Abstract
Matrix metalloproteinase enzymes, overexpressed in HT-1080 human fibrocarcinoma tumors, were used to guide the accumulation and retention of an enzyme-responsive nanoparticle in a xenograft mouse model. The nanoparticles were prepared as micelles from amphiphilic block copolymers bearing a simple hydrophobic block, and a hydrophilic peptide brush. The polymers were end-labeled with Alexa Fluor 647 dyes leading to the formation of labeled micelles upon dialysis of the polymers from DMSO to aqueous buffer. This dye-labeling strategy allowed the presence of the retained material to be visualized via whole animal imaging in vivo, and in ex vivo organ analysis following intratumoral injection into HT-1080 xenograft tumors. We propose that the material is retained by virtue of an enzyme-induced accumulation process whereby particles change morphology from 20 nm spherical micelles to micron-scale aggregates, kinetically trapping them within the tumor. This hypothesis is tested here via an unprecedented super resolution fluorescence analysis of ex vivo tissue slices confirming a particle size increase occurs concomitantly with extended retention of responsive particles compared to unresponsive controls.
In this paper we demonstrate enzyme-driven retention of a polymeric microscale scaffold within tumor tissue via the injection of nanoscale, matrix metalloproteinase-responsive micellar nanoparticles.1–12 In recent work from our laboratory,1 we described the first example of an enzyme-programmed tissue targeted nanoparticle probe and utilized a FRET (Förster resonance energy transfer) based assay for monitoring particle accumulation.1,13,14 Generation of a FRET signal provided evidence that the nanoparticles had undergone an enzyme directed aggregation process in tumor tissue generating a slow clearing, self-assembled “implant” of polymeric material within the tissue.1 Based on those results, we hypothesized that the materials had passively diffused into the tumors following injection, and then undergone a size increase, which trapped the material within the extracellular space within the tissue. To test this hypothesis, we synthesized a new set of polymeric micellar nanoparticles, prepared from the self-assembly of amphiphilic block copolymers consisting of a hydrophilic peptide brush generated via graft through polymerization of peptide-based monomers,15,16 and a simple hydrophobic block (Figure 1). We term these synthons for generating enzyme-responsive nanoparticles, peptide-polymer amphiphiles (PPAs). The PPAs in this study were labeled with Alex Fluor 647 to generate micelles labeled on their periphery with multiple dye molecules. This dye was chosen for two key reasons; 1) it is known that whole mouse imaging is facilitated by the long excitation and emission wavelength of the fluorophore (λex = 635 nm, λem = 670 nm), and 2) this photoswitching dye is amenable to analysis via super resolution fluorescence microscopy by employing stochastic optical reconstruction microscopy (STORM).17–19 The emergence of super resolution fluorescence microscopy techniques18,20,21 have allowed researchers to overcome the diffraction limit and enable the examination of various processes occurring at the sub-micron scale.22–24 Surprisingly, nanomaterials used as delivery therapeutics and diagnostics are rarely characterized via these useful super resolution techniques,25–27 in particular in cellular in vitro or in ex vivo tissue analysis studies. However, despite this lack of precedence, we determined that such an approach would be needed to confirm whether nanoscale particle accumulation into larger aggregates was occurring within the tumor tissue post-injection.
Figure 1.
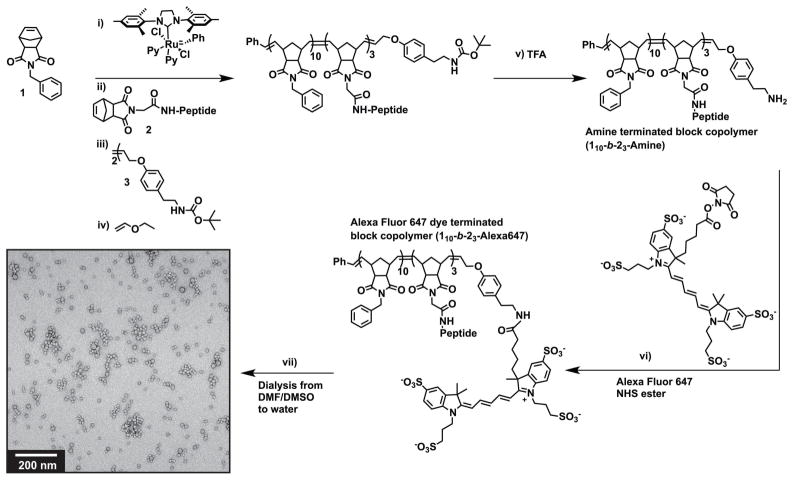
Preparation of enzyme-responsive Alexa Fluor 647 labeled micellar nanoparticles. L-amino acid based norbornyl-peptide substrates were polymerized to generate PPA-L (L-amino acid PPA) for assembly to give micelle, M. D-amino acid based peptides were utilized to generate PPA-D (D-amino acid PPA) for the preparation of non-enzyme responsive control micelle, MD. Block sizes were determined by SEC-MALS analysis and 1H NMR spectroscopy. Synthesis: i) 1 was mixed with Grubbs’ third generation, modified initiator for 30 min and an aliquot analyzed by SEC-MALS to confirm degree of polymerization. ii) Peptide monomer was added and stirred for 2 hr. Confirmation of 10:3 block copolymer ratio was again determined by SEC-MALS. iii) The polymer was terminated with amine termination agent for 1 hr followed by addition of ethyl vinyl ether (iv) to quench the catalyst. v) The Boc protecting group was removed by addition of 90% TFA/DMF for 1.5 hrs followed by precipitation with ether. v) 1.2 equiv. of Alexa Fluor 647 NHS ester was reacted with amine terminated polymers for 18 hrs followed by precipitation with ether. Vacuum dried polymers were then dissolved in 1:1 DMF/DMSO and dialyzed against PBS (pH 7.4) buffered water to generate micellar nanoparticles (M shown in TEM inset). Peptide sequences - L-amino acid sequence: GPLGLAGGWGERDGS. D-amino acid sequence: gplglagGWGERDGS (lower case indicates portion of D-amino acids).
Two micelles, M and MD were prepared from two different PPAs (Figure 1, and Supporting Information Figure 1S). M was generated from a PPA consisting of a peptide prepared with L-amino acids as an active substrate of matrix metalloproteinases (MMPs), known to be overexpressed in certain tumor tissues.2,3,6–8,10,12 MD was prepared from a PPA consisting of a sequence of D-amino acids to inhibit cleavage of the substrate by the protease. These two PPAs were synthesized by employing ring-opening metathesis polymerization (ROMP)28,29 because this technique is capable of the highly proficient, graft-through polymerization of peptide-monomers. Graft-through polymerization of this kind allows for the predictable synthesis of otherwise complex block copolymers in a single pot, not requiring post-polymerization modifications with the oligopeptides, that are unpredictable, and often low yielding.15,16 The polymerization reactions were terminated using a symmetrical olefinic termination agent consisting of a Boc-protected amino group. Subsequent deprotection, and reaction with the activated NHS-ester of Alexa Fluor 647 lead to the formation of Near-IR fluorescence-tagged PPAs. These are subsequently formulated into 25 nm spherical micelles via dialysis from DMSO into PBS buffered water over 24 hrs with three buffer changes.
The enzyme-responsive nature of M and MD was tested initially by mixing micelles with MMP-9 at 37 °C, followed by TEM analysis (Supporting Information, Figure 2S). These experiments confirmed that M and not MD underwent an accumulation process following cleavage of peptides in the shell of the micelles (Figure 1). These in vitro studies were followed by in vivo experiments conducted in mouse models inoculated with HT-1080 human cancer cells to generate xenografts known to contain elevated levels of MMPs (Figure 2, Figure 3S).2,3,6–8,10,12 Both M and MD were intratumorally injected into two different sets of mice, and imaged at eight time points: immediately (1 min) following injection, at 1 hr, 3 hrs, 6 hrs, 1 day, 3 days, 5 days and 7 days. That is, there were eight animals injected at T=0 with M and eight animals injected at T=0 with MD. Images are shown for each animal at given time points, immediately prior to that animal being sacrificed. These studies clearly reveal the retention of M within tumors, and rapid clearance of the D-amino acid control particle, MD. This is confirmed from the whole mouse scan in live mice (Figure 2 and Figure 3S), ex vivo organ analyses (Figure 4S), and ex vivo tumor analysis presented with different thresholds (Figures 5S–8S).
Figure 2.
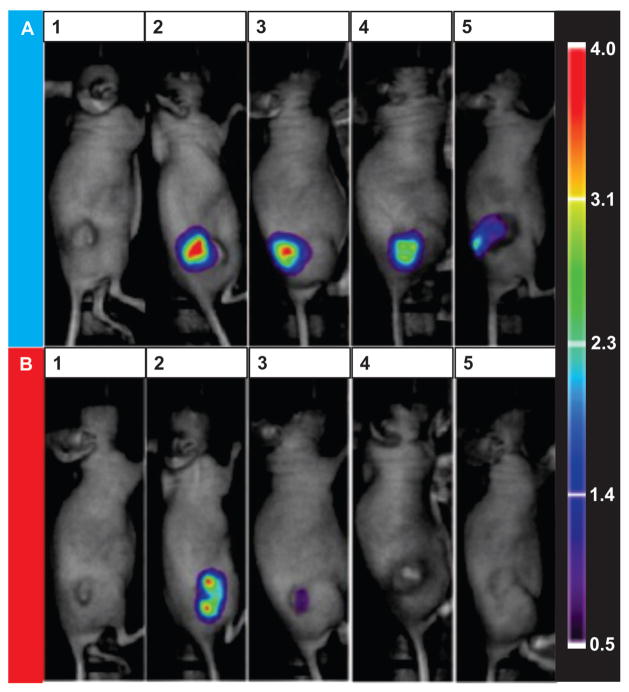
Intratumoral injection to determine relative levels of retention of enzyme-responsive nanoparticles vs control particles with HT-1080 tumors. A) M injected. B) MD injected. 1) Background prior to injection. 2) 1 min post-injection. 3) 1 hr post-injection. 4) 1 day post-injection. 5) 7 days post-injection. HT-1080 xenograft nude mice with tumor size approximately 150 mm3 (~ 0.5 nmoles of nanoparticles injected) were utilized for this study. The linear intensity scale bar from 0.5 to 4.0 is given in units of NC × 103, where NC is number of counts per second per microwatt. The lower threshold is equal to background intensity from control tissue. λex= 635 nm and λem= 670 nm See Supporting Information Figure 3S for additional time points.
To verify that particle activation and subsequent aggregate retention within the MMP-overexpressing HT-1080 tumor tissue2,3,6–8,10,12 occurs in conjunction with observable expression of MMP-9 in our hands, tumor tissue samples were measured via ELISA following imaging analysis (Figure 9S). A time-course study of the tumor tissues after injection shows constitutive expression of MMP-9. All M and MD injected tumor tissues possess MMP-9 concentrations with no statistical difference from that of control tumor tissue (no injection) (p > 0.05).
We propose the mechanism of retention is the assembly of nanoscale particles into larger, slower clearing particles upon reaction with MMPs. Whole mouse imaging reveals that there is a clear difference in behavior between the unresponsive, D-amino acid containing MD vs the L-amino acid, responsive M. In support of this interpretation, our previous studies1,13 have shown the onset of a FRET signal, unique to the formation of a new assembly in response to the enzymatic cleavage of the substrate, and reorganization of the micelles. However, neither of these studies actually allowed imaging of the assemblies themselves nor were they amenable to whole animal imaging. Therefore, we next analyzed tissue slices taken from mice over a range of time points following injection of both M and MD. These tissue slices were then analyzed via STORM to determine if retention could be correlated with a concomitant increase in average particle size within the tissue itself (Figure 3). We determined STORM was uniquely capable of imaging a nanoscale size increase within the tissues by breaking the diffraction limit through image reconstruction,18,22–24 and would reveal information not available in traditional fluorescence microscopy. Prior to STORM analysis, confocal tile scans were first conducted to visualize large areas of tissue (1.06 mm × 1.06 mm) to confirm in vivo imaging results for the tissue slices to be analyzed at higher magnification (see Supporting Information, Figure 10S). These scans revealed the same pattern of retention of M in tumor tissue for up to a week, and clearance of MD within an hour of injection. Next, an area was selected for imaging by traditional confocal fluorescence microscopy and overlaid on bright field images for context (Figure 3: LEFT columns). This process was conducted for tissues from M and MD injected mice (Figure 3; MIDDLE and TOP rows, respectively). Again, these images revealed undetectable signals from MD injected animals for all samples following the initial injection time point. This confirms successful injection, followed by rapid clearance within and hour. Therefore, subsequent analyses focused on STORM of MD injected samples taken immediately following injection (1 min), together with the entire time course of M injected samples (Figure 3). Selected regions (Figure 3: LEFT columns, black squares) were subjected to imaging by STORM (Figure 3: MIDDLE and RIGHT columns).
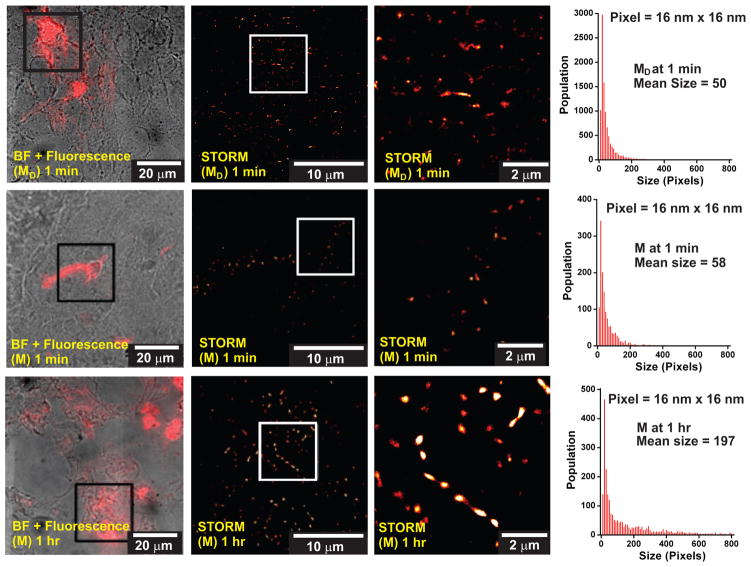
Figure 3.
Confocal and super resolution fluorescence microscopy images of tissue slices from M- and MD-intratumorally injected mice. TOP row: MD-injected mice sacrificed at t = 1 min post-injection. MIDDLE row: M-injected mice sacrificed at t = 1 min post-injection. BOTTOM row: M-injected mice sacrificed at t = 1 hr post-injection. Tumors were removed after sacrificing animals and tissue slices were prepared for imaging. LEFT column shows the overlay of bright field and fluorescence images, where the emission of Alexa Fluor 647-tagged particles is shown in red. The area outlined by the black square was imaged using STORM, as shown in the MIDDLE column, where the area outlined by the white square is enlarged in the RIGHT column. The sizes of the particles in the STORM images were measured and the distribution for each condition is shown in the histograms on the right.
Quantitative analysis of particle size in each STORM image (Figure 3: histograms) reveals the formation of larger particles at 1 hour post injection with M particles. As mentioned earlier, no fluorescence was detected at 1 hour post injection with MD particles (Supporting information, Figure 10S). This is consistent with what can be seen from image analysis of the size of the particles, which reveals an increase in size from initial injection to within one hour following injection (Figure 3: histograms). The size increase in pixels in M-injected tissue (Figure 3: compare MIDDLE and BOTTOM rows) corresponds to an increase from approximately 20–100 nm in diameter at 1 min, to over 200 nm on average after 1 hr. The increased brightness persists and then wanes at 7 days (Supporting Information, Figure 10S), again consistent with whole animal imaging, except here, we can directly observe this as the result of the formation of larger objects, with more dyes per unit area than initially observed immediately following injection. Again, this analysis reveals a similar sized MD particle at 1 min and note that this could not be done at 1 hr or subsequent time points, because of no detectable fluorescence in the tissue at those later time points. This provides evidence that enzymatically-induced aggregation of the materials within the tumor is responsible for their retention for extended periods of time.
In summary, we have utilized Alexa Fluor 647-conjugated peptide polymeric nanoparticles as probes for whole mouse imaging and show extended tumor retention via morphological aggregation in response to MMP enzyme cleavage.1 Furthermore, we provide compelling evidence that this accumulation process is due to assembly of nanometer particles into larger scale aggregates by employing STORM to study tumor tissue slices ex vivo. We observed fluorescent aggregates in targeted tumor tissues within an hour that were retained for at least a week via detailed tissue-slice analysis coupled with whole animal NIR-fluorescence imaging.1 Most importantly, particles designed to resist reaction with MMPs are cleared from tumor tissues within an hour as observed both in vivo and in ex vivo STORM and confocal fluorescence analysis of tissue slices. Together, these studies constitute a previously unexploited coupling of STORM with in vivo imaging. We assert that such an approach will be broadly applicable to other targeted materials and is potentially generalizable.
Supplementary Material
Acknowledgments
The authors acknowledge support for this work from NIH (NIBIB - 1R01EB011633). Furthermore, we thank NIH via a Director’s New Innovator Award (1DP2OD008724). N.C.G. acknowledges the Henry & Camille Dreyfus Foundation for a New Faculty Award and the Alfred P. Sloan Foundation for a fellowship. Part of this research was performed using EMSL, a national scientific user facility sponsored by the Department of Energy’s Office of Biological and Environmental Research and located at Pacific Northwest National Laboratory.
Footnotes
ASSOCIATED CONTENT
Supporting Information. Experimental details including additional timepoints for in vivo imaging data, ex vivo imaging of organs/tumors, and MMP-9 ELISAs are given. This information is available free of charge via the internet at http://pubs.acs.org.
References
- 1.Chien MP, Thompson MP, Barback CV, Ku TH, Hall DJ, Gianneschi NC. Adv Mat. 2013;25:3599. doi: 10.1002/adma.201300823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Galande AK, Hilderbrand SA, Weissleder R, Tung CH. J Med Chem. 2006;49:4715. doi: 10.1021/jm051001a. [DOI] [PubMed] [Google Scholar]
- 3.Jiang T, Olson ES, Nguyen QT, Roy M, Jennings PA, Tsien RY. Proc Natl Acad Sci. 2004;101:17867. doi: 10.1073/pnas.0408191101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kessenbrock K, Plaks V, Werb Z. Cell. 2010;141:52. doi: 10.1016/j.cell.2010.03.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Olson ES, Jiang T, Aguilera TA, Nguyen QT, Ellies LG, Scadeng M, Tsien RY. Proc Natl Acad Sci. 2010;107:4311. doi: 10.1073/pnas.0910283107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Querol M, Chen JW, Bogdanov AA., Jr Org Biomol Chem. 2006;4:1887. doi: 10.1039/b601540a. [DOI] [PubMed] [Google Scholar]
- 7.Scherer R, McIntyre J, Matrisian L. Cancer and Metastasis Rev. 2008;27:679. doi: 10.1007/s10555-008-9152-9. [DOI] [PubMed] [Google Scholar]
- 8.Simberg D, Duza T, Park JH, Essler M, Pilch J, Zhang L, Derfus AM, Yang M, Hoffman RM, Bhatia S, Sailor MJ, Ruoslahti E. Proc Natl Acad Sci. 2007;104:932. doi: 10.1073/pnas.0610298104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tarin D. Cancer Microenviron. 2012;5:95. doi: 10.1007/s12307-012-0099-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tung CH, Mahmood U, Bredow S, Weissleder R. Cancer Res. 2000;60:4953. [PubMed] [Google Scholar]
- 11.Vartak DG, Gemeinhart RA. J Drug Targ. 2007;15:1. doi: 10.1080/10611860600968967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.von Maltzahn G, Harris TJ, Park JH, Min DH, Schmidt AJ, Sailor MJ, Bhatia SN. J Am Chem Soc. 2007;129:6064. doi: 10.1021/ja070461l. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chien MP, Thompson MP, Lin EC, Gianneschi NC. Chem Sci. 2012;3:2690. doi: 10.1039/C2SC20165H. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ku TH, Chien MP, Thompson MP, Sinkovits RS, Olson NH, Baker TS, Gianneschi NC. J Am Chem Soc. 2011;133:8392. doi: 10.1021/ja2004736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hahn ME, Randolph LM, Adamiak L, Thompson MP, Gianneschi NC. Chem Commun. 2013;49:2873. doi: 10.1039/c3cc40472b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kammeyer JK, Blum AP, Adamiak L, Hahn ME, Gianneschi NC. Polymer Chem. 2013;4:3929. doi: 10.1039/C3PY00526G. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bates M, Huang B, Dempsey GT, Zhuang X. Science. 2007;317:1749. doi: 10.1126/science.1146598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Rust MJ, Bates M, Zhuang X. Nature Meth. 2006;3:793. doi: 10.1038/nmeth929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.van de Linde S, Loschberger A, Klein T, Heidbreder M, Wolter S, Heilemann M, Sauer M. Nat Protocols. 2011;6:991. doi: 10.1038/nprot.2011.336. [DOI] [PubMed] [Google Scholar]
- 20.Betzig E, Patterson GH, Sougrat R, Lindwasser OW, Olenych S, Bonifacino JS, Davidson MW, Lippincott-Schwartz J, Hess HF. Science. 2006;313:1642. doi: 10.1126/science.1127344. [DOI] [PubMed] [Google Scholar]
- 21.Hell SW, Wichmann J. Opt Lett. 1994;19:780. doi: 10.1364/ol.19.000780. [DOI] [PubMed] [Google Scholar]
- 22.Dani A, Huang B, Bergan J, Dulac C, Zhuang X. Neuron. 2010;68:843. doi: 10.1016/j.neuron.2010.11.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kanchanawong P, Shtengel G, Pasapera AM, Ramko EB, Davidson MW, Hess HF, Waterman CM. Nature. 2010;468:580. doi: 10.1038/nature09621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Willig KI, Rizzoli SO, Westphal V, Jahn R, Hell SW. Nature. 2006;440:935. doi: 10.1038/nature04592. [DOI] [PubMed] [Google Scholar]
- 25.Tantra RKA. Nanotoxicology. 2011;5:381. doi: 10.3109/17435390.2010.512987. [DOI] [PubMed] [Google Scholar]
- 26.Wang Z, Zhang X, Huang P, Zhao W, Liu D, Nie L, Yue X, Wang S, Ma Y, Kiesewetter D, Niu G, Chen X. Biomaterials. 2013;34:6194. doi: 10.1016/j.biomaterials.2013.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhu MQ, Zhang GF, Li C, Aldred MP, Chang E, Drezek RA, Li ADQ. J Am Chem Soc. 2010;133:365. doi: 10.1021/ja106895k. [DOI] [PubMed] [Google Scholar]
- 28.Sanford MS, Love JA, Grubbs RH. J Am Chem Soc. 2001;123:6543. doi: 10.1021/ja010624k. [DOI] [PubMed] [Google Scholar]
- 29.Smith D, Pentzer EB, Nguyen ST. Polymer Rev. 2007;47:419. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.