Background: LEV-9 is an extracellular scaffolding protein required for synaptic clustering of acetylcholine receptors in C. elegans.
Results: C-terminal cleavage of LEV-9 is necessary for its synaptic function.
Conclusion: LEV-9 is activated by proteolytic cleavage following the final tandem complement control protein (CCP) modules.
Significance: Many LEV-9-related factors are likely cleaved after CCP domains, suggesting that cleavage location can be conserved in distant architecturally related proteins.
Keywords: Extracellular Matrix Proteins, Nicotinic Acetylcholine Receptors, Post-translational Modification, Scaffold Proteins, Synapses, CCP Domain, Limited Proteolysis
Abstract
Correct positioning of neurotransmitter-gated receptors at postsynapses is essential for synaptic transmission. At Caenorhabditis elegans neuromuscular junctions, clustering of levamisole-sensitive acetylcholine receptors (L-AChRs) requires the muscle-secreted scaffolding protein LEV-9, a multidomain factor containing complement control protein (CCP) modules. Here we show that LEV-9 needs to be cleaved at its C terminus to exert its function. LEV-9 cleavage is not required for trafficking nor secretion but directly controls scaffolding activity. The cleavage site is evolutionarily conserved, and post-translational cleavage ensures the structural and functional decoupling between different isoforms encoded by the lev-9 gene. Data mining indicates that most human CCP-containing factors are likely cleaved C-terminally from CCP tandems, suggesting that not only domain architectures but also cleavage location can be conserved in distant architecturally related proteins.
Introduction
Scaffold-mediated receptor clustering at postsynapses is a key element of synaptic transmission. The activity of scaffold proteins can be regulated by several post-translational mechanisms including phosphorylation and degradation (1, 2), but also by limited proteolysis. For example, the rapid insertion of excitatory glutamate-gated AMPA receptors is regulated by cleaving the scaffolding protein GRIP (glutamate receptor-interacting protein) by Ca2+-dependent calpains (3). Calpains are also thought to regulate glutamate-gated NMDA receptor levels by acting upon the canonical cytoplasmic scaffolding protein PSD-95 (4). Extracellular proteolysis of scaffolding factors can also regulate synaptic receptor quantities. For instance, cleavage of the neuronal pentraxin receptor NRP by the extracellular metalloprotease TNF-converting enzyme (TACE) accelerates AMPA receptor endocytosis, resulting in a decreased content of AMPA receptors at synapses, a process necessary for mGluR1/5-dependent long-term depression in hippocampal and cerebellar synapses (5).
At excitatory neuromuscular junctions (NMJs)3 in Caenorhabditis elegans, an extracellular scaffold containing the LEV-9 and LEV-10 proteins is required for L-AChR clustering (6, 7). LEV-9 is a muscle-secreted factor composed by a WAP domain (whey acidic protein) followed by a tandem of eight consecutive CCP modules (also known as short consensus repeats or SUSHI repeats) (see Fig. 1A). It is architecturally related to several plasma proteins in vertebrates, including regulators of the complement system (8). LEV-10 is composed chiefly by CUB ectodomains (for complement C1r/C1s, Uegf, Bmp1) anchored to the muscle membrane via a transmembrane domain. LEV-10 belongs to a superfamily of ancillary proteins that regulate glutamate-gated receptors in vertebrates and invertebrates (8). Interaction between LEV-9, LEV-10, and L-AChRs is stabilized by OIG-4, a small secreted protein made of one immunoglobulin domain (9). In the absence of any of its components, including the L-AChRs themselves, the LEV-9-LEV-10 complex disassembles entirely. In lev-9 or lev-10 mutants, L-AChRs are declustered, but the total amount of functional receptors in the muscle membrane remains unchanged. However, these mutants survive chronic exposure to concentrations of the L-AChR agonist levamisole that are lethal to wild-type animals (6, 7, 9). Therefore, the L-AChR-associated complex is an attractive model for synaptic scaffolding by virtue of its domain architecture, constitutive activity, and involvement in downstream regulation of synaptic transmission.
FIGURE 1.
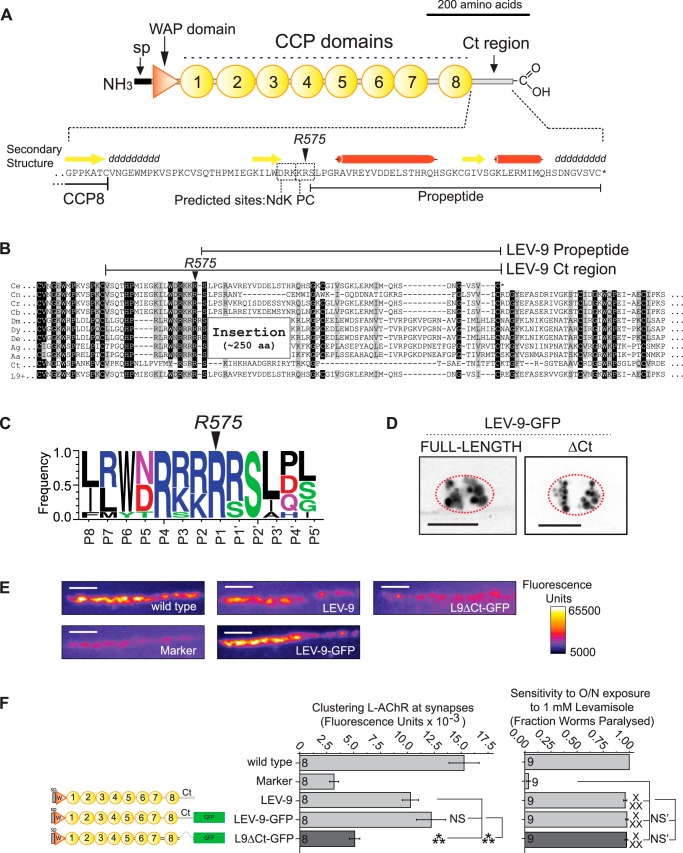
LEV-9 regulates the adaptation to the L-AChR agonist levamisole and is required to localize L-AChRs at NMJs in C. elegans. A, LEV-9 domain composition: signal peptide (sp), a WAP domain, CCP domains, and a Ct region devoid of predicted tertiary structure. The Ct region contains predicted cleavage sites for NdK (residues Asp-571/Arg-572/Lys-573) and PC (residues Lys-574/Arg-575/Ser-576). B, multiple sequence alignment at the Ct region of LEV-9 orthologs. Ce, C. elegans; Cn, Caenorhabditis brenneri; Cr, Caenorhabditis remanei; Cb, C. briggsae; Dm, D. melanogaster; Dy, Drosophila yakuba; De, Drosophila erecta; Ag, Anopheles gambiae; Aa; Aedes aegypti; Ct, Capitella teleta; L9+, lev-9:T07H6.4 fusion transcript. C, sequence logo of the PC predicted site (Arg-575 shown in position P1). D, detection of GFP endocytosed by coelomocytes after expression of GFP-tagged LEV-9 constructs in muscle cells and secretion into the extracellular space. Scale bars represent 10 μm. E, examples of NMJs used for quantification in lev-9 mutants rescued with different versions of LEV-9. Scale bars represent 5 μm. F, sensitivity to 18-h exposure to levamisole and synaptic clustering of L-AChR-tagRFPt in lev-9 mutants rescued with different versions of LEV-9 expressed under the control of the muscle specific promoter Pmyo-3. Columns and error bars represent averages and S.E., respectively; n numbers of worms (more than three lines) are shown at the base of columns. *** and NS equal p < 0.001 and p ≥ 0.05 in a t test, respectively. x x x and NS′ equal p < 0.001 and p ≥ 0.05 in a nonparametric Kolmogorov-Smirnov test, respectively.
In the present work, we show that LEV-9 is cleaved both in a heterologous expression system and in vivo. The cleaving site maps to a highly conserved set of residues present at the C-terminal end of LEV-9 that are predicted to be recognized by proprotein convertases. LEV-9 cleavage is required for its clustering activity but not for secretion nor traffic. We also show that the lev-9 locus encodes multiple isoforms, including a long polypeptide containing 17 CCP domains that is found in other species. We suggest that the post-translational cleavage of LEV-9 might serve to ensure its functional and structural decoupling from any C-terminal polypeptide attachments, as we provide some evidence suggesting that other CCP-containing proteins found in humans might undergo similar post-translational processing.
EXPERIMENTAL PROCEDURES
Genetics
Strains were maintained at 15 or 20 °C on regular Nematode Growth Medium plates. OP50 Escherichia coli was used as food source. Transgenic lines were obtained injecting construct at 10–20 ng/μl DNA and co-injection marker pEKL15 (lin-15+) at 80 ng/μl. Injections were done in the unc-29(kr208::tagRFPt); lev-9(ox177) lin-15(n765ts) strain, except for dominant-negative experiments that were done in unc-29(kr208::tagRFPt); lin-15(n765ts) strain. The strains used are compiled in the supplemental material.
Cell Culture
S2 Drosophila cells were maintained following the supplier recommendations (Invitrogen, catalog number R690-07) as described previously (7) and transiently transfected using calcium phosphate. Recombinant protein expression was driven by a metallothionein promoter and induced by copper sulfate.
Biochemistry
Media of transfected S2 cells were recovered and cleared by spinning. Cells were recovered, washed, and lysed with Triton X-100 and sonication. Worms were recovered from mixed staged plates, washed, and resuspended in β-mercaptoethanol-containing loading buffer and boiled. SDS-PAGE and immunoblots were done following standard procedures. The antibodies used were anti-GFP monoclonal JL-8 (Clontech, catalog number 632380, dilution 1:2500), anti-MYC (GeneTex, catalog number GTX29106, dilution 1:1000,), and anti-T7 (Novagen catalog number 69522, dilution 1/2000). The horseradish peroxidase-conjugated goat anti-rabbit (DAKO, catalog number K4002) and goat anti-mouse (DAKO, catalog number K4000) were used as secondary antibodies.
Fluorescent Microscopy and Quantification
Young adult worms were immobilized on agarose pads containing M9 medium/sodium azide. Images were acquired using a Leica DM5000B microscope, a CSU10 (Yokogawa) spinning disk head, a CoolSNAP HQ2 (Photometrics) CCD camera, and the software MetaMorph (v7.6.1.0, Molecular Devices). A cross-profile was obtained from z-projection of the most anterior 30 μm of the dorsal nerve cord using ImageJ. A background line was interpolated using the “tail” values, and the corresponding background values were subtracted from each point of the curve. The total intensity was obtained by summing the latter normalized values.
Bioinformatics
LEV-9 orthologues were identified with PSI-Blast, using C. elegans and Caenorhabditis briggsae orthologues as probes. The National Center for Biotechnology Information (NCBI) (ncbi.nlm.nih.gov), Joint Genome Institute (JGI), ENSEMBL, and Wellcome Trust Sanger Institute databases were analyzed. A subset of sequences was selected considering phylogenic distribution; alternative splice variants and redundant sequences were removed. Multiple sequence alignment was done using M-Coffee (10) and curated using BioEdit (v7.1.3, Ibis Biosciences). The Eukaryotic Linear Motif (ELM) resource (11) and ProP 1.0 Server (12) were used to identify protease sites. European Molecular Biology Laboratory (EMBL) SMART (13) was used to identify protein domains. WebLogo was used to create sequence logos. Secondary structure and disorder regions were predicted with Jpred3 (14) and PrDOS (15), respectively.
Molecular Biology
The primers used for the amplification of intron X:6286117–6286504 were: PrimerA (5′-AGTCCTTTAAGCTGCCACAATTGGC-3′), PrimerB (5′-GAGTGTTGCCGATGAGTTGA-3′), and PrimerC (5′-CAGGTTTCCATTTCCCGTTA-3′).
Behavioral and Pharmacological Assays
Chronic (>16 h) adaptation and acute response tests to levamisole (Sigma-Aldrich, catalog number L9756) were done as described previously (6). To analyze thrashing defects, single worms were transferred to wells of 48-well plates containing M9 medium and incubated there for 1 min; head swings were counted three times per animal for 15 s. Results were reported as swings per second.
Statistics
Levamisole adaptation tests were analyzed using a nonparametric Kolmogorov-Smirnov test. Receptor clustering quantification, which fitted a normal distribution (p < 0.05, normal distribution Kolmogorov-Smirnov test), was analyzed using an unpaired non-equal variance two-sample t test.
RESULTS AND DISCUSSION
The C-terminal Region of LEV-9 Is Required for AChR Clustering
LEV-9 orthologs are readily identified in ecdysozoans including nematodes and insects. Amino acid conservation is high in the region coding the WAP and the eight CCP domains but suddenly drops 33 residues after the last CCP domain. No protein domains are predicted to be fully contained within this 66-residue C-terminal (Ct) region of LEV-9. However, the Eukaryotic Linear Motif (ELM) on-line resource (11) predicted two adjacent recognition sites for convertases: one motif for a dibasic nardilysin convertase (residues Asp-571/Arg-572/Lys-573) and another motif for subtilisin/kexin2p-like proprotein convertases (PC) (residues Lys-574/Arg-575/Ser-576) (Fig. 1A). A predicted PC cleavage signal at the same site was confirmed by the PC-specific prediction software ProP (score 0.819; false-negative prediction <0.9% (12)) (Fig. 1A). Nardilysins (NdK) are zinc proteases of the M16 peptidase family that have been recently involved in axon maturation and myelination (16). PCs are related to subtilisin and yeast kexin2p protease families that play a major role in the proteolytic processing of both neuropeptides and peptide hormone precursors (17, 18). Both motifs were found present in most, if not all, identified LEV-9 orthologs (Fig. 1, B and C).
To test whether the Ct region was required for LEV-9 function, we expressed in muscle a LEV-9 version truncated after the eighth CCP domain and tagged with GFP. Like LEV-9-GFP, L9ΔCt-GFP was detected in coelomocytes, six scavenger cells that filter the fluid of the C. elegans pseudocoelomic cavity, arguing for correct folding and secretion into the extracellular medium (Fig. 1D). L-AChR clustering was assessed and quantified using a knock-in strain engineered by homologous recombination in which the subunit UNC-29 is tagged with tagRFPt (19)(Fig. 1E). As opposed to full-length LEV-9, L9ΔCt-GFP did not rescue L-AChR clustering defects of lev-9 null mutant. Surprisingly, worms expressing L9ΔCt-GFP showed a wild-type like sensitivity to 1 mm levamisole (p > 0.5, Kolmogorov-Smirnov test) despite considerable and significant (p < 0.001, t test) lack of L-AChRs at NMJs when compared with the wild type (Fig. 1, E and F).
These data indicated for the first time that it was possible to dissociate the levamisole effect, which kills the worms and requires the N-terminal part of the LEV-9 protein, and the clustering of the levamisole receptor, which requires the C-terminal region of LEV-9. There are two main models to account for this result. First, the N-terminal part of LEV-9 might couple L-AChRs to an actuator such as a channel or a kinase that would amplify L-AChR signaling and make levamisole lethal. Second, the N-terminal part of LEV-9 might stabilize L-AChRs in the plasma membrane independently from synaptic determinants. In fact, prolonged exposure to levamisole causes partial removal of L-AChRs from the muscle cell surface.4 The absence of LEV-9 in lev-9 mutants might enhance L-AChR removal, thus decreasing levamisole toxicity. In this model, L9ΔCt would be able to stabilize the levamisole receptors in the membrane and rescue levamisole sensitivity independently from L-AChR synaptic clustering.
A LEV-9 Predicted Cleavage Site Is Biochemically Active
To test whether the cleavage sites predicted in the Ct region are functional, we used S2 cells to express a LEV-9 construct tagged with a T7 epitope at its N terminus and with a MYC tag at its C terminus (T7-L9-MYC). The S2 (Schneider-2) cell line is derived from Drosophila embryonic cells and was successfully used in the past to express C. elegans proteins (7, 9). In cell extracts, T7-L9-MYC was detected by Western blot using anti-T7 or anti-MYC antibodies, whereas it was only detected by anti-T7 antibodies in cell medium, suggesting that the C-terminal MYC tag was cleaved off the secreted protein (Fig. 2, A and B). Similar results were obtained after removing the CCP-7 or the CCP-8 domain (T7-ΔCCP7-MYC and T7-ΔCCP8-MYC, respectively, in Fig. 2, A and B). However, the MYC tag remained detected in the medium of cells expressing a tagged version with a deletion of the Ct region, suggesting that the cleavage site was contained within this region.
FIGURE 2.
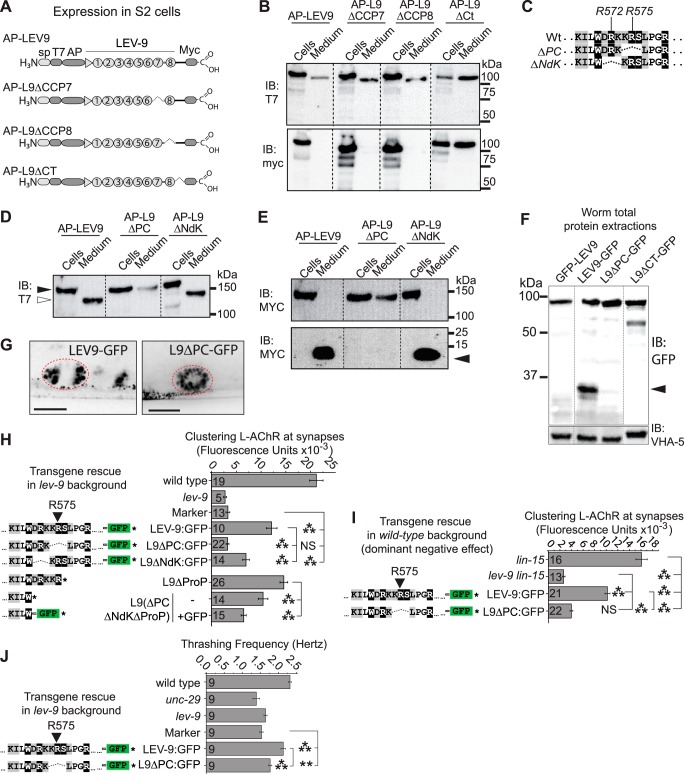
LEV-9 is cleaved within the predicted proprotein convertase site (residues Lys-574/Arg-575/Ser-576), and cleavage is required for clustering L-AChR at synapses. A, LEV-9 constructs transfected into Drosophila S2 cells. sp, signal peptide (sp); AP, alkaline phosphatase. B, immunoblots (IB) using anti-MYC and anti-T7 on samples of cell extracts and extracellular conditioned media recovered from transient transfections of S2 cells. C, sequence of constructs where the predicted nardilysin (ΔNdK) and PC (ΔPC) sites were removed. D and E, immunoblots of whole-cell extracts and extracellular media recovered from S2 cells expressing constructs lacking either of the predicted convertase sites. In D, the white arrow indicates the size shift between cells extracts (black arrow) and media. In E, the black arrow points to small cleaved C-terminal fragment. F, immunoblots using anti-GFP using whole protein extracts of C. elegans expressing different versions of LEV-9 from transgenes. The black arrowhead points to the proteolytic fragment found in LEV-9-GFP but not L9ΔPC-GFP. G, detection of GFP endocytosed by coelomocytes after expression of GFP-tagged LEV-9 constructs in muscle cells and secretion into the extracellular space. H, quantification of L-AChR-tagRFPt clustering in lev-9 worms rescued with different versions of LEV-9 expressed from transgenes. I, quantification of L-AChRs clustering in wild-type worms showing dominant-negative effect of wild-type and cleavage-resistant LEV-9 (ΔPC). J, locomotion in liquid reported as thrashing cycles per second (hertz). In all panels, columns and error bars represent averages and S.E., respectively; n numbers are shown at the base of columns. ***, **, and NS equal p < 0.001, p < 0.01, and p ≥ 0.05 in a t-Test, respectively.
To identify which of the predicted cleavage sites might be processed, we synthesized constructs lacking either the nardilysin (T7-ΔNdK-MYC) or the proprotein convertase (T7-ΔPC-MYC) sites (Fig. 2, C–E). In both ΔNdK and ΔPC, intracellular and secreted LEV-9 molecules could be detected through their N-terminal T7 epitope, but the apparent size of wild-type and ΔNdK variants decreased after secretion (Fig. 2D). No size shift could be found with ΔPC constructs (Fig. 2D). When the samples were run in a high percentage acrylamide gel (15%) and probed with the C-terminal MYC epitope, small proteolytic fragments could be found in the extracellular medium of wild-type and ΔNdK transfected cells, but not in ΔPC (Fig. 2E). The sizes of these fragments corresponded to the size expected if the constructs were cut in the Ct region. These results showed that in S2 cells LEV-9 was proteolytically processed at the predicted PC site, but processing of LEV-9 was not required for its secretion.
To verify that a similar post-translational maturation was also occurring in C. elegans, we expressed GFP-tagged LEV-9 with wild-type sequence or deletion of the protease predicted sites (L9ΔNdK-GFP and L9ΔPC-GFP in Fig. 2F). Whole protein extracts were analyzed by Western blot using anti-GFP antibodies (Fig. 2F). Partial proteolytic cleavage of the GFP was observed when LEV-9 was tagged at its C terminus, and the observed fragment corresponded to the size expected if the constructs would have been cut approximately at their Ct region. N-terminally GFP-tagged LEV-9 is also very likely cleaved at its C terminus; however, the little size difference (5 kDa) between the cleaved and uncleaved forms could not be resolved. Clipping off the C-end GFP tag was suppressed by removing either the whole Ct region or the predicted PC site, but not by removing the predicted NdK site (Fig. 2F and data not shown). In the construct where the whole Ct region was removed, an additional band of ∼50 kDa was observed; one possible explanation would be the presence of a cryptic cleaving site that could be favored when the Ct region is removed. Prevention of the cleavage did not impair secretion based on GFP detection in coelomocytes of all constructs (Fig. 2G).
LEV-9 Cleavage Is Required for L-AChR Synaptic Clustering
To test the functional significance of LEV-9 cleavage at the PC site, we tested the activity of L9ΔPC-GFP and demonstrated that it was no longer rescuing L-AChR clustering, whereas it restored levamisole sensitivity, similar to the truncated LEV-9 version lacking the entire Ct region (Figs. 1A and 2H and data not shown). Worms expressing LEV-9 lacking the nonfunctional NdK site (residues Asp-571/Arg-572/Lys-573 removed) partially rescued L-AChR clustering, but not as efficiently as wild-type LEV-9. This suggested that the last residues present just before the C terminus generated by cleavage at position Arg-575 were important for LEV-9 clustering function. Interestingly, a CCP7-CCP8-Ct construct only partially rescues AChR clustering, suggesting that the Ct functionally interacts with other LEV-9 regions.5 Alternatively, the short C terminus peptide release by cleavage might also have some biological activity. Expression of a truncated LEV-9 version ending at position Arg-575 (L9ΔProP) was as efficient as WT LEV-9 (Fig. 2H). Further removal of 5 residues reduced the clustering activity. Adding a GFP at the end of this truncated version completely abolished the activity of the protein. These data suggested that the C-terminal propeptide had no intrinsic activity and that unmasking the C-terminal residues before position Arg-575 is required to activate L-AChR clustering activity.
Because LEV-9 is part of multimolecular scaffold, we tested whether the processing-resistant LEV-9 variant would behave as a dominant-negative. In fact, overexpression of the processing-resistant LEV-9 variant in a wild-type background completely abrogated the clustering of L-AChRs (p < 0.001, t test) (Fig. 2I). Overexpression of wild-type LEV-9 partially reduced the number of L-AChR at NMJs (p < 0.001, t test), suggesting that LEV-9 expression or activity is tightly regulated to achieve proper L-AChR clustering.
LEV-9 Cleavage Is Required for Proper Function of the Neuromuscular Junction
Because LEV-9 cleavage is required for proper L-AChR positioning but can be inhibited without affecting the sensitivity to levamisole, we wondered whether LEV-9 processing modulated synaptic activity. We therefore quantified the coordinated movement of worms in liquid, commonly referred to as thrashing. Animals lacking L-AChRs, such as unc-29 mutants, or lacking LEV-9 have a significant thrashing frequency decrease as compared with wild-type worms (Fig. 2J). Thrashing rescue of lev-9 mutants was significantly more efficient with wild-type LEV-9 than with a cleavage-resistant LEV-9 version, indicating that LEV-9 processing is required for proper function of the neuromuscular junction. Altogether, these results suggest that cleaving LEV-9 at its C-end region is a limiting factor for L-AChR clustering and synaptic function.
Proteolytic Processing Generates a Constant LEV-9 Product out of Multiple lev-9 Transcripts
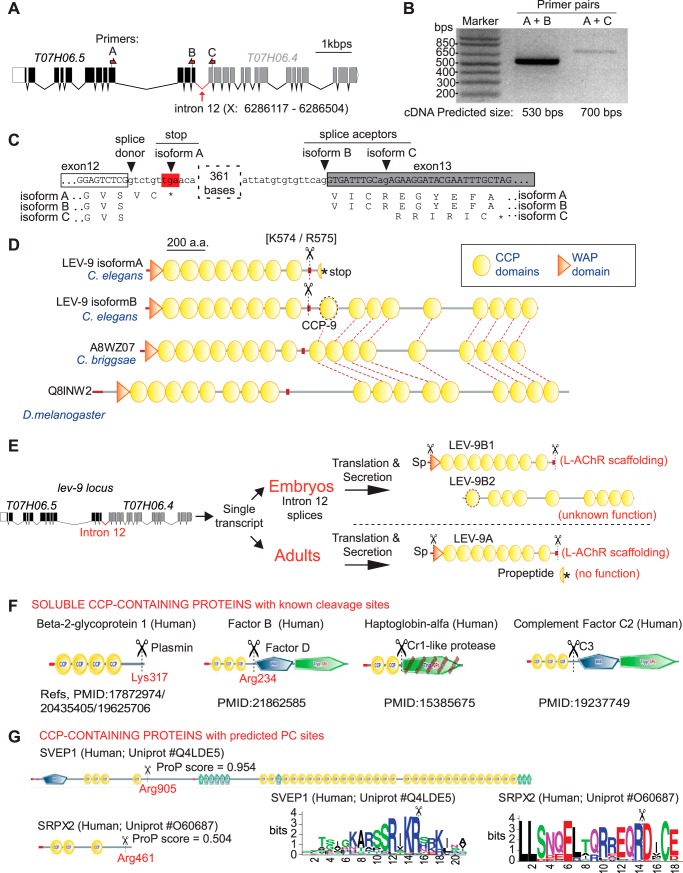
We previously demonstrated that lev-9 is encoded by the T07H6.5 gene because this genomic region rescues all synaptic defects of lev-9 mutants (7). A second gene, T07H6.4, located 300 bp downstream to T07H6.5, is predicted to code for a CCP-containing polypeptide. However, it lacks an identifiable signal peptide in contrast to LEV-9, and most CCP-containing proteins are usually extracellular. Disruption of T07H6.4 by a Tc1 transposon insertion causes no synaptic phenotype (7). Recent data from the modENCODE consortium annotated the genomic region between T07H6.5 and T07H6.4 as the intron of a putative T07H6.5-.4 joined transcript expressed in embryos, mid-L1 and dauer larvae (Fig. 3A) (modENCODE Project website) (20). Using RT-PCR on a mixed staged cDNA library, we were able to amplify a low abundance fragment containing both T07H6.5 last exon and T07H6.4 first exon (Fig. 3, A and B). Sequencing this RT-PCR product revealed that two distinct splice acceptors are used in T07H6.4 (Fig. 3C), generating two novel lev-9 transcripts. lev-9-B is a predicted full-length T07H6.5-T07H6.4 in-frame transcript, whereas in isoform C, a reading frameshift causes a nonsense mutation 18 bps downstream from the acceptor site. Interestingly, the lev-9B mRNA region covering the junction between T07H6.5 and T07H6.4 encodes a conserved cryptic CCP domain (CCP-9 in Fig. 3D). Such organization of the lev-9 locus is evolutionarily conserved. In Drosophila melanogaster and C. briggsae for example, a first region homologous to T07H6.5 encodes a predicted signal peptide, an N-terminal WAP domain, and eight CCP domains and is followed by a region homologous to T07H6.4 encoding nine CCP domains but no signal peptide (Fig. 3D). The proteolytic cleavage site after the eight CCP domain is readily detectable in all species. These results suggest that at least three distinct propeptides are encoded by the lev-9 locus and are able to generate the active LEV-9 scaffolding protein by cleavage at position Arg-575. The function of the C-terminal peptide of LEV-9B containing nine CCP domains is yet to be determined (Fig. 3E).
FIGURE 3.
LEV-9 cleavage ensures structural and functional decoupling of the multiple transcripts generated at lev-9 locus. A, genomic regions of T07H6.5 and T07H6.4; in embryos, the region between T07H6.5 and T07H6.4 is recognized as intron X:6286117–6286504. B, RT-PCR products from a mixed stage cDNA library using T07H6.5-only and T07H6.5/T07H6.4 annealing primers (shown in A as pairs A/B and A/C, respectively). C, isoforms B and C found by sequencing of amplicon A/C (in B). D, protein domain composition of LEV-9 isoforms A and B and LEV-9 orthologs in C. briggsae and D. melanogaster. A cryptic conserved CCP domain is encoded by T07H06.5 last and T07H06.4 first exons (dotted oval). Conserved cleavage-site is indicated by a small red box. Red dotted lines indicate homologous regions within orthologs. 200 a.a., 200 amino acids. E, model in which LEV-9 cleavage ensures structural and functional decoupling throughout development stages. Sp, signal peptide. F, examples of reported cases where C-terminal cleavage of CCP tandems regulate function in secreted plasma proteins in vertebrates. G, two examples in which PC predicted sites were found at the C-end of CCP-containing secreted proteins using ProP software.
Given our current results, we wondered to what extent the C-terminal cleavage of LEV-9 might be extended to other CCP-containing secreted factors. Interestingly, for several proteins containing tandems of CCP domains, it has been experimentally determined that cleavage is required. Examples include abundant factors secreted in human serum such as the 2-glycoprotein I or the complement factor C2, in addition to membrane-bound factors such as L-selectins and membrane cofactor protein CD46 (21–24) (Fig. 3F and supplemental Table 1). Cleavage serves either to activate function or to decouple CCP regulatory domains from catalytic domains (21–24). Interestingly, it occurs mostly C-terminally from tandems of CCP domains, as is the case for LEV-9 in C. elegans. Furthermore, we found conserved predicted PC sites C-terminally from CCP tandems in additional membrane-bound and secreted human factors (Fig. 3G and supplemental Table 1). This analysis therefore suggests that cleaving C-terminally from CCP domains to either control activity or structurally decouple CCP domains from C-terminal attachments is an architectural principle that has been conserved in proteins of unrelated function.
Supplementary Material
Acknowledgments
We are grateful to Marie Gendrel for sharing preliminary observations and Maelle Jospin and Lionel Pintard for critical reading of the manuscript.
This work was supported by INSERM, Agence Nationale de la Recherche (ANR) Grant 11-BSV4-019, and the Association Française contre les Myopathies.

This article contains supplemental text and Table 1.
L. Briseño-Roa and J.-L. Bessereau, unpublished results.
L. Briseño-Roa, M. Gendrel, and J.-L. Bessereau, manuscript in preparation.
- NMJ
- neuromuscular junction
- L-AChR
- levamisole-sensitive acetylcholine receptor
- CCP
- complement control protein
- PC
- subtilisin/kexin2p-like proprotein convertase(s)
- NdK
- nardilysin(s)
- Ct
- C-terminal
- NS
- not significant.
REFERENCES
- 1. Kim E., Sheng M. (2004) PDZ domain proteins of synapses. Nat. Rev. Neurosci. 5, 771–781 [DOI] [PubMed] [Google Scholar]
- 2. Tyagarajan S. K., Ghosh H., Yévenes G. E., Nikonenko I., Ebeling C., Schwerdel C., Sidler C., Zeilhofer H. U., Gerrits B., Muller D., Fritschy J.-M. (2011) Regulation of GABAergic synapse formation and plasticity by GSK3β-dependent phosphorylation of gephyrin. Proc. Natl. Acad. Sci. U.S.A. 108, 379–384 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Lu X., Wyszynski M., Sheng M., Baudry M. (2001) Proteolysis of glutamate receptor-interacting protein by calpain in rat brain: implications for synaptic plasticity. J. Neurochem. 77, 1553–1560 [DOI] [PubMed] [Google Scholar]
- 4. Vinade L., Petersen J. D., Do K., Dosemeci A., Reese T. S. (2001) Activation of calpain may alter the postsynaptic density structure and modulate anchoring of NMDA receptors. Synapse 40, 302–309 [DOI] [PubMed] [Google Scholar]
- 5. Cho R. W., Park J. M., Wolff S. B. E., Xu D., Hopf C., Kim J.-A., Reddy R. C., Petralia R. S., Perin M. S., Linden D. J., Worley P. F. (2008) mGluR1/5-dependent long-term depression requires the regulated ectodomain cleavage of neuronal pentraxin NPR by TACE. Neuron 57, 858–871 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Gally C., Eimer S., Richmond J. E., Bessereau J.-L. (2004) A transmembrane protein required for acetylcholine receptor clustering in Caenorhabditis elegans. Nature 431, 578–582 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Gendrel M., Rapti G., Richmond J. E., Bessereau J.-L. (2009) A secreted complement-control-related protein ensures acetylcholine receptor clustering. Nature 461, 992–996 [DOI] [PubMed] [Google Scholar]
- 8. Nakayama M., Hama C. (2011) Modulation of neurotransmitter receptors and synaptic differentiation by proteins containing complement-related domains. Neurosci. Res. 69, 87–92 [DOI] [PubMed] [Google Scholar]
- 9. Rapti G., Richmond J., Bessereau J.-L. (2011) A single immunoglobulin-domain protein required for clustering acetylcholine receptors in. C. elegans. EMBO J. 30, 706–718 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Wallace I. M., O'Sullivan O., Higgins D. G., Notredame C. (2006) M-Coffee: combining multiple sequence alignment methods with T-Coffee. Nucleic Acids Res. 34, 1692–1699 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Puntervoll P. (2003) ELM server: a new resource for investigating short functional sites in modular eukaryotic proteins. Nucleic Acids Res. 31, 3625–3630 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Duckert P., Brunak S., Blom N. (2004) Prediction of proprotein convertase cleavage sites. Protein Eng. Des. Sel. 17, 107–112 [DOI] [PubMed] [Google Scholar]
- 13. Schultz J., Milpetz F., Bork P., Ponting C. P. (1998) SMART, a simple modular architecture research tool: identification of signaling domains. Proc. Natl. Acad. Sci. U.S.A. 95, 5857–5864 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Cole C., Barber J. D., Barton G. J. (2008) The Jpred 3 secondary structure prediction server. Nucleic Acids Res. 36, W197–W201 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Ishida T., Kinoshita K. (2007) PrDOS: prediction of disordered protein regions from amino acid sequence. Nucleic Acids Res. 35, W460–W464 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Ohno M., Hiraoka Y., Matsuoka T., Tomimoto H., Takao K., Miyakawa T., Oshima N., Kiyonari H., Kimura T., Kita T., Nishi E. (2009) Nardilysin regulates axonal maturation and myelination in the central and peripheral nervous system. Nat. Neurosci. 12, 1506–1513 [DOI] [PubMed] [Google Scholar]
- 17. Fuller R. S., Brake A., Thorner J. (1989) Yeast prohormone processing enzyme (KEX2 gene product) is a Ca2+-dependent serine protease. Proc. Natl. Acad. Sci. U.S.A. 86, 1434–1438 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Seidah N. G. (2011) Proprotein Convertases (Mbikay M., Seidah N. G., eds), Humana Press, Totowa, NJ [Google Scholar]
- 19. Richard M., Boulin T., Robert V. J. P., Richmond J. E., Bessereau J.-L. (2013) Biosynthesis of ionotropic acetylcholine receptors requires the evolutionarily conserved ER membrane complex. Proc. Natl. Acad. Sci. U.S.A. 110, E1055–E1063 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Gerstein M. B., Lu Z. J., Van Nostrand E. L., Cheng C., Arshinoff B. I., Liu T., Yip K. Y., Robilotto R., Rechtsteiner A., Ikegami K., Alves P., Chateigner A., Perry M., Morris M., Auerbach R. K., Feng X., Leng J., Vielle A., Niu W., Rhrissorrakrai K., Agarwal A., Alexander R. P., Barber G., Brdlik C. M., Brennan J., Brouillet J. J., Carr A., Cheung M.-S., Clawson H., Contrino S., Dannenberg L. O., Dernburg A. F., Desai A., Dick L., Dosé A. C., Du J., Egelhofer T., Ercan S., Euskirchen G., Ewing B., Feingold E. A, Gassmann R., Good P. J., Green P., Gullier F., Gutwein M., Guyer M. S., Habegger L., Han T., Henikoff J. G., Henz S. R., Hinrichs A., Holster H., Hyman T., Iniguez a L., Janette J., Jensen M., Kato M., Kent W. J., Kephart E., Khivansara V., Khurana E., Kim J. K., Kolasinska-Zwierz P., Lai E. C., Latorre I., Leahey A., Lewis S., Lloyd P., Lochovsky L., Lowdon R. F., Lubling Y., Lyne R., MacCoss M., Mackowiak S. D., Mangone M., McKay S., Mecenas D., Merrihew G., Miller D. M., Muroyama A., Murray J. I., Ooi S.-L., Pham H., Phippen T., Preston E. a, Rajewsky N., Rätsch G., Rosenbaum H., Rozowsky J., Rutherford K., Ruzanov P., Sarov M., Sasidharan R., Sboner A., Scheid P., Segal E., Shin H., Shou C., Slack F. J., Slightam C., Smith R., Spencer W. C., Stinson E. O., Taing S., Takasaki T., Vafeados D., Voronina K., Wang G., Washington N. L., Whittle C. M., Wu B., Yan K.-K., Zeller G., Zha Z., Zhong M., Zhou X., Ahringer J., Strome S., Gunsalus K. C., Micklem G., Liu X. S., Reinke V., Kim S. K., Hillier L. W., Henikoff S., Piano F., Snyder M., Stein L., Lieb J. D., Waterston R. H. (2010) Integrative analysis of the Caenorhabditis elegans genome by the modENCODE project. Science 330, 1775–1787 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Nakagawa H., Yasuda S., Matsuura E., Kobayashi K., Ieko M., Kataoka H., Horita T., Atsumi T., Koike T. (2009) Nicked β2-glycoprotein I binds angiostatin 4.5 (plasminogen kringle 1–5) and attenuates its antiangiogenic property. Blood 114, 2553–2559 [DOI] [PubMed] [Google Scholar]
- 22. Krishnan V., Xu Y., Macon K., Volanakis J. E., Narayana S. V. L. (2009) The structure of C2b, a fragment of complement component C2 produced during C3 convertase formation. Acta Crystallogr. D. Biol. Crystallogr. 65, 266–274 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Hakulinen J., Junnikkala S., Sorsa T., Meri S. (2004) Complement inhibitor membrane cofactor protein (MCP; CD46) is constitutively shed from cancer cell membranes in vesicles and converted by a metalloproteinase to a functionally active soluble form. Eur. J. Immunol. 34, 2620–2629 [DOI] [PubMed] [Google Scholar]
- 24. Smalley D. M., Ley K. (2005) L-selectin: mechanisms and physiological significance of ectodomain cleavage. J. Cell. Mol. Med. 9, 255–266 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.