Abstract
We previously performed a lipid vesicle-based, high-throughput screen on a 26-residue combinatorial peptide library that was designed de novo to yield membrane permeabilizing peptides that fold into β-sheets. The most active and soluble library members that were identified permeabilized lipid vesicles detectably, but not with high potency. Nonetheless, they were broad-spectrum, membrane-permeabilizing antibiotics with minimum sterilizing activity at low μM concentrations. In an expansion of that work, we recently performed an iterative screen in which an active consensus sequence from that first generation library was used as a template to design a second generation library which was then screened against lipid vesicles at very high stringency. Compared to the consensus sequence from the first library, the most active second generation peptides are highly potent, equilibrium pore-formers in synthetic lipid vesicles. Here we use these first and second generation families of peptides to test the hypothesis that a large increase in potency in bacteria-like lipid vesicles will correlate with a large improvement in antimicrobial activity. The results do not support the hypothesis. Despite a 20-fold increase in potency against bacteria-like lipid vesicles, the second generation peptides are only slightly more active against bacteria, and at the same time, are also more toxic against mammalian cells. The results suggest that a “pipeline” strategy towards the optimization of antimicrobial peptides could begin with a vesicle-based screen for identifying families with broad-spectrum activity, but will also need to include screening or optimization steps that are done under conditions that are more directly relevant to possible therapeutic applications.
Introduction
The design of novel, membrane-permeabilizing antimicrobial peptides (AMP) that are potent against microbes but nontoxic to host cells has been a long-standing goal in bioengineering(1;2). The realization of this goal could help solve the growing crisis of morbidity and mortality caused by drug resistant bacteria(3). As an aid to rational design, researchers have examined the structure and function of natural and synthetic AMPs in lipid vesicles as surrogate biomembranes. Yet, despite decades of work, the molecular mechanism of action of antimicrobial peptides is not well understood(4;5). As a consequence of this roadblock, the rational design or optimization of new antimicrobial peptides has not been realized.
We and others have shown that combinatorial chemistry and high throughput screening are powerful tools when applied to design problems such as this one. Specifically, we showed that orthogonal high-throughput selection for peptide solubility and for the permeabilization of bacteria-like lipid vesicles holds promise as a method for identifying new AMPs(6–9). In one study, we described a high-throughput screen that was used to identify a family of 26-residue peptides, which have broad-spectrum antimicrobial activity at μM peptide concentration despite permeabilizing anionic synthetic lipid vesicles only at high peptide to lipid ratios(10). The members of this family also have moderately low toxicity against eukaryotic cells, suggesting that the vesicle-based screen may have utility for the discovery of novel, clinically useful AMPs, an idea that we have verified in other systems(6–8).
In a more recent study(11) we identified a family of peptides that were closely related to the 26-residue AMPs discussed above, but with significantly higher vesicle permeabilizing potency. To identify such gain-of-function sequences we used a synthetic molecular evolution-based approach in which an active consensus sequence from the first library was used as a template to design a second library in which we varied an alternate set of amino acid residues. The second library was screened for members that formed pores in lipid vesicles that are active at equilibrium (i.e., hours after peptide addition) and that are active at very high stringency using a new orthogonal high throughput assay(11). The most active sequences selected in the second-generation, iterative screen were significantly more potent against lipid vesicles than the consensus sequence from the first screen with membrane permeabilizing activity at peptide to lipid ratios (P:L) of 1:1000 where the first generation peptides are active at P:L=1:50(11). In fact, the second-generation pore-formers are as potent as the most active natural pore forming peptide toxins known, including melittin and alamethicin(11–13). In addition to the differences in activity level, the mechanism of membrane permeabilization was different. Specifically, the second-generation peptides formed equilibrium pores in vesicles that could still be detected hours after addition(11) whereas the first generation AMPs caused transient leakage events that were only present for 15–30 minutes after addition of peptide(10). Thus, we were successful in developing and using a novel and powerful approach, based on synthetic molecular evolution, to identify peptides with much higher membrane permeabilization in lipid vesicles(11).
For this work, we hypothesized that the second generation peptides would be better antimicrobial peptides than the first generation peptides because they are very similar to the first generation in secondary structure, molecular weight and hydrophobicity, but have much better membrane permeabilizing activity in bacteria-like anionic vesicles. Here we use these two generations of closely related peptides to ask, for the first time, a fundamental question about antimicrobial peptides: Does a large increase in vesicle permeabilizing potency within a closely homologous peptide family also lead to increased antimicrobial activity, increased cytotoxicity, or both? Our results show that the correlation between vesicle activity and biological activity is weak or nonexistent. Although vesicle-based screens clearly have utility as a first generation selection method, additional optimization or refinement of antimicrobial peptides will need to done with alternate methods that better represent the potential therapeutic applications.
Results
Membrane permeabilizing peptides
The seven peptides we used in this work are shown in Figure 1. The consensus sequence from the first iteration library, named FSKRGY for its variable residues, is 26 residues long, has a net charge of +7. Its Wimley-White interfacial hydrophobicity (bilayer to water) is +0.4 kcal/mol(14;15) indicating that its inherent hydrophobic interaction with bilayers is favorable, but very weak. Measurable binding occurs only because the peptide self-assembles into β-sheets in membranes, thereby reducing the cost of backbone partitioning(14). The other six “iteration” peptides, named It1-x, are from the second generation, iterative screen(11). It1-c is a consensus sequence created by combining the commonly observed residues. The other five sequences were actually observed in the screen(11). The iteration peptides are 25 or 26 residues long and share at least 20 residues in common with FSKRGY (identity ≥ 77%). The iteration peptides have net charges of +7 to +9, and are slightly more hydrophobic than FSKRGY by virtue of the fact that hydrophobic residues were selected in the six residues that were varied in the iterative library. Like FSKRGY, the iteration peptides bind strongly to membranes because they also self-assemble into β-sheets(11).
Figure 1.
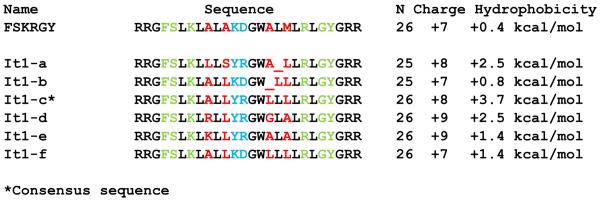

Sequences of the peptides synthesized for further study. FSKRGY is the 26 residue template sequence, identified in an earlier library and used as a template for the second generation library. The active peptides from the second generation library are the “iteration” peptides and are named It1-x. It1-c* is a consensus sequence, while the others were actually observed in the screen. The number of residues and the total charge are shown on the right, along with the Wimley-White interfacial hydrophobicity(14) for bilayer to water partitioning. The residues selected in the first generation screen are shown in green. Residues varied in the second generation screen are red (in the two putative diad repeat sequences) or blue (in the turn sequence).
Lipid vesicle permeabilization
The consensus peptide FSKRGY and the six iteration peptides, It1-a through It1-f, were tested for lipid vesicle permeabilization using three lipid compositions that are relevant to antimicrobial and cytotoxic activity: 90% phosphatidylglycerol (PC) + 10% phosphatidylcholine (PG), 100% PC and PC with 25% cholesterol (all are molar percentages). Many bacterial cell membranes are anionic because they are composed of a significant percentage of negatively charged phospholipids such as PG. Therefore, PC/PG vesicle are a rough mimic of a generic, anionic microbial membrane. On the other hand, in mammalian cells the outer leaflet of the plasma membrane contains mainly zwitterionic phospholipids such as PC plus variable amounts of the neutral sterol, cholesterol. Thus, 100% PC vesicles and PC with 25% cholesterol represents the extremes of a simple zwitterionic generic mammalian plasma membrane.
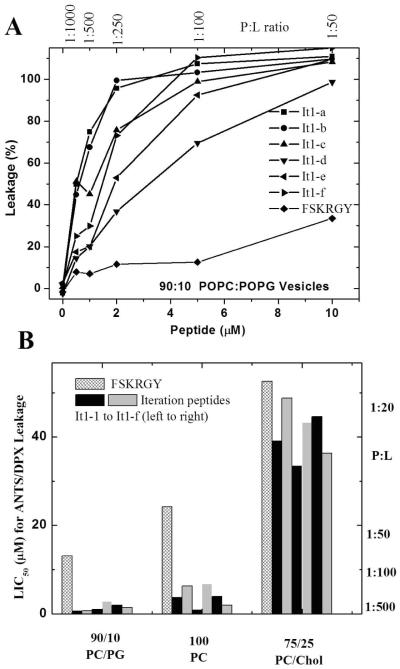
A typical set of leakage curves are shown in Figure 2A, and the summary data are shown in Fig, 2B. When expressed as the concentration that induces 50% leakage (LIC50) the iteration peptides are 5–20 fold more potent than FSKRGY against POPC/POPG vesicles and against POPC vesicles. This sharp difference in potency occurs despite the fact that they are about 80% identical to FSKRGY. The presence of 25% cholesterol inhibits the activity of the peptides substantially such that FSKRGY and the It1 peptides all have similar, low activity. While cholesterol is known to reduce the activity of membrane-permeabilizing peptides(16–18) it rarely has such a large effect. Here, the large effect is probably related to the effect of membrane fluidity on folding and self-assembly of β-sheets in membranes.
Figure 2.
Membrane permeabilization by template peptide FSKRGY and by the selected iteration peptides. Leakage of ANTS/DPX from 90:10 PC/PG vesicles was measured by fluorescence. In this experiment we used vesicles made from various lipid compositions at 1 mM lipid and measured the amount of leakage at three hours after the addition of peptides at various concentration. A: Leakage was measured at 3 hours after addition of peptide and was normalized to untreated vesicles (0% leakage) and Triton-solubilized vesicles (100% leakage). B: The leakage curves in panel A are hyperbolic. The peptide concentration that induces 50% leakage (LIC50 or Leakage Inducing Concentration for 50%) was obtained by curve fitting and plotted here. We measured leakage in POPC vesicles with 10% POPG, in plain POPC vesicles and in POPC vesicles with 25 mol% cholesterol. Measured values are means of 3–5 experiments. Standard errors of the LIC50 measurements are about 10% of the LIC50 value.
Antimicrobial activity
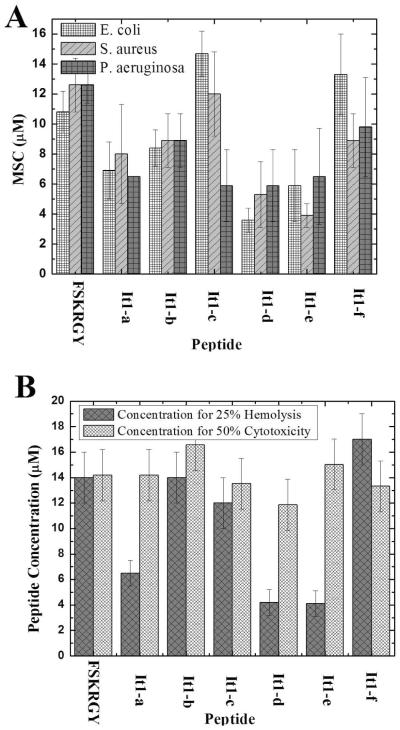
Our primary goal in this work was to directly compare antimicrobial, hemolytic and cytotoxic bioactivities in this set of closely related peptides that have very different membrane permeabilizing activity in lipid vesicles. To begin, we measured the minimum sterilizing concentration of FSKRGY and the iteration peptides against a panel of Gram negative and Gram positive microbes as described elsewhere(6;7;10). To increase the stringency and relevance of these experiments we increased the bacterial cell counts substantially from previous measurements. Despite the much higher potency of the iteration peptides in vesicle permeabilizing assays, there are only small differences in antimicrobial activity between them (Fig. 3A). The largest difference in minimum sterilizing concentration (MSC) between FSKRGY and the iteration peptides is about a factor of two, much less than the 20-fold increase we observed in vesicle leakage. By ANOVA, none of the pairwise comparisons are statistically significant.
Figure 3.
Bioactivities of the peptides studied. A: Minimum sterilizing concentrations (MSC) against three strains of bacteria were measured by incubating bacteria with peptide in a serial dilution experiment and then incubating the plates overnight to allow for recovery and growth of microbes. The values given are the mean of the Minimum sterilizing concentration +/− standard error of at least 3 experiments. B: Cytolysis and cytotoxicity. Cells were incubated with a series of peptide concentrations followed by measurement of hemolysis or toxicity. Hemolysis was quantified by measuring the fractional release of hemoglobin from human erythrocytes after 60 min incubation. Cytotoxicity was quantified my measuring the relative viability of CHO cells, using the MTT assay, after incubating them with peptide for two days. Values are mean +/− standard error for at least 3 experiments.
The peptide FSKRGY has broad-spectrum antimicrobial activity; it is similarly potent against Gram negative and Gram positive bacteria(10). This property is also conserved in the iteration peptides, as most of them have indistinguishable MSC values when comparing the activity against E. coli and Pseudomonas aeruginosa (Gram negative) and against Staph. aureus (Gram positive). We have shown that peptide libraries like the first and second generation libraries discussed in this work contain many species-specific antimicrobial peptides, but that broad-spectrum activity like this is rare(6). From this observation we have concluded that the vesicle-based screen in anionic POPC/POPG vesicles specifically selects for the rare broad-spectrum antimicrobial activity. Interestingly, the consensus iteration peptide It1-c, which was not actually selected in the screen, has the greatest variation in activity amongst the different microbes, further supporting the conclusion that the vesicle-based screen effectively selects for broad-spectrum antimicrobial activity.
Cytolytic and cytotoxic activity
We also studied the effect of FSKRGY and the iteration peptides on acute cytolysis of human erythrocytes, and on long term cytotoxicity against mammalian (Chinese Hamster ovary) cells in culture. Hemolysis of human erythrocytes was measured after incubation with peptide for 60 minutes as described elsewhere(7;10) and provides a direct measure of biomembrane permeabilization. Long term cytotoxicity was measured in living Chinese Hamster Ovary (CHO) cells by incubating cells in culture with peptide for 48 hours, followed by measurement of the cell viability with the MTT assay. The results of these measurements are shown together in Fig. 3B. All of the peptides are somewhat cytolytic and cytotoxic at microbicidal concentrations. Some of the iteration peptides are slightly more lytic and/or more toxic than the template sequence, FSKRGY. Others have the same toxicity as FSKRGY. However, even the largest differences are again much smaller than the 20 fold increase in potency observed in lipid vesicles.
Discussion
Insight into antimicrobial peptide design
In many studies it has been shown that the AMP-induced leakage of entrapped contents from liposomes happens in a rapid, transient burst of partial release that occurs for only a few minutes after the addition of peptide(4;5;12;19–21). Here we tested the hypothesis that it is possible to optimize membrane-permeabilizing antimicrobial peptides by using synthetic molecular evolution to identify peptides that are potent, equilibrium pore-formers in anionic, bacterial membrane-like vesicles. We found that the hypothesis is not correct. While we successfully used a de novo library screen to identify broad-spectrum antimicrobial peptides(9), and successfully used iterative library design and screening to identify very potent vesicle permeabilizing peptides(11), the improvements in vesicle permeabilizing potency were not strongly correlated with increased antimicrobial activity. Instead, we found that an increase in permeabilizing potency in lipid vesicles by up to 20-fold corresponds to only very small increases, if any, in antimicrobial activity. And the slight gain in antimicrobial activity is offset by a similar increase in cytolytic and cytotoxic activity against mammalian cells. These results support the growing belief that, while there are similarities that arise from an overlap in physical chemistry, the mechanism of antimicrobial of peptide activity in bacteria is not the same as the mechanism of permeabilization of lipid vesicles.
Optimization of antimicrobial peptides
A long-term goal of our lab is the optimization of antimicrobial peptides that could be used as drugs in the clinic to fight against drug-resistant bacterial infections. There is growing concern about the differences in the experimental conditions in assays of lipid vesicle permeabilization, microbial sterilization and host cell cytolytic activity or cytotoxicity. Moreover, the binding and interaction of antimicrobial peptides with microbial cells is clearly more complex that the simple math of peptide:lipid ratio(5). Antimicrobial activity in vivo is further complicated by the presence of mammalian cells to which some AMPs can bind. We have shown in published studies that selecting peptides from de novo designed combinatorial peptide libraries for vesicle permeabilizing activity leads to the identification of potent, broad-spectrum antimicrobial peptides with moderately low cytotoxic/cytolytic activity against mammalian cells(6–10). While their in vitro activity is promising, these peptides, and others like them, will need to be optimized further if they are to be considered as candidate anti-infective therapeutics. The work presented here showed that a synthetic molecular evolution-based approach using two iterative generations of libraries screened against lipid vesicles, did not result in the needed optimization.
Based on this outcome, we suggest that a “pipeline” strategy towards clinically useful antimicrobial peptides could begin with a vesicle-based screen to identify families of peptides with broad-spectrum in vitro activity, but will also need to include an additional screening/optimization step conducted in composite systems that better represent the ultimate therapeutic applications. For example, peptides could be optimized for broad-spectrum microbicidal activity in the presence of mammalian cells, and for simultaneous lack of binding to, or toxicity towards, host cells even at high concentration.
MATERIALS AND METHODS
ANTS/DPX leakage from lipid vesicles
ANTS at 5 mM and DPX at 12.5 mM were entrapped in 0.1 μm diameter extruded vesicles with various lipid compositions as described elsewhere(10;22;23). ANTS/DPX containing vesicles(24) were diluted to 1 mM total lipid concentration and peptide was added at concentrations between 0.5 and 20 μM, giving P:L between 0.0005 (1:2000) and 0.02 (1:50). Leakage was quantitated by the increase in ANTS fluorescence that occurs when the entrapped DPX quencher is diluted into the external buffer upon permeabilization. Leakage was measured after 3 hour incubation. The concentration dependence of leakage was fit with a hyperbolic/sigmoidal curve, and the peptide concentration that causes 50% leakage (LIC50) was determined from the curve fits.
Antimicrobial activity
E. coli, P. aeruginosa, and S. aureus were diluted to 106 CFU/ml. In sterile 96 well plates, eight columns were prepared for each peptide in 2/3 fold serial dilution starting at 10 μM. To each well a cell suspension in minimal media was added, wells were pre-incubated at 37°C for up to 3 hours and then 2X concentrated growth medium was added. After overnight recovery at 37 °C, wells were either opaque (OD600 > 0.5) indicating stationary phase growth or they were transparent (OD600 < 0.02) indicating no growth. Intermediate optical densities were rarely observed. For each experiment there were a number of sterile wells starting from the highest concentration. The lowest concentration of peptide that sterilized the cells is the minimum sterilizing concentration (MSC). All MSC measurements are the average of 3–5 independent experiments. Addition of rich growth medium to cells first followed by peptide gave very similar MSC values.
Hemolysis
Human red blood cells (RBC) were diluted to 7.7 × 106 cells/ml, and the assay was performed in 96 well plates in a final volume of 100 μl. Peptides, at a series of concentration up to 20 μM were incubated with RBC at 37°C for 1hr. The plates were then centrifuged at 4,000 g for 2 minutes, and the release of hemoglobin was monitored by measuring the absorbance of the supernatant at 540 nm. Baseline was defined by RBC incubated with PBS-EDTA pH 7.4. Complete lysis was normalized to measuring complete osmotic lysis in distilled water.
Cytotoxicity
Chinese Hamster Ovary cells were seeded into 96-well culture plates followed by addition of peptide at concentrations up to 20 μM. Following the addition of peptides, the cells were incubated for 48 hours at 37°C with 5% CO2 in a humidified incubator during which time the cells adhered to the plate. After the 48 hour incubation, 50 μg of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) was added to each well and incubated for six hours. In living cells, mitochondrial reductases convert the MTT tetrazolium to formazan which precipitates. Formazan crystals are dissolved using acidified isopropanol (90% isopropanol : 10% Triton-X-100: 0.4% HCl) and the optical density at 550 nm was measured using a plate spectrophotometer. Melittin at 5 μM was used as a positive control resulted in complete loss of cell viability.
Acknowledgements
We thank Andrew R Hoffmann and Andrew W. Wimley for peptide characterization. Funded by NIH grant GM060000 and NSF grant DMR-1003411.
References
- 1.Hancock RE, Sahl HG. Nat Biotechnol. 2006;24:1551–1557. doi: 10.1038/nbt1267. [DOI] [PubMed] [Google Scholar]
- 2.Hancock RE, Brown KL, Mookherjee N. Immunobiology. 2006;211:315–322. doi: 10.1016/j.imbio.2005.10.017. [DOI] [PubMed] [Google Scholar]
- 3.D'Agata EM. Infect Control Hosp Epidemiol. 2004;25:842–846. doi: 10.1086/502306. [DOI] [PubMed] [Google Scholar]
- 4.Wimley WC, Hristova K. J Membr Biol. 2011;239:27–34. doi: 10.1007/s00232-011-9343-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wimley WC. ACS Chem Biol. 2010;5:905–917. doi: 10.1021/cb1001558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Rathinakumar R, Wimley WC. FASEB J. 2010;24:3232–3238. doi: 10.1096/fj.10-157040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Rathinakumar R, Walkenhorst WF, Wimley WC. J Am Chem Soc. 2009;131:7609–7617. doi: 10.1021/ja8093247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Rathinakumar R, Wimley WC. J Am Chem Soc. 2008;130:9849–9858. doi: 10.1021/ja8017863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Rausch JM, Marks JR, Wimley WC. Proc Natl Acad Sci USA. 2005;102:10511–10515. doi: 10.1073/pnas.0502013102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rausch JM, Marks JR, Rathinakumar R, Wimley WC. Biochemistry. 2007;46:12124–12139. doi: 10.1021/bi700978h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Krauson AJ, He J, Hoffmann AR, Wimley AW, Wimley WC. ACS Chem Biol. 2013 doi: 10.1021/cb300598k. dx.doi.org/10.1021/cb300598k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Krauson AJ, He J, Wimley WC. Biochim Biophys Acta. 2012;1818:1625–1632. doi: 10.1016/j.bbamem.2012.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Krauson AJ, He J, Wimley WC. J Am Chem Soc. 2012;134:12732–12741. doi: 10.1021/ja3042004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.White SH, Wimley WC. Annu Rev Biophys Biomol Struct. 1999;28:319–365. doi: 10.1146/annurev.biophys.28.1.319. [DOI] [PubMed] [Google Scholar]
- 15.Wimley WC, White SH. Nature Struct Biol. 1996;3:842–848. doi: 10.1038/nsb1096-842. [DOI] [PubMed] [Google Scholar]
- 16.Ramamoorthy A, Lee DK, Narasimhaswamy T, Nanga RP. Biochim Biophys Acta. 2010;1798:223–227. doi: 10.1016/j.bbamem.2009.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Raghuraman H, Chattopadhyay A. Biophys J. 2007;92:1271–1283. doi: 10.1529/biophysj.106.088690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Raghuraman H, Chattopadhyay A. Chem Phys Lipids. 2005;134:183–189. doi: 10.1016/j.chemphyslip.2004.12.011. [DOI] [PubMed] [Google Scholar]
- 19.Gordon-Grossman M, Zimmermann H, Wolf SG, Shai Y, Goldfarb D. J Phys Chem B. 2012;116:179–188. doi: 10.1021/jp207159z. [DOI] [PubMed] [Google Scholar]
- 20.Shai Y. Biopolymers. 2002;66:236–248. doi: 10.1002/bip.10260. [DOI] [PubMed] [Google Scholar]
- 21.Shai Y, Oren Z. Peptides. 2001;22:1629–1641. doi: 10.1016/s0196-9781(01)00498-3. [DOI] [PubMed] [Google Scholar]
- 22.Ladokhin AS, Wimley WC, White SH. Biophys J. 1995;69:1964–1971. doi: 10.1016/S0006-3495(95)80066-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wimley WC, Selsted ME, White SH. Protein Sci. 1994;3:1362–1373. doi: 10.1002/pro.5560030902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ladokhin AS, Wimley WC, Hristova K, White SH. Methods Enzymol. 1997;278:474–486. doi: 10.1016/s0076-6879(97)78025-x. [DOI] [PubMed] [Google Scholar]