SUMMARY
Early vertebrate embryos must achieve totipotency and prepare for zygotic genome activation (ZGA). To understand this process, we determined the DNA methylation (DNAme) profiles of zebrafish gametes, embryos at different stages, and somatic muscle and compared them to gene activity and histone modifications. Sperm chromatin patterns are virtually identical to those at ZGA. Unexpectedly, the DNA of many oocyte genes important for germ-line functions (i.e., piwil1) or early development (i.e., hox genes) is methylated, but the loci are demethylated during zygotic cleavage stages to precisely the state observed in sperm, even in parthenogenetic embryos lacking a replicating paternal genome. Furthermore, this cohort constitutes the genes and loci that acquire DNAme during development (i.e., ZGA to muscle). Finally, DNA methyltransferase inhibition experiments suggest that DNAme silences particular gene and chromatin cohorts at ZGA, preventing their precocious expression. Thus, zebrafish achieve a totipotent chromatin state at ZGA through paternal genome competency and maternal genome DNAme reprogramming.
INTRODUCTION
Fertilization involves the joining of parental gametes to create a totipotent zygote. A central issue in developmental biology is to understand how totipotency is established—the enabling of all developmental decisions. Developmental decisions are often made via collaboration between signaling factors, transcription/ chromatin factors, and miRNAs, which need to be expressed at the proper time in early development, and avoid silencing by repressive chromatin and DNA methylation. One mechanism for transcriptional competence of developmental genes is their packaging in ‘‘bivalent’’ chromatin, bearing (simultaneously) histone modifications normally associated with transcriptional activity (i.e., H3K4me3) and silencing (H3K27me3), along with underlying DNA hypomethylation (Laurent et al., 2010; Lister et al., 2009; Zhou et al., 2011). Interestingly, in vertebrate sperm, the vast majority of developmental genes of importance in the early embryo are already packaged in bivalent chromatin (lacking DNA methylation), including virtually all HOX, SOX, FOX, TBX, PAX, CDX, and GATA family transcription factors (Arpanahi et al., 2009; Brykczynska et al., 2010; Farthing et al., 2008; Hammoud et al., 2009; Weber et al., 2007; Wu et al., 2011a). This raises important questions regarding the extent to which DNA methylation and chromatin structures important for totipotency are simply inherited or must be established or reestablished in the early embryo.
In mice, bulk DNA demethylation occurs at the one-cell stage, preferentially affects the male pronucleus (Hajkova et al., 2008; Mayer et al., 2000; Okada et al., 2010; Oswald et al., 2000), and likely involves a 5-hydroxymethylcytosine (5hmC) intermediate catalyzed by TET enzymes (Gu et al., 2011; Iqbal et al., 2011). Recent approaches with DNAme-IP or reduced representation bisulphite sequencing (RRBS) reveal separate phases of DNAme dynamics during preimplantation and postimplantation (Borgel et al., 2010; Smith et al., 2012), showing the lack of DNAme at key early developmental regulators in embryos and methylation of some of these early developmental genes following implantation. Curiously, a key germline gene (Piwil1) was methylated in the mouse oocyte yet was unmethylated in sperm (Borgel et al., 2010; Kobayashi et al., 2012), revealing an instance of parental DNAme asymmetry beyond imprinted genes. Additional work in Xenopus involved the sequencing of methyl-selected DNA to examine stages at and after zygotic genome activation (ZGA) (Bogdanovic et al., 2011). Surprisingly, promoter DNAme apparently did not confer silencing in Xenopus embryos (from ZGA through gastrulation), an issue revisited here in the zebrafish.
Zebrafish possess the basic enzymes shared in vertebrates for DNAme regulation (Dnmt1, Dnmt3a/b, TET family proteins, and MBD/MECP families), for pluripotency/self-renewal, and for chromatin regulation (Goll and Halpern, 2011; Vastenhouw and Schier, 2012; Wu et al., 2011b) but lack parental imprinting because they lack a Dnmt3L ortholog (McGowan and Martin, 1997). Zebrafish exhibit moderate bulk DNA demethylation following fertilization, with subsequent remethylation (to levels comparable to somatic cells) occurring before ZGA (~ 1,000 cells, blastula stage, ~3 hr postfertilization) (Mhanni and McGowan, 2004). For comparison, ZGA occurs at a much different stage in mice ( ~2 cell) or in humans ( ~4–8 cell) (Braude et al., 1988; Flach et al., 1982). Of technical importance, zebrafish generate large numbers of oocytes (~150/clutch) and demonstrate a delay before ZGA ( ~10 cell cycles), enabling examination of oocytes and early embryos prior to ZGA (Kane and Kimmel, 1993). Regarding chromatin, H3K4me3 and H3K27me3 are highly reduced in zebrafish prior to ZGA, though they are still detectable at the precise developmental genes that harbored them in sperm (Lindeman et al., 2011; Vastenhouw et al., 2010; Wu et al., 2011a). However, it remains unclear whether these low levels are indeed instructive for gene poising and/or deterring DNA methylation. Indeed, much remains to be learned, including (1) the status and dynamics of the entire genome at base-pair resolution in gametes and embryos prior to ZGA, (2) a better understanding of the need or roles for DNAme at/after ZGA, and (3) a better overall logic for the reprogramming process in relation to totipotency and germline specification. Here, we take multiple approaches to address these issues in zebrafish and relate our observations to those in mammals.
RESULTS
Zebrafish Genome Features and DNA Methylomes
We summarize briefly here the features of the zebrafish genome that impact DNAme. The zebrafish genome (1.5 GB) is parsed into 25 chromosomes, lacks a defined sex chromosome, and has ~24,800 nuclear genes and one mitochondrial chromosome. The nuclear genome is guanosine and cytosine (GC) poor (37%), relative to Xenopus (40%), mouse (42%), or human (42%; Table S1 available online), yet strikingly less CpG depleted (<2-fold) than mouse or human (both ~5-fold; Figure 1A). CpG islands (CGIs) are regions of high relative CpG density that are enriched near transcription start sites (TSS) of genes and are typically unmethylated; thus, the “CG content rule” is that regions with a high CpG ratio of observed over expected (obs/ exp) are (counterintuitively) unmethylated. The UCSC database algorithm/criteria overlapped well with our empirical hypomethylated loci (see Extended Experimental Procedures, section Table S3), though GC content adjustments can further increase this overlap (Long et al., 2013). Zebrafish CGIs (zCGIs, total 12,683, median 301 bp) have a mean obs/exp ratio of ~1.10, which is much higher than humans ( ~0.85) (Figure 1B and Table S1). zCGIs are highly enriched at TSS regions, reciprocal to repeat elements, prominent within genes (Figures 1C, 1D, and S1A), and correlated with gene ontology (GO) categories of metabolism and development (Table S7).
Figure 1. DNAme Features of the Zebrafish Genome.
(A) Zebrafish display modest CpG depletion relative to mammals. Observed and expected CpG fractions are displayed. (B) Zebrafish CGIs (zCGIs, UCSC) have extremely high obs/exp CpG frequencies. (C) Reciprocity of zCGIs and repeat types at TSS regions. Distribution of repeats and zCGIs plotted over TSS regions (±2 kb) of protein-coding genes (Ensembl, defined). (D) In zebrafish, fewer gene TSS regions (±250 bp) intersect with CGIs than in mammals. (E) Bulk DNAme levels: all mCG instances over all cytosines sequenced (C+mC). Bulk DNAme levels in oocyte and early embryos are greatly influenced due to the abundance of mitochondrial DNA (chrM), which we find unmethylated. In silico removal of chrM, left. Schematic of stages used in the experiment is shown on the bottom. (F) Regional methylation differences across development reflect bulk methylation changes. DNAme regions (≥ bp, ≥CpGs/region, with a minimum of five reads per CpG) were parsed into HypoM <0.2, PM ≥0.2 to ≤0.8, and HyperM ≥ 0.8. Note the high numbers of PM regions in oocyte and early embryos but exceptionally few in sperm and sphere.
See also Figure S1 and Tables S1, S2, and S7 for (A)–(D) and Figure S3 for (F).
To profile gametes and early development, we performed whole-genome “shotgun” bisulfite sequencing (Cokus et al., 2008; Lister et al., 2008) on mature sperm and oocytes as well as embryos following fertilization—including stages before (2–16 cell, 64 cell, and 256 cell), or just after (sphere) ZGA—along with adult skeletal muscle (schematic, Figure 1E) (see Extended Experimental Procedures on oocyte isolation). We utilized 101 bp paired-end sequencing formats and obtained 200– 400 M mapped filtered reads from each stage/tissue, yielding 10x to 24x genome coverage (Table 1).
Table 1.
Statistics for Bisulphite Sequencing of All Tissues and Stages
| Sperm | Egg | 2 to 16 | 64 | 256 | Sphere | Muscle | |
|---|---|---|---|---|---|---|---|
| Alignments | 481,791,530 | 486,728,832 | 912,758,760 | 512,417,692 | 696,177,617 | 405,903,980 | 681,903,486 |
| Alignments passing filters | 303,464,480 | 227,823,167 | 243,963,809 | 200,264,006 | 217,119,875 | 266,359,218 | 393,924,207 |
| Total base pairs passing all filters | 24,963,074,472 | 14,920,822,820 | 16,001,950,219 | 15,377,294,129 | 18,047,036,957 | 24,451,509,179 | 33,145,552,386 |
| Read coverage | 17.67 | 10.56 | 11.33 | 10.89 | 12.78 | 17.31 | 23.47 |
| Conversion rate, % | 98.99 | 98.67 | 99.70 | 99.76 | 96.58 | 99.82 | 99.57 |
| mCG/(C+mC), % | 18.65 | 10.77 | 10.18 | 9.01 | 13.20 | 17.08 | 12.63 |
| mNonCG/(C+mC), % | 1.39 | 1.98 | 0.40 | 0.35 | 6.85a | 0.39 | 0.60 |
| mCG/(C+mC) + mitochondrial DNA, % | 19.22 | 8.17 | 5.30 | 6.66 | 13.45 | 17.19 | 13.14 |
| mCG/(CG+mCG) | 0.95 | 0.75 | 0.83 | 0.83 | 0.91 | 0.94 | 0.90 |
The data describe, for each tissue/stage, the number of alignments (or base pairs) used in the analysis (rows 1–3). See Extended Experimental Procedures for information and thresholds used in filtering alignments. Read coverage (row 4) refers to genome equivalents. Read coverage calculations exclude chrM. Conversion rate (row 5) refers to the extent (in percent) of C > T conversion by bisulfite treatment. Unmethylated Lambda DNA was used as an internal control to determine bisulphite conversion rate. Rows 6–9 provide different measures (indicated) at calculating cytosine methylation in CG and non-CG contexts. See also Figure S2 and Table S3.
Majority due to nonconversion.
In general, we find CpG methylation higher (2- to 3-fold) in zebrafish than in mammals, reflecting their proportionally higher CpG density (Table 1, row 6). DNAme is much higher in sperm (95% of CpGs methylated) than in oocytes (75%; Table 1, row 9). Mitochondrial DNA was extremely abundant in oocytes, but not in sperm, and was almost entirely hypomethylated (Figure 1E and Table 1, row 8). Bulk DNAme of the nuclear genome reaches a minimum at the 64-cell stage, with remethylation prominent by the 256-cell stage, and is restored by sphere stage to levels observed in sperm (Figure 1E and Table 1, row 6). Non-CG methylation was low in germ cells ( ~1.5% in sperm and ~2% in oocyte) and was even lower in embryos and muscle (~0.5%) (Table 1, row 7). We then defined DNAme regions(≥500 bp), imposed quality filters (≥5 CpGs and ≥5 reads/ window), calculated mean fraction DNAme levels, and parsed regions into three types: hypomethylated (HypoM, < 0.2), partially methylated (PM, ≥ 0.2, ≤ 0.8), or hypermethylated (HyperM, > 0.8) (Figure 1F).
DNAme Comparison of TSS and zCGI Regions across Development
We initially compared sperm to oocyte to define all differentially methylated regions (DMRs), and we provide in the Supplemental Information our analysis of multiple types of loci, including TSS regions (Tables S2 and S7), zCGIs (Tables S2 and S7), genome-wide DMRs (Tables S4 and S7), intragenic features, repeat elements, noncoding RNAs (Figures S2A–S2C and Table S3), and hypomethylated regions (Figures S3A–S3C). Although these sperm-to-oocyte comparisons are interesting in isolation, the underlying logic for their differences is only clear when they are considered alongside the other stages, as described below.
Examination of TSS regions of all stages by k-means clustering analyses revealed four distinct clusters (Figure 2A). DNAme within the two largest clusters (3 and 4) was static. Here, the constitutively HypoM-TSS cohort (cluster 4) generated GO categories for metabolism, transcription, and early development (Table S7 and Figures 2M and S6B). Thus, many genes involved in early development and metabolism are already HypoM in both sperm and oocyte. In contrast, the constitutively HyperM-TSS cluster (cluster 3) was enriched for later developmental functions (neurogenesis, ion transport, and cell differentiation; Table S7 and Figures 2l and S6A), suggesting that many genes expressed later in development will undergo regulated demethylation after ZGA.
Figure 2. DNA Methylation Dynamics at TSS Regions.
(A) Four distinctive cohorts in regard to DNAme at TSS regions. k-means clustering (k = 4) of DNAme (mean fraction CG methylation, TSS ± 250 bp). (B–F) Class average DNAme plots were generated on TSS (±2 kb) for gene classes as defined by GO terms or those containing zCGIs. (G) Obs/exp frequency was calculated for each TSS cluster in (A). (H–L) Snapshots visualized on Integrated Genome Browser (IGB) for dnmt6, dnmt3, hoxd cluster, dazl, gata1a, and sox10 (DNAme scale: 0 to 1, mean base fraction CG methylation).
Dynamic TSS regions also partitioned into two groups. The first, cluster 1, was HyperM in sperm yet HypoM in oocyte, with those HypoM in oocyte transitioning to HyperM by sphere stage. These include factors expressed in mid- to late development (such as dnmt6, foxp1a, and sox12; Figure 2H) or that alternatively have a function during oogenesis (such as rxraa, cntn6, tdrd7, and stk38b; Figures S6J–S6M). The second, cluster 2, was PM or HyperM in oocytes yet HypoM in sperm and transitions to HypoM by sphere stage. These include genes involved in germ cells (dazl, piwi, and vasa; Figures 2K and S6F–S6H), a subset of developmental factors ntla, ddx43, irx3b, dnmt3, and many hox genes/clusters (Figures 2I, 2J, and S6C–S6E). Interestingly, this suggests that key genes for germline function/ specification as well as certain key genes for early development must be demethylated on the maternal allele in the early embryo prior to ZGA, perhaps to provide transcriptional competency prior to ZGA.
Examination of dynamic clusters (1 and 2) and genes reveals a particularly striking feature—upon arrival at sphere/ZGA, TSS loci in the oocyte are “reprogrammed” to the state observed in sperm (Figures 2H–2K and S6C–S6M) and can involve either DNAme acquisition (cluster 1) or loss (cluster 2). Here, DNAme gains are associated with loci/clusters with low CpG obs/exp ratios (median 0.5), whereas those that lost DNAme had intermediate/high ratios (median 0.9; Figure 2G); thus, reprogramming conforms oocyte TSS regions to standard CG content rules. Metagene analyses revealed DNA hypomethylation centered at/near the TSS at all stages (Figure 2B). Notably, genes linked to early development have a wider hypomethylated region (Figures 2C and 2D), a profile contrasted with genes linked to signaling (Figure 2E), which are more often transcribed in mid/ late development.
During early development, most zCGIs either remain hypomethylated (46%) or hypermethylated (45%), with about 10% of zCGIs undergoing DNAme changes (clusters 1 and 5) (Figure S1B). Those constitutively HypoM largely (64%) reside at TSS regions and are enriched at genes involved in metabolism, transcription, and embryo development (Figure S1D and Table S7). This is also evident in metagene analysis in which genes containing a zCGI have a wider hypomethylated TSS region (Figure 2F). Also, dynamic zCGIs clusters have slightly lower obs/ exp ratios than static clusters (Figure S1C). Notably, zCGIs at TSS regions are largely HypoM, whereas those at intragenics are largely HyperM (Figure S1D).
DNAme Comparison of All DMRs across Embryo Development
To define all DMRs genome wide between both germ cells and all stages (filters, 500 bp windows, ≥5 CpG, ≥5 reads per C; criteria, FDR ≥ 0.001 and absolute log2Ratio ≥ 1.5), we conducted sequential pairwise comparisons and summed all, yielding 9,013 total DMRs (Table S4). Clustering analysis revealed nine distinct patterns/clusters (Figure 3A), and their further analysis yielded five major observations.
Figure 3. Maternal DMRs Resolve to Resemble Paternal Status by Sphere Stage.
(A) Pairwise comparisons between developmental stages (summed) yielded differentially methylated regions (DMRs, 500 bp windows, ≥5 CpG, ≥5 reads per C; criteria, FDR R 0.001 and absolute log2Ratio ≥ 1.5, >20% change in fraction methylation). Combined unique regions were scored for mean fraction CG methylation across developmental stages and clustered (k-means, k = 9).(B) DMR locations were intersected with annotations (Ensembl). (C) Obs/exp frequency was calculated for each DMR cluster from (A). (D) DNAme reprogramming of the maternal genome occurs in maternal haploids. Assessment of whether DNAme reprogramming in early embryos relies on the continued presence of the paternal genome. The experimental setup (left) compares two types of sphere-stage embryos: (1) maternal/paternal diploid embryos derived from IVF (top) of wild-type oocytes and sperm containing a known mutation in the golden allele (gol b1, marking the paternal genome, yellow color), and (2) maternal haploid embryos derived from IVF of wild-type oocytes and UV-treated (yellow bolt, bottom) sperm from golden males, providing extensive DNA damage and rendering the genome incompetent for replication, rendering a sphere-stage embryo with only maternal DNA (red nucleus in outset). DNA was isolated from sphere-stage IVF embryos, and DNAme levels (mean fraction CG methylation) were assessed at each of 15 promoter regions (krt4, krt8, dnmt6, rarga, zgc:92231, zgc:101640, cpn1 hoxb1a, hoxb3a, pou5f1, dazl, vasa, irx3a, ntl, and dnmt3) using bisulfite sequencing of promoter amplicons in a high-throughput format (right). Here, the order of the genes at right (top to bottom) aligns with the order of the bars in the figure (left to right). Promoters representing sperm DMRs are depicted in red, and oocyte DMRs are depicted in blue. For comparisons, DNAme levels (mean fraction CG methylation) of these same 15 promoters from sperm, oocytes, and normal diploid sphere-stage embryos (converted from our genome-wide data) are provided.
See also Figures 4, S6, and S7 and Tables S4 and S7.
First, ~50% of DMRs reside within gene bodies—largely introns, with a preference for the first intron (Figures 3B and S6M–S6P), raising the possibility that DMRs might include enhancers. Interestingly, particular DMR clusters intersect significantly with known enhancer regions, defined from dome and later stages (data from Bogdanovic et al., 2012; Table S4 and Figures S4A–S4D). Second, DNAme changes very little between the 256-cell stage (~ 2 cell cycles before ZGA) and sphere ( ~2 cell cycles after ZGA), with only 40 total DMRs identified, suggesting virtually no refinement of DNAme while executing ZGA (Table S4). Third (as previewed above in our TSS analyses), reprogramming converts the maternal genome, prior to ZGA, to the status observed in sperm. Here, the nearidentical pattern of DNAme between sperm and sphere allowed us to attribute changes to the maternal genome without the need for parental-specific SNPs. However, we note that, for the minor subset of loci that are partially methylated at intermediate stages of reprogramming, we cannot attribute the changes at that time to a particular parental genome. The paternal genome is not simply static; loci in clusters 3 and 4 are demethylated by the 2- to 16- or 64-cell stage and are then remethylated (before ZGA) to strongly resemble their initial germline status (Figures 3A and S6Q–S6R). Loci in these clusters have a high overlap with DNA repeats (61%, p value < 0.001) and an intermediate obs/exp ratio ( 0.9) (Figure 3C and Table S4). Here, we speculate that their relatively high obs/exp ratio may promote their demethylation in the early embryo, whereas their repeat element density may promote remethylation by ZGA.
Fourth, the genes and behaviors of clusters 1 and 2 are arguably the most interesting and are methylated only in the oocyte and are demethylated by ZGA, yet are remethylated in muscle. Notably, when examining reads derived from individual loci undergoing demethylation, we observe a phase in which the majority of the reads show partial methylation, gradually shifting toward an unmethylated majority rather than a shift in proportion of two distinct populations, methylated and unmethylated. This behavior indicates a progressive and distributive process that is compatible with passive demethylation. Enriched GO categories for cluster 1 include transcription and early development (Table S7) and include most hox loci and many tbx and irx family transcription factors (Figures 2J and S6E). Cluster 2 enriches for transcription and signaling factors and contains key germlinespecific genes (i.e., dazl) and dnmt3 (Figures 2I, 2K, and S6F– S6H and Table S7). Interestingly, only clusters 1 and 2 have relatively high obs/exp ratio (~ 1.10) (Figure 3C). Notably, cluster 1 is relatively depleted of DNA and LTR repeats, is enriched for zCGIs (p values < 0.001, Table S4), and features typical of loci that are HypoM in other cell types (Borgel et al., 2010; Smith et al., 2012; Weber et al., 2007).
To test whether 5hmC might be an intermediate in cluster 1 and 2 demethylation, we mapped 5hmC levels in oocytes (limited coverage), 64-cell stage and sphere stage (Table S4). At 64-cell stage (during which some loci from clusters 1 and 2 are partially methylated), we obtained significant peaks (FDR ≥ 0.001, log2 ≥ 1), primarily at gene bodies (75%, p value < 0.001, Figure S7A). However, 5hmC enrichment was extremely rare (<2%, Table S4) within clusters 1 and 2, arguing against the general use of 5hmC as an intermediate at most DMRs. However, a few notable DMRs clearly overlap with 5hmC, including dnmt3 and a portion of the hoxc locus (Figures S7C and S7D), raising the possibility of 5hmC involvement at particular loci. Finally, levels of 5hmC are extremely low at sphere (Almeida et al., 2012), with residual levels enriched at genic regions (p value < 0.001, Figure S7A).
Fifth, certain loci (i.e., clusters 7–9) acquire DNAme even during the bulk demethylation phase, which is a behavior that may relate to their obs/exp ratios and repeat element properties (Table S4). Finally, the DNAme dynamics of other intragenic features, TSS regions, noncoding RNAs, and repeat elements (Figures S2A–S2C and Table S3 and S4) are provided in the Supplemental Information.
To summarize the GC content observations, clusters that lack a demethylation phase by sphere stage (clusters 7–9) have a low obs/exp ratio, those that demethylate then remethylate in early embryo (clusters 3 and 4) have intermediate ratios, and those that display maternal demethylation by ZGA (clusters 1 and 2) have high obs/exp ratio (Figure 3C). Thus, the logic for DNAme reprogramming is to achieve, prior to ZGA, (1) the demethylation of virtually all genes for early development and germline function and (2) the methylation of many genes for later development/ signaling. This reprogramming process largely involves conforming oocyte genes to GC content rules (high ratios, HypoM; low ratios, HyperM).
A Competent Paternal Genome Is Not Required for Maternal Genome DNAme Reprogramming
The observed “reprogramming” of DNAme of the maternal genome (by ZGA) to the state observed in the sperm raises the possibility of utilizing the sperm genome as a “template” for this process. To investigate this, we utilized parthenogenesis to create maternal haploids, a process that (in zebrafish) requires the paternal contribution of two centrioles, but not the paternal genome, to an acentriolar oocyte. To accomplish this, sperm were isolated from fish homozygous for the golden allele (gol b1/b1) and were either mock treated or UV treated, providing extensive DNA damage and chromatin/DNA crosslinking and thus rendering the paternal genome incompetent for chromatin repackaging or replication (Figure 3D, left). Mock-treated or UV-inactivated sperm were used to fertilize (via in vitro fertilization, IVF) wild-type oocytes, which were then grown to sphere-stage embryos and genotyped.
DNAme reprogramming at the following 15 loci was then assessed: seven loci where (in normal embryos) the maternal genome sharply gains DNAme by sphere stage (krt4, krt8, dnmt6, rarga, zgc:92231, zgc:101640, and cpn1, red bars, Figure 3D) and eight loci where the maternal genome sharply loses DNAme by sphere stage (hoxb1a, hoxb3a, pou5f1, dazl, vasa, irx3a, ntl, and dnmt3, blue bars, Figure 3D). Remarkably, all tested gene promoters in the maternal haploid embryos underwent DNAme reprogramming by sphere stage, displaying their respective sharp losses or gains in DNAme, to an extent that was indistinguishable from control IVF-fertilized embryos or embryos from normal matings (Figure 3D). This result strongly suggests that the paternal genome does not provide a competent and continual “template” for instructing maternal genome reprogramming and that the oocyte likely employs oocyte-derived factors for this purpose.
DMRs between Sphere and Muscle
Comparisons between sphere and muscle allowed us to contrast totipotency with terminal differentiation. We identify ~3,000 DMRs between sphere and muscle; ~2,000 gain DNAme, whereas ~1,000 DMRs lose DNAme (Table S4). DMRs map preferentially to genic regions, preferring introns (Figure 3B). However, few ( ~4%) TSS regions change (cutoff > 20%), with gains ( ~79%) outnumbering loss (21%) (Table S5). Interestingly, the majority involve transitions to a partially methylated TSS state (Table S5). TSS regions that gain methylation are linked to genes involved in transcription, metabolism, and germ-line/reproductive processes (Table S7), including hox genes, ntla, piwil1, dazl, and vasa. Those that lose methylation belong to gene classes involved in blood/vasculature morphogenesis, cytoskeleton organization, and muscle development (FDR < 0.08) (Table S7). Muscle-specific genes that become hypomethylated include musk, dystrophin, and several myosin component genes, whereas myod1, pbx2, pbx4, and certain myocyte enhancer factors are already hypomethylated at sphere; this implies a need for demethylation of factors involved in terminal differentiation of muscle. Again, we emphasize that DMRs that acquire DNAme in muscle are largely those that lose DNAme on the maternal genome by ZGA (clusters 1 and 2, Figure 3A), and loci that lose DNAme in muscle are a subset of those that gain DNAme on the maternal genome by ZGA (clusters 8 and 9, Figure 3A; see also Figures 2A and S1 for TSS and zCGIs).
Transcription-DNAme Relationships
To relate transcription to DNAme, we performed RNA sequencing (RNA-seq) of oocyte total RNA (Table S5) and compared to DNAme at TSS regions (RPKM > 2, classified as expressed). Overall, the vast majority of PM- and HyperM-TSS genes lacked expression (filtered for TSS misannotation; Table S5). Notably, an interesting set of Hyper- or PM-TSS genes were expressed, including many germ-cell-specific genes (dazl, piwil1, and the zona pellucida genes) as well as dnmt3 and dnmt8; however, we speculate that these germline genes and DNMTs were specifically methylated after their expression during oogenesis. HypoM genes that were not expressed yielded GO categories of transcription, developmental processes, and gastrulation (Table S7), which is consistent with “poising/competency” of selector transcription factors in oocytes. Expressed HypoM genes had housekeeping functions (Table S7). Lastly, as expected, 70% of all RNA-seq reads map to rRNA. However, DNAme at rDNA is strikingly lower in oocyte and early embryos compared to sperm, sphere-stage or differentiated muscle (Figures S2C and S6l), suggesting the active transcription of a much larger fraction of rDNA loci during oocyte maturation for sufficient maternal stores of rRNA.
To examine sphere/ZGA, we performed RNA-seq of total RNA (RPKM > 3 classified expressed) and compared to histone modifications and DNAme status (at TSS regions where high-quality histone modification data were available [40%]) (Lindeman et al., 2011). All findings were consistent with predicted chromatin relationships (Figure 4A and Table S7), though, importantly, 97% of HyperM genes were transcriptionally inactive at sphere stage, which is consistent with DNAme having a role in gene silencing at sphere (see Extended Experimental Procedures, Figure 4A section). Likewise, when comparing sphere and muscle, mean RPKM values decrease ( ~35-fold) or increase ( ~5-fold) at TSS regions that gain or lose DNAme, respectively (Table S5 and Figure S5; p value < 0.0001,), with notable examples in Figures 4B and 4C.
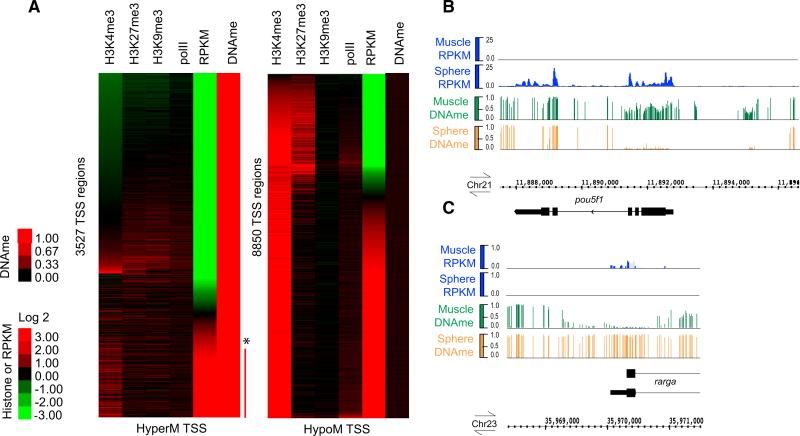
Figure 4. Relationship of DNAme and Histone Modifications at the TSS to Gene Expression during Sphere/PostMBT.
(A) TSS regions (±250bp) at sphere stage were separated into two groups based on their methylation status: either HyperM ≥ 0.8 or HypoM ≤ 0.2 (mean fraction CG methylation, scale 0 to 1) (note: TSSs with partial methylation are extremely rare at sphere). They were then subjected to separate k-means clustering with data sets for histone modifications and gene expression levels. Promoter histone modification status (mean log 2 ratio, array data; [Lindeman et al., 2011]) only available just after sphere/MBT (50% epiboly, 5.3 hpf). Gene expression RPKM levels (first exon, log 2 converted) from our total RNA-seq at sphere stage. Red bar and asterisk indicate loci with high DNAme and high RPKM, which are “false positives”as they mostly represent alternative or incorrectly annotated TSS with high DNAme (see Resultsand Supplemental Information).
(B and C) Example of developmentally regulated DNA demethylation or remethylation at the TSS upon differentiation into muscle with correlated gene expression. Browser snapshots of mean base fraction CG methylation tracks (scale 0 to 1) and relative RPKM values obtained from RNA-seq on total RNA from sphere stage (4 hpf) and adult muscle visualized on Integrated Genome Browser (IGB) for pou5f1 (RPKM scale 0–25) and rarga (RPKM scale 0 to 1).
See also Table S7 for (A) and Figure S5 and Tables S5 and S7 for (B) and (C).
Our genomic methods and thresholds (FDR ≥ 0.001, log2 ≥ 1) confirm earlier staining methods (Almeida et al., 2012) demonstrating that 5hmC is high in muscle but very low in sphere (Table S4). 5hmC is enriched in gene bodies in muscle (Figure S7A); 73% of 5hmC-enriched regions are within genes (p value < 0.001), and 29% of genes bear genic 5hmC (p value < 0.001). 5hmC enrichment increases with gene expression but only slightly and within a very narrow range (Figure S7B). Genes bearing 5hmC in muscle belong to a wide variety of gene classes, including metabolism, developmental processes, signaling, and muscle-specific functions (Table S7), and include the expressed mef2d and klhl30 locus and repressed hox loci (Figures S7C–S7G). However, as the proportion of highly expressed genes (23%) with 5hmC is similar to the proportion of nonexpressed genes (28%), the data do not reveal the logic for 5hmC placement.
Epigenome-Transcription Relationships across Early Development
Clustering analyses of available histone modification profiles (those flanking ZGA and also sperm) (Lindeman et al., 2011; Wu et al., 2011a) and DNAme (at sperm and sphere) yielded ten clusters (Figure 5A). Most clusters yielded expected GO categories (i.e., H3K27me3 at developmental genes), expected histone modification/DNAme relationships (except cluster 7), and expected relationships to gene expression (Tables S6 and S7). As reported previously, histone modification profiles in sperm are similar to those at/after ZGA (Lindeman et al., 2011; Wu et al., 2011a).
Figure 5. Relationship of DNAme Status to Histone Modifications across Development.
(A) Epigenetic features at TSS regions (±250 bp). Histone modifications (mean log 2 ratio, scale –1.5 to 1.5, array data: sperm [Wu et al., 2011a]; preZGA (2.5 hpf), ZGA (3.3 hpf), and postZGA (5.3 hpf) [Lindeman et al., 2011]). DNAme: mean faction CG methylation at sperm and sphere stage, with k-means clustering analysis. (B) Snapshots of the krt4 gene, as an example of a gene from cluster 7 whose expression is highly upregulated by 5-Aza-CyD treatment. DNAme (mean base fraction CG methylation, scale 0 to 1), gene expression (RPKM, scale 0–25, from RNA-seq of total RNA), histone modifications (probe log 2 ratio, scale 0–1.5 at postMBT; [Lindeman et al., 2011]) visualized on IGB.
Cluster 7 is remarkable; it bears low/moderate H3K4me3 and H3K27me3 and high H3K9me3 and is HyperM (after ZGA). This unusual combination of histone modifications at limited loci has been observed previously (Lindeman et al., 2011), but not their relationship to DNAme. Contrary to other clusters, its histone modification status deviates considerably (at ZGA) from sperm. Here, we reasoned that cluster 7 genes might have attributes both of bivalent genes and of genes repressed by an H3K9me3/DNAme axis, with the latter perhaps important for their repression at ZGA/sphere. If so, inhibition of DNAme might preferentially activate this cluster at sphere.
Inhibition of DNMTs Causes Precocious Transcription at Sphere
To assess the impact of DNAme on gene expression at ZGA/ sphere, we utilized 5-Aza-2′-deoxycytidine (5-AzadCyD), a covalent inhibitor of all vertebrate DNMTs, whose administration arrests zebrafish embryos during gastrulation (Martin et al., 1999). Fertilized embryos (one cell) were injected with 5-AzadCyD and grown in its presence for 4 hr (to sphere stage) when RNA was harvested and subjected to RNA-seq. Compared to control injected embryos, about 3,000 transcripts (9% of total transcripts) were either upregulated or downregulated (>2-fold, nearly equal split) following 5-AzadCyD treatment, and these changes in transcription likely represent a combination of primary and secondary effects. Interestingly, the ~1,500 upregulated transcripts were greatly enriched for GO categories of morphogenesis, motility, germ layer, and gastrulation (Table S7), strongly suggesting the precocious expression of gene sets at sphere that are normally expressed later in development. Notably, cluster 7 (Figure 5A) has both the highest fraction of upregulated genes (p values < 0.001) and the highest median level of upregulation (Table S6). Remarkably, most of these genes are highly expressed during gastrulation in the enveloping layer (EVL) (Thisse et al., 2001), such as keratin genes (krt4, 5, 8, 17, 18, 23, cyt1, cyt1l, and cki), claudins (cldnb, e, and f), capn9, sdprb, and cpn1 (Figure 5B). Thus, a subset of EVL-specific genes—those bearing a distinctive chromatin modification signature—may rely on DNAme for silencing in the early embryo.
DISCUSSION
A major issue in vertebrate developmental biology is how and when totipotency is achieved. Studies on embryonic stem (ES) cells, induced pluripotent stem (iPS) cells, and early development have together greatly informed the mechanisms underlying the pluripotent state in mammals (Takahashi et al., 2007; Zhou et al., 2011). However, we currently lack a clear mechanistic understanding of totipotency and its relationship to pluripotency and gametic states. Indeed, gametes represent an interesting node in development; they are derived from totipotent germline stem cells but also represent the terminal cell in a complex but unipotent developmental pathway—one highly asymmetric in the two genders. A key current issue is how and when totipotency is achieved, whether the mature gametes contain chromatin/DNAme patterns that promote totipotency, or alternatively, whether totipotency relies heavily on reprogramming in the zygote. This work addresses central aspects of these questions in zebrafish.
First, we observe a general correlation between DNAme status and timing of gene expression; genes normally transcribed at ZGA (for metabolism, protein synthesis) are HypoM, whereas most genes involved solely in later development and terminal differentiation are HyperM at ZGA (Figure 6). Second, the execution of ZGA is accompanied with very few changes in DNA methylation (Figure 3A). However, as development progresses, DNAme changes certainly occur, with recent data supporting clear changes as early as epiboly at promoters (Andersen et al., 2012) but showing very limited changes in gene bodies and repeat elements (Macleod et al., 1999).
Figure 6. Model Depicting the Logic of DNAme Reprogramming, ZGA Totipotency, and Differentiation.
(A) Oocyte DMRs are those bearing DNAme in oocyte, not sperm; these resolve to an unmethylated status and include key germline factors and early transcription factors. Sperm DMRs are those bearing DNAme in sperm, but not the oocyte; these resolve to a methylated status and include factors needed in mid/late embryo development.
(B) A logic is presented for the DNAme behavior of gene categories. Factors involved in germ layer specification/gastrulation are designated as “early,” progenitor cells of various stages are considered to be expressed at “mid” stages, and factors involved in terminal differentiation are considered “late.” Expression timing obtained from public sources (http://ZFIN.org). At ZGA/sphere, the DNAme status is consistent with totipotency and germline specification, as transcription factors and germline specification factors needed before or during gastrulation are DNA HypoM (green). Transcription factors utilized during mid/late embryo development can either be HypoM or HyperM (red). Importantly, a set of key factors needed in the early embryo for transcription or germline specification is HyperM in the oocyte, but not the sperm, and transition to demethylation in the embryo prior to ZGA (red to green, transition). Other aspects of these categories and their dynamics are self-evident or are described in the Results.
A key issue for totipotency and reprogramming is the similarity between DNAme patterns in gametes and those at ZGA. Recent work in zebrafish (using partial-genome array formats) observed similar DNAme patterns between sperm and sphere/ZGA at a majority of promoters, though ~40% of promoters reported methylated in sperm were reported demethylated by ZGA (Andersen et al., 2012). Our work extended those results and involved examination of the entire genome and instead revealed almost no differences in DNAme patterns between sperm and ZGA/sphere. Thus, our results strongly suggest genome-wide DNAme “competency” of the paternal genome for ZGA. Although DNAme patterns in the oocyte resemble the sperm (and the embryo at ZGA) at most loci, we defined a large number of parental DMRs. Notably, many key developmental (i.e., hox) and germline (piwil1, dazl, and vasa) genes were markedly HyperM in oocytes and selectively demethylated prior to ZGA (Figures 6A and 6B). Conversely, many genes utilized in mid/later development are HypoM in oocytes and methylated prior to ZGA. Remarkably, in both cases, we observe reprogramming of the maternal genome to the DNAme status observed in sperm, prior to ZGA. Here, we emphasize that the near-identical pattern of DNAme between sperm and sphere allows us to attribute observed changes to the maternal genome without the need for parental-specific SNPs.
The overall logic of reprogramming in the early embryo appears to align with standard CG content rules; when examining dynamic DMRs, those loci with “low obs/exp” ratios become methylated, whereas loci with “high obs/exp” ratios become demethylated. Notably, maternal haploid fish were able to reprogram their genome by ZGA (Figure 3D), establishing that the paternal genome is not used as a continual “template,” though we have not excluded other paternal contributions (i.e., small RNAs). Overall, this process provides parental genome equivalency at ZGA, which results in the execution of transcription on two uniform genomes, enabling the expression of metabolic genes and the competency to express all early developmental genes from both parental alleles (Figure 6B). Furthermore, the HyperM status at sphere of a considerable fraction of promoters for mid/late development suggests an effort to prevent their precocious expression, requiring subsequent cell-type-specific demethylation. Notably, our work does not support the widespread use of 5hmC either during DNAme reprogramming prior to ZGA or within pluripotent sphere-stage embryos, though the very earliest stages (one to two cell) have not been examined. This contrasts with work in mammals, in which 5hmC is observed in pluripotent ES cells (Tan and Shi, 2012).
Interestingly, the loci that acquire DNAme during later development (i.e., sphere to muscle) are the same cohort that undergo maternal-specific demethylation prior to ZGA (Figure 3A, clusters 1 and 2). In fact, oocyte DNAme profiles resemble muscle cells more than they resemble sperm or sphere-stage embryos. Here, we reason that oogenesis may be a developmental program akin to somatic differentiation, involving the production of myriad factors needed for progression through ZGA, a developmental process that may require maternal genome reprogramming prior to ZGA back to a totipotent state. Notably, the genes for germline specification and function (piwil1, dazl, and vasa) are HypoM and are largely bivalent at ZGA/sphere. We suggest that, although germ cell fate is specified by the germ plasm/nuage and is inherited by limited cells (Knaut et al., 2000), the genomes of all cells in the sphere-stage embryo are, from a DNAme standpoint, competent to become germline.
Here, we characterize a set of genes at sphere/ZGA with a distinctive chromatin signature, bearing H3K4me3 (albeit low), H3K27me3, H3K9me3, and HyperM (Figure 5A, cluster 7). Notably, this is the sole cluster that differs greatly between sphere and sperm; in sperm, this cohort lacks all three histone marks, yet remains HyperM. Interestingly, inhibition of DNMTs via 5-AzadCyD preferentially upregulated genes in cluster 7, genes which are normally upregulated well after sphere stage and are specifically expressed in the EVL. This suggests that DNAme functions to repress transcription during ZGA of genes enriching for a specific pathway bearing a distinct chromatin landscape. Interestingly, tfab2a, a transcription factor involved in EVL specification known to regulate the expression of krt18 and cki (Hoffman et al., 2007), was not found to be affected by 5-AzadCyD treatment, further supporting a role for chromatin in the upregulation of certain EVL genes. Furthermore, comparisons of DNAme to transcription show correlation of DNAme with silencing at the vast majority of genes. We suggest that, in zebra-fish, the presence of low/moderate H3K4me at the affected cohort may render them prone to transcription initiation, requiring DNAme to quell either initiation or elongation of Pol II until the appropriate time in development.
Certain observations in zebrafish DNAme dynamics resemble those in mice. First, sperm has higher DNAme levels than the oocyte (Howlett and Reik, 1991; Kobayashi et al., 2012; Mhanni and McGowan, 2004; Smith et al., 2012), and accordingly, the majority of DMRs are sperm DMRs (methylated only in sperm). Second, the mouse paternal genome displays reductions in bulk DNAme prior to implantation, arriving at levels comparable to the oocyte—a result similar to the 64-cell stage in zebrafish (though pre-ZGA) (Gu et al., 2011; Iqbal et al., 2011; Mayer et al., 2000; Okada et al., 2010; Oswald et al., 2000). Also, the rare oocyte HyperM DMRs in the mouse largely convert to the HypoM status later in development (Kobayashi et al., 2012; Smallwood et al., 2011; Smith et al., 2012), analogous to certain zebrafish loci (Figure 3, clusters 1 and 2). Notably, three characterized nonimprinted parental DMRs (germline DMRs) in the mouse (Dnmt3, Piwi1, and Pou6f) (Borgel et al., 2010; Kobayashi et al., 2012; Smallwood et al., 2011; Smith et al., 2012) are also parental DMRs in the zebrafish. Furthermore, certain early developmental genes acquire DNAme postimplantation, which is consistent conceptually with our results (Borgel et al., 2010; Hawkins et al., 2010; Laurent et al., 2010; Meissner et al., 2008; Mohn et al., 2008; Smith et al., 2012; Weber et al., 2007).
However, there are also major differences in reprogramming between mice and zebrafish. First, in zebrafish, parental DMRs are virtually all resolved prior to ZGA. Second, in zebrafish, this process involves converting virtually all oocyte loci to the DNAme status observed in sperm. Notably, this reprogramming involves the simultaneous demethylation of genes involved in early development/germline, alongside the methylation of genes involved in mid/late development (or oocyte development). Third, in mouse oocytes, most HOX genes and other early developmental regulators are already HypoM at their promoters. Furthermore, changes to both parental genomes occur in distinct phases in the mouse (zygotic, preimplantation, postim-plantation) (Borgel et al., 2010; Smith et al., 2012), with many occurring after ZGA.
Taken together, our results suggest that zebrafish and mice both largely achieve parental genome DNAme equivalence (imprinted genes excepted) but differ in regard to mechanisms and phasing. Zebrafish experience ten cell divisions between fertilization and ZGA, providing ample opportunity to utilize methylation and demethylation mechanisms (passive or active) to achieve parental equivalence. In contrast, the early specification of extraembryonic tissue and inner cell mass in the mouse may require an early ZGA, which is needed to help establish cell asymmetries and developmental decisions. Mice more distinctly separate DNAme reprogramming into phases, with an emphasis on bulk reductions in DNAme during early preimplantation phases and more gene-specific DNAme reprogramming during implantation and postimplantation, involving (for example) the demethylation of key germline genes. In contrast, zebrafish conduct ZGA at a later stage, lack an early trophectoderm decision, lack an implantation phase, and utilize an inherited germ plasm/nuage to define germ cells; thus, they can perform all needed DNAme reprogramming (involving both demethylation and remethylation) prior to ZGA to arrive at parental genome equivalency and a totipotent state at ZGA. During zebrafish ZGA, although chromatin modifications increase (Lindeman et al., 2011; Vastenhouw et al., 2010), almost no DNAme changes occur, which is consistent with prior DNAme grooming for ZGA, creating a transcriptionally active ~1,000 cell totipotent blastula fully competent for development and germ cell specification.
EXPERIMENTAL PROCEDURES
Zebrafish Stocks and Sample Collection
Tübingen zebrafish lines were utilized with standard germ cell and embryo collection procedures (Westerfield, 2000), with exceptional measures taken to assure the purity of oocytes (see Extended Experimental Procedures). Adult fish were euthanized with tricane (Sigma-Aldrich T0377) and, following skin elimination, skeletal muscle was carefully dissected from the trunk area and flash frozen (for DNA extraction) or used immediately (for RNA extraction). For DNA extraction from abundant tissues (sperm, sphere-stage embryos, and muscle), we used Gentra PUREGENE DNA Isolation Kit. DNA isolation from oocytes and early embryos involved nuclei isolation and phenol: chloroform (see Extended Experimental Procedures for details). For sperm, embryos, and adult muscle, 1–3 μg of DNA was sheared on COVARIS Adaptive Focused Acoustics S-series system (Woburn, MA) to a median 400 bp fragment size. Sheared DNA was purified using QIAquick PCR purification kit (QIAGEN, Valencia, CA) or was ethanol precipitated for less abundant DNA samples (64-cell and 256-cell stage). We observed overshearing of oocyte and 2- to 16-cell stage DNA with all modes of shearing (COVARIS, Diagenode, or probe sonication; even at low power/times), requiring the use of pooled enzymatic digestion. DNA from 30,000 oocytes and 15,000 2- to 16-cell stage embryos was equally divided and digested separately with 10 U of AluI, MseI, or MnlI restriction enzymes overnight at 37° C or DNA Fragmentase (NEBM0348S) for 60 min. After digestion, the DNA was combined and purified with phenol:chloroform extraction and ethanol precipitation. See Extended Experimental Procedures for library preparation.
RNA Extraction and Library Preparation
QIAGEN AllPrep DNA/RNA/Protein miniature kit (Catalog number [cat] 80004) was used to extract RNA from zebrafish samples (see Extended Experimental Procedures for details). Total RNA was DNase treated using TURBO DNA-free Kit, Ambion cat AM1907 according to manufacturer's procedure. RNA quality was assessed on Bioanalyzer RNA 6000 Nano Chip. Total RNA was then subjected to RiboMinus treatment (A10837-08, Eukaryote Kit) following manufacturer's procedure. Directional RNA library was performed according to Illumina's protocol and sequenced on a 50 bp single-end run on Illumina HiSeq 2000.
Generation of Maternal Haploid Embryos
Maternal haploid embryos were generated according to standard procedure (Westerfield, 2000) using sperm from “golden” homozygous males gol b1/b1 as a marker (see Extended Experimental Procedures for details). DNA from sphere-stage maternal haploid and sphere-stage control IVF diploids was extracted and bisulfite converted, followed by PCR amplification of 15 promoter regions, library preparation, and sequencing with a 150 bp paired-end format on an Illumina MiSeq.
5-Aza-2′-Deoxycytidine Treatment
1 nl of 1 mM 5-AzadCyD was injected into one-cell stage embryos, which were then incubated in 100 μM 5-AzadCyD until sphere stage, collected, and the RNA extracted. Embryos injected with embryo water served as control. The RNA-seq library was generated as described above.
5hmC Enrichment
A biotin-based enrichment technique (Active Motif, cat 55013) was used to detect the localization of 5-hmC mark in oocyte, 64-cell stage, sphere stage, and muscle followed by library preparation and sequencing on Illumina HiSeq 2000 (see Extended Experimental Procedures for experimental details, library preparation, and analysis).
Bioinformatics Analysis
Illumina Fastq files were aligned using Novoalign (Novocraft). Whole-genome bisulfite sequencing and RNA-seq samples were analyzed using the USEQ package (http://useq.sourceforge.net). Histone modification data were obtained from Lindeman et al. (2011) and Wu et al. (2011a) and reprocessed using the BioToolBox package (http://code.google.com/p/biotoolbox) for the Zv9 (danRer7 at UCSC) genome version. A detailed description of analysis is provided in the Extended Experimental Procedures.
Data Access
All data described in this paper may be downloaded from the Sequence Read Archive under the accession project number SRP020008. This includes raw fastq files for BisSeq, RNA-seq, and 5hmC enrichment experiments. An excel spreadsheet containing processed analysis data sets, complete GO category results, and fastq file information is available in the HCI's GNomEx: https://bioserver.hci.utah.edu/gnomex/gnomexFlex.jsp?topicNumber=10.
Supplementary Material
ACKNOWLEDGMENTS
We thank Brian Dally for genomics advice and Mengyao Tan, Meesha Last, and Minhtu Nguyen for help in oocyte collection. We thank Kazuyiki Hoshijima, Timothy Dahlem, and David Grunwald for homozygous golden fish and for advice regarding maternal haploid fish. We thank Saher Sue Hammoud, Shan-Fu Wu, Rod Stewart, David Jones, and Cairns lab members for advice. This work is supported by NICHD 5R01HD058506 (support of M.E.P. and supplies), HHMI (support of B.R.C. and T.J.P.), NHLBI U01HL0981 (support of T.J.P.), and CA24014 to the Huntsman Cancer Institute (for core facilities).
Footnotes
SUPPLEMENTAL INFORMATION
Supplemental Information includes Extended Experimental Procedures, seven figures, and seven tables and can be found with this article online at http://dx.doi.org/10.1016/j.cell.2013.04.030.
REFERENCES
- Almeida RD, Loose M, Sottile V, Matsa E, Denning C, Young L, Johnson AD, Gering M, Ruzov A. 5-hydroxymethyl-cytosine enrichment of non-committed cells is not a universal feature of vertebrate development. Epigenetics. 2012;7:383–389. doi: 10.4161/epi.19375. [DOI] [PubMed] [Google Scholar]
- Andersen IS, Reiner AH, Aanes H, Aleström P, Collas P. Developmental features of DNA methylation during activation of the embryonic zebrafish genome. Genome Biol. 2012;13:R65. doi: 10.1186/gb-2012-13-7-r65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arpanahi A, Brinkworth M, Iles D, Krawetz SA, Paradowska A, Platts AE, Saida M, Steger K, Tedder P, Miller D. Endonuclease-sensitive regions of human spermatozoal chromatin are highly enriched in promoter and CTCF binding sequences. Genome Res. 2009;19:1338–1349. doi: 10.1101/gr.094953.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bogdanovic O, Long SW, van Heeringen SJ, Brinkman AB, Gómez-Skarmeta JL, Stunnenberg HG, Jones PL, Veenstra GJ. Temporal uncoupling of the DNA methylome and transcriptional repression during embryogenesis. Genome Res. 2011;21:1313–1327. doi: 10.1101/gr.114843.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bogdanovic O, Fernandez-Miñán A, Tena JJ, de la Calle-Mustienes E, Hidalgo C, van Kruysbergen I, van Heeringen SJ, Veenstra GJ, Gómez-Skarmeta JL. Dynamics of enhancer chromatin signatures mark the transition from pluripotency to cell specification during embryogenesis. Genome Res. 2012;22:2043–2053. doi: 10.1101/gr.134833.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borgel J, Guibert S, Li Y, Chiba H, Schübeler D, Sasaki H, Forné T, Weber M. Targets and dynamics of promoter DNA methylation during early mouse development. Nat. Genet. 2010;42:1093–1100. doi: 10.1038/ng.708. [DOI] [PubMed] [Google Scholar]
- Braude P, Bolton V, Moore S. Human gene expression first occurs between the four- and eight-cell stages of preimplantation development. Nature. 1988;332:459–461. doi: 10.1038/332459a0. [DOI] [PubMed] [Google Scholar]
- Brykczynska U, Hisano M, Erkek S, Ramos L, Oakeley EJ, Roloff TC, Beisel C, Schübeler D, Stadler MB, Peters AH. Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nat. Struct. Mol. Biol. 2010;17:679–687. doi: 10.1038/nsmb.1821. [DOI] [PubMed] [Google Scholar]
- Cokus SJ, Feng S, Zhang X, Chen Z, Merriman B, Haudenschild CD, Pradhan S, Nelson SF, Pellegrini M, Jacobsen SE. Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature. 2008;452:215–219. doi: 10.1038/nature06745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farthing CR, Ficz G, Ng RK, Chan CF, Andrews S, Dean W, Hemberger M, Reik W. Global mapping of DNA methylation in mouse promoters reveals epigenetic reprogramming of pluripotency genes. PLoS Genet. 2008;4:e1000116. doi: 10.1371/journal.pgen.1000116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flach G, Johnson MH, Braude PR, Taylor RA, Bolton VN. The transition from maternal to embryonic control in the 2-cell mouse embryo. EMBO J. 1982;1:681–686. doi: 10.1002/j.1460-2075.1982.tb01230.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goll MG, Halpern ME. DNA methylation in zebrafish. Prog. Mol. Biol. Transl. Sci. 2011;101:193–218. doi: 10.1016/B978-0-12-387685-0.00005-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gu TP, Guo F, Yang H, Wu HP, Xu GF, Liu W, Xie ZG, Shi L, He X, Jin SG, et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature. 2011;477:606–610. doi: 10.1038/nature10443. [DOI] [PubMed] [Google Scholar]
- Hajkova P, Ancelin K, Waldmann T, Lacoste N, Lange UC, Cesari F, Lee C, Almouzni G, Schneider R, Surani MA. Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature. 2008;452:877–881. doi: 10.1038/nature06714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hammoud SS, Nix DA, Zhang H, Purwar J, Carrell DT, Cairns BR. Distinctive chromatin in human sperm packages genes for embryo development. Nature. 2009;460:473–478. doi: 10.1038/nature08162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hawkins RD, Hon GC, Lee LK, Ngo Q, Lister R, Pelizzola M, Edsall LE, Kuan S, Luu Y, Klugman S, et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell. 2010;6:479–491. doi: 10.1016/j.stem.2010.03.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffman TL, Javier AL, Campeau SA, Knight RD, Schilling TF. Tfap2 transcription factors in zebrafish neural crest development and ectodermal evolution. J. Exp. Zoolog. B Mol. Dev. Evol. 2007;308:679–691. doi: 10.1002/jez.b.21189. [DOI] [PubMed] [Google Scholar]
- Howlett SK, Reik W. Methylation levels of maternal and paternal genomes during preimplantation development. Development. 1991;113:119–127. doi: 10.1242/dev.113.1.119. [DOI] [PubMed] [Google Scholar]
- Iqbal K, Jin SG, Pfeifer GP, Szabó PE. Reprogramming of the paternal genome upon fertilization involves genome-wide oxidation of 5-methylcytosine. Proc. Natl. Acad. Sci. USA. 2011;108:3642–3647. doi: 10.1073/pnas.1014033108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kane DA, Kimmel CB. The zebrafish midblastula transition. Development. 1993;119:447–456. doi: 10.1242/dev.119.2.447. [DOI] [PubMed] [Google Scholar]
- Knaut H, Pelegri F, Bohmann K, Schwarz H, Nüsslein-Volhard C. Zebrafish vasa RNA but not its protein is a component of the germ plasm and segregates asymmetrically before germline specification. J. Cell Biol. 2000;149:875–888. doi: 10.1083/jcb.149.4.875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kobayashi H, Sakurai T, Imai M, Takahashi N, Fukuda A, Yayoi O, Sato S, Nakabayashi K, Hata K, Sotomaru Y, et al. Contribution of intragenic DNA methylation in mouse gametic DNA methylomes to establish oocyte-specific heritable marks. PLoS Genet. 2012;8:e1002440. doi: 10.1371/journal.pgen.1002440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laurent L, Wong E, Li G, Huynh T, Tsirigos A, Ong CT, Low HM, Kin Sung KW, Rigoutsos I, Loring J, Wei CL. Dynamic changes in the human methylome during differentiation. Genome Res. 2010;20:320–331. doi: 10.1101/gr.101907.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindeman LC, Andersen IS, Reiner AH, Li N, Aanes H, Østrup O, Winata C, Mathavan S, Müller F, Aleström P, Collas P. Prepatterning of developmental gene expression by modified histones before zygotic genome activation. Dev. Cell. 2011;21:993–1004. doi: 10.1016/j.devcel.2011.10.008. [DOI] [PubMed] [Google Scholar]
- Lister R, O'Malley RC, Tonti-Filippini J, Gregory BD, Berry CC, Millar AH, Ecker JR. Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell. 2008;133:523–536. doi: 10.1016/j.cell.2008.03.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo QM, et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature. 2009;462:315–322. doi: 10.1038/nature08514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Long HK, Sims D, Heger A, Blackledge NP, Kutter C, Wright ML, Grutzner F, Odom DT, Patient R, Ponting CP, et al. Epigenetic conservation at gene regulatory elements revealed by non-methylated DNA profiling in seven vertebrates. eLife. 2013;2:e00348. doi: 10.7554/eLife.00348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Macleod D, Clark VH, Bird A. Absence of genome-wide changes in DNA methylation during development of the zebrafish. Nat. Genet. 1999;23:139–140. doi: 10.1038/13767. [DOI] [PubMed] [Google Scholar]
- Martin CC, Laforest L, Akimenko MA, Ekker M. A role for DNA methylation in gastrulation and somite patterning. Dev. Biol. 1999;206:189–205. doi: 10.1006/dbio.1998.9105. [DOI] [PubMed] [Google Scholar]
- Mayer W, Niveleau A, Walter J, Fundele R, Haaf T. Demethylation of the zygotic paternal genome. Nature. 2000;403:501–502. doi: 10.1038/35000656. [DOI] [PubMed] [Google Scholar]
- McGowan RA, Martin CC. DNA methylation and genome imprinting in the zebrafish, Danio rerio: some evolutionary ramifications. Biochem. Cell Biol. 1997;75:499–506. [PubMed] [Google Scholar]
- Meissner A, Mikkelsen TS, Gu H, Wernig M, Hanna J, Sivachenko A, Zhang X, Bernstein BE, Nusbaum C, Jaffe DB, et al. Genomescale DNA methylation maps of pluripotent and differentiated cells. Nature. 2008;454:766–770. doi: 10.1038/nature07107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mhanni AA, McGowan RA. Global changes in genomic methylation levels during early development of the zebrafish embryo. Dev. Genes Evol. 2004;214:412–417. doi: 10.1007/s00427-004-0418-0. [DOI] [PubMed] [Google Scholar]
- Mohn F, Weber M, Rebhan M, Roloff TC, Richter J, Stadler MB, Bibel M, Schübeler D. Lineage-specific polycomb targets and de novo DNA methylation define restriction and potential of neuronal progenitors. Mol. Cell. 2008;30:755–766. doi: 10.1016/j.molcel.2008.05.007. [DOI] [PubMed] [Google Scholar]
- Okada Y, Yamagata K, Hong K, Wakayama T, Zhang Y. A role for the elongator complex in zygotic paternal genome demethylation. Nature. 2010;463:554–558. doi: 10.1038/nature08732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oswald J, Engemann S, Lane N, Mayer W, Olek A, Fundele R, Dean W, Reik W, Walter J. Active demethylation of the paternal genome in the mouse zygote. Curr. Biol. 2000;10:475–478. doi: 10.1016/s0960-9822(00)00448-6. [DOI] [PubMed] [Google Scholar]
- Smallwood SA, Tomizawa S, Krueger F, Ruf N, Carli N, Segonds-Pichon A, Sato S, Hata K, Andrews SR, Kelsey G. Dynamic CpG island methylation landscape in oocytes and preimplantation embryos. Nat. Genet. 2011;43:811–814. doi: 10.1038/ng.864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith ZD, Chan MM, Mikkelsen TS, Gu H, Gnirke A, Regev A, Meissner A. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature. 2012;484:339–344. doi: 10.1038/nature10960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, Yamanaka S. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861–872. doi: 10.1016/j.cell.2007.11.019. [DOI] [PubMed] [Google Scholar]
- Tan L, Shi YG. Tet family proteins and 5-hydroxymethylcytosine in development and disease. Development. 2012;139:1895–1902. doi: 10.1242/dev.070771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thisse B, Pflumio S, Fürthauer M, Loppin B, Heyer V, Degrave A, Woehl R, Lux A, Steffan T, Charbonnier XQ, Thisse C. Expression of the zebrafish genome during embryogenesis (NIH R01 RR15402) 2001 ZFIN Direct Data Submission ( http://zfin.org)
- Vastenhouw NL, Schier AF. Bivalent histone modifications in early embryogenesis. Curr. Opin. Cell Biol. 2012;24:374–386. doi: 10.1016/j.ceb.2012.03.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vastenhouw NL, Zhang Y, Woods IG, Imam F, Regev A, Liu XS, Rinn J, Schier AF. Chromatin signature of embryonic pluripotency is established during genome activation. Nature. 2010;464:922–926. doi: 10.1038/nature08866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weber M, Hellmann I, Stadler MB, Ramos L, Pääbo S, Rebhan M, Schübeler D. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat. Genet. 2007;39:457–466. doi: 10.1038/ng1990. [DOI] [PubMed] [Google Scholar]
- Westerfield M. The Zebrafish Book: A Guide for the Laboratory Use of Zebrafish (Danio rerio) University of Oregon Press; Eugene, OR: 2000. [Google Scholar]
- Wu SF, Zhang H, Cairns BR. Genes for embryo development are packaged in blocks of multivalent chromatin in zebrafish sperm. Genome Res. 2011a;21:578–589. doi: 10.1101/gr.113167.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu SF, Zhang H, Hammoud SS, Potok M, Nix DA, Jones DA, Cairns BR. DNA methylation profiling in zebrafish. Methods Cell Biol. 2011b;104:327–339. doi: 10.1016/B978-0-12-374814-0.00018-5. [DOI] [PubMed] [Google Scholar]
- Zhou VW, Goren A, Bernstein BE. Charting histone modifications and the functional organization of mammalian genomes. Nat. Rev. Genet. 2011;12:7–18. doi: 10.1038/nrg2905. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.