U S A. 1999;96:4011–6. doi: 10.1073/pnas.96.7.4011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [39].Van der Auwera P, Matsumoto T, Husson M. Intraphagocytic penetration of antibiotics. J Antimicrob Chemother. 1988;22:185–92. doi: 10.1093/jac/22.2.185. [DOI] [PubMed] [Google Scholar]
- [40].Hirata T, Saito H, Tomioka H, Sato K, Jidoi J, Hosoe K, et al. In vitro and in vivo activities of the benzoxazinorifamycin KRM-1648 against Mycobacterium tuberculosis. Antimicrob Agents Chemother. 1995;39:2295–303. doi: 10.1128/aac.39.10.2295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [41].Yamamoto T, Amitani R, Suzuki K, Tanaka E, Murayama T, Kuze F. In vitro bactericidal and in vivo therapeutic activities of a new rifamycin derivative, KRM-1648, against Mycobacterium tuberculosis. Antimicrob Agents Chemother. 1996;40:426–8. doi: 10.1128/aac.40.2.426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [42].Maggi N, Arioli V, Sensi P, Rifamycins XLI. A new class of active semisynthetic rifamycins. N-substituted aminomethyl derivatives of rifamycin SV 2. J Med Chem. 1965;8:790–3. doi: 10.1021/jm00330a013. [DOI] [PubMed] [Google Scholar]
- [43].Maggi N, Pallanza R, Sensi P. New derivatives of rifamycin SV. Antimicrob Agents Chemother (Bethesda) 1965;5:765–9. [PubMed] [Google Scholar]
- [44].Sensi P, Maggi N, Furesz S, Maffii G. Chemical modifications and biological properties of rifamycins. Antimicrob Agents Chemother (Bethesda) 1966;6:699–714. [PubMed] [Google Scholar]
- [45].Jin Y, Gill SK, Kirchhoff PD, Wan B, Franzblau SG, Garcia GA, et al. Synthesis and structure-activity relationships of novel substituted 8-amino, 8-thio, and 1,8-pyrazole congeners of antitubercular rifamycin S and rifampin. Bioorg Med Chem Lett. 2011;21:6094–9. doi: 10.1016/j.bmcl.2011.08.054. [DOI] [PubMed] [Google Scholar]
- [46].Gill SK, Garcia GA. Rifamycin inhibition of WT and Rif-resistant Mycobacterium tuberculosis and Escherichia coli RNA polymerases in vitro. Tuberculosis (Edinb) 2011;91:361–9. doi: 10.1016/j.tube.2011.05.002. [DOI] [PubMed] [Google Scholar]
- [47].Bolt HM, Remmer H. Implication of rifampicin-quinone in the irreversible binding of rifampicin to macromolecules. Xenobiotica. 1976;6:21–32. doi: 10.3109/00498257609151608. [DOI] [PubMed] [Google Scholar]
- [48].Gasymov OK, Abduragimov AR, Glasgow BJ. Intracavitary ligand distribution in tear lipocalin by site-directed tryptophan fluorescence. Biochemistry. 2009;48:7219–28. doi: 10.1021/bi9005557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [49].Breustedt DA, Chatwell L, Skerra A. A new crystal form of human tear lipocalin reveals high flexibility in the loop region and induced fit in the ligand cavity. Acta Crystallogr D Biol Crystallogr. 2009;65:1118–25. doi: 10.1107/S0907444909031011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [50].Abduragimov AR, Gasymov OK, Yusifov TN, Glasgow BJ. Functional cavity dimensions of tear lipocalin. Curr Eye Res. 2000;21:824–32. doi: 10.1076/ceyr.21.4.824.5551. [DOI] [PubMed] [Google Scholar]
- [51].Gasymov OK, Abduragimov AR, Glasgow BJ. Evidence for internal and external binding sites on human tear lipocalin. Arch Biochem Biophys. 2007;468:15–21. doi: 10.1016/j.abb.2007.09.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [52].Gasymov OK, Abduragimov AR, Glasgow BJ. Ligand binding site of tear lipocalin: contribution of a trigonal cluster of charged residues probed by 8-anilino-1-naphthalenesulfonic acid. Biochemistry. 2008;47:1414–24. doi: 10.1021/bi701955e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [53].Breustedt DA, Korndorfer IP, Redl B, Skerra A. The 1.8-A crystal structure of human tear lipocalin reveals an extended branched cavity with capacity for multiple ligands. J Biol Chem. 2005;280:484–93. doi: 10.1074/jbc.M410466200. [DOI] [PubMed] [Google Scholar]
- [54].Gasymov OK, Abduragimov AR, Yusifov TN, Glasgow BJ. Site-directed tryptophan fluorescence reveals the solution structure of tear lipocalin: evidence for features that confer promiscuity in ligand binding. Biochemistry. 2001;40:14754–62. doi: 10.1021/bi0110342. [DOI] [PubMed] [Google Scholar]
- [55].Marini A, Munoz-Losa A, Biancardi A, Mennucci B. What is solvatochromism? J Phys Chem B. 2010;114:17128–35. doi: 10.1021/jp1097487. [DOI] [PubMed] [Google Scholar]
- [56].Glasgow BJ, Gasymov OK, Abduragimov AR, Yusifov TN, Altenbach C, Hubbell WL. Side chain mobility and ligand interactions of the G strand of tear lipocalins by site-directed spin labeling. Biochemistry. 1999;38:13707–16. doi: 10.1021/bi9913449. [DOI] [PubMed] [Google Scholar]
- [57].Nanashima K, Mawatari T, Tahara N, Higuchi N, Nakaura A, Inamine T, et al. Genetic variants in antioxidant pathway: risk factors for hepatotoxicity in tuberculosis patients. Tuberculosis (Edinb) 2012;92:253–9. doi: 10.1016/j.tube.2011.12.004. [DOI] [PubMed] [Google Scholar]
- [58].Nakajima A, Fukami T, Kobayashi Y, Watanabe A, Nakajima M, Yokoi T. Human arylacetamide deacetylase is responsible for deacetylation of rifamycins: rifampicin, rifabutin, and rifapentine. Biochem Pharmacol. 2011;82:1747–56. doi: 10.1016/j.bcp.2011.08.003. [DOI] [PubMed] [Google Scholar]
- [59].Woo J, Cheung W, Chan R, Chan HS, Cheng A, Chan K. In vitro protein binding characteristics of isoniazid, rifampicin, and pyrazinamide to whole plasma, albumin, and alpha-1-acid glycoprotein. Clin Biochem. 1996;29:175–7. doi: 10.1016/0009-9120(95)02024-1. [DOI] [PubMed] [Google Scholar]
- [60].Ishii M, Ogata H. Determination of rifampicin and its main metabolites in human plasma by high-performance liquid chromatography. J Chromatogr. 1988;426:412–6. doi: 10.1016/s0378-4347(00)81972-6. [DOI] [PubMed] [Google Scholar]
- [61].O’Sullivan DM, McHugh TD, Gillespie SH. Analysis of rpoB and pncA mutations in the published literature: an insight into the role of oxidative stress in Mycobacterium tuberculosis evolution? J Antimicrob Chemother. 2005;55:674–9. doi: 10.1093/jac/dki069. [DOI] [PubMed] [Google Scholar]
- [62].Dietze R, Teixeira L, Rocha LM, Palaci M, Johnson JL, Wells C, et al. Safety and bactericidal activity of rifalazil in patients with pulmonary tuberculosis. Antimicrob Agents Chemother. 2001;45:1972–6. doi: 10.1128/AAC.45.7.1972-1976.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [63].Mae T, Hosoe K, Yamamoto T, Hidaka T, Ohashi T, Kleeman JM, et al. Effect of a new rifamycin derivative, rifalazil, on liver microsomal enzyme induction in rat and dog. Xenobiotica. 1998;28:759–66. doi: 10.1080/004982598239173. [DOI] [PubMed] [Google Scholar]
- [64].Burman WJ, Gallicano K, Peloquin C. Comparative pharmacokinetics and pharmacodynamics of the rifamycin antibacterials. Clin Pharmacokinet. 2001;40:327–41. doi: 10.2165/00003088-200140050-00002. [DOI] [PubMed] [Google Scholar]
- [65].Hosoe K, Mae T, Konishi E, Fujii K, Yamashita K, Yamane T, et al. Pharmacokinetics of KRM-1648, a new benzoxazinorifamycin, in rats and dogs. Antimicrob Agents Chemother. 1996;40:2749–55. doi: 10.1128/aac.40.12.2749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [66].Mross K, Richly H, Fischer R, Scharr D, Buchert M, Stern A, et al. First-in-Human Phase I Study of PRS-050 (Angiocal), an Anticalin Targeting and Antagonizing VEGF-A, in Patients with Advanced Solid Tumors. PLoS One. 2013;8:e83232. doi: 10.1371/journal.pone.0083232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [67].Kaneda K, Naito S, Imaizumi S, Yano I, Mizuno S, Tomiyasu I, et al. Determination of molecular species composition of C80 or longer-chain alpha-mycolic acids in Mycobacterium spp. by gas chromatography-mass spectrometry and mass chromatography. J Clin Microbiol. 1986;24:1060–70. doi: 10.1128/jcm.24.6.1060-1070.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [68].Baba T, Kaneda K, Kusunose E, Kusunose M, Yano I. Thermally adaptive changes of mycolic acids in Mycobacterium smegmatis. J Biochem. 1989;106:81–6. doi: 10.1093/oxfordjournals.jbchem.a122825. [DOI] [PubMed] [Google Scholar]
- [69].Gasymov OK, Abduragimov AR, Yusifov TN, Glasgow BJ. Binding studies of tear lipocalin: the role of the conserved tryptophan in maintaining structure, stability and ligand affinity. Biochim Biophys Acta. 1999;1433:307–20. doi: 10.1016/s0167-4838(99)00133-8. [DOI] [PubMed] [Google Scholar]
- [70].Halaas O, Steigedal M, Haug M, Awuh JA, Ryan L, Brech A, et al. Intracellular Mycobacterium avium intersect transferrin in the Rab11(+) recycling endocytic pathway and avoid lipocalin 2 trafficking to the lysosomal pathway. J Infect Dis. 2010;201:783–92. doi: 10.1086/650493. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.