Abstract
Objective
To determine if functional polymorphisms of folate/homocysteine pathway enzymes are associated with homocysteine concentrations and/or coronary artery calcification (CAC) scores in patients with systemic lupus erythematosus (SLE) and controls.
Methods
We investigated 163 SLE patients and 160 controls. Functional polymorphisms in 6 genes in the folate/homocysteine pathway were genotyped: 5,10-methylenetetrahydrofolate reductase (MTHFR) 677C>T, MTHFR 1298A>C, cystathionine ß-synthase (CBS) 844ins68, methionine synthase (MTR) 2756A>G, methionine synthase reductase (MTRR) 66A>G, thymidylate synthase (TYMS) 1494del6, and dihydrofolate reductase (DHFR) c.86+60_78.
Results
Homocysteine levels were higher in African American SLE patients than Caucasian patients and African American controls. Genotype distributions were significantly different in African American and Caucasian controls for 6 of the 7 polymorphisms. Genotype distributions for each polymorphism did not differ significantly between SLE patients and controls even after stratification by race. Glomerular filtration rate was strongly negatively correlated to homocysteine levels, and was therefore adjusted for as a covariate in the models of the effects of the polymorphisms on homocysteine levels. In SLE patients none of the 7 polymorphisms was associated with homocysteine concentrations. In Caucasian controls only MTHFR 677C>T and 1298A>C showed effects on homo-cysteine similar to what would be expected from the literature. There were no genotypic associations with median CAC scores in SLE patients or controls with and without stratification by race.
Conclusion
Polymorphisms in folate/homocysteine metabolizing enzymes do not predict higher homocysteine levels or CAC scores in patients with SLE.
Keywords: HOMOCYSTEINE, GENETIC POLYMORPHISMS, ATHEROSCLEROSIS, SYSTEMIC LUPUS ERYTHEMATOSUS, CORONARY ARTERY CALCIFICATION
Systemic lupus erythematosus (SLE) is a chronic inflammatory disease that occurs predominantly in women, and more commonly among African Americans1. Individuals with SLE have a greater risk of developing premature atheroscle rotic cardiovascular disease (ASCVD) than the general population2. Subclinical ASCVD can be measured by electron-beam computed tomography (EBCT), which is used to detect coronary artery calcification (CAC). CAC scores are related to risk of future cardiovascular events3.
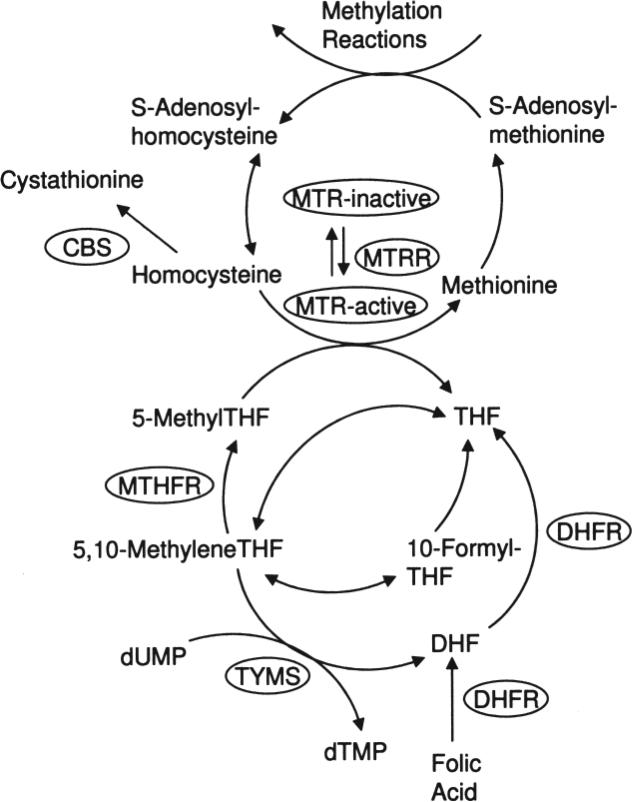
Atherosclerosis is a complex process and high homocysteine concentration is one component of that process. High circulating homocysteine concentrations are known to be associated with an increased risk of ASCVD in the general population4. Folate/homocysteine metabolism is important for DNA synthesis and for generating 5-methyltetrahydrofo-late, the source of methyl groups that are ultimately used in many methylation reactions (Figure 1). Homocysteine concentrations can be modified by several factors including dietary intake of folate and other B vitamins, lifestyle variables, and genetic polymorphisms in the enzymes of the folate/homocysteine pathway5. One of the most widely studied such enzymes is 5,10-methylenetetrahydrofolate reductase (MTHFR; EC 1.5.1.20), which converts 5,10-methylenetetrahydrofolate to 5-methyltetrahydrofolate that is in turn used in the remethylation of homocysteine to methion ine by methionine synthase (MTR; EC 2.1.1.13). The latter reaction requires methionine synthase reductase (MTRR; EC 2.1.1.135) to maintain the MTR cofactor cobalamin in its active form. The vitamin B6-dependent enzyme cystathionine ß-synthase (CBS; EC 4.2.1.22) irreversibly metabolizes homocysteine to cystathionine in the initial step of the transsulfuration pathway. Thymidylate synthetase (TYMS; EC 2.1.1.45) uses 5,10-methylenetetrahydrofolate to convert deoxyuridine monophosphate (dUMP) to deoxythymidine monophosphate (dTMP). Dihydrofolate reductase (DHFR; EC 1.5.1.3) catalyzes the conversion of dihydrofolate to tetrahydrofolate and is also important for converting synthetic folic acid from vitamin supplements and fortified food into a biologically relevant derivative that can be used by cells. All these enzymes play important roles in maintaining a healthy cell phenotype, and dysfunction in any could potentially cause atherogenic changes. This is evidenced by patients with homocystinuria due to CBS deficiency that have premature atherosclerosis and thrombosis6. All these enzymes have common functional polymorphisms (MTHFR 677C>T and 1298A>C, MTR 2756A>G, MTRR 66A>G, CBS 844ins68, TYMS 1494del6, and DHFR c.86+60_78) that are known to affect homocysteine concentrations in some populations7-16.
Figure 1.
The folate/homocysteine pathway. CBS: cystathionine ß-synthase; DHF: dihydrofolate; DHFR: dihydrofolate reductase; dTMP: deoxythymidine monophosphate; dUMP: deoxyuridine monophosphate; MTHFR: 5,10-methylenetetrahydrofolate reductase; MTR: methionine synthase; MTRR: methionine synthase reductase; THF: tetrahydrofolate; TYMS: thymidylate synthase.
Patients with SLE have elevated homocysteine levels compared to matched controls17-20, and high homocysteine in such patients is associated with the presence of CAC19. The extent, if any, to which polymorphisms in the above enzymes contribute to high homocysteine and CAC is unknown. We investigated to determine whether polymorphisms in key enzymes of the folate/homocysteine pathway are associated with increased homocysteine levels and/or CAC scores in patients with SLE.
MATERIALS AND METHODS
SLE and control subjects
Consecutive nonpregnant women over 18 years of age with SLE attending the University of Pennsylvania clinics and female controls matched for race and age (± 2 yrs) were invited to participate in the study; 163 patients and 160 controls were enrolled. SLE patients were required to fulfill at least 4 of the American College of Rheumatology revised criteria for the classification of SLE21. Controls were matched to patients with regard to key demographic features, and biochemical analyses indicated that they had similar traditional cardiovascular risk factor profiles but were otherwise healthy with no underlying inflammatory disease or exposure to corticosteroids. The study was approved by the University of Pennsylvania Institutional Review Board, and written informed consent was obtained from each participant.
Clinical assessments
A medical history was collected for all subjects. All participants had a physical examination and electrocardiography, gave a fasting blood sample, and underwent EBCT19. Clinical characteristics were as reported19,22. Framingham point scores were calculated based on a published method of scoring various cardiovascular risk factors23. Glomerular filtration rate (GFR) was calculated using the Modification of Diet in Renal Disease equation24.
Homocysteine measurement
Whole blood (5 ml) was drawn into EDTA and placed on ice until centrifuged at 2500 rpm for 5 min at room temperature. Plasma homocysteine concentrations were determined by fluorescence polarization immunoassay (AxSYM Homocysteine; Abbott Laboratories, Abbott Park, IL, USA). This assay was performed by the hospital's clinical laboratory, which used Abbott's homocysteine low and high controls daily and participated in CAP proficiency and linearity studies.
Genetic analysis
DNA was isolated using Generation Capture Column Kits (Gentra Systems, Minneapolis, MN, USA). MTHFR 677C>T (rs1801133) genotypes were analyzed using a heteroduplex generator method25, and a portion of samples were repeated using the TaqMan assay. MTHFR 677C>T, MTHFR 1298A>C (rs1801131), MTR 2756A>G (rs1805087), and MTRR 66A>G (rs1801394) were genotyped by TaqMan real-time polymerase chain reaction (PCR) assays on a DNA Engine Opticon 2 continuous fluorescence detection system (Bio-Rad, Hercules, CA, USA). PCR amplifications were performed using 20 ng genomic DNA with forward and reverse primers, allele-specific probes (at concentrations listed in Table 1), and TaqMan Universal PCR MasterMix (Applied Biosystems, Foster City, CA, USA). Some primer and probe sequences were derived from the SNP500Cancer website as designated in Table 126. MGB probes were custom-synthesized by Applied Biosystems. Each PCR was performed in 20 μl with an initial incubation at 50°C for 2 min, then 95°C for 10 min, followed by 50 cycles of denaturation and extension under the conditions listed in Table 1. Dual fluorescence was detected after each extension step. Genotype interpretations were performed using Opticon Monitor Analysis software, version 2.02 (Bio-Rad).
Table 1.
PCR primers, probes, concentrations, and conditions. Polymorphic site underlined in probe sequences.
| Gene and Polymorphism (dbSNP RS no.) | Primers and Probes, 5’–3’ Sequence | Concentration | Conditions |
|---|---|---|---|
| TaqMan | |||
| MTHFR 677C > T (1801133) | F: GCA GGG AGC TTT GAG GCT GAC C | 0.5 μM | 92°C 30 s |
| R: TGG GGC AAG TGA TGC CCA TGT | 0.5 μM | 56°C 1 min | |
| *T: 6FAM-ATG AAA TCG <U>A</U>CT CCC GC-MGBNFQ | 50 μM | 50 cycles | |
| *C: VIC-ATG AAA TCG <U>G</U>CT CCC GC-MGBNFQ | 100 μM | ||
| MTHFR 1298A > C (1801131) | F: GAG GAG CTG CTG AAG ATG T | 0.5 μM | 92°C 30 s |
| R: CGA GAG GTA AAG AAC GAA GA | 0.5 μM | 56°C 1 min | |
| *C: 6FAM-AGA CAC TT<U>G</U> CTT CAC T-MGBNFQ | 50 μM | 50 cycles | |
| *A: VIC-CAA AGA CAC TT<U>T</U> CTT C-MGBNFQ | 50 μM | ||
| MTR 2756A > G (1805087) | F: AGT GTT CCC AGC TGT TAG ATG A | 0.5 μM | 92°C 30 s |
| *R: TGT TTC TAC CAC TTA CCT TGA GAG ACT | 0.5 μM | 60°C 1 min | |
| *G: 6FAM-ACA GG<U>G</U> CCA TTA TG-MGBNFQ | 50 μM | 50 cycles | |
| *A: VIC-ATT AGA CAG G<U>A</U>C CAT TAT G-MGBNFQ | 100 μM | ||
| MTRR 66A > G (1801394) | *F: CAT GCC TTG AAG TGA TGA GG | 0.5 μM | 92°C 30 s |
| *R: GAT CTG CAG AAA ATC CAT GTA CCA | 0.5 μM | 60°C 1 min | |
| *G: 6FAM-CTT GCT CAC A<U>C</U>A TTT-MGBNFQ | 50 μM | 50 cycles | |
| *A: VIC-TGC TCA CA<U>T</U> ATT TC-MGBNFQ | 100 μM | ||
| Size Difference PCR | |||
| CBS 844ins68 | F: TAT TGG CCA CTC CCA TAA TAG A | 0.4 μM | 94°C 5 min |
| R: CGG CTC TGC GAG GAT GGA CCC TT | 0.4 μM | 35 cycles of | |
| 94°C 1 min | |||
| 55°C 1 min | |||
| 72°C 1 min | |||
| TYMS 1494de16 (16430) | F: CAT GAT GTA GAG TGT GGT TAT G | 0.4 μM | 94°C 2 min |
| R: GAA TGA ACA AAG CGT GGA | 0.4 μM | 35 cycles of | |
| 94°C 30 s | |||
| 51 °C 30 s | |||
| 72°C 30s then | |||
| 72°C 5 min | |||
| DHFR c.86+60_78 | F1: CCA CGG TCG GGG TAC CTG GG | 0.4 μM | 94°C 4 min |
| F2: ACG GTC GGG GTG GCC GAC TC | 0.4 μM | 35 cycles of | |
| R: AAA AGG GGA ATC CAG TCG G | 0.8 μM | 94°C 55s | |
| 62°C 55 s | |||
| 72°C 55 s | |||
| then 72°C 12 min |
Originated from SNP500Cancer website26.
Size-difference PCR methods were used to genotype CBS 844ins6825, TYMS 1494del6 (rs16430), and DHFR c.86+60_7827. PCR amplifications took place in 25 μl volumes that contained 50 ng genomic DNA, 0.4 μM of each forward and reverse primer (0.8 μM reverse primer for DHFR assay), 0.8 μM dNTPs, 10× PCR buffer (Applied Biosystems), 1.5 mM MgCl2, and 1 U AmpliTaq DNA polymerase (Applied Biosystems). Cycling conditions are listed in Table 1. PCR products were separated on 3% agarose gels, run at 140 V for 45 min, and stained with ethidium bromide.
Statistical analysis
SAS version 9.1 was used for all statistical analysis with Type I error rate set to 0.05. Homocysteine was log-transformed to better approximate normality in all analyses. Hardy-Weinberg equilibrium for each of the genotypes was assessed by chi-square test. Differences in genotype frequency distributions between African American and Caucasian controls and between case and control groups were assessed by chi-square and Fisher's exact test. Correlations with log homocysteine were assessed by Pearson's correlation coefficients for age, GFR, and Framingham point scores. Student's t-test was used for assessment of smoking status and use of B6, B12, and folic acid on log homocysteine. General linear modeling was used to assess the effect of the above correlated variables on homocysteine. When race was used as a classification variable in the models, this included 127 Caucasian and 163 African American SLE cases and controls. Any variable that significantly contributed to the model was used as a covariate in assessment of the effect of genotype on log homocysteine as well as an interaction term with genotype to assess effect modification. Results of log homocysteine analyses were back-transformed to report results in original measurement units (μmol/l).
RESULTS
Sample characteristics
The mean age of SLE patients was 43.3 ± 11.0 years (Table 2). The race distribution was 50.3% African American, 39.3% Caucasian, 4.9% Asian, 4.3% Hispanic, and 1.2% other, with controls having a similar distribution. Median CAC scores were significantly higher in SLE patients than controls (p = 0.0003). Homocysteine concentrations were also higher in patients than controls (10.4 vs 9.2 μmol/l; p < 0.0001). African American SLE patients had higher homocysteine levels than African American controls (12.1 vs 9.7 μmol/l, p < 0.0001), while Caucasian SLE patients did not differ significantly from Caucasian controls (10.0 vs 9.0 μmol/l, p = 0.12). African American controls did not differ significantly from Caucasian controls in homo-cysteine concentrations (p = 0.21), but African American SLE patients had higher homocysteine levels than Caucasian patients (p = 0.0009). SLE patients did not differ significantly from controls in terms of Framingham point scores or GFR.
Table 2.
Sample characteristics.
| Characteristic | SLE Patients | Controls | p |
|---|---|---|---|
| No. | 163 | 160 | ND |
| Age, yrs, mean ± SD | 43.3 ± 11.0 | 43.5 ± 10.5 | 0.90 |
| Race, % (n) | |||
| African American | 50.3 (82) | 50.6 (81) | 1.00 |
| Caucasian | 39.3 (64) | 39.4 (63) | |
| Asian | 4.9 (8) | 5.0 (8) | |
| Hispanic | 4.3 (7) | 3.8 (6) | |
| Other | 1.2 (2) | 1.2 (2) | |
| CAC, median (IQR) | 0 (0–7.2) | 0 (0–0) | 0.0003 |
| Homocysteine, μmol/l, mean ± SD (n) | 10.4 ± 1.4 | 9.2 ± 1.4 | < 0.0001 |
| African American* | 12.1 ± 1.4 (82) | 9.7 ± 1.3 (81) | < 0.0001 |
| Caucasian | 10.0 ± 1.3 (64) | 9.0 ± 1.5 (63) | 0.12 |
| Framingham point scores, median (IQR) (n) | 8 (3–13) (161) | 8 (2–12) (156) | 0.39 |
| GFR, ml/min/1.73m2, mean ± SD | 90.0 ± 32.5 | 94.5 ± 20.8 | 0.14 |
CAC: coronary artery calcification; GFR: glomerular filtration rate. ND: not determined; IQR: interquartile range.
African American case vs Caucasian case, p = 0.0009; African American control vs Caucasian control, p = 0.21.
Genotype frequency distributions of African American and Caucasian controls
SLE patients and controls were geno-typed for 7 polymorphisms in 6 enzymes in the folate/homo-cysteine pathway (MTHFR 677C>T and 1298A>C, MTR 2756A>G, MTRR 66A>G, CBS 844ins68, TYMS 1494del6, and DHFR c.86+60_78). All genotypes were in Hardy-Weinberg equilibrium for SLE patients and controls in the total study population and after stratification by race (data not shown). The genotype frequency distributions were significantly different between African American and Caucasian controls for all of the polymorphisms except for MTR 2756A>G by chi-square test (Table 3).
Table 3.
Distribution of genotype frequencies in African American and Caucasian controls.
| Polymorphism | Genotype | African American | Caucasians | p |
|---|---|---|---|---|
| MTHFR 677C>T | CC | 67.9 (55) | 39.7 (25) | 0.0003 |
| CT | 29.6 (24) | 41.3 (26) | ||
| TT | 2.5 (2) | 19.0 (12) | ||
| MTHFR 1298A>C | AA | 69.1 (56) | 55.6 (35) | 0.0347 |
| AC | 30.9 (25) | 38.1 (24) | ||
| CC | 0 | 6.3 (4) | ||
| MTHFR 677/1298 | CC/AA | 46.9 (38) | 11.1 (7) | < 0.0001 |
| CC/AC | 21.0 (17) | 22.2 (14) | ||
| CC/CC | 0 | 6.3 (4) | ||
| CT/AA | 19.7 (16) | 25.4 (16) | ||
| CT/AC | 9.9 (8) | 15.9 (10) | ||
| TT/AA | 2.5 (2) | 19.0 (12) | ||
| CBS 844ins68 | WW | 58.0 (47) | 87.3 (55) | 0.0005 |
| WI | 39.5 (32) | 12.7 (8) | ||
| II | 2.5 (2) | 0 | ||
| MTR 2756A>G | AA | 58.0 (47) | 47.6 (30) | 0.19 |
| AG | 33.3 (27) | 47.6 (30) | ||
| GG | 8.7 (7) | 4.8 (3) | ||
| MTRR 66A>G | AA | 48.2 (39) | 20.6 (13) | 0.0009 |
| AG | 40.7 (33) | 50.8 (32) | ||
| GG | 11.1 (9) | 28.6 (18) | ||
| TYMS 1494del6 | Ins/ins | 19.7 (16) | 39.7 (25) | 0.0012 |
| Ins/del | 53.1 (43) | 54.0 (34) | ||
| Del/del | 27.2 (22) | 6.3 (4) | ||
| DHFR c.86+60_78 | Ins/ins | 19.7 (16) | 38.1 (24) | 0.0068 |
| Ins/del | 45.7 (37) | 47.6 (30) | ||
| Del/del | 34.6 (28) | 14.3 (9) |
Genotype frequencies % (n). P values by chi-square test.
SLE and polymorphisms of folate/homocysteine-metabolizing enzymes
Neither the genotype frequency distributions of the 7 polymorphisms (data not shown) nor the carrier frequency distributions of each individual polymorphism stratified by race (Table 4) differed between SLE patients and controls by chi-square or Fisher's exact tests.
Table 4.
Distributions of carrier frequencies between SLE patients and controls.
| Genotype | SLE Patients | Controls | p |
|---|---|---|---|
| MTHFR 677T carriers | |||
| African American | 31.7 (26) | 32.1 (26) | 1.00 |
| Caucasian | 65.6 (42) | 60.3 (38) | 0.58 |
| MTHFR 1298C carriers | |||
| African American | 32.9 (27) | 30.9 (25) | 0.87 |
| Caucasian | 54.7 (35) | 44.4 (28) | 0.29 |
| CBS 844ins68 carriers | |||
| African American | 48.8 (40) | 42.0 (34) | 0.43 |
| Caucasian | 15.6 (10) | 12.7 (8) | 0.80 |
| MTR 2756G carriers | |||
| African American | 46.3 (38) | 42.0 (34) | 0.64 |
| Caucasian | 35.9 (23) | 52.4 (33) | 0.07 |
| MTRR 66G carriers | |||
| African American | 43.9 (36) | 51.9 (42) | 0.35 |
| Caucasian | 78.1 (50) | 79.4 (50) | 1.00 |
| TYMS 1494del6 ins carriers | |||
| African American | 63.4 (52) | 72.8 (59) | 0.24 |
| Caucasian | 89.1 (57) | 93.7 (59) | 0.53 |
| DHFR c.86+60_78 ins carriers | |||
| African American | 59.8 (49) | 65.4 (53) | 0.52 |
| Caucasian | 78.1 (50) | 85.7 (54) | 0.36 |
Carriers of alleles subset by race % (n). P values by Fisher's exact test.
Predictors of homocysteine levels
The continuous variables selected for correlation analysis with homocysteine were age, GFR, and Framingham point scores. Pearson coefficients and p values are given in Table 5; all 3 continuous variables were significantly correlated with homocysteine. Age and Framingham point scores were positively correlated while GFR was negatively correlated with homocysteine levels. Categorical variables selected for analysis by Student t-test for independent samples were folic acid use, B6 use, B12 use, and smoking status, which were uncontrolled sources of variation. None of the variables were significantly associated with homocysteine (p > 0.05). The results were similar when stratified by group (SLE patients vs controls) except for smoking status, which was significantly associated with mean homocysteine only in controls [smokers 11.8 ± 1.3 (n = 25) vs nonsmokers 9.5 ± 1.4 (n = 133); p = 0.0046].
Table 5.
Correlations with homocysteine levels.
| Variable | Pearson Correlation Coefficient | n | p |
|---|---|---|---|
| Age | 0.22917 | 323 | < 0.0001 |
| GFR | –0.42930 | 323 | < 0.0001 |
| Framingham point scores | 0.22249 | 317 | < 0.0001 |
GFR: glomerular filtration rate.
As discussed above there were differences in homocysteine levels by group and by race, which consisted of African Americans and Caucasians. From the above analysis the explanatory variables age, GFR, and Framingham point scores along with race and group were put into a general linear model with homocysteine as the dependent variable, with no interactions between any of the terms. GFR along with group and race were the only variables significantly associated with homocysteine (Table 6). These variables were used in the modeling of the effect of genotype on homocysteine.
Table 6.
Analysis of covariance for homocysteine.
| Source Variation | Sum of Squares | Df | F | p |
|---|---|---|---|---|
| GFR | 1.11 | 1 | 60.18 | < 0.0001 |
| Group* | 5.53 | 1 | 12.04 | 0.0006 |
| Race† | 2.14 | 1 | 23.28 | < 0.0001 |
Case and control groups.
African Americans and Caucasians.
Modeling the effects of polymorphisms of folate/homocysteine-metabolizing enzymes on homocysteine concentrations
After exploring more complex models we arrived at a clinically sound parsimonious model with variables that significantly contributed to the variation in homocysteine. A general linear model with homocysteine as the dependent variable and classified by race, group, and genotype and all possible interactions with the addition of the covariate GFR and its interaction with genotype was used for each polymorphism based on the above analysis, which determined these variables to be significantly associated with homocysteine concentrations. The 4 categories analyzed were African American SLE patients, African American controls, Caucasian SLE patients, and Caucasian controls. The least-square mean estimates of homocysteine were adjusted for the uncontrolled variable GFR. Two of the polymorphisms (MTHFR 1298 and CBS 844ins68) were modeled based on the combination of the heterozygotes with the homozygotes because the homozygotes for the polymorphism were not present in at least one of the 4 categories. Out of the 7 polymorphisms only 2 had significant results, both in Caucasian controls (Table 7). For MTHFR 677 CC vs CT and CC vs TT the homocysteine concentrations were 7.7 versus 9.4 (p = 0.0196) and 7.7 versus 9.8 μmol/l (p = 0.0275), respectively. For MTHFR 1298 AA versus AC/CC the homocysteine concentrations were 9.6 versus 7.8 μmol/l (p = 0.0083), respectively.
Table 7.
Analysis of covariance with 2 factors (group, race) and one covariate (glomerular filtration rate).
| Category | Genotype | Adjusted Means of Homocysteine | p* |
| Caucasian controls | MTHFR 677 CC | 7.7 | — |
| CT | 9.4 | 0.0196 | |
| TT | 9.8 | 0.0275 | |
| Caucasian controls | MTHFR 1298 AA | 9.6 | — |
| AC/CC† | 7.8 | 0.0083 |
p value for comparison to wild-type genotype.
These genotypes were combined because of low numbers of homozygotes.
CAC scores and polymorphisms of folate/homocysteine-metabolizing enzymes
None of the genotypes under test was associated with median CAC scores by Kruskal-Wallis test even after stratification by race (data not shown).
DISCUSSION
Our a priori hypothesis was that established functional polymorphisms of enzymes in the folate/homocysteine pathway would be associated with increased homocysteine concentrations in patients with SLE, and hence with CAC scores, a clinical finding for which elevated homocysteine concentrations are predictive19. The polymorphisms selected for this study (MTHFR 677C>T and 1298A>C, MTR 2756A>G, MTRR 66A>G, CBS 844ins68, TYMS 1494del6, and DHFR c.86+60_78) were those for which there is evidence for significant effects on homocysteine concentrations7-16. The differences in the distributions of genotype frequencies between African American and Caucasian controls mandated that associations of homocysteine with genotype be stratified by race. More specifically, it is known that MTHFR 677C>T and 1298A>C28, MTRR 66A>G28, CBS 844ins6829,30, and TYMS 1494del630 have different genotype frequencies in African Americans and Caucasians. Our study also found that frequencies of DHFR c.86+60_78 differed between races.
None of the genotypes for the polymorphisms under test differed significantly in distribution between SLE patients and controls, even after stratification by race, indicating that none are genetic risk factors for SLE per se. A study by Fijnheer, et al31 found that MTHFR 677C>T did not explain elevated homocysteine levels in SLE patients, which agrees with our findings. Our results contrast with a smaller Italian study reporting that SLE patients had a higher prevalence of the MTHFR 677TT genotype18. A Polish study found that frequencies of MTHFR 677C>T were not different between SLE patients and controls, but the authors found that the MTR 2756G allele was overrepresented in SLE patients32, which contrasts with our finding of lack of an association.
Although folate, B12, and B6 levels were not available for our study, use of folic acid, B12, and B6 supplements was studied as a surrogate, but was found to be not associated with homocysteine concentrations. GFR was correlated negatively with homocysteine, and other studies have found a similar relationship33. In SLE patients none of the 7 polymorphisms were associated with homocysteine levels. In controls there were 2 polymorphisms associated with homocysteine levels, but only in Caucasians. The MTHFR 677CT and TT genotypes were associated with an increase in homocysteine levels compared to CC, as expected from the literature7,8. Carriers of the MTHFR 1298C allele had lower homocysteine levels compared to AA. This finding is concordant with a study by Parle-McDermott, et al that found that MTHFR 1298AC and CC genotypes were associated with increased red-cell folate and a nonsignificant decrease in homocysteine within the MTHFR 677CC genotype in pregnant women34. A large study by Ulvik, et al used an approach similar to the Parle-McDermott study and stratified their analysis of biochemical variables based on both MTHFR 677 and 1298 genotypes. Ulvik, et al found that MTHFR 1298AC and CC genotypes were associated with higher homocysteine levels and lower serum folate levels9. The differences in findings among the studies on MTHFR 1298A>C may be due to the number of subjects studied or to the genetic variability of the populations studied.
Although our study has some limitations, the magnitude of the difference in homocysteine concentrations between SLE patients and controls was large enough to suggest that genetic factors might be responsible, at least in part, for the elevation of homocysteine in the patients. Such a genetic effect (i.e., the MTHFR 677C>T polymorphism) was observed in the controls, but not in patients with SLE. While we acknowledge that the size of our study population precludes a conclusion that there are no contributing genetic factors, our study suggests that if such factors are involved they are likely to have no more than a relatively small effect. Thus, the increase in homocysteine among SLE patients is probably due primarily to other variables that are components of the SLE disease process itself.
African American patients with SLE had elevated homo-cysteine levels compared to Caucasian patients and African American controls. None of the tested functional polymorphisms of enzymes in the folate/homocysteine pathway were associated with SLE. In addition, in SLE patients, none of the polymorphisms were associated with homocysteine concentrations even when adjusted for covariates, including GFR, and there were no associations with median CAC scores. It is unlikely that polymorphisms in folate/homocysteine-metabolizing enzymes contribute substantially to the elevated homocysteine levels observed in patients with SLE. Therefore the mechanism whereby SLE patients achieve elevated homocysteine concentrations and high CAC scores relative to controls is most likely due to inflammatory aspects of the disease process that dominate any genetic effects intrinsic to enzymes of the folate/homocysteine pathway.
Acknowledgments
Supported by National Institutes of Health grants AR47663 and ES013508 (A.S. Whitehead), by grants from the Lupus Research Institute (J.M. Von Feldt), and by the CTSA funded Clinical and Translational Research Center grant UL1RR024134 from the National Center for Research Resources.
REFERENCES
- 1.Simard JF, Costenbader KH. What can epidemiology tell us about systemic lupus erythematosus? Int J Clin Pract. 2007;61:1170–80. doi: 10.1111/j.1742-1241.2007.01434.x. [DOI] [PubMed] [Google Scholar]
- 2.Manzi S, Meilahn EN, Rairie JE, et al. Age-specific incidence rates of myocardial infarction and angina in women with systemic lupus erythematosus: comparison with the Framingham Study. Am J Epidemiol. 1997;145:408–15. doi: 10.1093/oxfordjournals.aje.a009122. [DOI] [PubMed] [Google Scholar]
- 3.Arad Y, Goodman KJ, Roth M, Newstein D, Guerci AD. Coronary calcification, coronary disease risk factors, C-reactive protein, and atherosclerotic cardiovascular disease events: the St. Francis Heart Study. J Am Coll Cardiol. 2005;46:158–65. doi: 10.1016/j.jacc.2005.02.088. [DOI] [PubMed] [Google Scholar]
- 4.Graham IM, Daly LE, Refsum HM, et al. Plasma homocysteine as a risk factor for vascular disease. The European Concerted Action Project. JAMA. 1997;277:1775–81. doi: 10.1001/jama.1997.03540460039030. [DOI] [PubMed] [Google Scholar]
- 5.Schneede J, Refsum H, Ueland PM. Biological and environmental determinants of plasma homocysteine. Semin Thromb Hemost. 2000;26:263–79. doi: 10.1055/s-2000-8471. [DOI] [PubMed] [Google Scholar]
- 6.Mudd SH, Skovby F, Levy HL, et al. The natural history of homocystinuria due to cystathionine beta-synthase deficiency. Am J Hum Genet. 1985;37:1–31. [PMC free article] [PubMed] [Google Scholar]
- 7.Harmon DL, Woodside JV, Yarnell JW, et al. The common ‘thermolabile’ variant of methylene tetrahydrofolate reductase is a major determinant of mild hyperhomocysteinaemia. QJM. 1996;89:571–7. doi: 10.1093/qjmed/89.8.571. [DOI] [PubMed] [Google Scholar]
- 8.Jacques PF, Bostom AG, Williams RR, et al. Relation between folate status, a common mutation in methylenetetrahydrofolate reductase, and plasma homocysteine concentrations. Circulation. 1996;93:7–9. doi: 10.1161/01.cir.93.1.7. [DOI] [PubMed] [Google Scholar]
- 9.Ulvik A, Ueland PM, Fredriksen A, et al. Functional inference of the methylenetetrahydrofolate reductase 677C > T and 1298A > C polymorphisms from a large-scale epidemiological study. Hum Genet. 2007;121:57–64. doi: 10.1007/s00439-006-0290-2. [DOI] [PubMed] [Google Scholar]
- 10.Dekou V, Gudnason V, Hawe E, Miller GJ, Stansbie D, Humphries SE. Gene-environment and gene-gene interaction in the determination of plasma homocysteine levels in healthy middle-aged men. Thromb Haemost. 2001;85:67–74. [PubMed] [Google Scholar]
- 11.Harmon DL, Shields DC, Woodside JV, et al. Methionine synthase D919G polymorphism is a significant but modest determinant of circulating homocysteine concentrations. Genet Epidemiol. 1999;17:298–309. doi: 10.1002/(SICI)1098-2272(199911)17:4<298::AID-GEPI5>3.0.CO;2-V. [DOI] [PubMed] [Google Scholar]
- 12.Fredriksen A, Meyer K, Ueland PM, Vollset SE, Grotmol T, Schneede J. Large-scale population-based metabolic phenotyping of thirteen genetic polymorphisms related to one-carbon metabolism. Hum Mutat. 2007;28:856–65. doi: 10.1002/humu.20522. [DOI] [PubMed] [Google Scholar]
- 13.Gaughan DJ, Kluijtmans LA, Barbaux S, et al. The methionine synthase reductase (MTRR) A66G polymorphism is a novel genetic determinant of plasma homocysteine concentrations. Atherosclerosis. 2001;157:451–6. doi: 10.1016/s0021-9150(00)00739-5. [DOI] [PubMed] [Google Scholar]
- 14.Gaughan DJ, Kluijtmans LA, Barbaux S, et al. Corrigendum to “The methionine synthase reductase (MTRR) A66G polymorphism is a novel genetic determinant of plasma homocysteine concentrations”. Atherosclerosis. 2002;167:373. doi: 10.1016/s0021-9150(00)00739-5. [Atherosclerosis 2001;157:451-6] [DOI] [PubMed] [Google Scholar]
- 15.Kealey C, Brown KS, Woodside JV, et al. A common insertion/deletion polymorphism of the thymidylate synthase (TYMS) gene is a determinant of red blood cell folate and homocysteine concentrations. Hum Genet. 2005;116:347–53. doi: 10.1007/s00439-004-1243-2. [DOI] [PubMed] [Google Scholar]
- 16.Gellekink H, Blom HJ, van der Linden IJ, den Heijer M. Molecular genetic analysis of the human dihydrofolate reductase gene: relation with plasma total homocysteine, serum and red blood cell folate levels. Eur J Hum Genet. 2007;15:103–9. doi: 10.1038/sj.ejhg.5201713. [DOI] [PubMed] [Google Scholar]
- 17.Bruce IN, Urowitz MB, Gladman DD, Ibanez D, Steiner G. Risk factors for coronary heart disease in women with systemic lupus erythematosus: the Toronto Risk Factor Study. Arthritis Rheum. 2003;48:3159–67. doi: 10.1002/art.11296. [DOI] [PubMed] [Google Scholar]
- 18.Afeltra A, Vadacca M, Conti L, et al. Thrombosis in systemic lupus erythematosus: congenital and acquired risk factors. Arthritis Rheum. 2005;53:452–9. doi: 10.1002/art.21172. [DOI] [PubMed] [Google Scholar]
- 19.Von Feldt JM, Scalzi LV, Cucchiara AJ, et al. Homocysteine levels and disease duration independently correlate with coronary artery calcification in patients with systemic lupus erythematosus. Arthritis Rheum. 2006;54:2220–7. doi: 10.1002/art.21967. [DOI] [PubMed] [Google Scholar]
- 20.Lee AB, Godfrey T, Rowley KG, et al. Traditional risk factor assessment does not capture the extent of cardiovascular risk in systemic lupus erythematosus. Intern Med J. 2006;36:237–43. doi: 10.1111/j.1445-5994.2006.01044.x. [DOI] [PubMed] [Google Scholar]
- 21.Hochberg MC. Updating the American College of Rheumatology revised criteria for the classification of systemic lupus erythematosus [letter]. Arthritis Rheum. 1997;40:1725. doi: 10.1002/art.1780400928. [DOI] [PubMed] [Google Scholar]
- 22.Brown KS, Nackos E, Morthala S, Jensen LE, Whitehead AS, Von Feldt JM. Monocyte chemoattractant protein-1: plasma concentrations and A(-2518)G promoter polymorphism of its gene in systemic lupus erythematosus. J Rheumatol. 2007;34:740–6. [PubMed] [Google Scholar]
- 23.National Cholesterol Education Program (NCEP) Expert Panel on Detection, Evaluation, and Treatment of High Blood Cholesterol in Adults (Adult Treatment Panel III). Third Report of the National Cholesterol Education Program (NCEP) Expert Panel on Detection, Evaluation, and Treatment of High Blood Cholesterol in Adults (Adult Treatment Panel III) final report. Circulation. 2002;106:3143–421. [PubMed] [Google Scholar]
- 24.Levey AS, Coresh J, Balk E, et al. National Kidney Foundation practice guidelines for chronic kidney disease: evaluation, classification, and stratification. Ann Intern Med. 2003;139:137–47. doi: 10.7326/0003-4819-139-2-200307150-00013. [DOI] [PubMed] [Google Scholar]
- 25.Barbaux S, Kluijtmans LA, Whitehead AS. Accurate and rapid “multiplex heteroduplexing” method for genotyping key enzymes involved in folate/homocysteine metabolism. Clin Chem. 2000;46:907–12. [PubMed] [Google Scholar]
- 26.Packer BR, Yeager M, Burdett L, et al. SNP500Cancer: a public resource for sequence validation, assay development, and frequency analysis for genetic variation in candidate genes. Nucl Acids Res. 2006;34:D617–21. doi: 10.1093/nar/gkj151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Johnson WG, Stenroos ES, Spychala JR, Chatkupt S, Ming SX, Buyske S. New 19 bp deletion polymorphism in intron-1 of dihydrofolate reductase (DHFR): a risk factor for spina bifida acting in mothers during pregnancy? Am J Med Genet A. 2004;124:339–45. doi: 10.1002/ajmg.a.20505. [DOI] [PubMed] [Google Scholar]
- 28.Shi M, Caprau D, Romitti P, Christensen K, Murray JC. Genotype frequencies and linkage disequilibrium in the CEPH human diversity panel for variants in folate pathway genes MTHFR, MTHFD, MTRR, RFC1, and GCP2. Birth Defects Res A Clin Mol Teratol. 2003;67:545–9. doi: 10.1002/bdra.10076. [DOI] [PubMed] [Google Scholar]
- 29.Pepe G, Vanegas OC, Rickards O, et al. World distribution of the T833C/844INS68 CBS in cis double mutation: a reliable anthropological marker. Hum Genet. 1999;104:126–9. doi: 10.1007/s004390050924. [DOI] [PubMed] [Google Scholar]
- 30.Ranganathan P, Culverhouse R, Marsh S, et al. Single nucleotide polymorphism profiling across the methotrexate pathway in normal subjects and patients with rheumatoid arthritis. Pharmacogenomics. 2004;5:559–69. doi: 10.1517/14622416.5.5.559. [DOI] [PubMed] [Google Scholar]
- 31.Fijnheer R, Roest M, Haas FJ, De Groot PG, Derksen RH. Homocysteine, methylenetetrahydrofolate reductase polymorphism, antiphospholipid antibodies, and thromboembolic events in systemic lupus erythematosus: a retrospective cohort study. J Rheumatol. 1998;25:1737–42. [PubMed] [Google Scholar]
- 32.Burzynski M, Duriagin S, Mostowska M, Wudarski M, Chwalinska-Sadowska H, Jagodzinski PP. MTR 2756 A > G polymorphism is associated with the risk of systemic lupus erythematosus in the Polish population. Lupus. 2007;16:450–4. doi: 10.1177/0961203307077988. [DOI] [PubMed] [Google Scholar]
- 33.Francis ME, Eggers PW, Hostetter TH, Briggs JP. Association between serum homocysteine and markers of impaired kidney function in adults in the United States. Kidney Int. 2004;66:303–12. doi: 10.1111/j.1523-1755.2004.00732.x. [DOI] [PubMed] [Google Scholar]
- 34.Parle-McDermott A, Mills JL, Molloy AM, et al. The MTHFR 1298CC and 677TT genotypes have opposite associations with red cell folate levels. Mol Genet Metab. 2006;88:290–4. doi: 10.1016/j.ymgme.2006.02.011. [DOI] [PubMed] [Google Scholar]