Abstract
Interleukin-1 (IL-1)-family cytokines are mediators of innate and adaptive immunity. They exert proinflammatory effects by binding a primary receptor that recruits a receptor accessory protein to form a signaling-competent heterotrimeric complex. Here we present the crystal structure of IL-1β bound to its primary receptor IL-1RI and its receptor accessory protein IL-1RAcP, providing insight into how IL-1–type cytokines initiate signaling and revealing an evolutionary relationship with the fibroblast growth factor receptor family.
Members of the IL-1 family of cytokines are important regulators of the innate and adaptive immune system1. Signal transduction by these cytokines is initiated by forming a heterotrimeric complex consisting of the ligand, a primary receptor subunit and a receptor accessory protein. Assembly of the ternary ligand–receptor complex juxtaposes the intracellular portions of the receptor chains containing Toll/IL-1 receptor (TIR) domains, instigating cytoplasmic signaling cascades. The pronounced proinflammatory activities of IL-1–type cytokines are tightly regulated at multiple levels through mechanisms that include decoy receptors and antagonistically acting ligands. Dysregulated IL-1 signaling has been shown to be involved in several human autoinflammatory diseases2. To gain insight into the assembly and architecture of the IL-1β–receptor signaling complex, we determined the crystal structure of IL-1β bound to IL-1RI and IL-1RAcP.
We determined the structure of human IL-1β in complex with the extracellular regions of IL-1RI and IL-1RAcP at a resolution of 3.1 Å (Fig. 1a, Supplementary Table 1 and Supplementary Methods). The overall architecture of the IL-1β–IL-1RI–IL-1RAcP signaling complex resembles the nonproductive decoy receptor complex containing IL-1RII3 (Supplementary Fig. 1a), which lacks the cytoplasmic TIR domain necessary for signal transduction. The r.m.s. deviation for all backbone Cα atoms is ∼2.1 Å. The three subdomains of IL-1RI wrap around the 12-stranded β-trefoil of IL-1β like a grasping hand4, and the IL-1β–IL-1RI binary complex forms a composite binding interface for the recruitment of IL-1RAcP. The structure of the IL-1β–IL-1RI binary complex is virtually unchanged by the binding of IL-1RAcP. The r.m.s. deviation for all backbone Cα atoms is ∼1.4 Å between the IL-1β–IL-1RI of the ternary complex and the previously published free binary complex4. The N-terminal subdomain of IL-1RAcP points away from the rest of the complex and forms no interactions. The crystal packing shows no evidence for higher-order complex formation that might involve this subdomain of IL-1RAcP.
Figure 1.
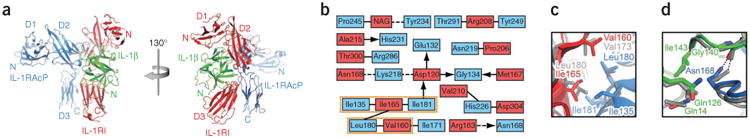
Structural features of the IL-1β signaling complex. (a) Ribbon diagram of IL-1β bound to the ectodomains of IL-1RI and IL-1RAcP in two different views, related to each other by a 130° rotation around the vertical axis. (b) Two-dimensional interaction map of the IL-1RI–IL-1RAcP interface. Amino acids are depicted as nodes. Interactions between side chains are represented by lines; interactions between side chains and backbone are depicted as arrows pointing toward the backbone. Van der Waals interactions and hydrophobic contacts are shown as solid lines, hydrogen bonds or electrostatic interactions as dashed lines. Interactions shared between the signaling and decoy receptor complex are indicated with an orange box. (c) Close-up view of the hydrophobic patch in the IL-1RI–IL-1RAcP interface that is shared between the signaling and decoy receptor complex. The decoy receptor complex is shown in gray. (d) Close-up view of selected residues in the IL-1β–IL-1RAcP interface. The hydrogen bond formed by Asn168 is depicted as a dashed line. The decoy receptor complex is shown in gray.
The IL-1RI–IL-1RAcP interface consists mainly of interactions that are distinct from those of the IL-1RII decoy receptor complex (Fig. 1b and Supplementary Figs. 2 and 3). The only interactions shared between the two complexes are found in a hydrophobic patch and are formed between Ile135, Ile181 and Leu180 in IL-1RAcP and Val160 (Val173 in IL-1RII) and Ile165 (Leu180 in IL-1RII) of IL-1RI (Fig. 1b,c and Supplementary Figs. 2 and 3). The unique interactions of the IL-1RI–IL-1RAcP interface include contacts between an N-acetylglucosamine moiety in IL-1RI (attached to Asn216) and Pro245 and Tyr234 in IL-1RAcP (Fig. 1b). In contrast to the IL-1R–IL-1RAcP interface, no significant differences between the signaling and decoy receptor complexes were observed in the interactions of the IL-1β– IL-1RAcP interface (Fig. 1d and Supplementary Figs. 1b and 3).
Despite low sequence homology, cytokines of the fibroblast growth factor (FGF) and IL-1β families share a common β-trefoil fold5. Structures of IL-1RI bound to IL-1β and IL-1Ra4,6 revealed a distinctive question mark–shaped architecture of the receptor grasping the ligands and provided a model for how the similar modular ectodomain of an FGF receptor (FGFR) would engage an FGF cytokine7. Although the FGF–FGFR binary complex does indeed resemble the IL-1RI structures, the FGFR signaling complex has a dimer-of-dimers design8 that contrasts with the heterotrimeric assembly of the IL-1 signaling complex, which contains a secondary receptor accessory protein that is incapable of cytokine binding9. Furthermore, in contrast to the IL-1 complexes, the N-terminal FGFR immunoglobulin domain is dispensable for cytokine binding, although it may retain an autoregulatory function in the absence of FGF10,11; in addition, formation of the FGF–FGFR complex depends on crossing heparin strands that act as molecular glue. Given the structural conservation of protein-protein contacts across divergent families of interacting molecules12—most strikingly observed in the complex structures of coevolving cytokines and receptors in the hematopoietic and TGF-β superfamilies13,14—can we consider FGF and IL-1 signaling complexes to be evolutionarily related?
The crystal structure of IL-1β bound to the decoy receptor IL-1RII and the accessory chain IL-1RAcP revealed a unique ternary-complex architecture, in which the convex face of IL-1RAcP is docked to one side of the IL-1β–IL-1RII protomer3. The present work showing the active IL-1RI–containing form of this complex proposes that this molecular design is paradigmatic for the IL-1 family of cytokines and receptors. Superimposing complexed IL-1β and one of the FGF ligands from a 2:2 FGF–FGFR complex8 reveals close parallels in receptor geometry and contacts between the ligand-aided IL-1RI–IL-1RAcP heterodimer and the FGFR homodimer (Fig. 2a,b and Supplementary Fig. 2). Visible differences between the two complexes include the cytokine-free IL-1RAcP chain on the first versus the FGF-bound second FGFR receptor chain on the second, and the role of the N-terminal immunoglobulin module of IL-1RI in IL-1β binding, which is in contrast with the dispensability of the same domain in FGFR (Fig. 2). Furthermore, the third immunoglobulin domain in IL-1RAcP is rotated relative to the FGF-engaged C-terminal immunoglobulin domain in the second FGFR protomer, which does not contact the first protomer. The rotation of the third immunoglobulin domain in IL-1RAcP is necessary for it to touch the composite face of IL-1RI and IL-1β3. Closer inspection of the IL-1β and FGF ligand interfaces with the common, C-terminal immunoglobulin domain pair of their respective IL-1RI and FGFR receptors reveals a core set of conserved site-I and site-II contacts (using the nomenclature in ref. 3; data not shown); a glancing site-III contact between bound cytokines and the second receptor chain is also preserved. The more compact site-IV footprint of FGFRs, which lacks the C-terminal immunoglobulin contacts, still superposes on the structurally equivalent IL-1RI–IL-1RAcP interface (data not shown).
Figure 2.
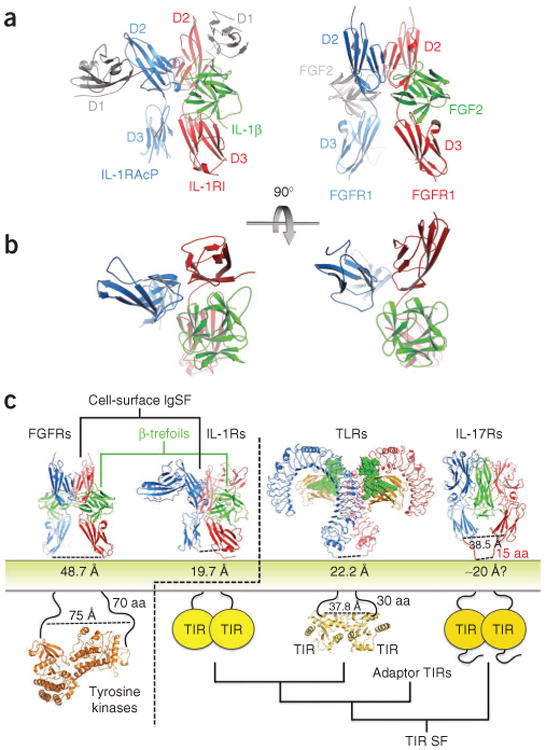
Comparison of IL-1β and FGF signaling complexes. (a) Side-by-side view of the active IL-1β–IL-1RI–IL-1RAcP heterocomplex and a representative FGF2–FGFR1 homodimeric complex. Non-equivalent N-terminal immunoglobulin domains in the IL-1 receptors, and a second bound FGF2 cytokine, are depicted in light gray. (b) Top view of the two complexes; the non-equivalent domains and second ligand have been deleted to highlight the similar interaction geometries of the three core receptors and β-trefoil ligand components. (c) Evolutionary landscape of TIR domain receptors. Structures of the FGF–FGFR complex and TIR domain receptor complexes (IL-1–IL-1RI–IL-1RAcP ternary complex, Toll-like receptor (TLR) complex and IL-17RA–IL-17F complex). Separations between receptor ectodomains as the chains enter the membrane are denoted below the respective structure. The IL-1R family architecture places the receptors at the evolutionary junction of two distinct signaling systems. FGF–FGFR homodimeric complex PDB 1FQ9 (ref. 8); FGFR kinase PDB 3CLY16; TLR complex PDB 3FXI; TLR10 TIR domain PDB 2J67 (ref. 17); IL-17RA–IL-17F PDB 3JVF18. SF, superfamily.
Divergent helical cytokines and receptors across the class-I and class-II divisions of the hematopoietic superfamily conserve a core binding structure that has sufficient plasticity to accommodate wide sequence divergence, promiscuous interactions and the influence of distinct extra-core domains14,15; likewise, the TGF-β and BMP clans of the TGF-β superfamily use a related heteromeric array of components to build architecturally and functionally similar signaling complexes13. In the case of the distantly related IL-1 and FGF cytokine systems, evolutionary divergence has given rise to two apparently distinct signaling complexes: (i) a homodimer of FGFRs bound equivalently to FGF cytokines and fused to intracellular kinase domains and (ii) a heteromer of IL-1R chains that are trained on a single IL-1 cytokine and signal through cytoplasmic TIR domains. Still, the extent of domain-level conservation of structure and interactions between the two active signaling complexes suggests an ancestral protomer of a β-trefoil ligand with an extracellular three-immunoglobulin domain receptor; whereas FGFRs have retained the homodimeric assembly of cytokine-loaded protomers, the loss of ligand binding by a divergent branch of IL-1 accessory receptors has created singly loaded heteromeric complexes (Fig. 3 and Supplementary Discussion). This evolutionary split manifests itself in very different separations between receptor ectodomains as the chains enter the membrane, leading to distinct FGFR and IL-1 receptor intracellular signaling machineries. FGFR C termini are nearly 49 Å apart, favoring the larger 75-Å gap between the N termini of their activating kinase dimers, whereas the smaller 20-Å gap between IL-1 receptors is similar to the 22-Å distance between the C termini of Toll-like receptors (TLRs); both the IL-1 receptor system and the Toll-like receptor system are linked to more closely packed TIR domains (Fig. 2c).
Figure 3.
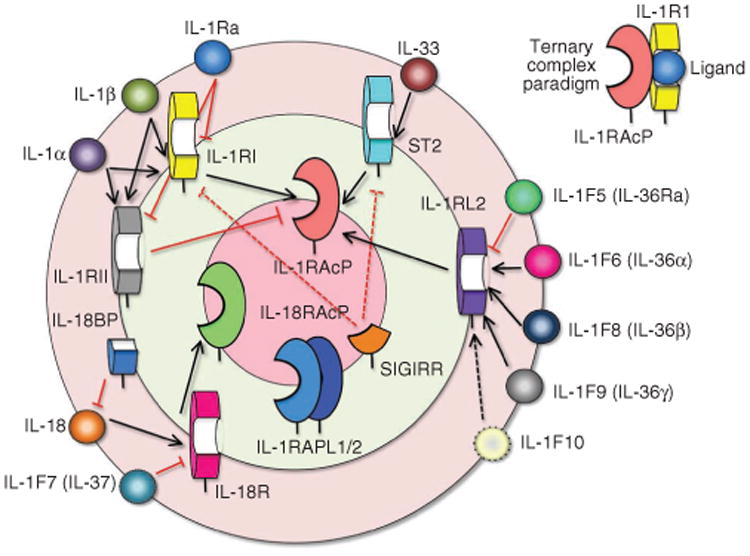
The IL-1 cytokine receptor family interaction wheel. The 11 IL-1 cytokines1,19 are drawn on the outer rim of the wheel and fall into four functional groups that correspond to different heterodimeric receptor signaling complexes. The five primary receptors are drawn on the middle wheel and the secondary or accessory receptors on the innermost wheel (for review, see ref. 20). To form the ternary complex paradigm (top right), the primary receptors first bind their corresponding cytokine ligands and then engage the accessory receptor (which is incapable of binding cytokines by itself). In the first binding event, IL-1 cytokine ligands connect to their (often promiscuous) primary receptors (shorter black arrows), which bind secondary receptors (longer black arrows) in the final step of ternary complex assembly. Red lines (solid for known interactions, dotted for predicted) mark decoy cytokine or receptor binding that creates nonfunctional partial receptor assemblies (as in the case of IL-1Ra, which binds IL-1RI or IL-1RII but does not engage IL-1RacP) or nonfunctional complete receptor assemblies (as with IL-1RII, which binds both cytokine and IL-1RacP but lacks an intracellular signaling domain). IL-1RAPL1/2, interleukin-1 receptor accessory protein–like 1/2; ST2, growth stimulation– expressed gene 2; SIGIRR, single immunoglobulin domain–containing IL-1R–related protein.
The structure of the heterotrimeric IL-1β–IL-1RI–IL-1RAcP complex presented here provides a general model for the signaling complex formed by the diverse IL-1 family of cytokines. Furthermore, it suggests a deep evolutionary relationship between the IL-1 and FGF receptor systems.
Supplementary Material
Acknowledgments
We thank the staff at the Advanced Light Source for their assistance. K.C.G. is an Investigator of the Howard Hughes Medical Institute and is supported by NIH grant R01-AI51321. C.T. was supported by a long-term postdoctoral fellowship from the International Human Frontier Science Program Organization.
Footnotes
Accession codes. Protein Data Bank: Coordinates and structure factors have been deposited under accession code 4DEP.
Note: Supplementary information is available on the Nature Structural & Molecular Biology website.
Author Contributions: K.C.G. conceived of the project, supervised the experiments and participated in writing the manuscript; J.F.B. contributed insights into the evolutionary relationships of the IL-1R activating complex to other receptors; C.T. executed the study and wrote the manuscript.
Competing Financial Interests: The authors declare no competing financial interests.
References
- 1.Sims JE, Smith DE. Nat Rev Immunol. 2010;10:89–102. doi: 10.1038/nri2691. [DOI] [PubMed] [Google Scholar]
- 2.Arend WP, Palmer G, Gabay C. Immunol Rev. 2008;223:20–38. doi: 10.1111/j.1600-065X.2008.00624.x. [DOI] [PubMed] [Google Scholar]
- 3.Wang D, et al. Nat Immunol. 2010;11:905–911. doi: 10.1038/ni.1925. [DOI] [PubMed] [Google Scholar]
- 4.Vigers GP, Anderson LJ, Caffes P, Brandhuber BJ. Nature. 1997;386:190–194. doi: 10.1038/386190a0. [DOI] [PubMed] [Google Scholar]
- 5.Murzin AG, Lesk AM, Chothia C. J Mol Biol. 1992;223:531–543. doi: 10.1016/0022-2836(92)90668-a. [DOI] [PubMed] [Google Scholar]
- 6.Schreuder H, et al. Nature. 1997;386:194–200. doi: 10.1038/386194a0. [DOI] [PubMed] [Google Scholar]
- 7.Venkataraman G, Raman R, Sasisekharan V, Sasisekharan R. Proc Natl Acad Sci USA. 1999;96:3658–3663. doi: 10.1073/pnas.96.7.3658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Schlessinger J, et al. Mol Cell. 2000;6:743–750. doi: 10.1016/s1097-2765(00)00073-3. [DOI] [PubMed] [Google Scholar]
- 9.Casadio R, et al. FEBS Lett. 2001;499:65–68. doi: 10.1016/s0014-5793(01)02515-7. [DOI] [PubMed] [Google Scholar]
- 10.Kiselyov VV, Kochoyan A, Poulsen FM, Bock E, Berezin V. Protein Sci. 2006;15:2318–2322. doi: 10.1110/ps.062206106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kalinina J, et al. Structure. 2012;20:77–88. doi: 10.1016/j.str.2011.10.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wass MN, David A, Sternberg MJ. Curr Opin Struct Biol. 2011;21:382–390. doi: 10.1016/j.sbi.2011.03.013. [DOI] [PubMed] [Google Scholar]
- 13.Ehrlich M, Horbelt D, Marom B, Knaus P, Henis YI. Cell Signal. 2011;23:1424–1432. doi: 10.1016/j.cellsig.2011.04.004. [DOI] [PubMed] [Google Scholar]
- 14.Wang X, Lupardus P, Laporte SL, Garcia KC. Annu Rev Immunol. 2009;27:29–60. doi: 10.1146/annurev.immunol.24.021605.090616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Thomas C, et al. Cell. 2011;146:621–632. doi: 10.1016/j.cell.2011.06.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chen H, et al. Proc Natl Acad Sci USA. 2008;105:19660–19665. doi: 10.1073/pnas.0807752105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Nyman T, et al. J Biol Chem. 2008;283:11861–11865. doi: 10.1074/jbc.C800001200. [DOI] [PubMed] [Google Scholar]
- 18.Ely LK, Fischer S, Garcia KC. Nat Immunol. 2009;10:1245–1251. doi: 10.1038/ni.1813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Dinarello C, et al. Nat Immunol. 2010;11:973. doi: 10.1038/ni1110-973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Boraschi D, Tagliabue A. Vitam Horm. 2006;74:229–254. doi: 10.1016/S0083-6729(06)74009-2. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


