Significance
Most clinically used antibiotics are derived from bacterial small molecules produced by dedicated biosynthetic gene clusters. Recent investigations have indicated that the majority of these biosynthetic genes are inactive or “silent.” Identifying the triggers that lead to activation of silent clusters would greatly expand our repertoire of therapeutic molecules. Described herein is a new, high-throughput approach for identifying activators of silent gene clusters. Application of this method to two bacterial clusters revealed that both could be efficiently activated and uncovered a new metabolite. Surprisingly, almost all elicitors discovered were antibiotics, suggesting they play an important role in modulating silent biosynthetic pathways. This approach promises to reveal useful small molecules and the biological regulatory mechanisms underlying silent gene clusters.
Keywords: natural products discovery, cryptic metabolites, malleilactone
Abstract
Over the past decade, bacterial genome sequences have revealed an immense reservoir of biosynthetic gene clusters, sets of contiguous genes that have the potential to produce drugs or drug-like molecules. However, the majority of these gene clusters appear to be inactive for unknown reasons prompting terms such as “cryptic” or “silent” to describe them. Because natural products have been a major source of therapeutic molecules, methods that rationally activate these silent clusters would have a profound impact on drug discovery. Herein, a new strategy is outlined for awakening silent gene clusters using small molecule elicitors. In this method, a genetic reporter construct affords a facile read-out for activation of the silent cluster of interest, while high-throughput screening of small molecule libraries provides potential inducers. This approach was applied to two cryptic gene clusters in the pathogenic model Burkholderia thailandensis. The results not only demonstrate a prominent activation of these two clusters, but also reveal that the majority of elicitors are themselves antibiotics, most in common clinical use. Antibiotics, which kill B. thailandensis at high concentrations, act as inducers of secondary metabolism at low concentrations. One of these antibiotics, trimethoprim, served as a global activator of secondary metabolism by inducing at least five biosynthetic pathways. Further application of this strategy promises to uncover the regulatory networks that activate silent gene clusters while at the same time providing access to the vast array of cryptic molecules found in bacteria.
The vast majority of antibiotic scaffolds that are in use today were found in an unusually productive period spanning the 1940s and 1950s, now referred to as the golden age of antibiotics discovery (1, 2). This period led to an explosion in the identification of new and biologically useful small molecules that has sustained our antibiotic arsenal to this day. Natural products were the key ingredients and as of 2010 accounted for >70% of all clinical antibiotics and >55% of antiviral remedies (3, 4). However, this reservoir is being challenged by two recent, alarming trends. No novel antibiotics scaffolds were discovered between 1962 and 2000, straining our reserve of useful antibiotics (1). This productivity crisis was further exacerbated by the rise of multidrug-resistant bacteria (5–8). The Center for Disease Control and Prevention recently reported that annually over 2 million Americans are infected by multidrug-resistant pathogens; at least 23,000 of these are fatal. Resistance to all major antibiotics classes has been observed and, given the steady rise of various multi-, pan-, or extensively drug-resistant pathogens, we would benefit from another exceedingly productive era of bioactive small molecule discovery (9, 10).
Where will the next generation of bioactive molecules come from? The answer may lie in recent genome sequencing efforts, which have revealed that our current discovery methods access, at best, 10% of the small molecule repertoire of bacteria (11). A detailed analysis of the sequenced genomes of actinomycetes, the group of bacteria responsible for over 50% of all antibiotics, has demonstrated that the great majority of biosynthetic gene clusters, the sets of genes responsible for production of bioactive compounds, remain inactive or silent for unknown reasons (11–13). Given the track record of natural products as therapeutics, these clusters, dubbed silent or cryptic gene clusters, harbor an extensive supply of potential drug candidates, and successful approaches that systematically awaken them would have a major impact on drug discovery.
The problem of silent gene clusters is challenging because an unknown signal activates an uncharacterized gene cluster leading to the production of a new metabolite (14). There are three variables in this process, two of which can be determined experimentally or computationally: bioinformatic methods allow for facile identification of genes that generate nonribosomal peptides, polyketides, and terpenes, and pinpointing gene assemblies of novel metabolites within these families can be performed with good fidelity (15). Once activated, the product of the gene cluster can be experimentally identified by differential metabolomics facilitating its isolation and structural elucidation via multidimensional NMR. Thus, the problem of crypticity may be reduced to the large variety of signals that may act as elicitors or activators of silent clusters.
Thus far, no method has been described that allows for identification of elicitors of a given silent gene cluster. An efficient platform that enables discovery of small molecule activators would allow scrutiny of the regulatory pathways that lead to induction of silent biosynthetic clusters as well as structural and functional elucidation of their products. Outlined herein is a high-throughput approach for uncovering elicitors of cryptic metabolites. In this method, genetic reporter fusions monitor the activity of the silent cluster of interest, while high-throughput screening of small molecule libraries provides the candidate elicitors. The approach has been applied to two gene clusters in the pathogenic model, Burkholderia thailandensis E264. One cluster is silent and produces, when activated, an important virulence factor required for infections, whereas the second is lowly expressed under typical laboratory growth conditions and gives rise to a histone deacetylase inhibitor. Using the screening approach, multiple elicitors for both clusters were discovered resulting in a 12–145-fold overproduction of their small molecule products. Surprisingly, most elicitors were antibiotics, implicating a role for these compounds in activating secondary metabolism, when supplied at subinhibitory concentrations. These results set the stage for understanding the regulatory cascades that induce silent gene clusters in response to exogenous elicitors and promise access to the extensive array of bioactive small molecules in bacterial cryptic metabolomes.
Results
A High-Throughput Screen for Elicitors of Silent Gene Clusters.
As a new approach to tackle the problem of crypticity, it was hypothesized that the complex environment in which bacteria grow and evolve, i.e., soil, contains the requisite, diffusible, small molecules that awaken silent gene clusters (16, 17). An approach was envisioned in which high-throughput screening of natural product libraries or extracts would mimic or represent candidate elicitors, and targeted GFP or lacZ reporter fusions would allow for a facile read-out of cryptic gene expression, as metabolic pathways in bacteria are typically regulated at the transcriptional level. Appearance of fluorescence or LacZ activity would thus signal activation or “awakening” of the chosen cryptic gene cluster.
To test these ideas, a known cryptic cluster in a genetically tractable organism, the malleilactone (mal) cluster in B. thailandensis E264 (hereafter E264), was targeted (Fig. 1). Within the Pseudomallei group pathogens––which include E264, Burkholderia mallei, and Burkholderia pseudomallei––malleilactone is a virulence factor required for nematode infections, and its structure was recently determined (1, Fig. 1A) (18–20). However, the elicitors that lead to activation of this silent virulence factor are not known. To monitor expression of this cluster, a translational lacZ fusion to malL, a gene essential for the biosynthesis of 1, was used (hereafter malL-lacZ) (21). malL-lacZ is not expressed under standard growth conditions and served as a negative control. A lacZ reporter in btaK, which is quorum-sensing-regulated and expressed at high cell densities, provided a positive control (Methods) (21–23). A number of growth parameters, including temperature, growth media, agitation rates, total volume, and initial inoculum were assessed and provided an optimized, robust screening assay with a Z′ value of 0.51 in a 384-well format (SI Appendix, Table S1). The positive control gave luminescence reads that were 16-fold larger than those for malL-lacZ, which exhibited low background activity.
Fig. 1.
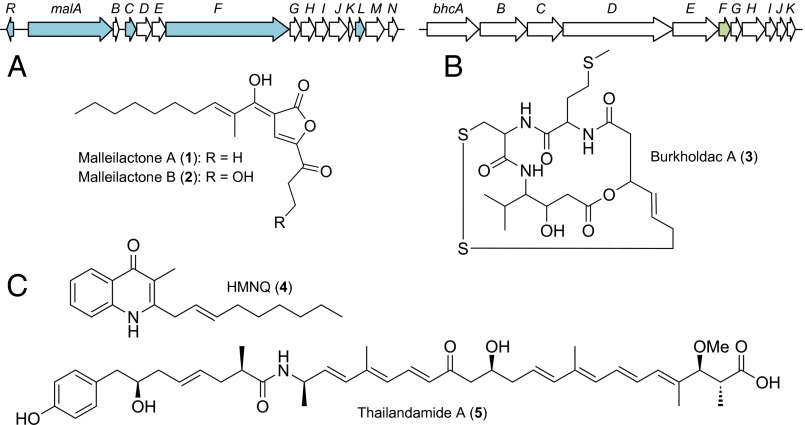
Cryptic biosynthetic gene clusters and their products modulated in this study. (A) Gene organization in the silent mal cluster (Upper). Gene transcripts that were monitored by RT-qPCR in this study are shown in aqua. A translational lacZ reporter fusion to malL was used as a read-out for malleilactone production. Structures of malleilactone A (1), determined previously (19, 20), and malleilactone B (2), elicited and solved in this study, are shown (Lower). (B) The bhc cluster (Upper); a lacZ reporter fusion to bhcF was used in this study. The structure of burkholdac A (3), the product of the bhc cluster is shown (Lower) (24). (C) Structures of two secondary metabolites produced by B. thailandensis and elicited in this study, HMNQ drawn as its quionolone tautomer (4) and thailandamide A (5), produced by the hmq and tha gene clusters, respectively (25–27).
A compound collection containing 640 members of well-characterized, biologically active metabolites was screened in duplicates against malL-lacZ in an attempt to find an elicitor that activates the mal biosynthetic cluster (SI Appendix, Table S1 and Fig. S1). The data can best be visualized by plotting the averaged total malL-lacZ activity, determined by end-point luminescence, as a function of each compound (Fig. 2). The luminescence outputs were further analyzed by calculating standard Z scores for each metabolite in the library (Methods). Lines corresponding to a Z score of 10 (Fig. 2, top blue line) and the averaged luminescence of the negative control (Fig. 2, bottom blue line), are shown. A total of nine hits with Z scores between 6 and 21 were observed (SI Appendix, Table S2). Four compounds, the β-lactam piperacillin (Fig. 2, 6), the dihydrofolate reductase inhibitor trimethoprim (7), and two cephalosporins, ceftazidime (8) and cefotaxime (9), had Z scores greater than 10 and appeared to efficiently activate the mal cluster. The next best five hits had Z scores between 6.4 and 8.1 and were all members of the fluoroquinolone antibiotics (Fig. 2, blue columns). These nine elicitors comprised 1.4% of the compounds in the library, suggesting that finding activators of the mal cluster by this method is fairly efficient. Interestingly, all nine elicitors are antibacterial compounds, almost all in current clinical use (see Discussion).
Fig. 2.
High-throughput screening as a strategy to awaken silent gene clusters. Two library plates (Upper and Lower) containing a total of 640 compounds were screened in duplicates against malL-lacZ. Each assay plate contained positive (btaK-lacZ) and negative (malL-lacZ with no elicitors) controls, which are shown in green bars. The averaged luminescence as a result of induced LacZ activity is shown for each compound and normalized to the positive control. The data are presented in clusters of 16 compounds. Horizontal blue lines correspond to a Z score of 10 (top line) and the averaged luminescence of the negative control (bottom line). Hits were defined as those with Z scores greater than 6. No hits were obtained on the second library plate (Lower). A total of nine hits were obtained on the first plate (Upper). The structures for the four best hits (orange and red bars), with Z scores greater than 10, are shown. These correspond to piperacillin (6), trimethoprim (7), ceftazidime (8), and cefotaxime (9). Five additional hits belonging to the family of fluoroquinolone antibiotics (tosufloxacin, sarafloxacin, ofloxacin, gatifloxacin, and ciprofloxacin) are shown in blue bars (SI Appendix, Tables S1 and S2).
Validating Elicitors of the mal Cluster.
Efforts were next focused on validating the top two hits. To this end, dose–response analyses were carried to establish a correlation between malleilactone production and elicitor concentrations. The results demonstrate a sigmoidal dependence of malL-lacZ activity on the concentrations of 6 and 7, yielding EC50 values, the elicitor concentration at which malleilactone production is half-maximal, of 1.4 and 18 μM, respectively (Fig. 3A). The ability of 6 and 7 to awaken the mal cluster was further validated using reverse-transcription quantitative PCR experiments (RT-qPCR). Wild-type E264 was cultured in the absence or presence of 6 or 7, and the mRNA levels of five genes, malR (luxR-type transcriptional regulator, btaR4), malA (polyketide synthase), malC (SyrP-like Fe-dependent hydroxylase), malF (a second polyketide synthase), and malL (acyltransferase), were determined (Fig. 1A). The data were normalized to the levels of these genes observed in wt E264 in the absence of elicitors (SI Appendix, Figs. S2 and S3 and Table S3). The results demonstrate that elicitor 6 affects a 5–10-fold up-regulation of the five target genes. Strikingly, an 8–45-fold increase was elicited by 7, indicating a pronounced effect on the expression of the mal biosynthetic genes (Fig. 3B).
Fig. 3.
Elicitors of cryptic metabolite production. (A) Hit validation by dose–response analysis for the ability of 6 (red dots) and 7 (blue dots) to turn on the silent mal cluster. Each data point is the average of two independent end-point luminescence measurements using malL-lacZ and varying concentrations of 6 or 7. Black lines describe fits to Eq. S3 (SI Appendix) and yield EC50 values of 1.4 μM (6) and 18 μM (7). (B) RT-qPCR analysis of mal genes in wt E264 (control, black bars) and as a function of 6 (red bars) or 7 (blue bars). The level of each gene is normalized to the E264 control and expressed as fold-change. Each bar is the average of two independent measurements, each determined in triplicates. (C) HPLC-MS analysis of E264 extracts containing no elicitor (black trace), 8.6 μM 6 (red trace), or 15.3 μM 7 (blue trace), which correspond to elicitor concentrations used in the screen. (Inset) Effect of saturating concentrations of 7 (30.6 μM) on malleilactone production. The starred peaks correspond to 1. (D) HPLC-MS analysis of E264 extracts containing no elicitor (black trace) or β-lapachone (green trace). Elution of the mass ion of 3 (starred peak) is shown as a function of retention time. The traces in C and D are normalized for OD600 nm of the corresponding cultures and vertically offset for clarity.
As further corroboration for the roles of 6 and 7 in inducing the mal cluster, wt E264 cultures were supplemented with 6, 7, or DMSO (control) and the supernatants were analyzed by HPLC-MS. The identity of 1 was verified by UV-vis spectroscopy (SI Appendix, Fig. S4), high-resolution (HR) HPLC-MS ([M+H]+calc 307.1909, [M+H]+obs 307.1907), and tandem HR-MS/MS (SI Appendix, Fig. S5 and Table S4). The results reveal a large induction of malleilactone production in the presence of both elicitors (Fig. 3C). The magnitude of this overproduction is difficult to determine because very little malleilactone is generated in the absence of elicitors. Using the mass-ion peak for 1, it is estimated that, at the concentrations used in the screen, 6 and 7 induce a 7–10-fold and a 40-fold overproduction of 1, respectively. At saturating concentrations, based on the results in Fig. 3A, 7 induced a remarkable 145-fold overproduction of 1 (Fig. 3C, Inset). Together, the results above show that trimethoprim and piperacillin are potent activators of the mal cluster, which is transcriptionally up-regulated in the presence of these antibiotics resulting in prominent production levels of an otherwise silent virulence factor.
A New Malleilactone Analog.
It was noticed that induction of malleilactone by 6 or 7 resulted in the appearance of two additional malleilactones that were not reported in previous studies (19, 20). These gave HR-MS data of [M+H]+obs 323.1855 and 365.1960 consistent with the addition of a hydroxyl group to 1 for the first analog ([M+H]+calc 323.1858), and a molecular formula of C20H28O6 for the second ([M+H]+calc 365.1964). Multiple attempts to purify the second analog were unsuccessful due to the instability of this group of metabolites (20). Isolation and structural elucidation by multidimensional NMR for the first compound revealed hydroxy-malleilactone (2, Fig. 1A), a new analog containing a primary alcohol on the short propionyl chain (SI Appendix, Figs. S4–S6 and Table S5). Tandem HR-MS/MS further corroborated the structure of 2 (SI Appendix, Table S4). The level of overproduction of 2 as a function of elicitor could not be quantified as no product was detected at all in the absence of elicitors, not even by sensitive MS methods.
Elicitors for the Burkholdac Cluster.
Having established an efficient screening method for finding elicitors of cryptic biosynthetic pathways, the generality of this approach was examined next by targeting the burkholdac (bhc) cluster. The bhc cluster is lowly expressed under typical growth conditions and produces the histone deacetylase inhibitor 3 (Fig. 1B) (24). In this case, a translational lacZ fusion to bhcF provided the read-out for burkholdac production. From a library of 800 compounds, 5 elicitors were found to induce the bhc cluster with Z scores of 15 or better: the Streptomycete-derived antitumor drug mitomycin, the plant-derived toxin, β-lapachone, the NF-κB inhibitor BAY 11–7082, and the antibiotics flumequine and trimethoprim (SI Appendix, Figs. S1 and S7 and Tables S6 and S7). To confirm induction of 3, E264 culture extracts were assessed in the presence and absence of elicitor. Analysis by HPLC-MS revealed a 12-fold overproduction of 3 in the presence of β-lapachone (Fig. 3D and SI Appendix, Table S4). The success in activating two different gene clusters––that is, turning on the silent mal cluster while increasing production of the lowly expressed bhc cluster––along with the efficiency in finding the appropriate elicitors, strongly indicates that this strategy will be generally applicable in activating and unearthing the products of silent biosynthetic pathways.
Small Molecule Pleiotropy.
To investigate whether the elicitors above were specifically up-regulating their intended targets or represented global activators of secondary metabolism, detailed metabolomic studies were undertaken. Whereas β-lapachone mainly affected burkholdac biosynthesis (SI Appendix, Fig. S8), trimethoprim appeared to activate multiple biosynthetic pathways (Fig. 4A). In addition to the mal cluster, at least four other groups of metabolites were significantly induced. Production of the thailandamides (Fig. 1, 5), a family of polyenes comprising multiple geometric isomers generated by the tha gene cluster (25, 26), was increased 8–36-fold, depending on the isomer (Fig. 4A). This marked effect was verified by RT-qPCR, which revealed a 26-fold and 17-fold up-regulation of thaH (polyketide synthase) and thaR (oxidoreductase), respectively, in the presence of elicitor 7 (Fig. 4B). The quinolone antibiotic and quorum-sensing modulator 4-hydroxy-3-methyl-2-(2-nonenyl)-quinoline (HMNQ) (Fig. 1, 4), was identified by HR-HPLC-MS ([M+H]+calc 284.2014, ([M+H]+obs 284.2015), tandem HR-MS/MS (SI Appendix, Table S4), and 1H-NMR (SI Appendix, Fig. S9), and was induced ∼sevenfold in the presence of trimethoprim (27). As mentioned above, trimethoprim was also discovered as an elicitor in screens for bhc induction and yielded a three- to fourfold overproduction of 3 (Fig. 4C). Finally, and perhaps most interestingly, a group of new, interrelated metabolites was also induced by trimethoprim by ∼10-fold (Fig. 4A, starred peaks). Structural elucidation of this class of compounds awaits further experiments. Collectively, the pleiotropic effects of trimethoprim highlight the potential for finding global small molecule activators of secondary metabolite production using the high-throughput screening strategy outlined above. They further exemplify the hormetic properties of antibiotics, which modulate bacterial secondary metabolism at low concentrations but result in cell death at high concentrations (28).
Fig. 4.
Trimethoprim activates multiple gene clusters in B. thailandensis. (A) Analysis of secondary metabolites in E264 extracts treated with DMSO (control, black trace) or with saturating concentrations of trimethoprim (30.6 μM, blue trace). Peaks corresponding to malleilactone (1), which does not absorb appreciably at 280 nm, HMNQ (4), the thailandamide isomers (5), and a set of new compounds elicited by trimethoprim (starred peaks) are indicated. (Inset) Characteristic absorption spectrum of thailandamide A (5). (B) RT-qPCR analysis of two genes within the thailandamide cluster (tha) in E264 cultures supplemented with DMSO (control, black bars) or with 30.6 μM trimethoprim (blue bars). The level of each gene is normalized to the E264 control and expressed as fold-change. Each bar is the average of two independent measurements, each determined in triplicates. (C) Induction of 3 by trimethoprim (30.6 μM, blue trace) compared with untreated samples (black trace). The traces in A and C are normalized for OD600 nm of the corresponding cultures and vertically offset for clarity.
Discussion
The realization that silent biosynthetic gene clusters harbor a wealth of new and potentially useful small molecules has motivated the development of methods to activate them. Whereas previous reports have focused on transplanting a specific cluster into a heterologous host or inserting artificial promoter elements in the endogenous producer, a different approach was taken in this study to examine a role for diffusible small molecules in signaling activation of cryptic pathways. In this approach, high-throughput screening was used, not to find a drug or an inhibitor as is usually the case, but rather to elicit production of drug-like molecules, while genetic reporter fusions provided a facile read-out for activation of the gene cluster of interest. The advantage of the approach herein lies in its capacity to unveil synthetic and natural elicitors of cryptic gene clusters, the endogenous regulatory networks that lead to their induction, and the products of these clusters, whereas existing methods have largely focused on the latter aspect.
By screening a collection of bioactive compounds, antibiotics were revealed as the major cues for induction of cryptic gene clusters. At high concentrations, many of these kill E264. In fact, current genetic approaches use trimethoprim (7) as a resistance marker in E264 (29). Lower concentrations of trimethoprim, however, caused a global activation of secondary metabolism by inducing at least five biosynthetic gene clusters (mal, bhc, tha, and hmq), including a new group of metabolites whose structures have yet to be determined. Likewise, the other elicitors discovered herein belong to the β-lactam, cephalosporin, fluoroquinolone, or DNA-alkylating antibiotic families, which at least at the low concentrations of the screen appear to exert transcriptional effects rather than growth inhibition. For some years, Davies and co-workers have argued that subinhibitory concentrations of antibiotics found in ecological bacterial niches are signaling agents, rather than growth inhibitors (28, 30–33). The results presented now show that the signaling functions of antibiotics can be extended to the activation of silent gene clusters, and in this case, even the production of a cryptic virulence factor (Fig. 1A, 1). As such, these results underline the prospect of exploiting high-throughput screening approaches to examine the natural roles of secondary metabolites.
With a method at hand to awaken silent gene clusters using the endogenous host’s regulatory pathways, the molecular mechanisms underlying this activation can now be unraveled. Although studies examining the mechanism of activation of the mal cluster are underway, it is noteworthy that both elicitors 6 and 7 led to an up-regulation of malR, suggesting a positive regulatory role for this gene. malR is a so-called orphan luxR, a transcriptional regulator that is not adjacent to its cognate luxI homoserine lactone synthase (34), and may therefore bind alternate small molecule elicitors, such as 6 or 7. In an alternative model, stress-induced mechanisms as a result of antibiosis may be responsible for activation of malR (9, 35–37). The fact that the synthetic antibiotics (7 and the fluoroquinolones) as well as derivatives of natural ones (6, 8, and 9) elicit production of 1 appears to be consistent with this latter model.
The approach herein can be easily modified to include multiple simultaneous reporters thus allowing activation of several gene clusters with a single screen. In addition, mass spectroscopic detection methods could replace the genetic reporter fusions in bacterial strains where genetic manipulations are difficult or not possible. Application of these strategies to other silent gene clusters promises to systematically reveal the structures of novel secondary metabolites, thus enabling future assessment of this large group of hidden molecules as therapeutic leads. These applications and variations of the method above are currently underway.
Methods
Bacterial Strains.
Four strains of B. thailandensis E264 were used in this study: the wild-type strain was obtained from the American Type Culture Collection, the btaK-lacZ variant (21) was kindly provided by Josephine Chandler in the laboratory of Pete Greenberg at the University of Washington, Seattle, WA, and reporter strains malL-lacZ and bhcF-lacZ, also derived from a sequence-defined transposon mutant library (21), were kindly provided by Beth Ramage in the laboratory of Colin Manoil at the University of Washington, Seattle, WA.
High-Throughput Screen.
The screen was performed at the Institute of Chemistry and Cell Biology Longwood Screening Facility (ICCB-L) at Harvard Medical School. Bacterial growth was carried out in LB supplemented with 50 mM Mops, pH 7.0 (hereafter, LB-Mops). To commence the screen, malL-lacZ from an LB agar plate was used to inoculate 5 mL of LB-Mops in a sterile 14-mL bacterial culture tube. The culture was grown overnight at 30 °C and 250 rpm. After 12–16 h, its OD600 nm was determined on an Ultraspec 5300 Pro Spectrophotometer (Amersham Biosciences). The culture was diluted into 100 mL of LB-Mops to give a final OD600 nm of 0.05. Subsequently a volume of 45 μL was dispensed into each of four sterile, white 384-well plates (Corning) using a WellMate automated dispenser (Matrix Technologies) at the ICCB-L. Using a Compound Transfer Robot (Seiko) equipped with a calibrated stainless steel pin transfer tool (V&P Scientific), each well was supplemented with 0.1 μL of a compound from the “Biomol-4” Library (Known Bioactives Collection, plates 2089 and 2090 at the ICCB-L acquired from Enzo Life Sciences, see SI Appendix), which comprises 640 molecules. The compounds were dispensed into columns 3 through 22 on each plate. Columns 1 and 2 contained the negative control (malL-lacZ in the absence of any library compounds) and columns 23 and 24 contained the positive control (btaK-lacZ). Between each dispense cycle, the pins were washed in MeOH, sonicated in a MeOH–water mixture, and dried with pressurized air. Each plate was covered with a Breathe-Easy sealing membrane (Sigma), vortexed briefly on a plate vortexer, and cultured at 30 °C and 230 rpm for 12 h in a Multitron Shaker (ATR) equipped with a green sealing tray. To maintain constant humidity, several 1-L Erlenmeyer flasks containing 200 mL of water were also placed inside the shaker. After 12 h, the plates were removed from the shaker and rested at room temperature for 5 min. The β-Glo reagent (Promega) was used to monitor LacZ activity. The reagent was diluted 2:1 with water. Then, each well was supplemented with 30 μL of the diluted β-Glo reagent using the WellMate automated dispenser, and subsequently vortexed, and incubated in the dark at room temperature for 45 min. Total end-point luminescence was then determined on an EnVision Multilabel Reader (Perkin-Elmer). The optimization wizard in the EnVision software was used to optimize plate dimensions and minimize cross-talk, yielding a software-derived correction factor of 0.27. The Z' score was calculated according to Eq. 1, where σp and σn correspond to the SD of the positive (btaK-lacZ) and negative (malL-lacZ in the absence of elicitors) controls, respectively, and μp and μn correspond to the mean LacZ activity for the positive and negative controls. Standard Z scores for each library compound in the actual screen (Fig. 2) were calculated using Eq. 2, where μc corresponds to the mean LacZ activity for each compound in the library, μn corresponds to the mean of the negative control on the same assay plate (columns 1 and 2), and σn corresponds to the SD of the negative control. High-throughput screens with bchF-lacZ were performed and analyzed in an identical fashion as described above. These assays also used compound libraries from the Known Bioactives Collection at the ICCB-L (plates 1921, 1989, 1990 acquired from Enzo Life Sciences and MicroSource Discovery Systems; see SI Appendix).
| [1] |
| [2] |
Dose–Response Analysis, RT-qPCR, HPLC-MS Assays, and Compound Isolation or Characterization.
Detailed procedures for additional experiments are given in SI Appendix.
Supplementary Material
Acknowledgments
The author thanks Dr. Josephine Chandler for many helpful discussions and for sharing strain btaK-lacZ; Beth Ramage and the Manoil lab for providing strains malL-lacZ and bchF-lacZ; and Dr. Emily Derbyshire for advice regarding high-throughput screening. Many thanks also to the personnel at the Harvard Medical School screening facility (ICCB-L), especially Dr. Su Chiang and Dr. Stewart Rudnicki, for technical assistance; and to Profs. Tom Muir, Jon Clardy, JoAnne Stubbe, and Roberto Kolter for helpful discussions and comments on the manuscript. The author acknowledges Princeton University start-up funds for support of this work.
Footnotes
The author declares no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1400019111/-/DCSupplemental.
References
- 1.Walsh C. Where will new antibiotics come from? Nat Rev Microbiol. 2003;1(1):65–70. doi: 10.1038/nrmicro727. [DOI] [PubMed] [Google Scholar]
- 2.Nathan C. Antibiotics at the crossroads. Nature. 2004;431(7011):899–902. doi: 10.1038/431899a. [DOI] [PubMed] [Google Scholar]
- 3.Newman DJ, Cragg GM. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J Nat Prod. 2012;75(3):311–335. doi: 10.1021/np200906s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Newman DJ, Cragg GM, Snader KM. The influence of natural products upon drug discovery. Nat Prod Rep. 2000;17(3):215–234. doi: 10.1039/a902202c. [DOI] [PubMed] [Google Scholar]
- 5.Clatworthy AE, Pierson E, Hung DT. Targeting virulence: A new paradigm for antimicrobial therapy. Nat Chem Biol. 2007;3(9):541–548. doi: 10.1038/nchembio.2007.24. [DOI] [PubMed] [Google Scholar]
- 6.Falagas ME, et al. Outcome of infections due to pandrug-resistant (PDR) Gram-negative bacteria. BMC Infect Dis. 2005;5:24–30. doi: 10.1186/1471-2334-5-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Klevens RM, et al. Invasive methicillin-resistant Staphylococcus aureus infections in the United States. JAMA. 2007;298(15):1763–1771. doi: 10.1001/jama.298.15.1763. [DOI] [PubMed] [Google Scholar]
- 8.Fischbach MA, Walsh CT. Antibiotics for emerging pathogens. Science. 2009;325(5944):1089–1093. doi: 10.1126/science.1176667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Clardy J, Fischbach MA, Walsh CT. New antibiotics from bacterial natural products. Nat Biotechnol. 2006;24(12):1541–1550. doi: 10.1038/nbt1266. [DOI] [PubMed] [Google Scholar]
- 10.Baltz RH. Renaissance in antibacterial discovery from actinomycetes. Curr Opin Pharmacol. 2008;8(5):557–563. doi: 10.1016/j.coph.2008.04.008. [DOI] [PubMed] [Google Scholar]
- 11.Nett M, Ikeda H, Moore BS. Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat Prod Rep. 2009;26(11):1362–1384. doi: 10.1039/b817069j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bentley SD, et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2) Nature. 2002;417(6885):141–147. doi: 10.1038/417141a. [DOI] [PubMed] [Google Scholar]
- 13.Oliynyk M, et al. Complete genome sequence of the erythromycin-producing bacterium Saccharopolyspora erythraea NRRL23338. Nat Biotechnol. 2007;25(4):447–453. doi: 10.1038/nbt1297. [DOI] [PubMed] [Google Scholar]
- 14.Zerikly M, Challis GL. Strategies for the discovery of new natural products by genome mining. ChemBioChem. 2009;10(4):625–633. doi: 10.1002/cbic.200800389. [DOI] [PubMed] [Google Scholar]
- 15.Blin K, et al. antiSMASH 2.0—a versatile platform for genome mining of secondary metabolite producers. Nucleic Acids Res. 2013;41(Web Server issue):W204-12. doi: 10.1093/nar/gkt449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Shank EA, Kolter R. New developments in microbial interspecies signaling. Curr Opin Microbiol. 2009;12(2):205–214. doi: 10.1016/j.mib.2009.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Seyedsayamdost MR, Traxler MF, Clardy J, Kolter R. Old meets new: Using interspecies interactions to detect secondary metabolite production in actinomycetes. Methods Enzymol. 2012;517:89–109. doi: 10.1016/B978-0-12-404634-4.00005-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Galyov EE, Brett PJ, DeShazer D. Molecular insights into Burkholderia pseudomallei and Burkholderia mallei pathogenesis. Annu Rev Microbiol. 2010;64:495–517. doi: 10.1146/annurev.micro.112408.134030. [DOI] [PubMed] [Google Scholar]
- 19.Biggins JB, Ternei MA, Brady SF. Malleilactone, a polyketide synthase-derived virulence factor encoded by the cryptic secondary metabolome of Burkholderia pseudomallei group pathogens. J Am Chem Soc. 2012;134(32):13192–13195. doi: 10.1021/ja3052156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Franke J, Ishida K, Hertweck C. Genomics-driven discovery of burkholderic acid, a noncanonical, cryptic polyketide from human pathogenic Burkholderia species. Angew Chem Int Ed Engl. 2012;51(46):11611–11615. doi: 10.1002/anie.201205566. [DOI] [PubMed] [Google Scholar]
- 21.Gallagher LA, et al. Sequence-defined transposon mutant library of Burkholderia thailandensis. MBio. 2013;4(6):e00604–e00613. doi: 10.1128/mBio.00604-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Duerkop BA, et al. Quorum-sensing control of antibiotic synthesis in Burkholderia thailandensis. J Bacteriol. 2009;191(12):3909–3918. doi: 10.1128/JB.00200-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Seyedsayamdost MR, et al. Quorum-sensing-regulated bactobolin production by Burkholderia thailandensis E264. Org Lett. 2010;12(4):716–719. doi: 10.1021/ol902751x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Biggins JB, Gleber CD, Brady SF. Acyldepsipeptide HDAC inhibitor production induced in Burkholderia thailandensis. Org Lett. 2011;13(6):1536–1539. doi: 10.1021/ol200225v. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Nguyen T, et al. Exploiting the mosaic structure of trans-acyltransferase polyketide synthases for natural product discovery and pathway dissection. Nat Biotechnol. 2008;26(2):225–233. doi: 10.1038/nbt1379. [DOI] [PubMed] [Google Scholar]
- 26.Ishida K, Lincke T, Hertweck C. Assembly and absolute configuration of short-lived polyketides from Burkholderia thailandensis. Angew Chem Int Ed Engl. 2012;51(22):5470–5474. doi: 10.1002/anie.201200067. [DOI] [PubMed] [Google Scholar]
- 27.Vial L, et al. Burkholderia pseudomallei, B. thailandensis, and B. ambifaria produce 4-hydroxy-2-alkylquinoline analogues with a methyl group at the 3 position that is required for quorum-sensing regulation. J Bacteriol. 2008;190(15):5339–5352. doi: 10.1128/JB.00400-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Davies J, Spiegelman GB, Yim G. The world of subinhibitory antibiotic concentrations. Curr Opin Microbiol. 2006;9(5):445–453. doi: 10.1016/j.mib.2006.08.006. [DOI] [PubMed] [Google Scholar]
- 29.Barrett AR, et al. Genetic tools for allelic replacement in Burkholderia species. Appl Environ Microbiol. 2008;74(14):4498–4508. doi: 10.1128/AEM.00531-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Goh E-B, et al. Transcriptional modulation of bacterial gene expression by subinhibitory concentrations of antibiotics. Proc Natl Acad Sci USA. 2002;99(26):17025–17030. doi: 10.1073/pnas.252607699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Davies J. Are antibiotics naturally antibiotics? J Ind Microbiol Biotechnol. 2006;33(7):496–499. doi: 10.1007/s10295-006-0112-5. [DOI] [PubMed] [Google Scholar]
- 32.Yim G, Wang HH, Davies J. Antibiotics as signalling molecules. Philos Trans R Soc Lond B Biol Sci. 2007;362(1483):1195–1200. doi: 10.1098/rstb.2007.2044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Romero D, Traxler MF, López D, Kolter R. Antibiotics as signal molecules. Chem Rev. 2011;111(9):5492–5505. doi: 10.1021/cr2000509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Patankar AV, González JE. Orphan LuxR regulators of quorum sensing. FEMS Microbiol Rev. 2009;33(4):739–756. doi: 10.1111/j.1574-6976.2009.00163.x. [DOI] [PubMed] [Google Scholar]
- 35.Craney A, Ozimok C, Pimentel-Elardo SM, Capretta A, Nodwell JR. Chemical perturbation of secondary metabolism demonstrates important links to primary metabolism. Chem Biol. 2012;19(8):1020–1027. doi: 10.1016/j.chembiol.2012.06.013. [DOI] [PubMed] [Google Scholar]
- 36.Yoon V, Nodwell JR. Activating secondary metabolism with stress and chemicals. J Ind Microbiol Biotechnol. 2014;41(2):415–424. doi: 10.1007/s10295-013-1387-y. [DOI] [PubMed] [Google Scholar]
- 37.Traxler MF, Watrous JD, Alexandrov T, Dorrestein PC, Kolter R. Interspecies interactions stimulate diversification of the Streptomyces coelicolor secreted metabolome. MBio. 2013;4(4):00459–00513. doi: 10.1128/mBio.00459-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.