Abstract
In the title compound, C30H31N3O2S, the fused pyrrolidine ring bearing three substituents adopts an envelope conformation with the C atom bearing the benzoyl group as the flap. The other fused pyrrolidine ring adopts a twisted conformation about one of its C—C bonds. The dihedral angle between the isatin ring system and the methylthiazole ring is 25.95 (8)°. An intramolecular C—H⋯O interaction closes an S(8) ring. In the crystal, molecules are linked by C—H⋯O interactions, generating C(11) chains propagating in [001].
Related literature
For general background to spiro compounds and their biological activity, see: Pradhan et al. (2006 ▶); Saeedi et al. (2010 ▶); Dandia et al. (2011 ▶). For uses of oxindole derivatives, see: Rajeswaran et al. (1999 ▶) and of pyrrolidine derivatives, see: Suzuki et al. (1994 ▶). For the biological activity of pyrrolidine derivatives, see: Cuzzocrea et al. (2002 ▶); Obniska et al. (2002 ▶); Amal Raj et al. (2003 ▶).
Experimental
Crystal data
C30H31N3O2S
M r = 497.64
Monoclinic,

a = 14.5718 (4) Å
b = 9.7218 (2) Å
c = 18.2609 (5) Å
β = 94.604 (1)°
V = 2578.57 (11) Å3
Z = 4
Mo Kα radiation
μ = 0.16 mm−1
T = 298 K
0.35 × 0.20 × 0.10 mm
Data collection
Bruker APEXII CCD diffractometer
Absorption correction: multi-scan (SADABS; Bruker, 2004 ▶) T min = 0.947, T max = 0.984
14973 measured reflections
4494 independent reflections
3647 reflections with I > 2σ(I)
R int = 0.017
Refinement
R[F 2 > 2σ(F 2)] = 0.039
wR(F 2) = 0.112
S = 1.06
4494 reflections
328 parameters
H-atom parameters constrained
Δρmax = 0.48 e Å−3
Δρmin = −0.27 e Å−3
Data collection: APEX2 (Bruker, 2004 ▶); cell refinement: SAINT-Plus (Bruker, 2004 ▶); data reduction: SAINT-Plus; program(s) used to solve structure: SHELXS2013 (Sheldrick, 2008 ▶); program(s) used to refine structure: SHELXL2013 (Sheldrick, 2008 ▶); molecular graphics: ORTEP-3 for Windows (Farrugia, 2012 ▶); software used to prepare material for publication: SHELXL2013.
Supplementary Material
Crystal structure: contains datablock(s) global, I. DOI: 10.1107/S1600536814006990/hb7204sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S1600536814006990/hb7204Isup2.hkl
CCDC reference: 994375
Additional supporting information: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| C21—H21⋯O2 | 0.93 | 2.43 | 3.191 (2) | 138 |
| C30—H30A⋯O2i | 0.96 | 2.57 | 3.533 (3) | 178 |
Symmetry code: (i)  .
.
Acknowledgments
The authors acknowledge the Department of Chemistry, IIT Madras, for the X-ray data collection and Orchid Chemicals and Pharmaceuticals, Chennai, India, for providing necessary laboratory facilities.
supplementary crystallographic information
1. Comment
Spirooxindoles are a class of oxindoles with a 3,3-spirocyclic junction and these compounds are extremely common in nature as part of natural products as well as many synthetic drugs (Pradhan et al., 2006); Oxindole derivatives help to treat and prevent diabetic complications arising from elevated levels of sorbitol, and act as aldose reductase inhibitors (Rajeswaran et al.,1999). Thus more and more novel spiroheterocycle compounds have been prepared and characterized (Saeedi et al., 2010); Dandia et al., 2011). In addition, the pyrrolidine group occurs in many families of biologically important compounds. Derivatives of pyrrolidine have anticonvulsant (Obniska et al., 2002), antimicrobial and antifungal activity against various pathogens, except Bacillus subtilis (Amal Raj et al., 2003). Pyrrolidine dithiocarbamate attenuates the development of acute and chronic inflammation (Cuzzocrea et al., 2002). Optically active pyrrolidine derivatives have been used as intermediates in controlled asymmetric synthesis (Suzuki et al., 1994). As spiro pyrrolidine compounds are of great medicinal properties, we have undertaken the three dimensional structure of the title compound. In view of these importance and continuation of our work on the crystal structure analysis of spiropyrrolidine derivatives, the crystal structure of the title compound has been carried out and the results are presented here.
The title compound, C30H31N3O2S, the pyrrolidine ring (N2/C9—C12)is twisted and the other pyrrolidine ring (N1/C1/C6—C8) is almost planar. In one of the pyrrolidine rings (N2/C9—C12) carbon atom C11 deviates by 0.537 Å from the plane. The dihedryl angle between the isatin ring and methylthiazol ring is 25.95 (8)°. The crystal structure features a C—H···O interaction which is connected along the c-axis forming a chain.
2. Experimental
Equimolar quantities of dimethyl acetophenone (0.02 mol) and methyl thiazole aldehyde (0.02 mol) were dissolved in 15 mL of ethanol, and aqueous NaOH (50% 12 mL) was added in dropwise. The reaction mixture was stirred at room temperature the pure compound precipitated from the solution filtered and dried. The dried thiazole aldehyde chalcone product (0.02 mol) reacted with of L-Proline (0.02 mol) and substituted isatins (0.02 mol) in ethanol solvent and the reaction mixture refluxed for two hours to form novel spiro compounds. The completion of the reaction monitored by TLC, the reaction mixture was cooled to room temperature and the solvent evaporated by vacuo the resulting reaction mass purified by column chromatography to isolate the pure compound. Colourless blocks of the title compound were obtained from ethanol solution by slow evaporation at room temperature.
3. Refinement
All hydrogen atoms were fixed geometrically and allowed to ride on the parent carbon atoms with aromatic C—H = 0.93 Å, methine C—H = 0.98 Å me thylene C—H = 0.97 Å and methyl C—H = 0.96 Å. The displacement parameters were set for phenyl H atoms at Uiso(H) = 1.2Ueq(C) and for methine,methylene and methyl H atoms at Uiso(H) =1.5Ueq(C).
Figures
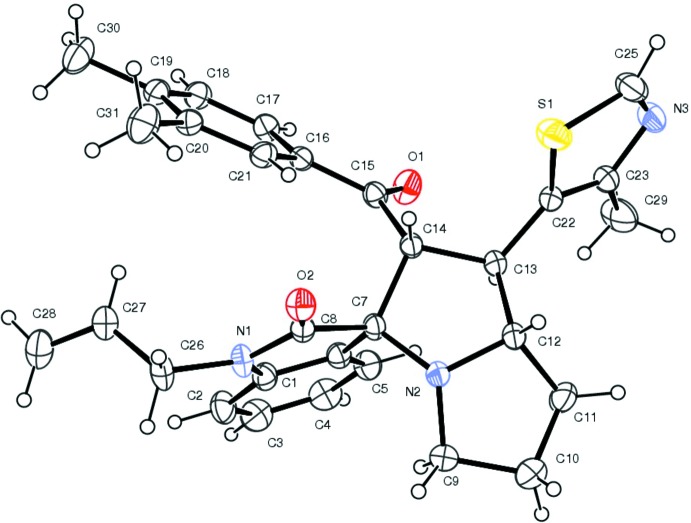
Fig. 1.
ORTEP of the molecule with atoms represented as 30% probability ellipsoids.
Fig. 2.

Packing diagram showing the C—H···O interaction along the C-axis
Crystal data
| C30H31N3O2S | F(000) = 1056 |
| Mr = 497.64 | Dx = 1.282 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| a = 14.5718 (4) Å | Cell parameters from 7315 reflections |
| b = 9.7218 (2) Å | θ = 2.5–28.3° |
| c = 18.2609 (5) Å | µ = 0.16 mm−1 |
| β = 94.604 (1)° | T = 298 K |
| V = 2578.57 (11) Å3 | Block, colourless |
| Z = 4 | 0.35 × 0.20 × 0.10 mm |
Data collection
| Bruker APEXII CCD diffractometer | 3647 reflections with I > 2σ(I) |
| phi and ω scans | Rint = 0.017 |
| Absorption correction: multi-scan (SADABS; Bruker, 2004) | θmax = 25.0°, θmin = 1.4° |
| Tmin = 0.947, Tmax = 0.984 | h = −17→17 |
| 14973 measured reflections | k = −11→10 |
| 4494 independent reflections | l = −17→21 |
Refinement
| Refinement on F2 | 0 restraints |
| Least-squares matrix: full | Hydrogen site location: inferred from neighbouring sites |
| R[F2 > 2σ(F2)] = 0.039 | H-atom parameters constrained |
| wR(F2) = 0.112 | w = 1/[σ2(Fo2) + (0.0482P)2 + 1.2211P], where P = (Fo2 + 2Fc2)/3 |
| S = 1.06 | (Δ/σ)max = 0.001 |
| 4494 reflections | Δρmax = 0.48 e Å−3 |
| 328 parameters | Δρmin = −0.27 e Å−3 |
Special details
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| C1 | 0.65413 (13) | 0.39883 (19) | 0.85262 (10) | 0.0383 (4) | |
| C2 | 0.58768 (14) | 0.4613 (2) | 0.80571 (11) | 0.0518 (5) | |
| H2 | 0.5986 | 0.5458 | 0.7841 | 0.062* | |
| C3 | 0.50416 (14) | 0.3940 (3) | 0.79186 (12) | 0.0581 (6) | |
| H3 | 0.4582 | 0.4339 | 0.7605 | 0.070* | |
| C4 | 0.48832 (14) | 0.2689 (2) | 0.82378 (12) | 0.0552 (6) | |
| H4 | 0.4323 | 0.2246 | 0.8132 | 0.066* | |
| C5 | 0.55540 (13) | 0.2081 (2) | 0.87182 (11) | 0.0461 (5) | |
| H5 | 0.5441 | 0.1241 | 0.8938 | 0.055* | |
| C6 | 0.63861 (12) | 0.27365 (18) | 0.88649 (10) | 0.0350 (4) | |
| C7 | 0.72224 (11) | 0.23936 (17) | 0.93863 (9) | 0.0318 (4) | |
| C8 | 0.78884 (12) | 0.35889 (17) | 0.92293 (9) | 0.0325 (4) | |
| C9 | 0.63441 (14) | 0.3191 (2) | 1.04689 (11) | 0.0466 (5) | |
| H9A | 0.5829 | 0.3301 | 1.0103 | 0.056* | |
| H9B | 0.6563 | 0.4093 | 1.0629 | 0.056* | |
| C10 | 0.60697 (16) | 0.2348 (2) | 1.11132 (13) | 0.0564 (6) | |
| H10A | 0.6448 | 0.2578 | 1.1558 | 0.068* | |
| H10B | 0.5428 | 0.2497 | 1.1196 | 0.068* | |
| C11 | 0.62314 (16) | 0.0887 (2) | 1.08836 (12) | 0.0547 (6) | |
| H11A | 0.5721 | 0.0547 | 1.0560 | 0.066* | |
| H11B | 0.6320 | 0.0285 | 1.1307 | 0.066* | |
| C12 | 0.71102 (13) | 0.09952 (18) | 1.04812 (10) | 0.0376 (4) | |
| H12 | 0.7650 | 0.0896 | 1.0833 | 0.045* | |
| C13 | 0.72035 (12) | 0.00419 (18) | 0.98251 (10) | 0.0345 (4) | |
| H13 | 0.6581 | −0.0167 | 0.9612 | 0.041* | |
| C14 | 0.76642 (12) | 0.09562 (17) | 0.92753 (9) | 0.0315 (4) | |
| H14 | 0.8324 | 0.1013 | 0.9427 | 0.038* | |
| C15 | 0.75471 (12) | 0.03912 (18) | 0.84903 (10) | 0.0351 (4) | |
| C16 | 0.81039 (12) | 0.09678 (18) | 0.79174 (9) | 0.0330 (4) | |
| C17 | 0.78684 (13) | 0.06349 (19) | 0.71827 (10) | 0.0393 (4) | |
| H17 | 0.7378 | 0.0045 | 0.7060 | 0.047* | |
| C18 | 0.83632 (14) | 0.1180 (2) | 0.66390 (10) | 0.0435 (5) | |
| H18 | 0.8206 | 0.0938 | 0.6152 | 0.052* | |
| C19 | 0.90914 (13) | 0.2083 (2) | 0.68006 (10) | 0.0407 (4) | |
| C20 | 0.93371 (12) | 0.24228 (19) | 0.75315 (10) | 0.0379 (4) | |
| C21 | 0.88487 (12) | 0.18377 (19) | 0.80783 (10) | 0.0362 (4) | |
| H21 | 0.9027 | 0.2036 | 0.8567 | 0.043* | |
| C22 | 0.76718 (13) | −0.12999 (18) | 1.00085 (10) | 0.0374 (4) | |
| C23 | 0.72805 (15) | −0.25530 (19) | 1.00826 (11) | 0.0456 (5) | |
| N3 | 0.78804 (15) | −0.36140 (18) | 1.02743 (10) | 0.0574 (5) | |
| C25 | 0.87117 (18) | −0.3172 (2) | 1.03383 (12) | 0.0587 (6) | |
| H25 | 0.9209 | −0.3750 | 1.0460 | 0.070* | |
| C26 | 0.78500 (16) | 0.5701 (2) | 0.84570 (11) | 0.0510 (5) | |
| H26A | 0.8461 | 0.5823 | 0.8701 | 0.061* | |
| H26B | 0.7483 | 0.6495 | 0.8569 | 0.061* | |
| C27 | 0.79208 (15) | 0.5635 (2) | 0.76434 (12) | 0.0545 (6) | |
| H27 | 0.8124 | 0.4820 | 0.7445 | 0.065* | |
| C28 | 0.77137 (17) | 0.6652 (3) | 0.72003 (13) | 0.0625 (6) | |
| H28A | 0.7508 | 0.7480 | 0.7384 | 0.075* | |
| H28B | 0.7771 | 0.6554 | 0.6699 | 0.075* | |
| C29 | 0.62774 (17) | −0.2887 (3) | 0.99541 (17) | 0.0746 (8) | |
| H29A | 0.6055 | −0.3229 | 1.0400 | 0.112* | |
| H29B | 0.6191 | −0.3575 | 0.9578 | 0.112* | |
| H29C | 0.5943 | −0.2072 | 0.9802 | 0.112* | |
| C30 | 0.96004 (16) | 0.2666 (3) | 0.61853 (12) | 0.0646 (7) | |
| H30A | 0.9340 | 0.2307 | 0.5725 | 0.097* | |
| H30B | 1.0238 | 0.2413 | 0.6255 | 0.097* | |
| H30C | 0.9547 | 0.3650 | 0.6183 | 0.097* | |
| C31 | 1.00940 (15) | 0.3431 (3) | 0.77401 (13) | 0.0595 (6) | |
| H31A | 1.0671 | 0.3062 | 0.7612 | 0.089* | |
| H31B | 1.0123 | 0.3596 | 0.8260 | 0.089* | |
| H31C | 0.9972 | 0.4281 | 0.7482 | 0.089* | |
| N1 | 0.74354 (10) | 0.44675 (15) | 0.87448 (8) | 0.0393 (4) | |
| N2 | 0.70847 (10) | 0.23905 (14) | 1.01728 (8) | 0.0353 (3) | |
| O1 | 0.69856 (10) | −0.05143 (15) | 0.83445 (8) | 0.0535 (4) | |
| O2 | 0.86754 (9) | 0.37134 (13) | 0.94943 (7) | 0.0409 (3) | |
| S1 | 0.88481 (4) | −0.14445 (6) | 1.01858 (3) | 0.05538 (18) |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| C1 | 0.0442 (10) | 0.0379 (10) | 0.0324 (10) | −0.0008 (8) | 0.0013 (8) | 0.0026 (8) |
| C2 | 0.0567 (12) | 0.0521 (13) | 0.0457 (12) | 0.0046 (10) | −0.0018 (10) | 0.0158 (10) |
| C3 | 0.0466 (12) | 0.0744 (16) | 0.0514 (13) | 0.0104 (11) | −0.0084 (10) | 0.0065 (12) |
| C4 | 0.0405 (11) | 0.0676 (15) | 0.0562 (14) | −0.0054 (10) | −0.0048 (10) | −0.0061 (11) |
| C5 | 0.0438 (10) | 0.0441 (11) | 0.0501 (12) | −0.0068 (9) | 0.0017 (9) | −0.0002 (9) |
| C6 | 0.0379 (9) | 0.0337 (10) | 0.0331 (10) | −0.0010 (8) | 0.0019 (7) | 0.0000 (7) |
| C7 | 0.0374 (9) | 0.0277 (9) | 0.0303 (9) | −0.0041 (7) | 0.0032 (7) | 0.0019 (7) |
| C8 | 0.0420 (10) | 0.0284 (9) | 0.0274 (9) | −0.0042 (7) | 0.0036 (7) | −0.0019 (7) |
| C9 | 0.0562 (12) | 0.0392 (11) | 0.0459 (12) | 0.0077 (9) | 0.0131 (9) | 0.0006 (9) |
| C10 | 0.0591 (13) | 0.0564 (14) | 0.0562 (14) | 0.0026 (11) | 0.0204 (11) | 0.0020 (11) |
| C11 | 0.0724 (14) | 0.0442 (12) | 0.0514 (13) | −0.0075 (10) | 0.0296 (11) | 0.0017 (10) |
| C12 | 0.0495 (10) | 0.0308 (9) | 0.0331 (10) | 0.0022 (8) | 0.0075 (8) | 0.0047 (8) |
| C13 | 0.0407 (9) | 0.0283 (9) | 0.0351 (10) | −0.0003 (7) | 0.0069 (8) | 0.0041 (7) |
| C14 | 0.0358 (9) | 0.0282 (9) | 0.0308 (9) | −0.0012 (7) | 0.0038 (7) | 0.0027 (7) |
| C15 | 0.0398 (9) | 0.0298 (9) | 0.0353 (10) | −0.0014 (8) | 0.0009 (8) | −0.0002 (7) |
| C16 | 0.0386 (9) | 0.0292 (9) | 0.0312 (9) | 0.0035 (7) | 0.0032 (7) | 0.0004 (7) |
| C17 | 0.0459 (10) | 0.0357 (10) | 0.0354 (10) | −0.0010 (8) | −0.0018 (8) | −0.0015 (8) |
| C18 | 0.0546 (11) | 0.0484 (12) | 0.0272 (9) | 0.0077 (9) | 0.0013 (8) | −0.0001 (8) |
| C19 | 0.0411 (10) | 0.0446 (11) | 0.0372 (10) | 0.0101 (9) | 0.0077 (8) | 0.0063 (8) |
| C20 | 0.0346 (9) | 0.0397 (10) | 0.0400 (11) | 0.0038 (8) | 0.0060 (8) | 0.0023 (8) |
| C21 | 0.0393 (9) | 0.0390 (10) | 0.0301 (9) | 0.0023 (8) | 0.0023 (7) | −0.0031 (8) |
| C22 | 0.0483 (10) | 0.0331 (10) | 0.0321 (10) | 0.0043 (8) | 0.0103 (8) | 0.0035 (7) |
| C23 | 0.0635 (13) | 0.0310 (10) | 0.0439 (11) | 0.0046 (9) | 0.0147 (10) | 0.0032 (8) |
| N3 | 0.0843 (14) | 0.0359 (10) | 0.0530 (11) | 0.0144 (9) | 0.0116 (10) | 0.0062 (8) |
| C25 | 0.0804 (17) | 0.0487 (13) | 0.0464 (13) | 0.0279 (12) | 0.0009 (12) | 0.0014 (10) |
| C26 | 0.0656 (13) | 0.0370 (11) | 0.0497 (12) | −0.0152 (10) | 0.0008 (10) | 0.0130 (9) |
| C27 | 0.0633 (13) | 0.0458 (13) | 0.0562 (14) | −0.0030 (10) | 0.0160 (11) | 0.0126 (10) |
| C28 | 0.0760 (16) | 0.0611 (15) | 0.0510 (14) | −0.0116 (12) | 0.0082 (12) | 0.0139 (11) |
| C29 | 0.0684 (16) | 0.0407 (13) | 0.117 (2) | −0.0092 (11) | 0.0203 (15) | 0.0028 (14) |
| C30 | 0.0615 (14) | 0.0898 (19) | 0.0442 (13) | −0.0014 (13) | 0.0154 (11) | 0.0134 (12) |
| C31 | 0.0477 (12) | 0.0712 (16) | 0.0601 (14) | −0.0150 (11) | 0.0069 (11) | 0.0050 (12) |
| N1 | 0.0468 (9) | 0.0334 (8) | 0.0368 (9) | −0.0089 (7) | −0.0016 (7) | 0.0093 (7) |
| N2 | 0.0459 (8) | 0.0285 (8) | 0.0323 (8) | 0.0006 (6) | 0.0078 (7) | 0.0018 (6) |
| O1 | 0.0681 (9) | 0.0484 (9) | 0.0446 (8) | −0.0245 (7) | 0.0081 (7) | −0.0083 (6) |
| O2 | 0.0408 (7) | 0.0399 (7) | 0.0413 (7) | −0.0082 (6) | −0.0012 (6) | −0.0010 (6) |
| S1 | 0.0504 (3) | 0.0509 (3) | 0.0646 (4) | 0.0101 (2) | 0.0029 (3) | 0.0022 (3) |
Geometric parameters (Å, º)
| C1—C2 | 1.381 (3) | C15—C16 | 1.484 (2) |
| C1—C6 | 1.392 (3) | C16—C21 | 1.389 (2) |
| C1—N1 | 1.411 (2) | C16—C17 | 1.396 (2) |
| C2—C3 | 1.387 (3) | C17—C18 | 1.379 (3) |
| C2—H2 | 0.9300 | C17—H17 | 0.9300 |
| C3—C4 | 1.376 (3) | C18—C19 | 1.390 (3) |
| C3—H3 | 0.9300 | C18—H18 | 0.9300 |
| C4—C5 | 1.391 (3) | C19—C20 | 1.394 (3) |
| C4—H4 | 0.9300 | C19—C30 | 1.506 (3) |
| C5—C6 | 1.377 (3) | C20—C21 | 1.393 (3) |
| C5—H5 | 0.9300 | C20—C31 | 1.502 (3) |
| C6—C7 | 1.522 (2) | C21—H21 | 0.9300 |
| C7—N2 | 1.466 (2) | C22—C23 | 1.357 (3) |
| C7—C8 | 1.555 (2) | C22—S1 | 1.7244 (19) |
| C7—C14 | 1.559 (2) | C23—N3 | 1.379 (3) |
| C8—O2 | 1.214 (2) | C23—C29 | 1.497 (3) |
| C8—N1 | 1.362 (2) | N3—C25 | 1.282 (3) |
| C9—N2 | 1.468 (2) | C25—S1 | 1.716 (2) |
| C9—C10 | 1.514 (3) | C25—H25 | 0.9300 |
| C9—H9A | 0.9700 | C26—N1 | 1.459 (2) |
| C9—H9B | 0.9700 | C26—C27 | 1.499 (3) |
| C10—C11 | 1.506 (3) | C26—H26A | 0.9700 |
| C10—H10A | 0.9700 | C26—H26B | 0.9700 |
| C10—H10B | 0.9700 | C27—C28 | 1.298 (3) |
| C11—C12 | 1.530 (3) | C27—H27 | 0.9300 |
| C11—H11A | 0.9700 | C28—H28A | 0.9300 |
| C11—H11B | 0.9700 | C28—H28B | 0.9300 |
| C12—N2 | 1.468 (2) | C29—H29A | 0.9600 |
| C12—C13 | 1.529 (3) | C29—H29B | 0.9600 |
| C12—H12 | 0.9800 | C29—H29C | 0.9600 |
| C13—C22 | 1.498 (2) | C30—H30A | 0.9600 |
| C13—C14 | 1.535 (2) | C30—H30B | 0.9600 |
| C13—H13 | 0.9800 | C30—H30C | 0.9600 |
| C14—C15 | 1.532 (2) | C31—H31A | 0.9600 |
| C14—H14 | 0.9800 | C31—H31B | 0.9600 |
| C15—O1 | 1.217 (2) | C31—H31C | 0.9600 |
| C2—C1—C6 | 121.97 (18) | C21—C16—C17 | 118.18 (16) |
| C2—C1—N1 | 128.02 (18) | C21—C16—C15 | 122.89 (16) |
| C6—C1—N1 | 110.01 (15) | C17—C16—C15 | 118.93 (16) |
| C1—C2—C3 | 117.8 (2) | C18—C17—C16 | 120.03 (18) |
| C1—C2—H2 | 121.1 | C18—C17—H17 | 120.0 |
| C3—C2—H2 | 121.1 | C16—C17—H17 | 120.0 |
| C4—C3—C2 | 120.95 (19) | C17—C18—C19 | 121.61 (18) |
| C4—C3—H3 | 119.5 | C17—C18—H18 | 119.2 |
| C2—C3—H3 | 119.5 | C19—C18—H18 | 119.2 |
| C3—C4—C5 | 120.6 (2) | C18—C19—C20 | 119.08 (17) |
| C3—C4—H4 | 119.7 | C18—C19—C30 | 119.51 (18) |
| C5—C4—H4 | 119.7 | C20—C19—C30 | 121.41 (19) |
| C6—C5—C4 | 119.3 (2) | C21—C20—C19 | 118.87 (17) |
| C6—C5—H5 | 120.4 | C21—C20—C31 | 119.46 (17) |
| C4—C5—H5 | 120.4 | C19—C20—C31 | 121.65 (17) |
| C5—C6—C1 | 119.34 (17) | C16—C21—C20 | 122.17 (17) |
| C5—C6—C7 | 131.98 (17) | C16—C21—H21 | 118.9 |
| C1—C6—C7 | 108.61 (15) | C20—C21—H21 | 118.9 |
| N2—C7—C6 | 116.59 (14) | C23—C22—C13 | 128.09 (18) |
| N2—C7—C8 | 108.57 (13) | C23—C22—S1 | 109.12 (14) |
| C6—C7—C8 | 101.61 (13) | C13—C22—S1 | 122.77 (14) |
| N2—C7—C14 | 102.47 (13) | C22—C23—N3 | 115.7 (2) |
| C6—C7—C14 | 115.69 (14) | C22—C23—C29 | 126.31 (19) |
| C8—C7—C14 | 112.06 (14) | N3—C23—C29 | 117.93 (19) |
| O2—C8—N1 | 126.11 (16) | C25—N3—C23 | 110.32 (19) |
| O2—C8—C7 | 125.75 (15) | N3—C25—S1 | 115.74 (17) |
| N1—C8—C7 | 108.14 (14) | N3—C25—H25 | 122.1 |
| N2—C9—C10 | 104.31 (16) | S1—C25—H25 | 122.1 |
| N2—C9—H9A | 110.9 | N1—C26—C27 | 112.65 (17) |
| C10—C9—H9A | 110.9 | N1—C26—H26A | 109.1 |
| N2—C9—H9B | 110.9 | C27—C26—H26A | 109.1 |
| C10—C9—H9B | 110.9 | N1—C26—H26B | 109.1 |
| H9A—C9—H9B | 108.9 | C27—C26—H26B | 109.1 |
| C11—C10—C9 | 103.73 (17) | H26A—C26—H26B | 107.8 |
| C11—C10—H10A | 111.0 | C28—C27—C26 | 123.7 (2) |
| C9—C10—H10A | 111.0 | C28—C27—H27 | 118.2 |
| C11—C10—H10B | 111.0 | C26—C27—H27 | 118.2 |
| C9—C10—H10B | 111.0 | C27—C28—H28A | 120.0 |
| H10A—C10—H10B | 109.0 | C27—C28—H28B | 120.0 |
| C10—C11—C12 | 102.96 (16) | H28A—C28—H28B | 120.0 |
| C10—C11—H11A | 111.2 | C23—C29—H29A | 109.5 |
| C12—C11—H11A | 111.2 | C23—C29—H29B | 109.5 |
| C10—C11—H11B | 111.2 | H29A—C29—H29B | 109.5 |
| C12—C11—H11B | 111.2 | C23—C29—H29C | 109.5 |
| H11A—C11—H11B | 109.1 | H29A—C29—H29C | 109.5 |
| N2—C12—C13 | 105.10 (14) | H29B—C29—H29C | 109.5 |
| N2—C12—C11 | 104.56 (15) | C19—C30—H30A | 109.5 |
| C13—C12—C11 | 117.70 (16) | C19—C30—H30B | 109.5 |
| N2—C12—H12 | 109.7 | H30A—C30—H30B | 109.5 |
| C13—C12—H12 | 109.7 | C19—C30—H30C | 109.5 |
| C11—C12—H12 | 109.7 | H30A—C30—H30C | 109.5 |
| C22—C13—C12 | 114.85 (15) | H30B—C30—H30C | 109.5 |
| C22—C13—C14 | 115.80 (14) | C20—C31—H31A | 109.5 |
| C12—C13—C14 | 103.44 (14) | C20—C31—H31B | 109.5 |
| C22—C13—H13 | 107.4 | H31A—C31—H31B | 109.5 |
| C12—C13—H13 | 107.4 | C20—C31—H31C | 109.5 |
| C14—C13—H13 | 107.4 | H31A—C31—H31C | 109.5 |
| C15—C14—C13 | 112.50 (14) | H31B—C31—H31C | 109.5 |
| C15—C14—C7 | 115.27 (14) | C8—N1—C1 | 111.42 (14) |
| C13—C14—C7 | 103.13 (13) | C8—N1—C26 | 123.80 (16) |
| C15—C14—H14 | 108.6 | C1—N1—C26 | 124.68 (15) |
| C13—C14—H14 | 108.6 | C7—N2—C12 | 112.11 (13) |
| C7—C14—H14 | 108.6 | C7—N2—C9 | 121.47 (15) |
| O1—C15—C16 | 120.99 (16) | C12—N2—C9 | 110.19 (14) |
| O1—C15—C14 | 119.13 (16) | C25—S1—C22 | 89.08 (11) |
| C16—C15—C14 | 119.87 (15) | ||
| C6—C1—C2—C3 | −1.1 (3) | C15—C16—C17—C18 | 178.67 (16) |
| N1—C1—C2—C3 | 179.66 (19) | C16—C17—C18—C19 | −1.1 (3) |
| C1—C2—C3—C4 | −0.1 (3) | C17—C18—C19—C20 | 1.3 (3) |
| C2—C3—C4—C5 | 1.1 (4) | C17—C18—C19—C30 | −179.07 (19) |
| C3—C4—C5—C6 | −0.8 (3) | C18—C19—C20—C21 | 0.5 (3) |
| C4—C5—C6—C1 | −0.4 (3) | C30—C19—C20—C21 | −179.18 (18) |
| C4—C5—C6—C7 | 175.96 (19) | C18—C19—C20—C31 | −177.71 (19) |
| C2—C1—C6—C5 | 1.4 (3) | C30—C19—C20—C31 | 2.6 (3) |
| N1—C1—C6—C5 | −179.25 (17) | C17—C16—C21—C20 | 2.6 (3) |
| C2—C1—C6—C7 | −175.73 (18) | C15—C16—C21—C20 | −176.85 (16) |
| N1—C1—C6—C7 | 3.6 (2) | C19—C20—C21—C16 | −2.5 (3) |
| C5—C6—C7—N2 | −63.4 (3) | C31—C20—C21—C16 | 175.78 (18) |
| C1—C6—C7—N2 | 113.23 (17) | C12—C13—C22—C23 | 101.5 (2) |
| C5—C6—C7—C8 | 178.8 (2) | C14—C13—C22—C23 | −137.95 (19) |
| C1—C6—C7—C8 | −4.58 (18) | C12—C13—C22—S1 | −76.69 (19) |
| C5—C6—C7—C14 | 57.1 (3) | C14—C13—C22—S1 | 43.9 (2) |
| C1—C6—C7—C14 | −126.20 (16) | C13—C22—C23—N3 | −178.37 (17) |
| N2—C7—C8—O2 | 60.6 (2) | S1—C22—C23—N3 | 0.0 (2) |
| C6—C7—C8—O2 | −175.91 (17) | C13—C22—C23—C29 | 3.7 (3) |
| C14—C7—C8—O2 | −51.8 (2) | S1—C22—C23—C29 | −178.0 (2) |
| N2—C7—C8—N1 | −119.33 (15) | C22—C23—N3—C25 | −0.5 (3) |
| C6—C7—C8—N1 | 4.11 (18) | C29—C23—N3—C25 | 177.7 (2) |
| C14—C7—C8—N1 | 128.21 (15) | C23—N3—C25—S1 | 0.8 (2) |
| N2—C9—C10—C11 | −32.1 (2) | N1—C26—C27—C28 | 136.2 (2) |
| C9—C10—C11—C12 | 37.9 (2) | O2—C8—N1—C1 | 177.76 (17) |
| C10—C11—C12—N2 | −29.4 (2) | C7—C8—N1—C1 | −2.3 (2) |
| C10—C11—C12—C13 | −145.56 (18) | O2—C8—N1—C26 | 1.3 (3) |
| N2—C12—C13—C22 | 152.65 (15) | C7—C8—N1—C26 | −178.70 (17) |
| C11—C12—C13—C22 | −91.5 (2) | C2—C1—N1—C8 | 178.46 (19) |
| N2—C12—C13—C14 | 25.51 (18) | C6—C1—N1—C8 | −0.8 (2) |
| C11—C12—C13—C14 | 141.35 (17) | C2—C1—N1—C26 | −5.1 (3) |
| C22—C13—C14—C15 | 72.1 (2) | C6—C1—N1—C26 | 175.59 (18) |
| C12—C13—C14—C15 | −161.31 (14) | C27—C26—N1—C8 | 115.9 (2) |
| C22—C13—C14—C7 | −163.02 (15) | C27—C26—N1—C1 | −60.1 (3) |
| C12—C13—C14—C7 | −36.48 (17) | C6—C7—N2—C12 | 108.89 (17) |
| N2—C7—C14—C15 | 156.57 (14) | C8—C7—N2—C12 | −137.19 (15) |
| C6—C7—C14—C15 | 28.6 (2) | C14—C7—N2—C12 | −18.49 (18) |
| C8—C7—C14—C15 | −87.22 (17) | C6—C7—N2—C9 | −24.4 (2) |
| N2—C7—C14—C13 | 33.57 (16) | C8—C7—N2—C9 | 89.54 (19) |
| C6—C7—C14—C13 | −94.39 (17) | C14—C7—N2—C9 | −151.77 (16) |
| C8—C7—C14—C13 | 149.78 (14) | C13—C12—N2—C7 | −4.19 (19) |
| C13—C14—C15—O1 | 13.8 (2) | C11—C12—N2—C7 | −128.77 (16) |
| C7—C14—C15—O1 | −104.05 (19) | C13—C12—N2—C9 | 134.38 (16) |
| C13—C14—C15—C16 | −167.26 (15) | C11—C12—N2—C9 | 9.8 (2) |
| C7—C14—C15—C16 | 74.9 (2) | C10—C9—N2—C7 | 147.77 (17) |
| O1—C15—C16—C21 | −169.49 (18) | C10—C9—N2—C12 | 13.7 (2) |
| C14—C15—C16—C21 | 11.6 (3) | N3—C25—S1—C22 | −0.64 (18) |
| O1—C15—C16—C17 | 11.0 (3) | C23—C22—S1—C25 | 0.32 (15) |
| C14—C15—C16—C17 | −167.86 (15) | C13—C22—S1—C25 | 178.81 (16) |
| C21—C16—C17—C18 | −0.8 (3) |
Hydrogen-bond geometry (Å, º)
| D—H···A | D—H | H···A | D···A | D—H···A |
| C21—H21···O2 | 0.93 | 2.43 | 3.191 (2) | 138 |
| C30—H30A···O2i | 0.96 | 2.57 | 3.533 (3) | 178 |
Symmetry code: (i) x, −y+1/2, z+1/2.
Footnotes
Supporting information for this paper is available from the IUCr electronic archives (Reference: HB7204).
References
- Amal Raj, A., Raghunathan, R., Sridevi Kumari, M. R. & Raman, N. (2003). Bioorg. Med. Chem. 11, 407–409. [DOI] [PubMed]
- Bruker (2004). APEX2, SAINT-Plus and SADABS Bruker AXS Inc., Madison, Wisconsin, USA.
- Cuzzocrea, S., Chatterjee, P. K., Mazzon, E., Dugo, L., Serraino, I., Britti, D., Mazzullo, G., Caputi, A. P. & Thiemermann, C. (2002). Br. J. Pharmacol. 135, 496–510. [DOI] [PMC free article] [PubMed] [Retracted]
- Dandia, A., Singh, R., Bhaskarana, S. & Samant, S. D. (2011). Green Chem. 13, 1852–1859.
- Farrugia, L. J. (2012). J. Appl. Cryst. 45, 849–854.
- Obniska, J., Zeic, A. & Zagorska, A. (2002). Acta Pol. Pharm. 59, 209–213. [PubMed]
- Pradhan, R., Patra, M., Behera, A. K. & Behera, R. K. (2006). Tetrahedron, 62, 779–828.
- Rajeswaran, W. G., Labroo, R. B. & Cohen, L. A. (1999). J. Org. Chem. 64, 1369–1371.
- Saeedi, M., Heravi, M. M., Beheshtiha, Y. S. & Oskooie, H. A. (2010). Tetrahedron, 66, 5345–5348.
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Suzuki, H., Aoyagi, S. & Kibayashi, C. (1994). Tetrahedron Lett. 35, 6119–6122.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablock(s) global, I. DOI: 10.1107/S1600536814006990/hb7204sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S1600536814006990/hb7204Isup2.hkl
CCDC reference: 994375
Additional supporting information: crystallographic information; 3D view; checkCIF report