Abstract
Genomic data now allow the large-scale manual or semi-automated reconstruction of metabolic networks. A network reconstruction represents a highly curated organism-specific knowledge base. A few genome-scale network reconstructions have appeared for metabolism in the baker’s yeast Saccharomyces cerevisiae. These alternative network reconstructions differ in scope and content, and further have used different terminologies to describe the same chemical entities, thus making comparisons between them difficult. The formulation of a ‘community consensus’ network that collects and formalizes the ‘community knowledge’ of yeast metabolism is thus highly desirable. We describe how we have produced a consensus metabolic network reconstruction for S. cerevisiae. Special emphasis is laid on referencing molecules to persistent databases or using database-independent forms such as SMILES or InChI strings, since this permits their chemical structure to be represented unambiguously and in a manner that permits automated reasoning. The reconstruction is readily available via a publicly accessible database and in the Systems Biology Markup Language, and we describe the manner in which it can be maintained as a community resource. It should serve as a common denominator for system biology studies of yeast. Similar strategies will be of benefit to communities studying genome-scale metabolic networks of other organisms.
Introduction
One of the goals of integrative systems biology is the accurate representation of biochemical, metabolic and signaling networks by mathematical models. This undertaking can be divided into four stages 1. The first is a qualitative stage in which are listed all the reactions that are known to occur in the system or organism of interest; in the modern era, and especially for metabolic networks, these reaction lists are often derived in part from genomic annotations 2, 3 with curation based on literature (‘bibliomic’) data 4. A second stage, again qualitative, adds known effectors, while the third and fourth stages – essentially amounting to molecular enzymology – include the known kinetic rate equations and the values of their parameters. Armed with such information it is then possible to provide a stochastic or ordinary differential equation model of the entire metabolic network of interest. An attractive feature of metabolism, for the purposes of modeling, is that, in contrast to signaling pathways, metabolism is subject to direct thermodynamic and (in particular) stoichiometric constraints 3. Our focus here is on the first two stages of the reconstruction process, especially as it pertains to the mapping of experimental metabolomics data onto metabolic network reconstructions.
Besides being an industrial workhorse for a variety of biotechnological products, baker’s yeast (Saccharomyces cerevisiae) is a highly developed model organism for biochemical, genetic, pharmacological and post-genomic studies (e.g. 5). It is especially attractive because of its sequenced genome 6, the availability of a whole series of bar-coded deletion 7, 8 and other 9 strains, extensive experimental omics data (e.g. 10–14), and the ability to grow it for extended periods in highly controlled conditions 15. The scientific community that works on S. cerevisiae is very active and has a history of collaborative research projects that have led to significant advances in our understanding of eukaryotic biology (e.g. 6, 8, 13, 16, 17). Furthermore, yeast metabolic physiology has been the subject of intensive study and most of the components of the yeast metabolic network are relatively well characterized. Taken together, these factors make yeast metabolism an attractive topic to test a community approach to model building in systems biology.
A number of groups (e.g. 18–21) have reconstructed the metabolic network of yeast from genomic and literature data and made the reconstructions freely available. However, due to different approaches utilized in the reconstruction as well as different interpretations of the literature, the existing reconstructions have many differences. Additionally, the naming of metabolites and enzymes in the existing reconstructions was, at best, inconsistent, and there were no systematic annotations of the chemical species in the form of links to external databases that store chemical compound information. This lack of model annotation complicated the use of the models for data analysis and integration. Members of the yeast systems biology community therefore recognized that a single ‘consensus’ reconstruction and annotation of the metabolic network was highly desirable as a starting point for further investigations.
A crucial factor that enabled the building of a consensus network reconstruction is the ability to describe and exchange biochemical network models in a standard format, the Systems Biology Markup Language (SBML; www.sbml.org) 22. The SBML format is utilized by most commonly used software applications for visualization, simulation, and analysis of biochemical networks, and also in pathway databases. SBML also provides the necessary standardized means (MIRIAM 23) to annotate models with information that is required to identify uniquely network components, including metabolites, proteins, and genes. Representing the consensus metabolic network reconstruction in a MIRIAM-compatible SBML format allows widespread use of the reconstruction as well as assisting in its continued curation, expansion and revision.
We developed this consensus reconstruction using a ‘jamboree’ approach, a large, focused work meeting, where we defined the protocol for the curation process as well as resolving the majority of discrepancies between the existing reconstructions. The jamboree event was followed by an extended process of curation of remaining discrepancies and careful annotation of components of the reconstructions by a smaller group of people. The overall goal of the effort was, by careful curation and comprehensive annotation of the network model and its components, to make the consensus reconstruction useful for the broadest possible set of users. The general reconstruction could then be used directly in bioinformatics applications aimed at integration of, for example, metabolomics and proteomics data or as a starting point for building predictive models using a number of different approaches (e.g. 24, 25), and for other purposes outlined below.
Here we describe how an initial ‘community consensus’ reconstruction of the yeast metabolic network was carried out. We make some further proposals for how this reconstruction of the yeast metabolic network may evolve as more information is acquired. We also discuss the possibility of using a similar approach to build consensus models of metabolic and other networks in other organisms.
Consensus reconstruction process
As a starting point for the development of a consensus reconstruction, we chose two separately developed freely available metabolic network reconstructions, iMM904 (http://gcrg.ucsd.edu/In_Silico_Organisms/Yeast) and iLL672 20, containing 904 and 672 yeast genes, respectively. Both of these reconstructions were derived from the first genome-scale metabolic network reconstruction for yeast iFF708 18 (for the basis of this terminology see 26), but the process of curating the original reconstruction was significantly different for the two derived reconstructions. The iMM904 reconstruction has eight different compartments and it was developed by curating and expanding an earlier reconstruction, iND750 19. In contrast, the iLL672 reconstruction 20 was directly derived from iFF708 by extensively curating the reconstruction in order to improve the ability of the flux balance model derived from the reconstruction to predict gene deletion phenotypes 27. It should be noted that yeast metabolic pathways in the KEGG 28 and SGD (see e.g. 29, 30) databases were used to establish the starting point for building the original iFF708 reconstruction and also for curating the iLL672 and iMM904 reconstructions. Hence, the information from early versions of these two reconstructions is included implicitly in the consensus reconstruction.
Due to the lack of common metabolite names and annotations, the comparison of the two starting-point reconstructions required first manually defining the correspondences between metabolites. After these had been assigned, the overall metabolite and reaction content of the two reconstructions could be compared (Table 1). The majority of metabolites (444) were found in both reconstructions, whereas 8 were only in iLL672 and 269 only in iMM904. In terms of reactions, 566 were in both reconstructions, 177 reactions were only found in iLL672, and 836 only in iMM904. The large number of additional reactions in iMM904 is mostly due to the expanded number of compartments represented in this reconstruction.
Table 1.
Comparison of starting point-reconstructions.
| iMM9 04 |
iLL672 | Common | iMM904 only |
iLL672 only |
|
|---|---|---|---|---|---|
| Metabolites | 713 | 452 | 444 | 269 | 8 |
| Reactions | 1402 | 743 | 566* | 836 | 177 |
| Genes | 904 | 659 | 646 | 258 | 13 |
| Compartments | 8 | 2 | 2 | 6 | 0 |
Reaction comparisons were done by considering every reaction to be reversible and without taking into account water and extracellular or intracellular protons (explicitly accounted for in iMM904).
The actual reconstruction jamboree was held at The University of Manchester, UK, in April 2007. The comparison between the iLL672 and iMM904 reconstructions shown in Table 1, proposed at a meeting of the YSBN (www.ysbn.eu/) in Helsinki, Finland in June 2006, formed the starting point for the reconstruction. The three-day Manchester event concentrated on three separate areas: (1) defining standards for curation as well as for representation of the annotated reconstruction in SBML, (2) annotating the metabolites with reference to external compound databases, and (3) resolving discrepancies between the reaction/metabolite sets in the two reconstructions. The presence of experts in relevant fields such as yeast genetics and physiology, systems modeling, metabolomics, standards (SBML/MIRIAM/metabolomics), and database/ontology development allowed the group to make significant progress in all three areas during the event. The annotation/curation was aided by a version of the B-Net database 31, and is provided as SBML (see Supplementary Table 1). After the jamboree, a sub-group of the authors verified the curation/annotation and resolved the remaining discrepancies between models. In the following we discuss in detail some of the major components of the curation and annotation processes.
Metabolite naming conventions
The initial comparison made it very clear that the naming conventions used in the two models were completely different, such that it was hard in some cases even for experts to know which chemical entities were meant. In addition, some of the reactions involved ‘generic’ structures (molecules with R-groups or so-called ‘Markush’ structures) which are not effectively represented in stoichiometric metabolic models, while some of the named entities represented ‘composite’ substances such as mixtures of different lipids or ‘biomass’. Without standardized names, it is extremely hard to enable computer software to reason about the similarities and differences between different models 32–37; this problem is even more significant in the case of reconstructions of the larger human metabolic network (e.g. 4, 38).
However, SBML allows one to annotate species such as metabolites with external references, and this has permitted us to relate them to molecules in ChEBI 39, KEGG 28, and PubChem 40, and to identify them definitively using database-independent representations of small molecules such as SMILES 41 and InChI 36, 42. We took advantage of this aspect of SBML to identify and annotate (manually) which chemical species were being described. In general the procedure we followed was to search these databases with the contents of the species’ name attribute field in the SBML representation or by the chemical formula of the compound sought. The order of annotation was such that we annotated metabolite species using ChEBI identifiers and InchI strings (if possible). If these did not exist or could not be resolved, then we used KEGG IDs (or in two cases HMDB identifiers 43) followed by PubChem IDs and finally PubMed references. This activity generated, for the first time, a representation that allows computational comparisons to be performed.
Because some individual molecules have multiple states (e.g. because of acid-base reactions), it would be desirable to use the chemical entities believed to be most common at the pH of the relevant compartment. However, in this version of the consensus reconstruction, all species are assumed to be in the form that corresponds to the most common protonation state at pH 7.2. Whenever possible, the metabolites were annotated with a database entry with the correct protonation state but, in a number of cases, the databases only contained the metabolite in a neutral form or otherwise in an incorrect or incorrectly annotated protonation state.
Annotation of large-scale metabolic models in SBML
While large-scale metabolic network reconstructions and models are now commonly represented in SBML, there has not thus far been a standard way to annotate these models. As part of the consensus reconstruction effort, we tried to develop such a standard that is compliant with MIRIAM 23. While the annotation of metabolites is quite straightforward, standardized annotation of the reaction content (molecules and reactions) of the reconstructed network proved to be more involved.
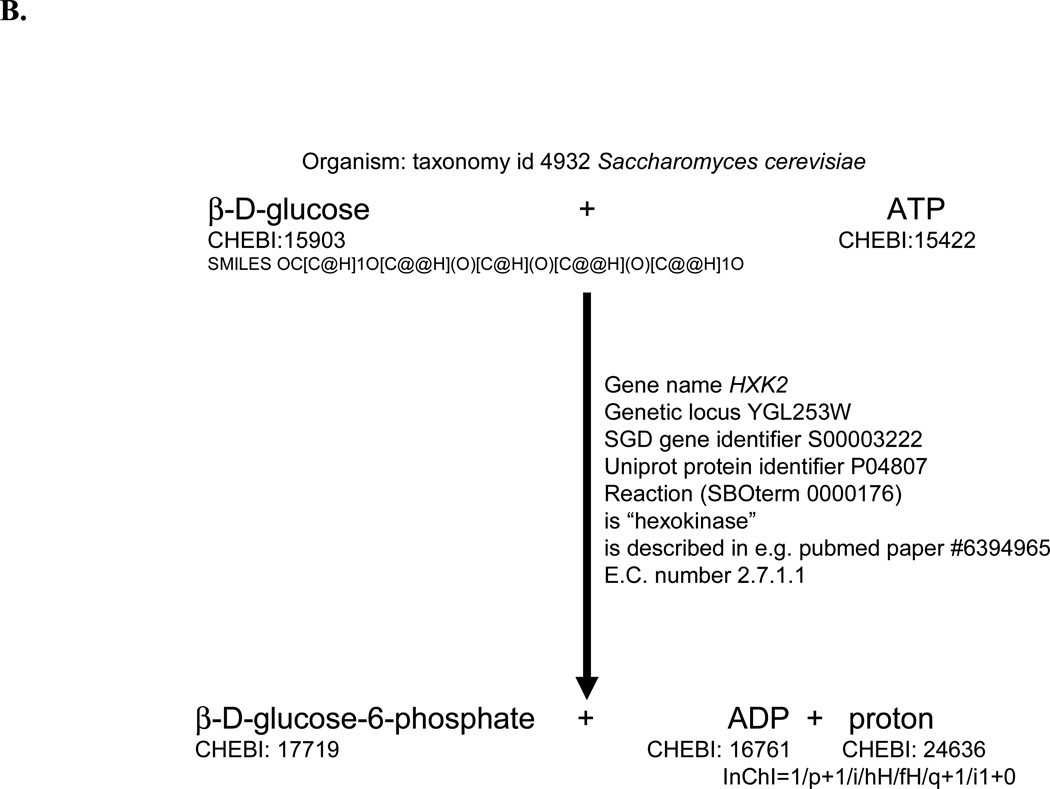
Where possible, we annotated reactions using literature references encoded as PubMed IDs, using the MIRIAM- and SBML-compliant "isDescribedBy" RDF annotation tag. In addition, reaction annotations include modifiers (enzymes/enzyme complexes) where possible. If a given reaction can be catalyzed by two or more isozymes, we generated an individual reaction for each isozyme (or complex). We represented the formation of protein complexes by separate reactions. Proteins and genes were finally annotated by references to SGD 29 and UniProt 44. In addition, we annotated cellular compartments using GO terms 45. In all cases where annotations were used, the MIRIAM 23 web services (http://www.ebi.ac.uk/compneur-srv/miriam-main/mdb?section=ws) were consulted to ensure correct annotation. Examples of fully annotated species and reaction entries are shown in Figure 2 and in Supplementary Figure 1.
Figure 2.
An example of the SBML annotation of a metabolite species using the example of ATP, as used in the reconstruction of the consensus network, illustrating its use of the Systems Biology Ontology (http://www.ebi.ac.uk/sbo/) and its MIRIAM-compliance. A. Relevant parts of the SBML code. B. An indication of the kinds of annotations included (for clarity not all are shown).
Contents of the consensus reconstructions
In all, the resulting consensus network consists of 2153 species (1168 metabolites, 832 genes, 888 proteins, and 96 catalytic protein complexes) and 1857 reactions (1761 metabolic reactions and 96 complex formation reactions). Reactions and species can be localized to 15 compartments (see Table 2) including membrane compartments. The network contains 664 distinct chemical entities (i.e. ATP is present in the nucleus, cytoplasm, Golgi, mitochondrion, peroxisome and vacuole and as such represents 6 metabolites but here we used just one chemical species, ignoring, e.g., Mg++ liganding). Of these distinct chemical entities, 554 are annotated with ChEBI identifiers, 564 with InChI identifiers, 78 with KEGG identifiers, 10 with PubChem identifiers, two with HMDB identifiers, and five only have PubMed references. In addition, 26 compounds are currently unannotated in this way – the majority of these are fatty acyl CoA’s or Acyl carrier proteins (ACPs) where the corresponding fatty acid is in public databases, but the fatty acyl CoA or ACP is currently not (but will be submitted to them).
Table 2.
Summary of the consensus reconstruction by compartment
| Compartment | Reactions | Metabolites |
|---|---|---|
| Cytoplasm | 835 | 590 |
| Extracellular | 15 | 158 |
| Golgi | 2 | 13 |
| Mitochondrion | 188 | 235 |
| Nucleus | 30 | 42 |
| Endoplasmic reticulum | 32 | 28 |
| Vacuole | 2 | 22 |
| Peroxisome | 77 | 80 |
| Mitochondrial membrane | 142 | 0 |
| Plasma membrane | 311 | 0 |
| Peroxisomal membrane | 44 | 0 |
| ER membrane | 17 | 0 |
| Vacuolar membrane | 35 | 0 |
| Golgi membrane | 5 | 0 |
| Nuclear membrane | 26 | 0 |
The network includes 1312 unique chemical transformations, of which 911 occur within a single compartment and the remaining 401 are transport reactions. The overall distribution of metabolites and reactions between the various compartments in the consensus network is given in Table 2. EC number and PubMed reference annotations are provided for 738 and 478 unique transformations in the network, respectively. Each reaction includes all of its cofactors (sometimes known as ‘currency metabolites’) such as ATP, NADH and CoA. In addition, and while we recognize that there is a certain arbitrariness about this, we have assigned pathway names for each reaction in the network.
We have removed various reactions from the initial networks, especially where they contained Markush structures or ambiguities. This has led to the under-representation of lipids, where there are many combinatorial issues (e.g. 46). We anticipate that lipid pathways will be added in the future, but “lipidomics” experiments will eventually be necessary to define the full complement of different lipid species present in S. cerevisiae. In a similar vein, composite items such as ‘biomass’ are excluded; while these are required for flux balance analysis (FBA), our purpose here is to provide the basic inventory of metabolites and network structure that can be used e.g. to compare the network with experimental metabolomics data. This inventory can then form the basis for setting up flux balance models using different assumptions required for setting up these kinds of model, e.g. assumptions on the biomass composition, reversibility of reactions and lumping of the reactions into fewer compartments.
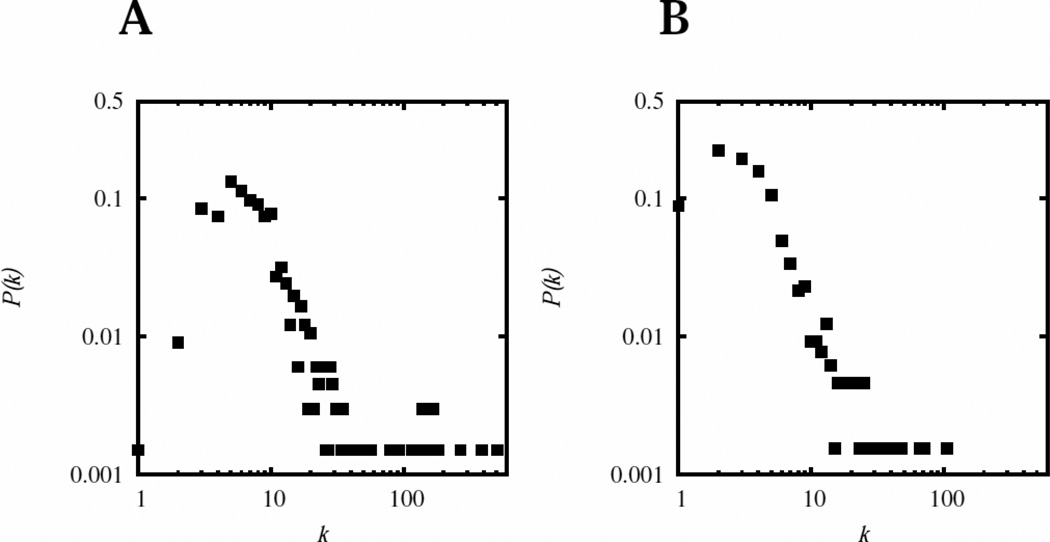
Figure 1 depicts the degree distribution (see e.g. 47) of the complete metabolite network, and a version where the currency metabolites were ignored as described earlier by Wagner and Fell 48. The complete network (Fig 1A) has an average clustering coefficient of 0.742, average node degree of 13.166, characteristic path length of 2.186, and betweenness centralization of 0.3897. The network without currency metabolites (Fig 1B) has an average clustering coefficient of 0.421, average node degree of 5.138, characteristic path length of 4.178, and betweenness centralization of 0.2329. In the full network the largest value for the shortest distance between any two metabolites (‘diameter’) is only 4 reaction steps while in the one without “currency” metabolites it is 11 reaction steps (between dTTP and Heme A). These statistics indicate that the “currency” metabolites should not be ignored as is sometimes done; without them the network is considerably less connected and several unconnected subnets appear, thus leaving some areas of metabolism unconnected from the rest. The center metabolite in the complete network is the proton, while in the smaller one it is coenzyme A. Table 3 lists the top 15 most-connected metabolites of each network.
Figure 1. Degree distribution of the metabolic network.
The metabolic reaction network was first summarized in a metabolite network, where metabolites are the nodes and one edge links two metabolites that co-occur in a reaction (in any role as substrates or products) as described in 48. For this analysis transport steps were not considered, nor were protein-protein binding reactions. The figures plot the distribution of the degree of connectivity, P(k), expressed as the fraction of metabolites that have k links out of the total number of metabolites plotted against the number of links (k) (A) in the complete network and, (B) in a network where the following metabolites were not considered: {water, proton, carbon dioxide, dioxygen, phosphate3−, diphosphate4−, ammonium, ATP, ADP, AMP, NAD+, NADH, NADP+, NADPH} (to be comparable with the analysis in 48).
Table 3.
Most connected nodes in the metabolite network. The degree is the number of metabolites that co-occur in metabolic reactions; the betweeness quantifies the number of paths between any two pairs of metabolites in the network that this one mediates (a global property)
| Complete metabolite network (as in Fig 1A) | Abbreviated metabolite network (as in Fig 1B) | ||||
|---|---|---|---|---|---|
| Metabolite | Degree | Betweeness | Metabolite | Degree | Betweeness |
| Proton | 506 | 0.391 | Coenzyme A | 106 | 0.237 |
| Water | 390 | 0.226 | L-glutamate1− | 71 | 0.232 |
| ATP | 268 | 0.099 | Acetyl-CoA | 66 | 0.065 |
| Diphosphate4− | 181 | 0.062 | 2-oxoglutarate2− | 48 | 0.078 |
| Phosphate3− | 178 | 0.041 | Hydrogen peroxide | 45 | 0.070 |
| ADP | 173 | 0.026 | Pyruvate | 38 | 0.070 |
| NADPH | 166 | 0.021 | Glycine | 31 | 0.041 |
| NADP+ | 166 | 0.021 | S-adenosyl-L-methionine | 28 | 0.063 |
| NAD+ | 143 | 0.018 | Acetate | 26 | 0.021 |
| Carbon dioxide | 139 | 0.020 | S-adenosyl-L-homocysteine | 25 | 0.039 |
| NADH | 139 | 0.017 | (6S)-5,6,7,8-tetrahydrofolic acid | 25 | 0.031 |
| AMP | 128 | 0.029 | L-glutamine | 25 | 0.025 |
| Coenzyme A | 119 | 0.021 | Succinate2− | 24 | 0.029 |
| Dioxygen | 116 | 0.014 | Acyl-carrier protein | 23 | 0.013 |
| Ammonium | 92 | 0.011 | L-cysteine | 22 | 0.023 |
Dissemination and future curation of the reconstruction
An SBML-encoded version of the base model (with and without compartments) is available at http://www.comp-sys-bio.org/yeastnet. Specifically, the SBML representation of the model is made available under the Creative Commons Attribution-Share Alike 3.0 Unported License (see www.creativecommons.org/). This is the preferred source for using the complete model with systems biology software. We have tested the SBML using various XML validators, and shown that it loads successfully into the COPASI biochemical software simulator 49. COPASI shows that there are a total of 307 mass conservation relations, which were calculated from the stoichiometry matrix using the method of Vallabhajosyula 50, which is now standard in COPASI 49. We have also loaded the model successfully into some versions of Cytoscape 51 and CellDesigner 52. The software is fully libSBML-compliant (see e.g. http://sbml.org/software/libsbml/ and 53, 54).
Recognizing that for many applications only subsets of this model are going to be relevant, we also make it available in an online database that facilitates searching the model. We used the database schema B-Net 31 which already supported all of the features required for our SBML model, including a structured mechanism for MIRIAM annotations. This B-Net representation of the model can be searched using synonyms and it also allows the user to navigate through the network, for example going from a metabolite to all its reactions, then to the genes that encode the enzymes catalyzing those reactions, etc. The database is also available at http://www.comp-sys-bio.org/yeastnet.
The B-Net database provides another important function as it is also the preferred means by which the community will be able to edit the model. It will thus be the primary source for the model. The advantages of using a database for managing data are well known and need not be reiterated here; suffice to say that, since there is no redundancy in the database, any change in any component immediately becomes global. For the time being, editing the model is limited to a few curators to ensure that the current standards are maintained. However, community annotation has major benefits 55, 56. Thus we also have included at the database a mechanism that collects annotations from anyone who wishes to communicate corrections or additions to the model. These annotations will then be reviewed and incorporated into the model for future releases of new versions.
Discussion
We have brought together a large segment of the community engaged in research involving genome-scale metabolic networks of yeast to create a consensus network that is freely available without restrictions and that can form the basis for future improvements. The SBML representation of the reconstruction is freely available under a Creative Commons License, and representations of the network were designed to facilitate future improvements.
Although annotation was semi-automated, a considerable element of manual annotation was still required, especially the parsing of the starting models. One of the biggest problems was the use of non-standard and often arcane synonyms for referring to the same chemical entity. A number of commentators have recognized the difficulties caused by issues of synonyms (e.g. 4, 33, 38). For these purposes, we believe and strongly recommend that the best solution to the synonym problem is to reference chemical entities in persistent databases and with database-independent representations such as SMILES 41 and InChI 42. Referencing the true chemical entity intended requires detailed consideration of its stereochemistry and the anomeric specificity of reactions in which it is involved, and not all databases have the required level of precision. We would also recommend that one should first build these networks in an assumption-free manner, and only then introduce (and annotate) extra features or assumptions that may be required for specific purposes – for instance adding composite compounds for flux balance analyses. A further benefit of the jamboree approach is the access to experts necessary to annotate details such as the precise gene-protein relationships underlying specific reactions.
The reconstruction presented here is currently the most comprehensive and consistent stoichiometric representation of yeast metabolism, from which predictive (sub)models, for example for genome-scale flux balance analysis, can be extracted and deployed. Presently, the reconstruction lacks information on effectors, reaction kinetics and parametrization. However, the basic framework of B-Net coupled to SBML models that can easily be populated with such data enables these to be added as they become available, and thus kinetic models that can be directly linked to the genome-scale metabolic network can be built. Some parameters are already available at SABIO-RK (http://sabio.villa-bosch.de/SABIORK/).
Network reconstruction approaches have developed rapidly in recent years. When they reach the genome-scale, they can be viewed as systems-level genome annotations 57. Genome-annotation is produced by a community-driven process to reach a consensus annotation that represents the state of knowledge about the genome of the target organism. Annotations are then updated based on new information and they serve as a common denominator for genome science studies of the target organism. The yeast metabolic reconstruction presented here represents an analogous process for systems biology studies of a target organism. With the successful achievement of the first consensus reconstruction the systems biology community can look forward to similar two-dimensional annotation jamborees for other organisms.
The metabolite nomenclature proposed here will hopefully become the standard terminology for metabolic models because the compounds themselves are essentially identical in all species. In the present case, we believe that the semantically annotated reconstruction we here provide will have especial utility in a number of areas. First is the basic exploration of metabolic pathways and well-curated connections between gene products. Further, the reconstruction will allow the automated interpretation and visualization of metabolomics data as well as data on metabolic proteins, genes and transcripts. The network can form the basis of phenotype predictions (including product yield) in response to genetic/environmental perturbations using a variety of methods (including FBA and logical approaches 58), and it can be used in metabolic flux estimation based on isotopomer data (e.g. 59), for filling gaps in metabolic pathways, and for exploring questions related to comparative metabolomics 60 and of metabolic pathway evolution. The widespread use of a consensus starting point will make both the comparison and the integration of such studies considerably easier.
Supplementary Material
Acknowledgments
The Manchester groups thank the UK BBSRC and EPSRC for financial support including for the Manchester Centre for Integrative Systems Biology (www.mcisb.org/). The UCSD participants thank NIH for financial support (NIH R01 GM071808). We thank Sarah Keating and Norman Paton for many useful discussions. The Jamboree was held under the auspices, and with the sponsorship of, the Yeast Systems Biology Network (EC Contract: LSHG-CT-2005-018942).
Footnotes
Author contributions. All authors conceived the idea of the consensus reconstruction, the majority were present during the jamboree itself, and all contributed to the writing of, and approved, the manuscript.
REFERENCES
- 1.Kell DB. Metabolomics, modelling and machine learning in systems biology: towards an understanding of the languages of cells. The 2005 Theodor Bücher lecture. FEBS J. 2006;273:873–894. doi: 10.1111/j.1742-4658.2006.05136.x. [DOI] [PubMed] [Google Scholar]
- 2.Arakawa K, Yamada Y, Shinoda K, Nakayama Y, Tomita M. GEM System: automatic prototyping of cell-wide metabolic pathway models from genomes. BMC Bioinformatics. 2006;7:168. doi: 10.1186/1471-2105-7-168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Palsson BØ. Systems biology: properties of reconstructed networks. Cambridge: Cambridge University Press; 2006. [Google Scholar]
- 4.Duarte NC, et al. Global reconstruction of the human metabolic network based on genomic and bibliomic data. Proc Natl Acad Sci. 2007;104:1777–1782. doi: 10.1073/pnas.0610772104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Mager WH, Winderickx J. Yeast as a model for medical and medicinal research. Trends Pharmacol Sci. 2005;26:265–273. doi: 10.1016/j.tips.2005.03.004. [DOI] [PubMed] [Google Scholar]
- 6.Goffeau A, et al. Life With 6000 Genes. Science. 1996;274:546–567. doi: 10.1126/science.274.5287.546. [DOI] [PubMed] [Google Scholar]
- 7.Winzeler EA, et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science. 1999;285:901–906. doi: 10.1126/science.285.5429.901. [DOI] [PubMed] [Google Scholar]
- 8.Giaever G, et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature. 2002;418:387–391. doi: 10.1038/nature00935. [DOI] [PubMed] [Google Scholar]
- 9.Yen K, Gitsham P, Wishart J, Oliver SG, Zhang N. An improved tetO promoter replacement system for regulating the expression of yeast genes. Yeast. 2003;20:1255–1262. doi: 10.1002/yea.1037. [DOI] [PubMed] [Google Scholar]
- 10.Hughes TR, et al. Functional discovery via a compendium of expression profiles. Cell. 2000;102:109–126. doi: 10.1016/s0092-8674(00)00015-5. [DOI] [PubMed] [Google Scholar]
- 11.Allen JK, et al. High-throughput characterisation of yeast mutants for functional genomics using metabolic footprinting. Nature Biotechnol. 2003;21:692–696. doi: 10.1038/nbt823. [DOI] [PubMed] [Google Scholar]
- 12.Zhu H, et al. Global analysis of protein activities using proteome chips. Science. 2001;293:2101–2105. doi: 10.1126/science.1062191. [DOI] [PubMed] [Google Scholar]
- 13.Castrillo JI, et al. Growth control of the eukaryote cell: a systems biology study in yeast. J. Biol. 2007;6:4. doi: 10.1186/jbiol54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Delneri D, et al. Identification and characterization of high-flux-control genes of yeast through competition analyses in continuous cultures. Nat Genet. 2008;40:113–117. doi: 10.1038/ng.2007.49. [DOI] [PubMed] [Google Scholar]
- 15.Wu J, Zhang N, Hayes A, Panoutsopoulou K, Oliver SG. Global analysis of nutrient control of gene expression in Saccharomyces cerevisiae during growth and starvation. Proc Natl Acad Sci U S A. 2004;101:3148–3153. doi: 10.1073/pnas.0308321100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Oliver SG. A network approach to the systematic analysis of gene function. Trends Genet. 1996;12:241–242. doi: 10.1016/0168-9525(96)30053-x. [DOI] [PubMed] [Google Scholar]
- 17.Suter B, Auerbach D, Stagljar I. Yeast-based functional genomics and proteomics technologies: the first 15 years and beyond. Biotechniques. 2006;40:625–644. doi: 10.2144/000112151. [DOI] [PubMed] [Google Scholar]
- 18.Förster J, Famili I, Fu P, Palsson BØ, Nielsen J. Genome-scale reconstruction of the Saccharomyces cerevisiae metabolic network. Genome Res. 2003;13:244–253. doi: 10.1101/gr.234503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Duarte NC, Herrgard MJ, Palsson BØ. Reconstruction and validation of Saccharomyces cerevisiae iND750, a fully compartmentalized genome-scale metabolic model. Genome Res. 2004;14:1298–1309. doi: 10.1101/gr.2250904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kuepfer L, Sauer U, Blank LM. Metabolic functions of duplicate genes in Saccharomyces cerevisiae. Genome Res. 2005;15:1421–1430. doi: 10.1101/gr.3992505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Caspi R, et al. MetaCyc: a multiorganism database of metabolic pathways and enzymes. Nucleic Acids Res. 2006;34:D511–D516. doi: 10.1093/nar/gkj128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hucka M, et al. The systems biology markup language (SBML): a medium for representation and exchange of biochemical network models. Bioinformatics. 2003;19:524–531. doi: 10.1093/bioinformatics/btg015. [DOI] [PubMed] [Google Scholar]
- 23.Le Novère N, et al. Minimum information requested in the annotation of biochemical models (MIRIAM) Nat Biotechnol. 2005;23:1509–1515. doi: 10.1038/nbt1156. [DOI] [PubMed] [Google Scholar]
- 24.Çakir T, et al. Integration of metabolome data with metabolic networks reveals reporter reactions. Mol Syst Biol. 2006;2:50. doi: 10.1038/msb4100085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kümmel A, Panke S, Heinemann M. Putative regulatory sites unraveled by network-embedded thermodynamic analysis of metabolome data. Mol Syst Biol. 2006;2 doi: 10.1038/msb4100074. 2006 0034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Reed JL, Vo TD, Schilling CH, Palsson BØ. An expanded genome-scale model of Escherichia coli K12 (iJR904 GSM/GPR) Genome Biol. 2003;4:R54. doi: 10.1186/gb-2003-4-9-r54. (open access at http://genomebiology.com/2003/2004/2009/R2054) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Förster J, Famili I, Palsson BØ, Nielsen J. Large-scale evaluation of in silico deletions in Saccharomyces cerevisiae. Omics. 2003;7:193–202. doi: 10.1089/153623103322246584. [DOI] [PubMed] [Google Scholar]
- 28.Kanehisa M, et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 2006;34:D354–D357. doi: 10.1093/nar/gkj102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Nash R, et al. Expanded protein information at SGD: new pages and proteome browser. Nucleic Acids Res. 2007;35:D468–D471. doi: 10.1093/nar/gkl931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Caspi R, et al. The MetaCyc Database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases. Nucleic Acids Res. 2008;36:D623–D631. doi: 10.1093/nar/gkm900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Li XJ, et al. In: Metabolic profiling: its role in biomarker discovery and gene function analysis. Harrigan GG, Goodacre R, editors. Boston: Kluwer Academic Publishers; 2003. pp. 293–309. [Google Scholar]
- 32.Goble C, Wroe C. The Montagues and the Capulets. Comp Func Genomics. 2004;5:623–632. doi: 10.1002/cfg.442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ananiadou S, Kell DB, Tsujii J-i. Text Mining and its potential applications in Systems Biology. Trends Biotechnol. 2006;24:571–579. doi: 10.1016/j.tibtech.2006.10.002. [DOI] [PubMed] [Google Scholar]
- 34.Poolman MG, Bonde BK, Gevorgyan A, Patel HH, Fell DA. Challenges to be faced in the reconstruction of metabolic networks from public databases. IEE Proc Syst Biol. 2006;153:379–384. doi: 10.1049/ip-syb:20060012. [DOI] [PubMed] [Google Scholar]
- 35.Spasić I, et al. Facilitating the development of controlled vocabularies for metabolomics with text mining. BMC Bioinformatics. 2008;9:S5. doi: 10.1186/1471-2105-9-S5-S5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Williams AJ. Internet-based tools for communication and collaboration in chemistry. Drug Discov Today. 2008;13:502–506. doi: 10.1016/j.drudis.2008.03.015. [DOI] [PubMed] [Google Scholar]
- 37.Williams AJ. A perspective of publicly accessible/open-access chemistry databases. Drug Discov Today. 2008;13:495–501. doi: 10.1016/j.drudis.2008.03.017. [DOI] [PubMed] [Google Scholar]
- 38.Ma H, et al. The Edinburgh human metabolic network reconstruction and its functional analysis. Mol Syst Biol. 2007;3:135. doi: 10.1038/msb4100177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Brooksbank C, Cameron G, Thornton J. The European Bioinformatics Institute's data resources: towards systems biology. Nucleic Acids Res. 2005;33:D46–D53. doi: 10.1093/nar/gki026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wheeler DL, et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2007;35:D5–D12. doi: 10.1093/nar/gkl1031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Weininger D. SMILES, a chemical language and information system.1. Introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 1988;28:31–36. [Google Scholar]
- 42.Coles SJ, Day NE, Murray-Rust P, Rzepa HS, Zhang Y. Enhancement of the chemical semantic web through the use of InChI identifiers. Organic & Biomolecular Chemistry. 2005;3:1832–1834. doi: 10.1039/b502828k. [DOI] [PubMed] [Google Scholar]
- 43.Wishart DS, et al. HMDB: the Human Metabolome Database. Nucleic Acids Res. 2007;35:D521–D526. doi: 10.1093/nar/gkl923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.The UniProt Consortium The universal protein resource (UniProt) Nucleic Acids Res. 2008;36:D190–D195. doi: 10.1093/nar/gkm895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Ashburner M, et al. Gene Ontology: tool for the unification of biology. Nature Genetics. 2000;25:25–29. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Sud M, et al. LMSD: LIPID MAPS structure database. Nucleic Acids Res. 2007;35:D527–D532. doi: 10.1093/nar/gkl838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Barabási A-L, Oltvai ZN. Network biology: understanding the cell's functional organization. Nat. Rev. Genet. 2004;5:101–113. doi: 10.1038/nrg1272. [DOI] [PubMed] [Google Scholar]
- 48.Wagner A, Fell DA. The small world inside large metabolic networks. Proc R Soc Lond B Biol Sci. 2001;268:1803–1810. doi: 10.1098/rspb.2001.1711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Hoops S, et al. COPASI: a COmplex PAthway SImulator. Bioinformatics. 2006;22:3067–3074. doi: 10.1093/bioinformatics/btl485. [DOI] [PubMed] [Google Scholar]
- 50.Vallabhajosyula RR, Chickarmane V, Sauro HM. Conservation analysis of large biochemical networks. Bioinformatics. 2006;22:346–353. doi: 10.1093/bioinformatics/bti800. [DOI] [PubMed] [Google Scholar]
- 51.Shannon P, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–2504. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Funahashi A, Tanimura N, Morohashi M, Kitano H. CellDesigner: a process diagram editor for gene-regulatory and biochemical networks. Biosilico. 2003;1:159–162. [Google Scholar]
- 53.Li P, Oinn T, Stoiland S, Kell DB. Automated manipulation of systems biology models using libSBML within Taverna workflows. Bioinformatics. 2008;24:287–289. doi: 10.1093/bioinformatics/btm578. [DOI] [PubMed] [Google Scholar]
- 54.Bornstein BJ, Keating SM, Jouraku A, Hucka M. LibSBML: an API library for SBML. Bioinformatics. 2008;24:880–881. doi: 10.1093/bioinformatics/btn051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Surowiecki J. The wisdom of crowds: why the many are smarter than the few. London: Abacus; 2004. [Google Scholar]
- 56.Tapscott D, Williams A. Wikinomics: how mass collaboration changes everything. New Paradigm; 2007. [Google Scholar]
- 57.Palsson B. Two-dimensional annotation of genomes. Nat Biotechnol. 2004;22:1218–1219. doi: 10.1038/nbt1004-1218. [DOI] [PubMed] [Google Scholar]
- 58.Whelan KE, King RD. Using a logical model to predict the growth of yeast. BMC Bioinformatics. 2008;9 doi: 10.1186/1471-2105-9-97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Blank LM, Kuepfer L, Sauer U. Large-scale 13C-flux analysis reveals mechanistic principles of metabolic network robustness to null mutations in yeast. Genome Biol. 2005;6:R49. doi: 10.1186/gb-2005-6-6-r49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Kell DB. Systems biology, metabolic modelling and metabolomics in drug discovery and development. Drug Disc Today. 2006;11:1085–1092. doi: 10.1016/j.drudis.2006.10.004. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.