Abstract
Background
Specialized interactions help structure communities, but persistence of specialized organisms is puzzling because a generalist can occupy more environments and partake in more beneficial interactions. The “Jack-of-all-trades is a master of none” hypothesis asserts that specialists persist because the fitness of a generalist utilizing a particular habitat is lower than that of a specialist adapted to that habitat. Yet, there are many reasons to expect that mutualists will generalize on partners.
Plant-soil feedbacks help to structure plant and microbial communities, but how frequently are soil-based symbiotic mutualistic interactions sufficiently specialized to influence species distributions and community composition? To address this question, we quantified realized partner richness and phylogenetic breadth of four wild-grown native legumes (Lupinus bicolor, L. arboreus, Acmispon strigosus and A. heermannii) and performed inoculation trials to test the ability of two hosts (L. bicolor and A. strigosus) to nodulate (fundamental partner richness), benefit from (response specificity), and provide benefit to (effect specificity) 31 Bradyrhizobium genotypes.
Results
In the wild, each Lupinus species hosted a broader genetic range of Bradyrhizobium than did either Acmispon species, suggesting that Acmispon species are more specialized. In the greenhouse, however, L. bicolor and A. strigosus did not differ in fundamental association specificity: all inoculated genotypes nodulated both hosts. Nevertheless, A. strigosus exhibited more specificity, i.e., greater variation in its response to, and effect on, Bradyrhizobium genotypes. Lupinus bicolor benefited from a broader range of genotypes but averaged less benefit from each. Both hosts obtained more fitness benefit from symbionts isolated from conspecific hosts; those symbionts in turn gained greater fitness benefit from hosts of the same species from which they were isolated.
Conclusions
This study affirmed two important tenets of evolutionary theory. First, as predicted by the Jack-of-all-trades is a master of none hypothesis, specialist A. strigosus obtained greater benefit from its beneficial symbionts than did generalist L. bicolor. Second, as predicted by coevolutionary theory, each test species performed better with partner genotypes isolated from conspecifics. Finally, positive fitness feedback between the tested hosts and symbionts suggests that positive plant-soil feedback could contribute to their patchy distributions in this system.
Keywords: Mutualism, Symbiosis, Specialization, Coevolution, Lupinus bicolor (dove lupine), Acmispon strigosus (strigose trefoil), Lotus, Alpha-proteobacteria, Legumes, Rhizobia
Background
Specialized biotic interactions contribute to processes that structure communities [1] and adapt populations to local partners and environments [2,3]. But, why do specialists exist if a generalist organism can occupy more environments and partake in more beneficial interactions [4,5]? The “Jack-of-all-trades is a master of none” hypothesis asserts that specialists persist in a heterogeneous environment because the fitness of a generalist utilizing a particular habitat is lower than that of a specialist adapted to that habitat [5-7]. This hypothesis is based on the assumptions that adapting to new habitats involves fitness costs and that traits adaptive in one subset of environments are negatively genetically correlated with traits adaptive in other environments [4,8-11], leading to local adaptation [2,3]. When the habitat is a host or interaction partner, this trade-off could lead to co-adaptation [12].
Fitness feedbacks between plants and specialized soil-based microbes significantly affect plant productivity, community composition, and the distribution and abundance of plants [13-21]. Strong plant-soil feedbacks are often negative; due to antagonistic interactions between plants and pathogens [13,21-27]. However, many plants engage in mutualistic interactions with soil-borne symbionts [6]. Is there sufficient specialization among plant hosts to soil-borne mutualistic symbionts to impact plant community structure and diversity via positive feedbacks?
Multiple arguments predict that mutualists interacting with partners acquired from the environment are likely to be generalists [28]. Since mutualists have higher fitness when partnered than when not, any partner should be better than no partner. If so, then selection should disfavor specialized mutualists (i.e., those with reduced interaction breadth relative to the breadth of available partners) that resist associating with common partners [29]. Similarly, specialized mutualistic lineages should be vulnerable to perturbations in partner populations [30-36] and the geographic distribution and spread of specialized mutualists could be limited by partner availability [37-44]. Generalized mutualists might also access a wider range of environmental conditions by utilizing partners with differing ecological tolerances [45-48]. Finally, evolutionary convergence of traits among mutualists within an interaction group [49-51], but see [52], could reduce variance in partner benefit and thereby weaken the effectiveness of selection to specialize [53,54].
Specialized mutualists nonetheless exist, e.g. [55-57]. Indeed, meta-analysis of bipartite interactions found that mutualistic webs are more specialized than antagonistic ones [1]. Further, mutualism theory predicts that, when available partners vary in quality, selection favors mechanisms by which mutualists can select partners [58-63], which can lead to specialization. However, if more-beneficial partners are rare or distributed unpredictably, then selection for specialization via partner choice might be weak [45,64,65]. How frequently symbiotic mutualistic interactions are specialized is therefore an open question.
Here, we examined specialization in a group of wild legumes that interact with Bradyrhizobium genotypes in coastal sand dunes of Sonoma County, California. Bradyrhizobium is a genus of rhizobia that can infect legume roots and fix nitrogen endosymbiotically within root nodules in an interaction that is generally mutualistic [66,67]. Dominant genes that either restrict nodulation with or alter the effectiveness of particular rhizobial genotypes have been described in both natural and managed plant populations [57,68], but the effect of such genes on fitness feedbacks has been little studied. Moreover, little is known about whether legume-rhizobium interactions are sufficiently specialized to contribute to structuring communities via plant-soil feedbacks or coevolution.
We quantified the realized richness and phylogenetic breadth of rhizobial symbionts collected from plants growing in situ, and measured the frequency with which each host species associated with each of the identified Bradyrhizobium genotypes. Association frequencies quantify the interaction [69] or link [70] strength of each pairwise interaction and were used to calculate the Paired Differences Index PDI [71], for each host. This index summarizes variation in a partner’s link strengths [69,70] with all available partners without making any assumptions about their statistical distribution.
Realized association specificity [72,73] is strongly influenced by the local sampling environment [74]. Therefore, we experimentally paired partners under controlled greenhouse conditions to examine fundamental association frequency [4,75] of two host species. We also used the greenhouse experiment to quantify the fitness effect of each host on the inoculated Bradyrhizobium genotypes, which is a measure of the host’s potential impact on the rhizobium population and therefore its functional role within the community [76].
Finally, we used these data to test whether this system supports the “Jack-of-all-trades is a master of none” hypothesis, as well as its underlying assumption that, as frequently interacting partners become co-adapted, they become less adapted to other partners and consequently exchange weaker benefits.
Results
Isolate collection and sequence datasets
We isolated Bradyrhizobium bacteria from nodules of four native California legumes growing on the Bodega Dunes at Bodega Marine Reserve and Sonoma Coast State Park, Sonoma County. Two species, Acmispon strigosus and Lupinus bicolor, are fast-growing annuals that are patchily distributed across the Bodega Dunes. In contrast, A. heermannii is a decumbent suffrutescent perennial and L. arboreus is a large upright perennial shrub, both widely distributed across the dunes.
Two DNA regions (NifD and ITS) were PCR amplified and sequenced from two to eight isolates from each plant, which yielded 84 sequenced isolates, including 81 NifD amplicons and 82 ITS amplicons (Genbank accession numbers in Additional file 1: Table S1). Sequences that differed by one nucleotide were considered distinct, resulting in 22 ITS genotypes, of which 13 occurred in multiple nodules, and 22 NifD genotypes, of which 13 occurred in multiple nodules. There were 39 unique genotype combinations.
Realized in situ association specificity (realized niche breadth)
The observed PDIs of the four hosts (L. arboreus: 0.264, A. strigosus: 0.301, A. heermannii: 0.395, L. bicolor: 0.433) all differed significantly from the expected joint PDI (0.069 ± 95% confidence limit of 0.002) and from each other, which indicates that each of the four hosts specialized on a subset of the available NifD types, but differed in degree of specialization; L. bicolor was most specialized.
Genetic breadth of symbiotic partners
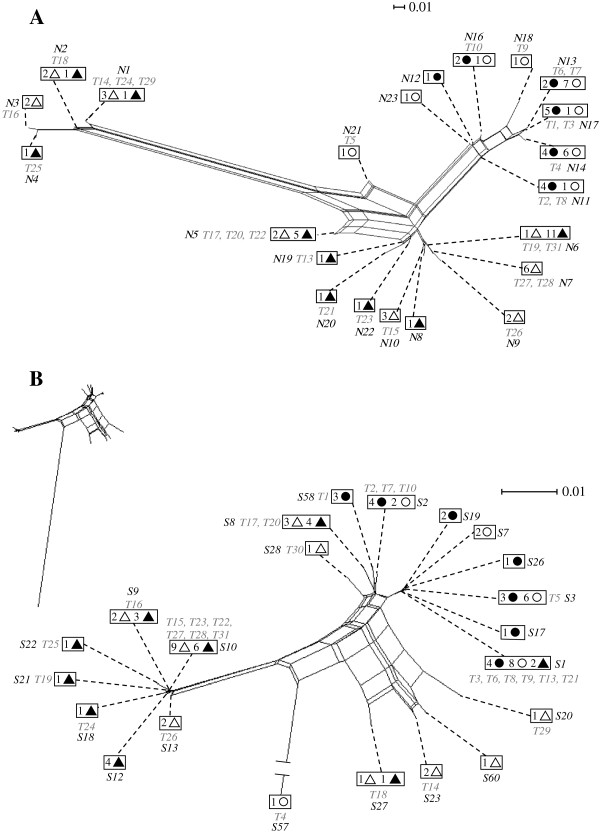
In both gene networks, Lupinus hosts associated with a genetically broader range of bacteria than did Acmispon hosts. Isolates from within a host genus tended to cluster together (Figure 1) and exhibited little overlap between host genera. Host genera shared no genotypes in the NifD network (Figure 1A). A single common ITS genotype (S1) was shared between host genera and one genotype from Lupinus (S8) clustered closely to most genotypes from Acmispon (Figure 1B).
Figure 1.
Genotype networks generated by the neighbor-net algorithm. (A)NifD locus, based on the distance calculated by MODELTEST, using a GTR + I model of sequence evolution [121]. (B)ITS region, based on the distance calculated by MODELTEST using a HKY + I + G model of sequence evolution [122]. Dotted lines represent the position of a genotype on the network. Each box represents one genotype, each symbol represents a host species (filled circles represent bacteria harvested from nodules of A. strigosus; open circles, A. heermannii; filled triangles, L. bicolor; open triangles, L. arboreus), and numbers indicate the number of strains representing a particular genotype; labels T1 to T31 in italics represent the Bradyrhizobium genotypes used in the greenhouse experiment (as listed in Additional file 1: Table S1). One genotype, T30, appears only in the ITS network, because we were unable to sequence its NifD locus. Genotypes T11 and T12 do not figure in either network because they were addenda from another study, used for reference, but not part of the original sample.
Genetic distance matrices among bacterial communities
Bacterial populations differed significantly between the two host genera but not between species within a genus. For both DNA regions, significantly large Φst values and corrected between-population mismatch rates occurred between Bradyrhizobium populations isolated from different host genera, but not between populations isolated from different host species within a genus (Table 1). However, bacteria isolated from Lupinus were more genetically variable than those isolated from Acmispon. For both DNA regions, the average pairwise sequence divergences between Bradyrhizobium genotypes from Lupinus were two- to five-fold greater than they were between genotypes from Acmispon (Table 1).
Table 1.
Analyses of population structure of Bradyrhizobium sampled from in situ nodules
| (a) ITS region | ||||
|---|---|---|---|---|
| |
A. strigosus |
A. heermannii |
L. arboreus |
L. bicolor |
|
A. strigosus |
6.6601 |
0.0422ns |
0.5814* |
0.4405* |
|
A. heermannii |
0.4506ns |
13.1696 |
0.5459* |
0.4126* |
|
L. arboreus |
24.6912* |
23.4825* |
26.2134 |
0.0058ns |
|
L. bicolor |
18.1281* |
17.9398* |
0.0078ns |
35.922 |
|
(b) nifD locus | ||||
| |
A. strigosus |
A. heermannii |
L. arboreus |
L. bicolor |
|
A. strigosus |
5.1895 |
-0.0040ns |
0.5649* |
0.6024* |
|
A. heermannii |
-0.0217ns |
7.4971 |
0.5830* |
0.6223* |
|
L. arboreus |
21.6199* |
22.0285* |
25.1368 |
0.0406ns |
| L. bicolor | 19.3541* | 19.6343* | 1.0162ns | 17.0395 |
Significance, obtained from 1023 permutations: ns, not significant; *, p < 0.0001; significant values in boldface.
Sequence divergence (along diagonal), corrected between-population differences (below diagonal), and between-population Φst (above diagonal).
Distribution of bacterial genetic variance
In a two-level hierarchical AMOVA (Table 2), neither ITS nor NifD sequence explained a significant component of variance between host species within genus. Most genetic variance occurred among plants within species (50.29% for ITS and 40.84% for NifD) and between host genera (49.14% for ITS and 57.92% for NifD). Permutations suggest that the variance component due to host genus was not significant (Table 2); even though genera harbored distinct genotypes in both networks (Figure 1) and all inter-genus values of pairwise genetic differences and Φst were statistically significant for both DNA regions (Table 1). This discrepancy probably arises from a lack of power in the AMOVA, which treats genus as a random effect, even though it is associated with only one degree of freedom.
Table 2.
AMOVA of (a) ITS and (b) NifD sequences from Bradyrhizobium from in situ nodules
| (a) ITS region | |||||
|---|---|---|---|---|---|
| Sources of variation |
df |
Sum of squares |
variance component |
% of variance |
P |
| Between genera |
1 |
440.898 |
10.5300 |
49.14 |
ns |
| Between species w/in genus |
2 |
26.594 |
0.1229 |
0.57 |
ns |
| Within species |
78 |
840.667 |
10.7778 |
50.29 |
*** |
| Total |
81 |
1308.159 |
21.4308 |
|
|
|
(b) NifD locus | |||||
| Sources of variation |
df |
Sum of squares |
variance component |
% of variance |
P |
| Between genera |
1 |
290.41 |
10.0195 |
57.92 |
ns |
| Between species w/in genus |
2 |
20.031 |
0.2149 |
1.24 |
ns |
| Within species |
77 |
416.246 |
7.06457 |
40.84 |
*** |
| Total | 80 | 980.938 | 17.2989 | ||
Significance: ns, not significant; ***, p < 0.001.
Sampled host species: A. strigosus, A. heermannii, L. bicolor and L. arboreus.
Fundamental association specificity (fundamental niche breadth)
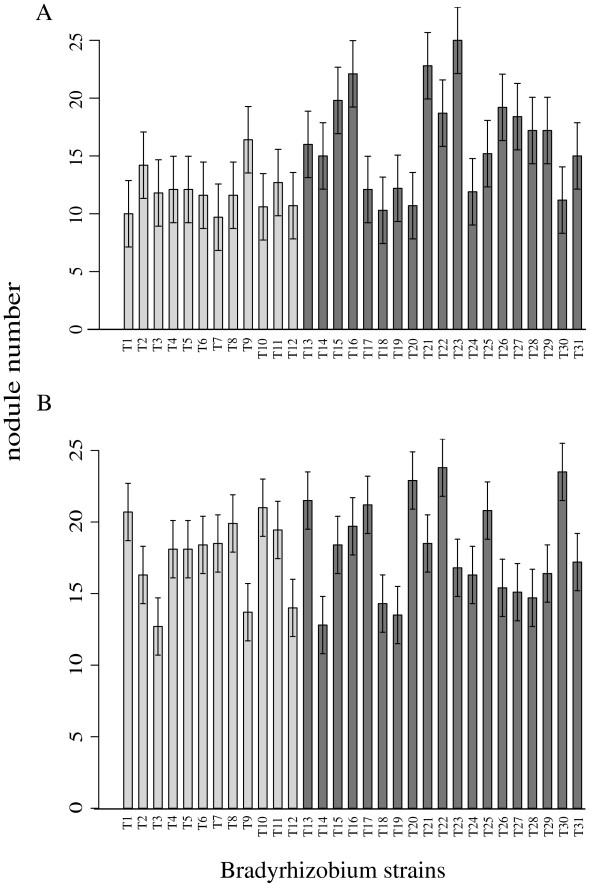
In the greenhouse inoculation experiment, neither test host exhibited fundamental association specificity; each could nodulate all tested Bradyrhizobium genotypes (Figures 2 and 3).
Figure 2.
Average number of nodules produced by each Bradyrhizobium genotype on each host species. Untransformed data. A)A. strigosus. B)L. bicolor. Error bars represent ± 1 standard error. Solid grey bars represent bacteria originally isolated from A. strigosus and A. heermannii nodules (A bacterial group); solid black bars represent bacteria harvested from L. bicolor and L. arboreus nodules (L bacterial group).
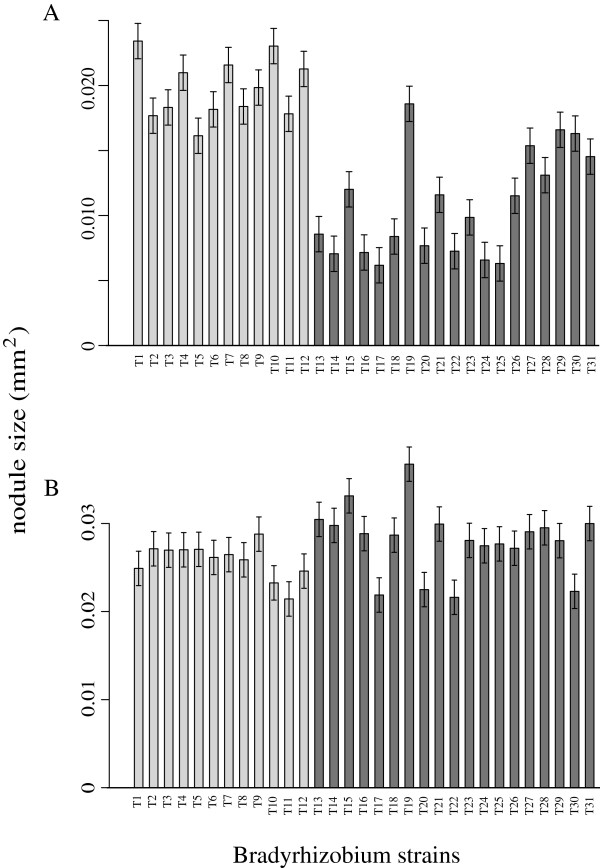
Figure 3.
Average nodule size produced by each Bradyrhizobium genotype on each host species. Untransformed data (cm2). A)A. strigosus. B)L. bicolor. Error bars represent ± 1 standard error. Solid grey bars represent bacteria originally isolated from A. strigosus and A. heermannii nodules (A bacterial group); solid black bars represent bacteria harvested from L. bicolor and L. arboreus nodules (L bacterial group).
G × G interaction (Response specificity)
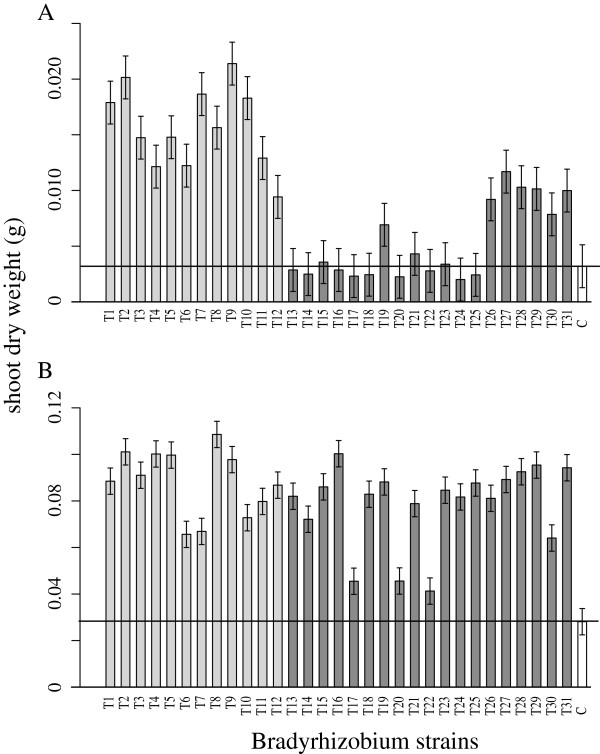
On average, inoculation increased host shoot dry weight over that of uninoculated control plants. The average increase was 2.8-fold for A. strigosus (F1, 318 = 64.261, p value < 0.0001) and 2.9-fold for L. bicolor (F1, 314 = 387.191, p value < 0.0001). However, bacterial genotypes differed significantly in their effects on shoot dry weight (Figure 4, Additional file 2: Table S2 and Additional file 3: Table S3). Notably, several genotypes failed to improve host growth beyond that of uninoculated controls (17 for A. strigosus and three for L. bicolor, using the Tukey-Kramer honestly significant difference test; 13 for A. strigosus and two for L. bicolor, using the less conservative Student’s t-test).
Figure 4.
Mean shoot dry weight of each host species when inoculated by each Bradyrhizobiumgenotype. Untransformed data (g). (A) A. strigosus. (B) L. bicolor. Error bars represent ± 1 standard error. Horizontal reference lines represent mean dry weight of uninoculated control plants. Light grey bars represent bacterial group A, originally isolated from A. strigosus and A. heermannii nodules; dark grey bars represent bacterial group L, originally isolated from L. bicolor and L. arboreus nodules; solid white bars represent uninoculated control plants. Genotype order recapitulates that found in Figures 2 and 3.
The conservative analysis presented above does not test for between-host differences in response specificity, which is typically detected as a significant interaction between test host species and Bradyrhizobium genotype G × G interaction [77], in an ANOVA including data from both host species. In a less conservative analysis, host by genotype (G × G) interactions were prominent in the best fitting model (model 16 in Additional file 4: Table S4). A significant G × G interaction of test host with Bradyrhizobium groups isolated from different host species (test host by species of origin interaction, F3,25 = 19.03, p > F < 0.0001; Additional file 5: Table S5), occurs because variation in response to different Bradyrhizobium genotypes was large in A. strigosus and almost absent in L. bicolor (Additional file 6: Table S6). Further, the G × G interaction due to test host species by Bradyrhizobium genotype nested within species of origin (covariance parameter estimate ± 1 S. E. = 0.07235 ± 0.024; Additional file 7: Table S7) accounted for 7.24% ± 2.28% of the variance in plant dry weight. This significant interaction indicates that even within a Bradyrhizobium group isolated from the same host species, genotypes differed in the symbiotic benefits they provided to the two different test host species.
As expected from the conservative analysis, test host species differed significantly in dry weight (least square means [logits] of shoot dry weight ± 1 S. E. for A. strigosus = -4.80 ± 0.10, L. bicolor = -2.40 ± 0.11; t9 = -18.36, p < 0.0001) and groups of Bradyrhizobium genotypes isolated from different host species differed significantly in average effect on test host shoot dry weight (species of origin effect F3,25 = 24.56, p > F < 0.0001; Additional file 8: Table S8). In particular, shoot dry weight produced by genotypes isolated from L. bicolor differed from that produced by genotypes isolated from other species (Additional file 6: Table S6 and Additional file 8: Table S8). When averaged across the test hosts, there was no significant variance in shoot dry weight due to inoculation with different Bradyrhizobium genotypes isolated from the same host species (strain(sp_orig) covariance parameter estimate ± 1 S. E. = 0.01929 ± 0.020; Additional file 7: Table S7).
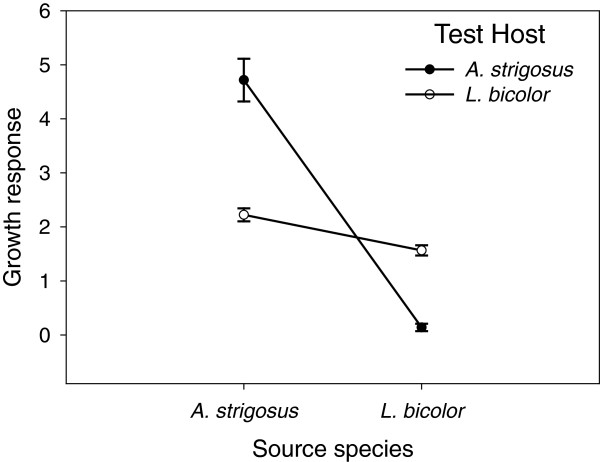
The fold increase in growth of inoculated plants over uninoculated controls was strongly affected by the interaction of test host with source host (Figure 5; χ2d.f. = 1 = 25.61, p < 0.0001). Further, the planned contrast comparing the fold increase in fitness obtained from Bradyrhizobium genotypes isolated from conspecifics versus heterospecifics was highly significant (likelihood ratio χ2d.f. = 2 = 28.76, p < 5 × 10-7). Both main effects were statistically significant (test host species χ2d.f. = 1 = 5.059, p < 0.03; host species of origin χ2d.f. = 1 = 5.059, p < 0.03), but the mean growth effect on each host species changed rank between conspecific and heterospecific pairings (χ2d.f. = 1. = 25.62, p = 4 × 10-7).
Figure 5.
Mean shoot dry weight response to Bradyrhizobium genotypes isolated from conspecific versus allospecific hosts. Response = fold increase over uninoculated control plants. Closed circles indicate growth of A. strigosus test plants; open circles indicate growth of L. bicolor test plants. Error bars represent ± 1 standard error.
Response specificity, as indicated by the paired difference indices calculated from within-block growth responses stimulated by the different Bradyrhizobium genotypes, differed significantly between host species and was more than three-fold greater in A. strigosus (mean PDI ± 1 S. E. = 6.6 ± 0.7) than in L. bicolor (mean PDI ± 1 S. E. = 1.7 ± 0.1).
G × G interaction (Effect specificity)
Bradyrhizobium fitness was estimated from nodule number, total nodule area, and area of the average nodule. Both nodule number and total nodule area differed significantly among Bradyrhizobium genotypes on each host (Additional file 2: Table S2, Figures 2 and 3).
On A. strigosus, over 30% of variance in total nodule area was due to Bradyrhizobium genotype: genotypes isolated from congeners produced significantly greater total nodule area than did those isolated from lupines. Ten percent of variance in nodule number was due to Bradyrhizobium genotype: genotypes isolated from congeners produced slightly fewer nodules than did those isolated from lupines. Seventy percent of variance in average individual nodule area was due to Bradyrhizobium genotype: genotypes isolated from congeners produced significantly larger nodules.
On L. bicolor, only 15% of variance in total nodule area was explained by Bradyrhizobium genotype. Although genotypes isolated from congeners produced significantly greater nodule area, the difference was slight. Bradyrhizobium genotype explained almost 15% of variance in nodule number, but there was no significant difference between genotypes isolated from congeners versus Acmispon species. Less than 20% of variance in the average individual nodule area on L. bicolor was due to Bradyrhizobium genotype. Genotypes isolated from congeners produce marginally significantly larger nodules than did those isolated from Acmispon hosts.
Paired difference indices (PDI) calculated from host effect on Bradyrhizobium fitness under non-competitive conditions of single inoculations is presented in Table 3. When measured in terms of nodule number, A. strigosus was significantly more specialized than L. bicolor, whereas PDI of the two hosts did not differ when calculated in terms of total nodule area.
Table 3.
PDI values for two Bradyrhizobium fitness components
|
Fitness component |
L. bicolor
|
A. strigosus
|
Difference |
||||||
|---|---|---|---|---|---|---|---|---|---|
| Mean PDI | LCL | UCL | Mean PDI | LCL | UCL | Diff PDI | LCL | UCL | |
| Nodule area* |
0.217 |
0.189 |
0.246 |
0.228 |
0.186 |
0.271 |
0.102 |
-0.034 |
0.056 |
| Nodule number† | 0.672 | 0.605 | 0.739 | 1.067 | 0.832 | 1.302 | 0.395 | 0.183 | 0.607 |
*Natural log transformed.
†Square-root transformed.
For each statistic, LCL = lower 95% confidence limit and UCL = upper 95% confidence limit; Diff PDI = PDI difference between species. Variables that differed significantly between hosts are in bold face.
Association between bacterial genotypes and phenotypes
The matrices of pairwise phenotypic differences in A. strigosus shoot dry weight and average nodule area were each significantly associated with pairwise genetic differences at the NifD locus; association with the ITS region was not significant for shoot dry weight and marginal for average nodule area. Traits measured on L. bicolor exhibited no significant associations (Table 4).
Table 4.
Mantel tests of the correlation between genetic distance matrices and phenotypic difference matrices
| r | P | |
|---|---|---|
|
(a) A. strigosus dry weight | ||
|
NifD |
0.3190 |
** |
|
ITS |
0.0659 |
ns |
|
(b) A. strigosus average nodule area | ||
|
NifD |
0.413 |
*** |
|
ITS |
0.128 |
* |
|
(c) L. bicolor dry weight | ||
|
NifD |
-0.006 |
ns |
|
ITS |
-0.038 |
ns |
|
(d) L. bicolor average nodule area | ||
|
NifD |
-0.0996 |
ns |
| ITS | -0.0280 | ns |
Significance, estimated from 50,000 permutations: ns, not significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001.
Fitness feedback between host and symbiont
For A. strigosus, there was an overall positive relationship between total nodule area and host dry weight (genotype mean correlation between total nodule area and host dry weight = 0.77, prob(H0: r = 0) < 0.0001). Bradyrhizobium genotypes that were more beneficial for A. strigosus also produced on that host fewer nodules (genotype mean correlation between nodule number and host dry weight = -0.35, p < 0.0001) that were larger (genotype mean correlation between average nodule area and host dry weight = 0.71, p < 0.0001).
A weak positive relationship between total nodule area and host dry weight (genotype mean correlation between natural log of nodule area and host dry weight = 0.36, prob(H0: r = 0) < 0.05) suggests a weaker but still positive fitness feedback between L. bicolor and Bradyrhizobium symbionts. In L. bicolor, there was no relationship between host dry weight and nodule number (genotype mean correlation between natural log of nodule number and host dry weight = 0.15, prob(H0: r = 0) > 0.4).
Missing samples and cross-contamination
Nodule data were lost from 14 L. bicolor plants. Seven out of 100 control plants had nodules, indicating cross-contamination (perhaps water draining from containers was splashed from the greenhouse bench). These plants (two A. strigosus and five L. bicolor) were removed from analyses. Cross-contamination probably occurred late in the experiment because the nodulated control plants were still very small and had formed only a few small nodules. We are confident that these potential cross contaminations do not undermine our conclusions, since any broader cross contamination would only reduce phenotypic differences observed between Bradyrhizobium genotypes.
Discussion
Association richness and phylogenetic niche breadth
When growing wild, all four legume species that we examined specialized on subsets of the nodulating community of Bradyrhizobium genotypes. However, accounting for genetic breadth of symbionts in wild-collected nodules shifted the ranking of host specialization. When Bradyrhizobium genotype was ignored (i.e., by simply counting the total proportion of symbionts that were non-identical at the focal loci), L. bicolor ranked as most specialized and L. arboreus as most generalized. However, Bradyrhizobium isolates from wild-collected nodules were genetically clustered by host genus and each Lupinus species hosted a phylogenetically broader group of symbionts than did either Acmispon species. Clustering was strongest when isolates were categorized by NifD genotypes, which suggests that hosts respond to trait loci located on the Bradyrhizobium symbiosis island. The observation that species in Acmispon exhibit greater realized association specificity than do those in Lupinus agrees with previous surveys of wild-collected nodules from plants in these genera [66,78-80]. However, neither L. bicolor nor A. strigosus exhibited fundamental association specificity in greenhouse inoculation tests: both hosts could nodulate all tested Bradyrhizobium strains.
A gap between fundamental and realized association frequency is not uncommon [6] and indicates that environmental context strongly influences nodulation. The complex biotic community within natural soil could influence a genotype’s nodulation success [81-83]. For example, soil pathogens could trigger systemic acquired resistance in the host, which might affect nodulation by some rhizobial genotypes but not others [84].
The availability in soil of multiple rhizobial genotypes also provides scope for plants to actively prefer certain genotypes. Host-symbiont recognition signaling is a well-studied feature of legume-rhizobium interactions [85]. In particular, it has been hypothesized that, during nodule formation and prior to nitrogen fixation, plants can recognize and prefer more-beneficial genotypes via pre-infection partner choice; reviewed in [62]. It is unclear however, whether this host ability would be evolutionarily durable, as it requires genetic linkage between at least three sets of genes in two different organisms (nitrogen fixation genes and signaling genes in the bacterial genome and receptors to the bacterial signals in the host plant genome). Such linkage could be stable under very specific conditions, but could also be broken by any incident of recombination [86]. In several legume-rhizobium systems, including A. strigosus[87], multiple-genotype inoculation experiments have found that more-beneficial and/or co-evolved genotypes are more likely to occupy nodules [81,88,89]. However, none of these studies could definitively distinguish plant choice from interactions among rhizobia, which could be due to varying abilities to compete for soil resources [90], withstand direct interference by other genotypes [91], colonize the rhizosphere [92], and utilize a range of rhizosphere resources [93,94]. In experiments with Bradyrhizobium strains collected from our site, neither in vitro growth rate nor interstrain interference was correlated with nodulation rate on A. strigosus[87], but these and other hypotheses should be further tested in more complex conditions where bacterial traits not directly related to symbiont effectiveness might influence nodulation ability.
Realized association frequency might also be restricted by patchily distributed plant and symbiont genotypes [95-97], which could be caused by coevolution [98] and/or soil heterogeneity. In the Bodega Dunes, two episodes of Holocene dune advance [99,100] left contrasting soils that might structure both plant and Bradyrhizobium populations: L. bicolor is restricted to mid-Holocene dunes whereas A. strigosus and A. heermannii occur only on poorly stabilized late-Holocene dunes (E. L. S. and T. J. M., personal observation). In contrast, L. arboreus occurs across dunes of both ages [101]. We specifically sampled nodules from L. arboreus across its habitat range to control for potential confounding of host and Bradyrhizobium distribution; indeed, L. arboreus nodules harbored genotypes from across each gene network. However, the genotypic composition of isolates from L. bicolor did not differ significantly from those isolated from L. arboreus, which suggests that soil habitat is not the only determinant of genotype in wild-collected nodules of Lupinus.
Response and effect specificity, fitness trade-offs, and fitness feedback
The “Jack-of-all-trades is a master of none” hypothesis was supported by evidence that specialist A. strigosus obtained more fitness benefit from its genetically narrower group of beneficial symbionts than generalist L. bicolor obtained from its genetically broader group of compatible symbionts.
As predicted by the co-adaptation hypothesis, each host received greater fitness benefit from genotypes isolated from congeners or conspecifics (genotypes isolated from L. bicolor provided particularly poor benefit to A. strigosus). Although the sample sizes used to estimate fitness are not large, this pattern of response specificity has been detected in several studies of unmanaged legumes [102-104] and suggests that symbiont effectiveness may be increased by host-symbiont coevolution [45,89,98,105].
A. strigosus plants were highly specialized in their fitness effect on the tested Bradyrhizobium genotypes, which is consistent with previous evidence that this host imposes absolute sanctions [64] on less effective genotypes [87]. There was much less variance among Bradyrhizobium genotypes in the benefits they obtained from L. bicolor.
Also supporting the co-adaptation hypothesis, genotypes of Bradyrhizobium were better adapted to hosts related to those from which their ancestors were isolated. This pattern was strongest on A. strigosus: genotypes isolated from congeneric hosts produced greater total nodule area and therefore likely yielded more progeny than did genotypes isolated from lupines.
As a consequence, symbiont benefits to A. strigosus fitness positively fed back to symbiont fitness via increased allocation to nodules: genotypes beneficial to this host realized greater fitness benefits than did those that were not. An overall positive correlation between NifD genotypic mean values of total nodule area and host shoot dry weight on A. strigosus suggests that strong positive fitness feedback between mutualists in this partnership [67] is driven by traits encoded in the symbiosis island. This feedback pattern was much weaker on the generalist, L. bicolor, and was not significantly correlated with ITS genotype.
Positive partner feedback can produce positive frequency dependence and drive the most beneficial partners to local fixation [49,106], a process that has been predicted for other legumes that exhibit symbiont specificity [83].
Much stronger positive fitness feedback might allow specialist A. strigosus to outcompete generalist L. bicolor in local patches [107]. However, if L. bicolor were a better colonizer, which is expected of generalists [108], then it might persist in a spatially patchy environment through a competition-colonization trade-off [109]. In that case, A. strigosus and L. bicolor would not coexist at a local scale, but both might persist at a larger spatial scale via patch dynamics. This process could also spatially structure Bradyrhizobium populations [12]. Indeed, wild A. strigosus plants were found in association with a genetically narrow range of Bradyrhizobium genotypes that were most beneficial to their growth, suggesting that these genotypes dominate Bradyrhizobium populations in areas inhabited by this host. In contrast, as predicted from the weaker correlations between plant and rhizobial fitness components on L. bicolor, wild L. bicolor plants associated with a genetically wider range of Bradyrhizobium genotypes that were not necessarily the most beneficial.
Conclusions
A field survey detected differences in partner specificity among four sampled host species and greenhouse experiments revealed that at this site A. strigosus clearly specializes on a genetically narrower range of symbionts than does L. bicolor. The fitness benefits that these two hosts received from symbiotic partners affirmed two important tenets of evolutionary theory. First, as predicted by the Jack-of-all-trades is a master of none hypothesis, specialist A. strigosus obtained greater benefit from its genetically narrow group of beneficial symbionts than generalist L. bicolor obtained from a genetically more diverse group. Second, as predicted by the co-adaptation hypothesis [2,3,50], each test species performed better with partner genotypes isolated from conspecifics. Further, host fitness benefit translated directly into symbiont fitness via increased allocation to nodules: nodules occupied by genotypes isolated from conspecific hosts received the greatest benefit. These patterns were strongest in A. strigosus. Positive fitness feedback between this specialized host and its co-adapted symbionts could drive positive plant-soil feedback and contribute to natural distribution patterns observed in the field.
Methods
Collection of wild Bradyrhizobium isolates
During the end of March and the beginning of April 2007, we isolated Bradyrhizobium bacteria from nodules of four native California legumes growing on the Bodega Dunes at Bodega Marine Reserve (38°19′01″N, 123°04′18″W) and Sonoma Coast State Park (38°20′34″N, 123°03′32″W). Portions of these dunes were stabilized in the mid-twentieth century by planting European beach grass, Ammophila arenaria[110,111], but no samples were collected in areas occupied by A. arenaria. The four host legumes belong to two genera, Acmispon (A. heermannii Greene [112], and A. strigosus Nutt. Ex Torr. & Gray [112], previously Lotus heermannii and Lo. strigosus, respectively) and Lupinus (L. bicolor and L. arboreus), which are from distantly related clades of papilionoid legumes [113].
We harvested and cultured isolates from up to 20 nodules from each of four to six seedlings of each host species using the procedures of Sachs and colleagues [66]. Briefly, intact plants were excavated, the root systems washed in tap water; nodules excised, surface sterilized, rinsed, crushed and individually streaked onto two replica plates of solid modified arabinose-gluconate medium (MAG). Successful cultures were archived in 25% glycerol-MAG at -80°C.
Molecular methods and analysis
Genomic DNA was purified from 25 μl of each frozen isolate using Gentra Puregene Yeast/Bacteria kits, (Qiagen, Valencia, CA). We PCR amplified the intragenic spacer between 16 s and 23 s ribosomal subunits (ITS, 1256 nt) [114] and portions of the nitrogenase α-subunit gene (NifD, 756 nt) [115] as previously described [66]. Amplification products were sequenced in both directions using an Applied Biosystems 96 capillary 3730xl DNA Analyzer (Foster City, CA) at the University of California, Berkeley, Sequencing Facility. Two to eight isolates from each plant were successfully amplified and sequenced, which ultimately yielded a sample of 84 sequenced isolates (Additional file 1: Table S1).
Sequences were aligned using MAFFT [116] with default parameters. Genotypes were identified in MacClade 4.05 [117]. Some portions of the ITS included indels and these regions were removed from the analysis.
Quantifying realized in situ association specificity
Specialization is most easily quantified as the number of taxa with which a taxon interacts [118], termed “partner richness”. However, this measure accounts for neither association frequency [72,73], phylogenetic distance among partners, e.g., phylogenetic breadth [28], nor fitness effects of interactions. Resurging interest has stimulated new methods for quantifying ecological specialization [1,71,75]; we used the Paired Differences Index.
From field collected samples, we estimated the in situ link strength of each pairwise interaction, Pij, as the proportion of nodules sampled from the ith host species occupied by the jthBradyrhizobium NifD type:
| (1) |
where ni,j = the number of nodules on the ith host species occupied by the jthBradyrhizobium NifD type and ni = the total number of nodules sampled from the ith host (nACHE = 19, nACST = 18, nLUAR = 21, nLUBI = 23).
Observed specificity of each of the four hosts was then measured as the differential frequency of interaction with the 22 different available NifD types, using the Paired Differences Index (PDI) measure [71]:
| (2) |
where Pi,1 is the strength of the strongest link with the ith host. The PDI values were not normally distributed; therefore, to compare PDIs of the four hosts, we estimated confidence limits from 1000 simulated 20-nodule populations generated as follows. Using the RAND (“TABLE”,) function of SAS® 9.3, we specified the frequency with which each genotype appeared in each simulated population, assuming that hosts randomly sampled rhizobia and therefore the observed frequencies of the 22 NifD genotypes in the total sample of 81 nodules represented their availability in the Bradyrhizobium population. From each simulated population, we calculated the joint PDI for all four hosts together:
| (3) |
where Pj is the link strength averaged across the four hosts of Bradyrhizobium strain j, P1 is the average link strength of the Bradyrhizobium strain with the strongest link averaged over the four host species, and N = 20 nodules, then used these values to estimate the mean and variance of PDI and calculate 95% confidence limits.
Genetic breadth of partners isolated from wild-grown hosts
We constructed a molecular network for each DNA region with the neighbor-net algorithm implemented in SplitsTree 4.8 [119], using the Akaike Information Criterion (AIC) implemented in Modeltest (version 3.4) to choose a suitable model of sequence evolution [120]. The NifD genotype network was estimated using the GTR + I model [121] of sequence evolution and the ITS network was estimated with the HKY + I + G model [122]. When there were multiple isolates with the same sequence, we randomly picked one for analysis.
A molecular network approach was favored over a phylogenetic tree approach, as previous studies have repeatedly found incongruence between phylogenies of different gene loci in Bradyrhizobium bacterial strains [66,79,123-125]. Moreover, in our sample, SplitsTree found significant recombination in the ITS region and marginally significant recombination within the NifD locus. Genetic networks are standard tools for the analysis of populations in which genomes recombine; such events will induce a reticulated representation of genealogies, which violates the bifurcating model of phylogenetic trees, as reviewed in [126].
For each DNA region, we computed the matrix of average number of differences between pairs of sequences (pairwise differences) within and between host genera to test whether bacterial communities harbored by the two genera differed in genetic diversity. We further computed matrices of within and between host-species pairwise Φst[127] to test whether pairs of host species harbored genetically distinct groups of bacteria. The Φst statistic for nucleotide diversity is analogous to Wright’s FST, but accounts for sequence divergence as well as genotype frequency. Finally, each DNA region was analyzed with a two-level hierarchical AMOVA model with the following levels: within host species, between species within host genus, and between host genera. Significance tests for each level of the AMOVA, the Φst values, and the pairwise differences were obtained by permuting genotypes between levels or populations. These analyses were performed with Arlequin 3.0b [128].
Greenhouse experiment
Seed source
Seeds of A. strigosus and L. bicolor were obtained from ripe fruits of many plants growing at Bodega Marine Reserve in May-June 2008, pooled, mixed and stored dry at room temperature. In mid-July, seeds were surface-sterilized and scarified [66,129], then germinated individually in wells of 96-well culture plates containing 200 ml of sterile ddH2O (incubated at 15°C, 5–7 days). Seedlings of A. strigosus were transferred to bleach-sterilized 38-mm × 140-mm Conetainers (Stuewe & Sons, Corvallis, OR, USA) filled with autoclaved quartzite sand [66]. One-week old seedlings of L. bicolor were transferred to bleach-sterilized 64-mm × 250-mm Deepots (Stuewe & Sons, Corvallis, OR, USA) filled with autoclaved calcine-clay (Turface MVP®, Profile Products, Buffalo Grove, IL, USA). Media were chosen to maximize plant survival. Neither medium provides nitrogen; we have no reason to expect that differences in media would affect symbiont specificity or fitness benefits to hosts.
All transplants were placed in a greenhouse under ca. 50% shade for seven days of hardening, with misting twice daily, and subsequently exposed to full sunlight, watered daily, and fertilized weekly with 10-ml Jensens’s nitrogen-free solution [130] until treatments were applied one month after germination (see below).
Preparation of bacterial inocula
For the greenhouse inoculation experiment, we chose 29 Bradyrhizobium strains that represented the range of a preliminary concatenated nifD and ITS network of the sequenced isolates. For reference to previously described Bradyrhizobium genotypes, we added two strains (T11 and T12) from among those described by Sachs and colleagues [66], for a total of 31 strains. A culture of each strain was initiated from ~50 μl of original stock archived in glycerol, plated onto solid MAG medium [78] and incubated at room temperature until bacteria covered the plate. Bacteria were washed from plates into 50-ml polypropylene tubes containing sterile dd H2O and vortexed. Bacterial concentration of each inoculum was estimated via light absorbance at 600 nm with a spectrophotometer and a previously established standard growth curve; then adjusted to 108 cells ml-1 by dilution in sterile dd H2O. Each treatment plant received 10 ml of bacterial suspension and control plants received 10 ml of sterile dd H2O.
Experimental design
The two host species were planted and inoculated at the same time, but grown in different substrates and separated on the greenhouse bench to prevent shading of A. strigosus by the larger L. nanus. Each host species was arranged into ten 36-seedling blocks: 31 test seedlings, each randomly assigned to a Bradyrhizobium strain, and five seedlings that received the sterile water control inoculum. All treatments were spatially randomized within each block and the 10 blocks were randomly located on the greenhouse bench, for a total of 360 plants per host species. Seven to eight weeks after inoculation, plants were harvested to obtain oven-dried aboveground (shoot) dry weight (a component of plant fitness). We counted, excised, and photographed nodules with a stage micrometer to obtain their total projected area using ImageJ64 [131]. Because symbiotic rhizobia reproduce clonally and are ultimately released from legume nodules into the soil, nodule number and mass or projected area have been treated as multiplicative components of rhizobial fitness [67,129,132-134]. When obtained using a standard-density, single-genotype inoculum, nodule number indexes the probability that an individual rhizobium cell can initiate a nodule [88,135]. Nodule size, measured as biomass or projected area, is a good proxy for the number of viable rhizobia in a nodule [129,132,136]. Since most nodules are initiated by a single rhizobium cell [88,137], nodule size indexes rhizobium fecundity. Like many fitness components, nodule number and individual nodule size are often negatively correlated [138]. Nonetheless, when all nodules on a host are occupied by a single genotype, the ratio of total nodule area to nodule number estimates average fecundity on that host for cells of that genotype.
Analysis
Fundamental association specificity
No analysis was required because all hosts nodulated all Bradyrhizobium genotypes.
Host response specificity
Because the experimental design technically precludes treating test host as an independent variable in the ANOVA, it was most conservative to analyze each host species separately. These models (Additional file 2: Table S2) included random effects of spatial block and Bradyrhizobium genotype, with genotype nested within host species from which it was isolated and further nested within host genus of isolation, along with all appropriate interactions. A planned one degree-of-freedom contrast compared the growth of inoculated versus uninoculated test plants and a posterior 1-df contrast compared the growth of plants inoculated with bacterial genotypes that had been isolated from the each of the two plant genera (i.e., Lupinus, L, versus Acmispon, A). Bacterial genotype effects on dry weight (Additional file 3: Table S3) were compared using a Tukey-Kramer test of honestly significant pairwise differences [139].
Although the experimental design technically precludes treating test host as an independent variable in the ANOVA, if the different growth conditions (different growth media and locations in the greenhouse) of the two test host species did not interact with the effect of Bradyrhizobium genotype, then both species might be analyzed with a single model. Lack of replication of Bradyrhizobium genotype within blocks precluded testing for this interaction within each species. Nonetheless, we cautiously tried this less conservative approach, in which a significant interaction between test host species and Bradyrhizobium genotype G × G interaction [77], would indicate differences in response specificity between L. bicolor and A. strigosus.
We analyzed the entire data set (both host species) by adding the fixed factor of test host and all its interactions to a nested hierarchy of generalized linear mixed models to examine how host species, Bradyrhizobium genotypes, the species or genus of plant from which they were isolated, and two-way interactions affected plant benefit. The two hosts differed so much in response specificity that they exhibited very different distributions of shoot weight. For this reason, all models employed a gamma probability density function and a logit link function. Corrected Akaike Information Criteria (AICc) were used to find the best model (i.e. the one with the lowest AICc). Relative likelihoods of other models were compared using e(AICmin-AICi)/2, where AICmin is the corrected AICc of the best-fitting model and AICi is the AICc of the model being compared [140]. This analysis was performed with SAS Version 9.3 (SAS Institute Inc., 2002–2010, Cary, NC, USA). The best fitting model (model 16 in Additional file 4: Table S4) included the fixed effects of host species (host), host species from which the Bradyrhizobium genotype was isolated (sp_orig), and the host species by species of origin interaction, random effects of block (block) and Bradyrhizobium genotype nested within the host species from which the genotype was isolated (strain(ge_orig)), and the interaction of host species with Bradyrhizobium genotype nested within host species of origin (host* strain(sp_orig)). Thus, the G × G interaction was split between a fixed effect interaction (Additional file 5: Table S5) and a random effect interaction (Additional file 7: Table S7).
We also compared test host growth response, relative to uninoculated control, to Bradyrhizobium genotypes isolated from conspecifics versus response to those isolated from allospecifics. To calculate growth response (fold increase over uninoculated control), we used shoot weight of the control plants in each block to calculate the mean growth effect on each test host of the ith genotype in each block: (wi – w0)/w0, where wi = shoot dry weight of the plant inoculated with genotype i and w0 = shoot dry weight of the uninoculated conspecific control plant. This variable was analyzed with a generalized linear model including test host, host species of origin, and their interaction, with an exponential distribution and canonical (reciprocal) link function. A planned contrast was used to compare the fitness benefit obtained from Bradyrhizobium genotypes isolated from conspecific versus heterospecific hosts. This analysis was performed with JMP Version 10.0.0 (SAS Institute Inc, Cary, NC, 1989–2007).
To quantify specialization using the host fitness responses, we calculated the fitness effect of each Bradyrhizobium genotype on each test host species separately for each of the k = 10 blocks,
| (4) |
where wijk = shoot dry weight of the ith host species inoculated with the jth strain in the kth block, and wiCk = shoot dry weight of the uninoculated conspecific control plant in the same block. The Paired Differences Index (PDI) [71] of the ith test host can then be calculated for each block by ranking within-block values of pijk from highest (j = 1) to lowest (j = N), where N = 30 tested strains, to obtain:
| (5) |
Variance among blocks was used to calculate 95% confidence intervals for the PDI of each test host species.
Specificity of host effect on symbionts
The effect of the two test hosts on the fitness of each bacterial genotype was examined by univariate ANOVAs on nodule number, total nodule area per plant, and average nodule size. All dependent variables were log transformed to meet the ANOVA normality assumptions [141], pp. 185, 202–204. Standard deviations were not equal among factor levels; so we performed a Welch ANOVA, which allows for heteroscedasticity among factors [141], pg. 183.
Specificities of effect of the two test hosts on Bradyrhizobium genotype fitness under conditions of individual inoculation were calculated from equation (4), above, by defining pijk in terms of each of two rhizobial fitness components, nodule number and total projected area of nodules. To meet the assumptions of normality and homoscedasticity between populations, nodule number was natural log transformed and nodule area was square root transformed. Variance among blocks was used to calculate 95% confidence intervals.
Test of association between bacterial genotypes and phenotypes
The groups of Bradyrhizobium genotypes isolated from congeneric versus heterogeneric hosts were identified by the population genetic analysis. The A group (genotypes T1 - T12) were originally isolated from Acmispon; the L group (genotypes T13 - T31) were originally isolated from Lupinus. Having been classified by host association, the bacteria within one group may not be more related to each other than to bacteria in the other group. Thus, although this contrast compares the average effects of the two bacterial groups, it does not test for a specific association between bacterial genotype and phenotype. To explicitly test for an association between bacterial genotype and symbiotic phenotypes, we used Mantel tests of non-random association between matrices. We computed six matrices: two matrices of pairwise genetic distances between all pairs of genotypes, one for NifD and one for ITS, and four matrices of pairwise mean phenotypic differences between all pairs of bacterial genotypes, two for A. strigosus and two for L. bicolor. Each genetic distance matrix was then compared with the four different phenotype matrices. Under the null hypothesis of these tests, a random association between the elements of the phenotype matrix and those of the genotype matrix would indicate that across Bradyrhizobium genotype pairs, the two genotypes within a pair shared phenotypes independently of the genetic distance between them. In contrast, a positive, non-random association would indicate that the phenotypic difference between the genotypes in a pair increases in concert with an increase in their genetic distance. The two reference genotypes, T11 and T12, were omitted during computation of the matrices, as was genotype T30, which we were unable to sequence at the NifD locus. The ANOVAs were performed with JMP®, Version 7. (SAS Institute Inc., Cary NC, 1989 – 2007); the Mantel tests were performed with the R statistical software [142].
Fitness feedback between host and symbiont
To estimate fitness feedback between host and symbiont, the JMP statistical package was used to calculate the Pearson product mean correlation between host fitness, measured as shoot dry mass, and symbiont fitness, measured as nodule number, total nodule area per plant, and average nodule size.
Availability of supporting data
The sequence datasets supporting the results of this article are available from Genbank (http://www.ncbi.nlm.nih.gov/genbank/); accession numbers JQ230720 – JQ230882.
Abbreviations
χ2: Chi-square test statistic; A: Acmispon; ACHE: Acmispon heermannii; ACST: Acmispon strigosus; AIC: Akaike information criterion; AMOVA: Analysis of molecular variance; ANOVA: Analysis of variance; d.f.: degrees of freedom; dd H2O: double-distilled water; DNA: Deoxyribonucleic acid; F: F-test statistic; G × G: Genotype by genotype interaction; GTR + I: Generalized time-reversible model of DNA evolution with invariant sites (I); HKY + I + G: Hasegawa, Kishino and Yano model of DNA evolution with invariant sites (I) and Gamma-distributed among site rate variation (G); ITS: 16S-23S rRNA internal transcribed spacer; L: Lupinus; LUAR: Lupinus arboreus; LUBI: Lupinus bicolor; MAG: Modified arabinose gluconate; nt: nucleotides; NifD: Gene coding for the alpha subunit of dinitrogenase; p: probability of obtaining a test statistic at least as extreme as the one observed, assuming a true null hypothesis; PDI: Paired differences index; Φst: The correlation of haplotypic diversity at different levels of hierarchical subdivision.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
With advice from ELS and assistance from JLS and TJM, ME collected, isolated, archived and genotyped all but two of the samples (which had been collected by JLS), conducted the experiments and analyzed the genetic data. ELS and SSP calculated the paired difference indices, and ELS and ME analyzed the phenotypic data. ME wrote the manuscript with advice and assistance from ELS, who subsequently extensively revised the final manuscript. All authors read and approved the final manuscript.
Supplementary Material
Title of data: Field-sampled nodules, their source host, identifying symbols, and accession numbers for NifD and ITS sequences submitted to GenBank. Description of data: * host = host species, pln = plant within species, nod = nodule on plant, strain = strain identifier, T-type = inoculation type in greenhouse experiment (blank cells indicate strains not used in that experiment), g’type NifD and g’type ITS identify the NifD and ITS genotypes, respectively, included on Figure 1. Respective NifD and ITS GenBank sequence IDs and accession number are indicated as seq nifD, seq ITS, accession # NifD, and accession # ITS. Blank cells in these columns indicate isolates that did not amplify.
Title of data: Separate ANOVA for each host species of plant and bacterial fitness-components. Description of data: Significance: ns, not significant; **, p<0.01; ***, p<0.001. 1 Degrees of freedom for F-test of random block effect: numerator = 9; denominator for (a) = 271, for (b) = 256. 2 Degrees of freedom for F-test of random genotype effect: numerator = 30, denominator for (a) = 271, for (b) = 256. 3 Tests of contrast between Bradyrhizobium groups A (isolated from Acmispon) and L (isolated from Lupinus); degrees of freedom: numerator = 1, denominator for (a) = 271, for (b) = 256.
Title of data: Tukey-Kramer honestly significant differences in trait values between Bradyrhizobium genotypes inoculated on two host species, (a) A. strigosus, (b) L. bicolor. Description of data: Genotypes sharing the same capital letter are not statistically significantly different at p < 0.05. * shoot dry weight.
Title of data: Generalized linear mixed models fitted to variance in host shoot dry weight. Description of data:+ estimated G-matrix not positive definite. Host = test host species, block = spatial block, strain = Bradyrhizobium genotype, sp_orig = host species from which Bradyrhizobium genotypes were isolated, ge_orig = host genus from which Bradyrhizobium genotypes were isolated. All models assumed a gamma distribution with a logit link function. Model 16 (bold face) provided the best fit (i.e., minimized AICc). Adding the block by host species interaction improved the fit (e(AICmin-AICi)/2 = 5.21 x 10–13) but the proportion of variance explained by this component (4.91% + 2.42%, Additional file 7: Table S7) was only marginally significant. The next best fitting model (model 1) included the fixed effect of host and the random effects of block and Bradyrhizobium genotype, with interactions of host with strain and with block. Residuals plots suggested that this model did not perform much worse than the best model, but nesting Bradyrhizobium genotype into host species from which it was isolated greatly improved the relative likelihood that the model fit the data (model 16 versus model 1; e(AICmin-AICi)/2 = 1.28 × 10–12)). Examination of Q-Q plots revealed that all other models provided much poorer fits to the data.
Title of data: ANOVA table of Type III tests of fixed effects on test host shoot dry weight. Description of data: From model 16 of Additional file 4: Table S4.
Title of data: Differences of least square means (logits) of shoot dry weight of test hosts by host of origin. Description of data: From model 16 in Additional file 4: Table S4. Test host species: ACST = A.strigosus; LUBI = L. bicolor. Origin host species: ache = A. heermannii; acst = A.strigosus; luar = L. arboreus; lubi = L. bicolor.
Title of data: Covariance parameter estimates for random factors associated with test host shoot dry weight. Description of data: From model 16 of Additional file 4: Table S4.
Title of data: Least squares means (logits) of shoot dry weight of test plants inoculated with genotypes isolated from the four hosts. Description of data: From model 16 of Table 3.
Contributor Information
Martine Ehinger, Email: maehinger@gmail.com.
Toni J Mohr, Email: tjmohr@berkeley.edu.
Juliana B Starcevich, Email: juli.starcevich@gmail.com.
Joel L Sachs, Email: joels@ucr.edu.
Stephanie S Porter, Email: stephanie.simone.porter@gmail.com.
Ellen L Simms, Email: esimms@berkeley.edu.
Acknowledgments
We thank Barbara Rotz and the Oxford Tract greenhouse staff for their valuable help; Michiel Vos for insights on genetic analysis; and the Bodega Marine Reserve of the U. C. Natural Reserve System and its outstanding staff. Seed and tissue collections were approved by the Bodega Marine Reserve Director and by permit from State of California Department of Parks and Recreation. This research was funded by grant DEB-0645791 to E. L. S, from the USA NSF. During some of this work, M. E. was supported by a grant from the Swiss NSF.
References
- Poisot T, Bever JD, Nemri A, Thrall PH, Hochberg ME. A conceptual framework for the evolution of ecological specialisation. Ecol Lett. 2011;14(9):841–851. doi: 10.1111/j.1461-0248.2011.01645.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams GC. Adaptation and Natural Selection: A Critique of Some Current Evolutionary Thought. Princeton, New Jersey, USA: Princeton University Press; 1966. [Google Scholar]
- Kawecki TJ, Ebert D. Conceptual issues in local adaptation. Ecol Lett. 2004;7(12):1225–1241. doi: 10.1111/j.1461-0248.2004.00684.x. [DOI] [Google Scholar]
- Futuyma DJ, Moreno G. The evolution of ecological specialization. Annu Rev Ecol Syst. 1988;19:207–234. doi: 10.1146/annurev.es.19.110188.001231. [DOI] [Google Scholar]
- Wilson DS, Yoshimura J. On the coexistence of specialists and generalists. Am Nat. 1994;144(4):692–707. doi: 10.1086/285702. [DOI] [Google Scholar]
- Burdon JJ, Gibson AH, Searle SD, Woods MJ, Brockwell J. Variation in the effectiveness of symbiotic associations between native rhizobia and temperate Australian Acacia: within-species interactions. J Appl Ecol. 1999;36(3):398–408. doi: 10.1046/j.1365-2664.1999.00409.x. [DOI] [Google Scholar]
- Straub CS, Ives AR, Gratton C. Evidence for a trade-off between host-range breadth and host-use efficiency in Aphid parasitoids. Am Nat. 2011;177(3):389–395. doi: 10.1086/658177. [DOI] [PubMed] [Google Scholar]
- Agrawal AA, Conner JK, Rasmann S. In: Evolution After Darwin: The First 150 Years. Bell MA, Futuyma DJ, Eanes WF, Levinton JS, editor. Sunderland, MA, USA: Sinauer; 2010. Tradeoffs and negative correlations in evolutionary ecology; pp. 243–268. [Google Scholar]
- Kassen R. The experimental evolution of specialists, generalists, and the maintenance of diversity. J Evol Biol. 2002;15:173–190. doi: 10.1046/j.1420-9101.2002.00377.x. [DOI] [Google Scholar]
- Levins R. Evolution in Changing Environments. Princeton, N.J.: Princeton University Press; 1968. [Google Scholar]
- MacArthur RH. Geographical Ecology: Patterns in the Distribution of Species. New York, USA: Harper & Row; 1972. [Google Scholar]
- Bever JD. Dynamics within mutualism and the maintenance of diversity: inference from a model of interguild frequency dependence. Ecol Lett. 1999;2:52–61. doi: 10.1046/j.1461-0248.1999.21050.x. [DOI] [Google Scholar]
- van der Putten WH, Van Dijk C, Peters BAM. Plant-specific soil-borne diseases contribute to succession in foredune vegetation. Nature. 1993;362:53–56. doi: 10.1038/362053a0. [DOI] [Google Scholar]
- Connell JH. In: Dynamics of Populations. Boer PJ, Gradwell GR, editor. Wageningen: Centre for Agricultural Publications and Documentation; 1971. On the Role of Natural Enemies in Preventing Competitive Exclusion in Some Marine Animals and in Rain Forest Trees; pp. 298–310. [Google Scholar]
- Janzen DH. Herbivores and the number of tree species in tropical forests. Am Nat. 1970;104:501–528. doi: 10.1086/282687. [DOI] [Google Scholar]
- Augspurger CK. Seed dispersal of the tropical tree, Platypodium elegans, and the escape of its seedlings from fungal pathogens. J Ecol. 1983;71(3):759–771. doi: 10.2307/2259591. [DOI] [Google Scholar]
- Blomqvist MM, Olff H, Blaauw MB, Bongers T, Van Der Putten WH. Interactions between above- and belowground biota: importance for small-scale vegetation mosaics in a grassland ecosystem. Oikos. 2000;90(3):582–598. doi: 10.1034/j.1600-0706.2000.900316.x. [DOI] [Google Scholar]
- Bever JD, Dickie IA, Facelli E, Facelli JM, Klironomos J, Moora M, Rillig MC, Stock WD, Tibbett M, Zobel M. Rooting theories of plant community ecology in microbial interactions. Trends Ecol Evol. 2010;25(8):468–478. doi: 10.1016/j.tree.2010.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friesen ML, Porter SS, Stark SC, Wettberg EJ, Sachs JL, Martinez-Romero E. Microbially-mediated plant functional traits. Annu Rev Ecol Evol Syst. 2011;43:23–46. [Google Scholar]
- Bever JD, Broadhurst LM, Thrall PH. Microbial phylotype composition and diversity predicts plant productivity and plant–soil feedbacks. Ecol Lett. 2013;16(2):167–174. doi: 10.1111/ele.12024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klironomos JN. Feedback with soil biota contributes to plant rarity and invasiveness in communities. Nature. 2002;417(6884):67–70. doi: 10.1038/417067a. [DOI] [PubMed] [Google Scholar]
- Kulmatiski A, Beard KH, Stevens JR, Cobbold SM. Plant–soil feedbacks: a meta-analytical review. Ecol Lett. 2008;11(9):980–992. doi: 10.1111/j.1461-0248.2008.01209.x. [DOI] [PubMed] [Google Scholar]
- Liu X, Liang M, Etienne RS, Wang Y, Staehelin C, Yu S. Experimental evidence for a phylogenetic Janzen–Connell effect in a subtropical forest. Ecol Lett. 2012;15(2):111–118. doi: 10.1111/j.1461-0248.2011.01715.x. [DOI] [PubMed] [Google Scholar]
- Li R, Yu S, Wang Y, Staehelin C, Zang R. Distance-dependent effects of soil-derived biota on seedling survival of the tropical tree legume Ormosia semicastrata. J Veg Sci. 2009;20(3):527–534. doi: 10.1111/j.1654-1103.2009.01058.x. [DOI] [Google Scholar]
- Florence RG. Decline of old-growth redwood forests in relation to some soil microbiological processes. Ecology. 1965;46(1/2):52–64. doi: 10.2307/1935257. [DOI] [Google Scholar]
- Webb LJ, Tracey JG, Haydock KP. A factor toxic to seedlings of the same species associated with living roots of the non-gregarious subtropical rain forest tree Grevillea robusta. J Appl Ecol. 1967;4(1):13–25. doi: 10.2307/2401406. [DOI] [Google Scholar]
- Mordecai EA. Pathogen impacts on plant communities: unifying theory, concepts, and empirical work. Ecol Monogr. 2011;81(3):429–441. doi: 10.1890/10-2241.1. [DOI] [Google Scholar]
- Taylor DL, Bruns TD. Community structure of ectomycorrhizal fungi in a Pinus muricata forest: Minimal overlap between the mature forest and resistant propagule communities. Mol Ecol. 1999;8(11):1837–1850. doi: 10.1046/j.1365-294x.1999.00773.x. [DOI] [PubMed] [Google Scholar]
- Law R. In: The Biology of Mutualism: Ecology and Evolution. Boucher D, editor. New York, NY: Oxford University Press; 1985. Evolution in a Mutualistic Environment. [Google Scholar]
- van der Pijl L. Ecological aspects of flower evolution. I. Phyletic evolution. Evolution. 1960;14(4):403–416. doi: 10.2307/2405990. [DOI] [Google Scholar]
- Pauw A. Collapse of a pollination web in small conservation areas. Ecology. 2007;88(7):1759–1769. doi: 10.1890/06-1383.1. [DOI] [PubMed] [Google Scholar]
- Geerts S, Pauw A. The cost of being specialized: pollinator limitation in the endangered geophyte Brunsvigia litoralis (Amaryllidaceae) in the Cape Floristic Region of South Africa. S Afr J Bot. 2012;78:159–164. [Google Scholar]
- Fettell NA, O’Connor GE, Carpenter DJ, Evans J, Bamforth I, Oti-Boateng C, Hebb DM, Brockwell J. Nodulation studies on legumes exotic to Australia: the influence of soil populations and inocula of Rhizobium leguminosarum bv. viciae on nodulation and nitrogen fixation by field peas. Appl Soil Ecol. 1997;5(3):197–210. doi: 10.1016/S0929-1393(96)00146-1. [DOI] [Google Scholar]
- Coventry DR, Hirth JR, Reeves TG, Jones HR. Development of populations of rhizobium trifolii and nodulation of subterranean clover following the cropping phase in crop-pasture rotations in southeastern Australia. Soil Biol Biochem. 1985;17(1):17–22. doi: 10.1016/0038-0717(85)90084-7. [DOI] [Google Scholar]
- Del Papa MF, Pistorio M, Balagué LJ, Draghi WO, Wegener C, Perticari A, Niehaus K, Lagares A. A microcosm study on the influence of pH and the host-plant on the soil persistence of two alfalfa-nodulating rhizobia with different saprophytic and symbiotic characteristics. Biol Fertil Soils. 2003;39(2):112–116. doi: 10.1007/s00374-003-0690-6. [DOI] [Google Scholar]
- Hagen M, Puhler A, Selbitschka W. The persistence of bioluminescent Rhizobium meliloti strains L1 (RecA-) and L33 (RecA+) in non-sterile microcosms depends on the soil type, on the co-cultivation of the host legume alfalfa and on the presence of an indigenous R. meliloti population. Plant Soil. 1997;188:257–266. doi: 10.1023/A:1004229819988. [DOI] [Google Scholar]
- Simonet P, Navarro E, Rouvier C, Reddell P, Zimpfer J, Dommergues Y, Bardin R, Combarro P, Hamelin J, Domenach A-M, Gourbière F, Prin Y, Dawson JO, Normand P. Co-evolution between Frankia populations and host plants in the family Casuarinaceae and consequent patterns of global dispersal. Environ Microbiol. 1999;1(6):525–533. doi: 10.1046/j.1462-2920.1999.00068.x. [DOI] [PubMed] [Google Scholar]
- Béna G, Lyet A, Huguet T, Olivieri I. Medicago-Sinorhizobium symbiotic specificity evolution and the geographic expansion of Medicago. J Evol Biol. 2005;18(6):1547–1558. doi: 10.1111/j.1420-9101.2005.00952.x. [DOI] [PubMed] [Google Scholar]
- Parker MA. Mutualism as a constraint on invasion success for legumes and rhizobia. Divers Distrib. 2001;7:125–136. doi: 10.1046/j.1472-4642.2001.00103.x. [DOI] [Google Scholar]
- Parker MA, Malek W, Parker IM. Growth of an invasive legume is symbiont limited in newly occupied habitats. Divers Distrib. 2006;12(5):563–571. doi: 10.1111/j.1366-9516.2006.00255.x. [DOI] [Google Scholar]
- Rodríguez-Echeverría S, Crisóstomo JA, Nabais C, Freitas H. Belowground mutualists and the invasive ability of Acacia longifolia in coastal dunes of Portugal. Biol Invasions. 2009;11(3):651–661. doi: 10.1007/s10530-008-9280-8. [DOI] [Google Scholar]
- Rodríguez-Echeverría S. Rhizobial hitchhikers from down under: invasional meltdown in a plant-bacteria mutualism? J Biogeog. 2010;37(8):1611–1622. [Google Scholar]
- Stanton-Geddes J, Anderson CG. Does a facultative mutualism limit species range expansion? Oecologia (Berlin) 2011;167:149–155. doi: 10.1007/s00442-011-1958-4. [DOI] [PubMed] [Google Scholar]
- Reinhart KO, Callaway RM. Soil biota and invasive plants. New Phytol. 2006;170(3):445–457. doi: 10.1111/j.1469-8137.2006.01715.x. [DOI] [PubMed] [Google Scholar]
- Douglas AE. Host benefit and the evolution of specialization in symbiosis. Heredity. 1998;81:599–603. doi: 10.1046/j.1365-2540.1998.00455.x. [DOI] [Google Scholar]
- Buddemeier RW, Fautin DG. Coral bleaching as an adaptive mechanism. Bioscience. 1993;43(5):320–326. doi: 10.2307/1312064. [DOI] [Google Scholar]
- Lundgren JG, Lehman RM. Bacterial gut symbionts contribute to seed digestion in an omnivorous beetle. PLoS ONE. 2010;5(5):e10831. doi: 10.1371/journal.pone.0010831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mueller UG, Mikheyev AS, Hong E, Sen R, Warren DL, Solomon SE, Ishak HD, Cooper M, Miller JL, Shaffer KA, Juenger TE. Evolution of cold-tolerant fungal symbionts permits winter fungiculture by leafcutter ants at the northern frontier of a tropical ant–fungus symbiosis. Proc Natl Acad Sci. 2011;108(10):4053–4056. doi: 10.1073/pnas.1015806108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiester AR, Lande R, Schemske DW. Models of coevolution and speciation in plants and their pollinators. Am Nat. 1984;124(2):220–243. doi: 10.1086/284265. [DOI] [Google Scholar]
- Thompson JN. The Geographic Mosaic of Coevolution. Chicago: University of Chicago Press; 2005. [Google Scholar]
- Rathke B. In: Pollination Biology. Real L, editor. New York: Academic Press; 1983. Competition and Facilitation Among Plants for Pollination; pp. 305–329. [Google Scholar]
- Yoder JB, Nuismer SL. When does coevolution promote diversification? Am Nat. 2010;176(6):802–817. doi: 10.1086/657048. [DOI] [PubMed] [Google Scholar]
- Fleming TH, Venable DL, Herrera LG. Opportunism vs specialization: the evolution of dispersal strategies in fleshy-fruited plants. Vegetatio. 1993;107/108:107–120. [Google Scholar]
- Waser NM, Chittka L, Price MV, Williams NM, Ollerton J. Generalization in pollination systems, and why it matters. Ecology. 1996;77(4):1043–1060. doi: 10.2307/2265575. [DOI] [Google Scholar]
- Barrett LG, Broadhurst LM, Thrall PH. Geographic adaptation in plant-soil mutualisms: tests using Acacia spp. and rhizobial bacteria. Funct Ecol. 2011;26(2):457–468. [Google Scholar]
- Bruns TD, Bidartondo MI, Taylor DL. Host specificity in ectomycorrhizal communities: what do the exceptions tell us? Integr Comp Biol. 2002;42(2):352–359. doi: 10.1093/icb/42.2.352. [DOI] [PubMed] [Google Scholar]
- Yang SM, Tang F, Gao MQ, Krishnan HB, Zhu HY. R gene-controlled host specificity in the legume-rhizobia symbiosis. Proc Natl Acad Sci U S A. 2010;107(43):18735–18740. doi: 10.1073/pnas.1011957107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bull JJ, Rice WR. Distinguishing mechanisms for the evolution of cooperation. J Theor Biol. 1991;149(1):63–74. doi: 10.1016/S0022-5193(05)80072-4. [DOI] [PubMed] [Google Scholar]
- Noë R, Hammerstein P. Biological markets: supply and demand determine the effect of partner choice in cooperation, mutualism and mating. Behav Ecol Sociobiol. 1994;35:1–11. doi: 10.1007/BF00167053. [DOI] [Google Scholar]
- Johnstone RA, Bshary R. From parasitism to mutualism: partner control in asymmetric interactions. Ecol Lett. 2002;5(5):634–639. doi: 10.1046/j.1461-0248.2002.00358.x. [DOI] [Google Scholar]
- Sachs JL, Mueller UG, Wilcox TP, Bull JJ. The evolution of cooperation. Q Rev Biol. 2004;79:135–160. doi: 10.1086/383541. [DOI] [PubMed] [Google Scholar]
- Simms EL, Taylor DL. Partner choice in nitrogen-fixation mutualisms of legumes and rhizobia. Integr Comp Biol. 2002;42(2):369–380. doi: 10.1093/icb/42.2.369. [DOI] [PubMed] [Google Scholar]
- Bronstein JL. The exploitation of mutualisms. Ecol Lett. 2001;4(3):277–287. doi: 10.1046/j.1461-0248.2001.00218.x. [DOI] [Google Scholar]
- West SA, Kiers ET, Simms EL, Denison RF. Sanctions and mutualism stability: why do rhizobia fix nitrogen? Proc R Soc Lond Ser B. 2002;269(1492):685–694. doi: 10.1098/rspb.2001.1878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Foster KR, Kokko H. Cheating can stabilize cooperation in mutualisms. Proc R Soc B Biol Sci. 2006;273(1598):2233–2239. doi: 10.1098/rspb.2006.3571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sachs JL, Ehinger MO, Simms EL. Origins of cheating and loss of symbiosis in wild Bradyrhizobium. J Evol Biol. 2010;23(5):1075–1089. doi: 10.1111/j.1420-9101.2010.01980.x. [DOI] [PubMed] [Google Scholar]
- Friesen ML. Widespread fitness alignment in the legume-rhizobium symbiosis. New Phytol. 2012;194(4):1096–1111. doi: 10.1111/j.1469-8137.2012.04099.x. [DOI] [PubMed] [Google Scholar]
- Tirichine L, De Billy F, Huguet T. Mtsym6, a gene conditioning Sinorhizobium strain-specific nitrogen fixation in Medicago truncatula. Plant Physiol (Rockville) 2000;123(3):845–852. doi: 10.1104/pp.123.3.845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paine RT. Food webs - Linkage, interaction strength and community infrastructure - the 3rd Tansley Lecture. J Anim Ecol. 1980;49(3):667–685. [Google Scholar]
- Berlow EL, Neutel A-M, Cohen JE, De Ruiter PC, Ebenman B, Emmerson M, Fox JW, Jansen VAA, Iwan Jones J, Kokkoris GD, Logofet DO, McKane AJ, Montoya JM, Petchey O. Interaction strengths in food webs: issues and opportunities. J Anim Ecol. 2004;73(3):585–598. doi: 10.1111/j.0021-8790.2004.00833.x. [DOI] [Google Scholar]
- Poisot T, Canard E, Mouquet N, Hochberg ME. A comparative study of ecological specialization estimators. Methods Ecol Evol. 2012;3(3):537–544. doi: 10.1111/j.2041-210X.2011.00174.x. [DOI] [Google Scholar]
- Schemske DW, Horvitz CC. Variation among floral visitors in pollination ability: a precondition for mutualism specialization. Science. 1984;225:519–521. doi: 10.1126/science.225.4661.519. [DOI] [PubMed] [Google Scholar]
- Reynolds HL, Vogelsang KM, Hartley AE, Bever JD, Schultz PA. Variable responses of old-field perennials to arbuscular mycorrhizal fungi and phosphorus source. Oecologia. 2006;147(2):348–358. doi: 10.1007/s00442-005-0270-6. [DOI] [PubMed] [Google Scholar]
- Harley JL, Smith SE. Mycorrhizal Symbiosis. New York, USA: Academic Press; 1983. [Google Scholar]
- Devictor V, Clavel J, Julliard R, Lavergne S, Mouillot D, Thuiller W, Venail P, Villéger S, Mouquet N. Defining and measuring ecological specialization. J Appl Ecol. 2010;47(1):15–25. doi: 10.1111/j.1365-2664.2009.01744.x. [DOI] [Google Scholar]
- Chase JM, Leibold MA. Ecological Niches: Linking Classical and Contemporary Approaches. Chicago, IL, USA: University of Chicago Press; 2003. [Google Scholar]
- Heath KD. Intergenomic epistasis and coevolutionary constrain in plants and rhizobia. Evolution. 2009;64(5):1446–1458. doi: 10.1111/j.1558-5646.2009.00913.x. [DOI] [PubMed] [Google Scholar]
- Sachs JL, Kembel SW, Lau AH, Simms EL. In situ phylogenetic structure and diversity of wild Bradyrhizobium communities. Appl Environ Microbiol. 2009;75(14):4727–4735. doi: 10.1128/AEM.00667-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stępkowski T, Hughes CE, Law IJ, Markiewicz L, Gurda D, Chlebicka A, Moulin L. Diversification of lupine Bradyrhizobium strains: evidence from nodulation gene trees. Appl Environ Microbiol. 2007;73(10):3254–3264. doi: 10.1128/AEM.02125-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarabo-Lorenzo A, Perez-Galdona R, Donate-Correa J, Rivas R, Velazquez E, Hernandez M, Temprano F, Martinez-Molina E, Ruiz-Argueso T, Leon-Barrios M. Genetic diversity of bradyrhizobial populations from diverse geographic origins that nodulate Lupinus spp. and Ornithopus spp. Syst Appl Microbiol. 2003;26(4):611–623. doi: 10.1078/072320203770865927. [DOI] [PubMed] [Google Scholar]
- Aguilar OM, Riva O, Peltzer E. Analysis of Rhizobium etli and of its symbiosis with wild Phaseolus vulgaris supports coevolution in centers of host diversification. Proc Natl Acad Sci U S A. 2004;101(37):13548–13553. doi: 10.1073/pnas.0405321101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brockwell J, Bottomley PJ, Thies JE. Manipulation of rhizobia microflora for improving legume productivity and soil fertility: a critical assessment. Plant Soil. 1995;174(1–2):143–180. [Google Scholar]
- Rangin C, Brunel B, Cleyet-Marel J-C, Perrineau M-M, Béna G. Effects of Medicago truncatula genetic diversity, rhizobial competition, and strain effectiveness on the diversity of a natural Sinorhizobium species community. Appl Environ Microbiol. 2008;74(18):5653–5661. doi: 10.1128/AEM.01107-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saeki K. Rhizobial measures to evade host defense strategies and endogenous threats to persistent symbiotic nitrogen fixation: a focus on two legume-rhizobium model systems. Cell Mol Life Sci. 2011;68(8):1327–1339. doi: 10.1007/s00018-011-0650-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oldroyd GED, Murray JD, Poole PS, Downie JA. The rules of engagement in the legume-rhizobial symbiosis. Annu Rev Genet. 2011;45(1):119–144. doi: 10.1146/annurev-genet-110410-132549. [DOI] [PubMed] [Google Scholar]
- Rousset F, Roze D. Constraints on the origin and maintenance of genetic kin recognition. Evolution. 2007;61(10):2320–2330. doi: 10.1111/j.1558-5646.2007.00191.x. [DOI] [PubMed] [Google Scholar]
- Sachs JL, Russell JE, Lii YE, Black KC, Lopez G, Patil AS. Host control over infection and proliferation of a cheater symbiont. J Evol Biol. 2010;23(9):1919–1927. doi: 10.1111/j.1420-9101.2010.02056.x. [DOI] [PubMed] [Google Scholar]
- Heath KD, Tiffin P. Stabilizing mechanisms in a legume-rhizobium mutualism. Evolution. 2009;63(3):652–662. doi: 10.1111/j.1558-5646.2008.00582.x. [DOI] [PubMed] [Google Scholar]
- Lie TA, Göktan D, Engin M, Pijnenborg J, Anlarsal E. Co-evolution of the legume-Rhizobium association. Plant Soil. 1987;100:171–181. doi: 10.1007/BF02370940. [DOI] [Google Scholar]
- Wielbo J. Rhizobial communities in symbiosis with legumes: genetic diversity, competition and interactions with host plants. Cent Eur J Biol. 2012;7(3):363–372. doi: 10.2478/s11535-012-0032-5. [DOI] [Google Scholar]
- Robleto EA, Kmiecik K, Oplinger ES, Nienhuis J, Triplett EW. Trifolitoxin production increases nodulation competitiveness of Rhizobium etli CE3 under agricultural conditions. Appl Environ Microbiol. 1998;64(7):2630–2633. doi: 10.1128/aem.64.7.2630-2633.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bittinger MA, Milner JL, Saville BJ, Handelsman J. rosR, a determinant of nodulation competitiveness in Rhizobium etli. Mol Plant Microbe Interact. 1997;10(2):180–186. doi: 10.1094/MPMI.1997.10.2.180. [DOI] [PubMed] [Google Scholar]
- Wielbo J, Marek-Kozaczuk M, Kubik-Komar A, Skorupska A. Increased metabolic potential of Rhizobium spp. is associated with bacterial competitiveness. Can J Microbiol. 2007;53(8):957–967. doi: 10.1139/W07-053. [DOI] [PubMed] [Google Scholar]
- Hynes MF, Oconnell MP. Host plant effect on competition among strains of Rhizobium leguminosarum. Can J Microbiol. 1990;36(12):864–869. doi: 10.1139/m90-150. [DOI] [Google Scholar]
- Chatel DL, Parker CA. The colonization of host-root and soil by Rhizobia—I. Species and strain differences in the field. Soil Biol Biochem. 1973;5(4):425–432. doi: 10.1016/0038-0717(73)90069-2. [DOI] [Google Scholar]
- Wollum AG, Cassel DK. Spatial variability of Rhizobium japonicum in Two North Carolina Soils. Soil Sci Soc Am J. 1984;48(5):1082–1086. doi: 10.2136/sssaj1984.03615995004800050026x. [DOI] [Google Scholar]
- Vlassak KM, Vanderleyden J. Factors influencing nodule occupancy by inoculant rhizobia. Crit Rev Plant Sci. 1997;16(2):163–229. doi: 10.1080/07352689709701948. [DOI] [Google Scholar]
- Parker MA. Mutualism in metapopulations of legumes and rhizobia. Am Nat. 1999;153:S48–S60. doi: 10.1086/303211. [DOI] [PubMed] [Google Scholar]
- Orme AR, Tchakerian VP. In: Aeolian Geomorphology Proceedings of the 17th Annual Binghamton Geomorphology Symposium, September 1986. Nickling WG, editor. Boston: Allen & Unwin; 1986. Quaternary Dunes of the Pacific Coast of the Californias; pp. 149–175. [Google Scholar]
- Cooper WS. Coastal Dunes of California, Volume 104. Boulder, CO: Geological Society of America; 1967. [Google Scholar]
- Barbour MG, Craig RB, Drysdale FR, Ghiselin MT. Coastal Ecology: Bodega Head. Berkeley, CA: University of California Press; 1973. [Google Scholar]
- Wilkinson HH, Parker MA. Symbiotic specialization and the potential for genotypic coexistence in a plant-bacterial mutualism. Oecologia. 1996;108:361–367. doi: 10.1007/BF00334662. [DOI] [PubMed] [Google Scholar]
- Thrall PH, Burdon JJ, Woods MJ. Variation in the effectiveness of symbiotic associations between native rhizobia and temperate Australian legumes: Interactions within and between genera. J Appl Ecol. 2000;37(1):52–65. doi: 10.1046/j.1365-2664.2000.00470.x. [DOI] [Google Scholar]
- Murray BR, Thrall PH, Woods MJ. Acacia species and rhizobial interactions: Implications for restoration of native vegetation. Ecol Manage Restor. 2001;2(3):213–219. doi: 10.1046/j.1442-8903.2001.00086.x. [DOI] [Google Scholar]
- Martinez-Romero E. Coevolution in rhizobium-legume symbiosis? DNA Cell Biol. 2009;28(8):361–370. doi: 10.1089/dna.2009.0863. [DOI] [PubMed] [Google Scholar]
- Akçay E, Simms EL. Negotiation, sanctions, and context dependency in the legume-rhizobium mutualism. Am Nat. 2011;178(1):1–14. doi: 10.1086/659997. [DOI] [PubMed] [Google Scholar]
- Bever JD. Soil community feedback and the coexistence of competitors: conceptual frameworks and empirical tests. New Phytol. 2003;157(3):465–473. doi: 10.1046/j.1469-8137.2003.00714.x. [DOI] [PubMed] [Google Scholar]
- Kisdi Ã. Dispersal: risk spreading versus local adaptation. Am Nat. 2002;159(6):579–596. doi: 10.1086/339989. [DOI] [PubMed] [Google Scholar]
- Hastings A. Disturbance, coexistence, history, and competition for space. Theor Popul Biol. 1980;18(3):363–373. doi: 10.1016/0040-5809(80)90059-3. [DOI] [Google Scholar]
- Buell AC, Pickart AJ, Stuart JD. Introduction history and invasion patterns of Ammophila arenaria on the North Coast of California. Conserv Biol. 1995;9(6):1587–1593. doi: 10.1046/j.1523-1739.1995.09061587.x. [DOI] [Google Scholar]
- Danin A, Rae S, Barbour M, Jurjavcic N, Connors P, Uhlinger E. Early primary succession on dunes at bodega head. California. Madrono. 1998;45(2):101–109. [Google Scholar]
- Brouillet L. The taxonomy of North American loti (Fabaceae: Loteae): New names in Acmispon and Hosackia. J Bot Res Inst Tex. 2008;2(1):387–394. [Google Scholar]
- Wojciechowski MF, Lavin M, Sanderson MJ. A phylogeny of legumes (Leguminosae) based on analysis of the plastid matK gene resolves many well-supported subclades within the family. Am J Bot. 2004;91(11):1846–1862. doi: 10.3732/ajb.91.11.1846. [DOI] [PubMed] [Google Scholar]
- van Berkum P, Fuhrmann JJ. Evolutionary relationships among the soybean bradyrhizobia reconstructed from 16S rRNA gene and internally transcribed spacer region sequence divergence. Int J Syst Evol Microbiol. 2000;50:2165–2172. doi: 10.1099/00207713-50-6-2165. [DOI] [PubMed] [Google Scholar]
- Parker MA. Divergent Bradyrhizobium symbionts on Tachigali versicolor from Barro Colorado Island, Panama. Syst Appl Microbiol. 2000;23(4):585–590. doi: 10.1016/S0723-2020(00)80034-X. [DOI] [PubMed] [Google Scholar]
- Katoh K, Misawa K, Kuma K, Miyata T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002;30(14):3059–3066. doi: 10.1093/nar/gkf436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maddison DR, Maddison WP. MacClade 4: Analysis of Phylogeny and Character Evolution. Sunderland, MA: Sinauer Associates; 2008. [Google Scholar]
- Fox LR, Morrow PA. Specialization: species property or local phenomenon. Science. 1981;211(4485):887–893. doi: 10.1126/science.211.4485.887. [DOI] [PubMed] [Google Scholar]
- Huson DH, Bryant D. Application of phylogenetic networks in evolutionary studies. Mol Biol Evol. 2006;23(2):254–267. doi: 10.1093/molbev/msj030. [DOI] [PubMed] [Google Scholar]
- Posada D, Crandall KA. MODELTEST: testing the model of DNA substitution. Bioinformatics. 1998;14(9):817–818. doi: 10.1093/bioinformatics/14.9.817. [DOI] [PubMed] [Google Scholar]
- Rodríguez F, Oliver JL, Marín A, Medina JR. The general stochastic model of nucleotide substitution. J Theor Biol. 1990;142(4):485–501. doi: 10.1016/S0022-5193(05)80104-3. [DOI] [PubMed] [Google Scholar]
- Hasegawa M, Kishino H, Yano T-a. Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol. 1985;22(2):160–174. doi: 10.1007/BF02101694. [DOI] [PubMed] [Google Scholar]
- Parker MA, Lafay B, Burdon JJ, van Berkum P. Conflicting phylogeographic patterns in rRNA and nifD indicate regionally restricted gene transfer in Bradyrhizobium. Microbiology (UK) 2002;148:2557–2565. doi: 10.1099/00221287-148-8-2557. [DOI] [PubMed] [Google Scholar]
- Parker MA. Genetic markers for analysing symbiotic relationships and lateral gene transfer in Neotropical bradyrhizobia. Mol Ecol. 2003;12(9):2447–2455. doi: 10.1046/j.1365-294X.2003.01908.x. [DOI] [PubMed] [Google Scholar]
- Vinuesa P, Silva C, Werner D, Martínez-Romero E. Population genetics and phylogenetic inference in bacterial molecular systematics: the roles of migration and recombination in Bradyrhizobium species cohesion and delineation. Mol Phylogenet Evol. 2005;34:29–54. doi: 10.1016/j.ympev.2004.08.020. [DOI] [PubMed] [Google Scholar]
- Posada D, Crandall KA. Evaluation of methods for detecting recombination from DNA sequences: computer simulations. Proc Natl Acad Sci U S A. 2001;98(24):13757–13762. doi: 10.1073/pnas.241370698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Excoffier L, Smouse PE, Quattro JM. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics. 1992;131(2):479–491. doi: 10.1093/genetics/131.2.479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Excoffier L, Laval G, Schneider S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol Bioinform. 2005;1:47–50. [PMC free article] [PubMed] [Google Scholar]
- Simms EL, Povich J, Shefferson RP, Taylor DL, Sachs JL, Tausczik Y, Urbina M. An empirical test of partner choice in a legume-rhizobium mutualism. Proc R Soc B Biol Sci. 2006;273:77–81. doi: 10.1098/rspb.2005.3292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Somasegaran P, Hoben HJ. Handbook for Rhizobia: Methods in Legume-Rhizobium Technology. New York: Springer-Verlag; 1994. [Google Scholar]
- Abramoff MD, Magelhaes PJ, Ram SJ. Image processing with ImageJ. Biophoton Int. 2004;11(7):36–42. [Google Scholar]
- Kiers ET, Rousseau RA, West SA, Denison RF. Host sanctions and the legume-rhizobium mutualism. Nature. 2003;425(6953):78–81. doi: 10.1038/nature01931. [DOI] [PubMed] [Google Scholar]
- Oono R, Denison RF, Kiers ET. Controlling the reproductive fate of rhizobia: how universal are legume sanctions? New Phytol. 2009;183(4):967–979. doi: 10.1111/j.1469-8137.2009.02941.x. [DOI] [PubMed] [Google Scholar]
- Porter SS, Stanton ML, Rice KJ. Mutualism and adaptive divergence: co-invasion of a heterogeneous grassland by an exotic legume-rhizobium symbiosis. PLoS ONE. 2011;6(12):e27935. doi: 10.1371/journal.pone.0027935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friesen ML, Heath KD. One hundred years of solitude: integrating single-strain inoculations with community perspectives in the legume–rhizobium symbiosis. New Phytol. 2013;198(1):7–9. doi: 10.1111/nph.12173. [DOI] [PubMed] [Google Scholar]
- Heath KD, Tiffin P. Context dependence in the coevolution of plant and rhizobial mutualists. Proc R Soc B Biol Sci. 2007;274(1620):1905–1912. doi: 10.1098/rspb.2007.0495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moawad M, Schmidt EL. Occurrence and nature of mixed infections in nodules of field-grown soybeans (Glycine max) Biol Fertil Soils. 1987;5(2):112–114. [Google Scholar]
- Laguerre G, Depret G, Bourion V, Duc G. Rhizobium leguminosarum bv. viciae genotypes interact with pea plants in developmental responses of nodules, roots and shoots. New Phytol. 2007;176(3):680–690. doi: 10.1111/j.1469-8137.2007.02212.x. [DOI] [PubMed] [Google Scholar]
- Ramsey PH, Ramsey PP. Power of pairwise comparisons in the equal variance and unequal sample size case. Br J Math Stat Psychol. 2008;61(1):115–131. doi: 10.1348/000711006X153051. [DOI] [PubMed] [Google Scholar]
- Burnham KP, Anderson DR. Model Selection and Multimodel Inference: A Practical Information-Theoretic Approach. 2. Berlin: Springer-Verlag; 2002. [Google Scholar]
- Zar JH. Biostatistical Analysis. 4. Englewood Cliffs, NJ: Prentice-Hall; 1999. [Google Scholar]
- R_Development_Core_Team: R. A Language and Environment for Statistical Computing. Vienna, Austria: R Foundation for Statistical Computing; 2008. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Title of data: Field-sampled nodules, their source host, identifying symbols, and accession numbers for NifD and ITS sequences submitted to GenBank. Description of data: * host = host species, pln = plant within species, nod = nodule on plant, strain = strain identifier, T-type = inoculation type in greenhouse experiment (blank cells indicate strains not used in that experiment), g’type NifD and g’type ITS identify the NifD and ITS genotypes, respectively, included on Figure 1. Respective NifD and ITS GenBank sequence IDs and accession number are indicated as seq nifD, seq ITS, accession # NifD, and accession # ITS. Blank cells in these columns indicate isolates that did not amplify.
Title of data: Separate ANOVA for each host species of plant and bacterial fitness-components. Description of data: Significance: ns, not significant; **, p<0.01; ***, p<0.001. 1 Degrees of freedom for F-test of random block effect: numerator = 9; denominator for (a) = 271, for (b) = 256. 2 Degrees of freedom for F-test of random genotype effect: numerator = 30, denominator for (a) = 271, for (b) = 256. 3 Tests of contrast between Bradyrhizobium groups A (isolated from Acmispon) and L (isolated from Lupinus); degrees of freedom: numerator = 1, denominator for (a) = 271, for (b) = 256.
Title of data: Tukey-Kramer honestly significant differences in trait values between Bradyrhizobium genotypes inoculated on two host species, (a) A. strigosus, (b) L. bicolor. Description of data: Genotypes sharing the same capital letter are not statistically significantly different at p < 0.05. * shoot dry weight.
Title of data: Generalized linear mixed models fitted to variance in host shoot dry weight. Description of data:+ estimated G-matrix not positive definite. Host = test host species, block = spatial block, strain = Bradyrhizobium genotype, sp_orig = host species from which Bradyrhizobium genotypes were isolated, ge_orig = host genus from which Bradyrhizobium genotypes were isolated. All models assumed a gamma distribution with a logit link function. Model 16 (bold face) provided the best fit (i.e., minimized AICc). Adding the block by host species interaction improved the fit (e(AICmin-AICi)/2 = 5.21 x 10–13) but the proportion of variance explained by this component (4.91% + 2.42%, Additional file 7: Table S7) was only marginally significant. The next best fitting model (model 1) included the fixed effect of host and the random effects of block and Bradyrhizobium genotype, with interactions of host with strain and with block. Residuals plots suggested that this model did not perform much worse than the best model, but nesting Bradyrhizobium genotype into host species from which it was isolated greatly improved the relative likelihood that the model fit the data (model 16 versus model 1; e(AICmin-AICi)/2 = 1.28 × 10–12)). Examination of Q-Q plots revealed that all other models provided much poorer fits to the data.
Title of data: ANOVA table of Type III tests of fixed effects on test host shoot dry weight. Description of data: From model 16 of Additional file 4: Table S4.
Title of data: Differences of least square means (logits) of shoot dry weight of test hosts by host of origin. Description of data: From model 16 in Additional file 4: Table S4. Test host species: ACST = A.strigosus; LUBI = L. bicolor. Origin host species: ache = A. heermannii; acst = A.strigosus; luar = L. arboreus; lubi = L. bicolor.
Title of data: Covariance parameter estimates for random factors associated with test host shoot dry weight. Description of data: From model 16 of Additional file 4: Table S4.
Title of data: Least squares means (logits) of shoot dry weight of test plants inoculated with genotypes isolated from the four hosts. Description of data: From model 16 of Table 3.