Background: Oxysterols are a class of emerging signaling molecules whose cell biology is poorly understood.
Results: A click chemistry-based imaging strategy shows that 20(S)-hydroxycholesterol accumulates in Golgi membranes in a process that depends on ATP and lysosome function.
Conclusion: 20(S)-Hydroxycholesterol is transported through a vesicular pathway to the Golgi.
Significance: Specific transport pathways may regulate the oxysterol content of cellular membranes.
Keywords: Imaging, Lipids, Lysosomes, Signaling, Trafficking, Oxysterol
Abstract
Oxysterols, oxidized metabolites of cholesterol, are endogenous small molecules that regulate lipid metabolism, immune function, and developmental signaling. Although the cell biology of cholesterol has been intensively studied, fundamental questions about oxysterols, such as their subcellular distribution and trafficking pathways, remain unanswered. We have therefore developed a useful method to image intracellular 20(S)-hydroxycholesterol with both high sensitivity and spatial resolution using click chemistry and fluorescence microscopy. The metabolic labeling of cells with an alkynyl derivative of 20(S)-hydroxycholesterol has allowed us to directly visualize this oxysterol by attaching an azide fluorophore through cyclo-addition. Unexpectedly, we found that this oxysterol selectively accumulates in the Golgi membrane using a pathway that is sensitive to ATP levels, temperature, and lysosome function. Although previous models have proposed nonvesicular pathways for the rapid equilibration of oxysterols between membranes, direct imaging of oxysterols suggests that a vesicular pathway is responsible for differential accumulation of oxysterols in organelle membranes. More broadly, clickable alkynyl sterols may represent useful tools for sterol cell biology, both to investigate the functions of these important lipids and to decipher the pathways that determine their cellular itineraries.
Introduction
Charting the spatial compartmentalization of signal transduction in cells has required the development of probes to reveal the subcellular localization of both signaling proteins and small molecule second messengers. The chemical diversity of endogenous regulatory small molecules precludes the use of a single strategy for their visualization and has necessitated the development of specific probes for each class (1), such as cAMP sensors based on FRET (2), calcium indicator dyes (3), and fluorescent protein-tagged domains that recognize inositol phospholipids (4). In this study, we describe a method to visualize the subcellular localization of an emerging class of signaling lipids called oxysterols, oxidized metabolites of cholesterol.
In addition to being intermediates in the synthesis of bile acids and steroids, oxysterols have diverse regulatory roles in lipid metabolism, immune responses, and developmental signaling (5). Oxysterols have three properties distinct from cholesterol that make them ideally suited to function as second messengers in signaling pathways as follows: cellular concentrations 3 orders of magnitude lower than cholesterol (6, 7); transfer rates between membranes that are ∼50-fold higher than cholesterol (8–10), and shorter half-lives compared with cholesterol (11).
Oxysterols have been most intensively studied for their signaling role in regulating cholesterol homeostatic pathways. They regulate SREBP (12) and LXR (13, 14) proteins, two classes of transcription factors that control a suite of genes involved in sterol uptake and metabolism. More recently, oxysterols have been implicated in several other biological processes. They can regulate midbrain neurogenesis by acting as direct ligands for LXR proteins (15), and they are agonists for two 7-pass transmembrane receptors, Epstein-Barr virus-induced gene 2 (EBI2) protein (16, 17) and Smoothened (SMO) (18). Gradients of oxysterols in lymphoid tissues guide activated B-cells through EBI2 during humoral responses (19). Oxysterols can activate Hedgehog (Hh) transcriptional responses and cell differentiation decisions by inducing the accumulation of SMO in the primary cilium (20–22). A family of oxysterol-binding proteins called the ORP family (oxysterol-binding protein-related protein) has been implicated in vesicle trafficking (23, 24), cancer cell physiology (25), and intracellular lipid transport (26, 27). Oxysterols also have effects in other contexts that are less well understood. For example, oxysterols, synthesized during the interferon response, can have significant antiviral effects (28, 29).
In addition to acting through protein-mediated transcriptional pathways, oxysterols exert their effects through modulation of the biophysical properties of membranes due to their greater polarity and poorer packing with phospholipids in comparison with cholesterol (30–32). Consequently, the oxysterol content of membranes can influence the activity of embedded proteins and lipids (33, 34). These “nongenomic” effects have been specifically implicated in the regulation of cholesterol homeostatic pathways but are likely to be generally important for oxysterol signaling (35, 36).
In contrast to the case for cholesterol, the cell biology of oxysterols remains poorly understood (37). Remarkably, the subcellular localization of oxysterols is unknown. We do not know whether oxysterols distribute uniformly among cellular membranes (as has been suggested based on their proclivity for rapid inter-membrane transfer) or whether they localize in specific membrane compartments or form microdomains. The mechanisms by which oxysterols are imported into cells and then distributed to various organelles also remain obscure.
Here, we report a method to study the subcellular localization of oxysterols with both high sensitivity and high spatial resolution. We focused on 20(S)-hydroxycholesterol (20(S)-OHC),3 an agonist of SMO. The labeling of cells with an alkynyl derivative 20(S)-OHC, called 20(S)-yne, allowed us to visualize this oxysterol by attaching a fluorophore using a bioorthogonal click reaction, alkyne-azide cycloaddition. The fluorescent adduct could be visualized in cells with high precision using confocal fluorescence microscopy. Pulse-chase analysis in combination with quantitative fluorescence microscopy revealed an unexpected active transport pathway, dependent on lysosome function, which culminates in the accumulation of 20(S)-yne in the Golgi apparatus.
EXPERIMENTAL PROCEDURES
Cell Culture and 20(S)-yne Labeling
NIH/3T3 cells were cultured in high glucose DMEM supplemented with 10% fetal bovine serum (FBS), and CHO cells were cultured in F-12K with 10% FBS. Cells were plated at subconfluent densities on coverslips for 20(S)-yne labeling and subsequent imaging. Sterol incubation and media chases were performed in culture media with 0.5% FBS. Unless otherwise noted, standard conditions were 1 μm 20(S)-yne for 1 h, with or without a subsequent 1-h chase with media lacking 20(S)-yne. For competition treatment, cells were incubated in 20(S)-yne and 20(S)-OHC or ent-20(S)-OHC (18) simultaneously at a 1:10 ratio, for a total of 11 μm oxysterol. Cyclodextrin conjugates of sterols were generated using previously described methods (38). A 10 mm sterol stock solution in ethanol was diluted to 1 mm in PBS containing 10 mm methyl-β-cyclodextrin (MBCD; Sigma) and rotated overnight at room temperature. The solution was filtered through a 0.22- μm syringe filter prior to use.
For small molecule inhibitors, cells were pretreated for 30 min with drug (or a matched solvent control) prior to the assessment of 20(S)-yne uptake as described above. Drugs were maintained throughout the experiment. The concentration of drugs used are as follows: 10 μg/ml brefeldin A, 10 mm NH4Cl, 100 μm chloroquine, and 10–20 mm NaN3 + 5–10 mm deoxyglucose for ATP depletion. To measure 20(S)-yne uptake at various temperatures, cells were cooled for 10 min to the indicated temperature prior to 20(S)-yne labeling.
Click Chemistry
After 20(S)-yne labeling, cells were fixed in 4% paraformaldehyde for 10 min at room temperature, washed with TBS (50 mm Tris, pH 7.6, 150 mm NaCl), and subjected to a click reaction on the coverslip. Except when explicitly stated, the click reaction was performed with 1 μm Alexa 594/488-azide (Invitrogen) in 100 mm Tris, pH 8.8, 1 mm CuSO4, 100 mm ascorbic acid. The click reaction was performed for 30 min in the dark, followed by one 5-min wash each with 40 mg/ml BSA in TBS, 0.5 m NaCl, and TBS. Coverslips were mounted onto slides with Prolong Gold Antifade Reagent with DAPI (Invitrogen) for imaging.
Inhibitor Treatment
Cells were pretreated 30 min with inhibitors or solvent control and then incubated with 20(S)-yne plus inhibitors, sometimes followed by media chase including inhibitors but not 20(S)-yne. Inhibitors used are as follows: 10 μg/ml brefeldin A, 10 mm NH4Cl, 100 μm chloroquine, and 10–20 mm NaN3 + 5–10 mm deoxyglucose for ATP depletion. For temperature dependence, cells were cooled for 10 min at room temperature (20–22 °C) or 15 °C prior to 20(S)-yne labeling at each temperature.
Immunofluorescence
Cells were transfected with FP-tagged constructs using FuGENE HD (Promega). After several hours for expression of FP-tagged proteins, cells were re-plated to coverslips for 20(S)-yne labeling. For immunofluorescence, cells were permeabilized after the click reaction and subsequent washes. For Golgi staining in NIH/3T3 cells, cells were permeabilized with 0.003% digitonin during the 30-min blocking step (TBS with 1% normal donkey serum and 10 mg/ml BSA). For lysosome staining in CHO cells, cells were permeabilized with 0.03% saponin during the 30 min blocking step, and saponin was present in all following steps. Coverslips were incubated in primary antibody diluted in blocking buffer for 2 h at room temperature, followed by three 5-min washes with TBS. Secondary antibody was then added (also in blocking buffer) and incubated at room temperature for 1 h. The coverslips were once again washed three times in TBS and then mounted using Prolong Gold Antifade reagent. Antibody dilutions were as follows: anti-giantin-488 1:500, mouse anti-LGP95 sera undiluted, anti-mouse-Alexa 488 1:500.
Data Acquisition and Analysis
Images were acquired on a Leica SP5 laser scanning confocal microscope using a HyD hybrid detector for 20(S)-yne imaging. For quantification of 20(S)-yne uptake, images were acquired with identical laser power, gain, and offset settings for all experimental conditions from 4 to 5 disparate areas of the coverslip and managed identically in ImageJ or Adobe Photoshop. Cell Profiler 2.0 (39) was used to identify nuclei and cells and to measure the total fluorescence intensity of 20(S)-yne label per cell. The upper and lower 10% of values from each image were discarded to remove software error in identifying and segmenting objects. For graphs and statistical analysis, GraphPad Prism software was used. In most cases a 1-way ANOVA with a Bonferroni multiple comparison test was used to determine statistical significance. In Fig. 6D an unpaired t test was used.
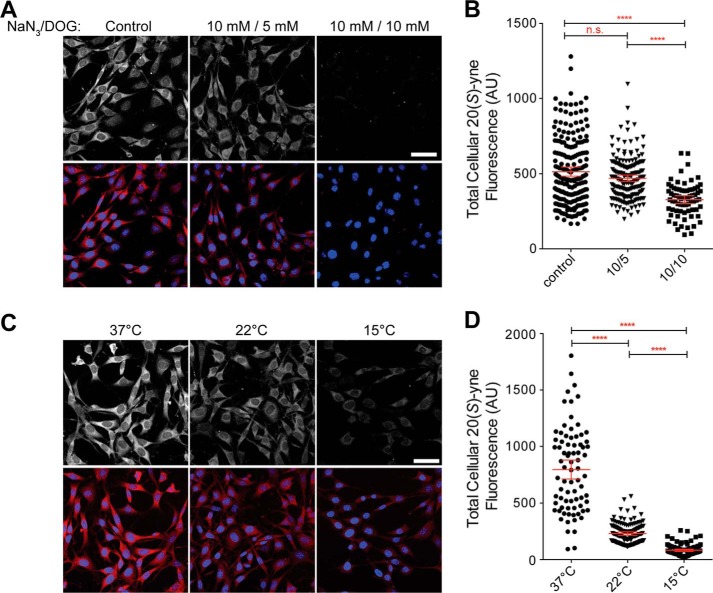
FIGURE 6.
20(S)-yne accumulation is dependent on cellular ATP levels and on temperature. 20(S)-yne accumulation (red) in cells treated with the indicated concentrations of azide and deoxyglucose to reduce ATP levels (A and B) or in cells cooled to the indicated temperatures (C and D). Scale bar, 50 μm. Representative images are shown in A and C, and quantitative analysis of total cellular 20(S)-yne fluorescence measured from individual cells across multiple images is presented in B and D. Each point depicts the total 20(S)-yne level in an individual cell, and red bars show the mean level across the population (±95% confidence interval). In both B and D, statistical significance was determined by 1-way ANOVA plus Bonferroni multiple comparison test (****, p < 0.0001, n.s., not significant, n >70 for each condition). AU, arbitrary units.
Throughout the figures, unless otherwise indicated, images are shown after acquisition with identical settings to allow direct comparison of intensities. However, in some panels (bottom rows of the panels in Figs. 2B, and 5, A–C) the images are acquired with optimal (and thus different) settings to best show subcellular distribution of 20(S)-yne.
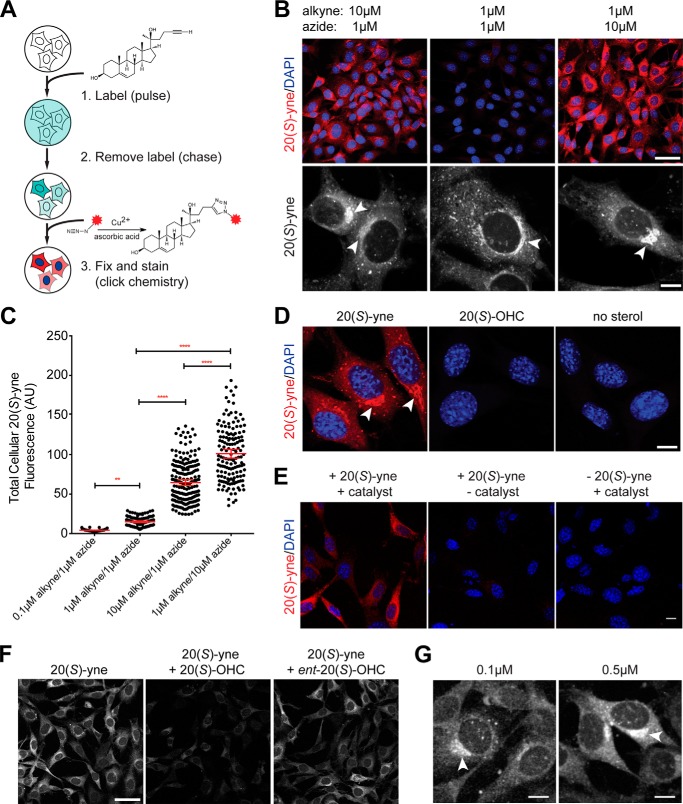
FIGURE 2.
20(S)-yne labels specific membranes in NIH/3T3 cells. A, pulse-chase scheme for measuring the uptake of 20(S)-yne. After the chase, cells were fixed, and the 20(S)-yne was detected by conjugating it to Alexa 594-azide (red asterisk). B, Alexa 594 staining intensity (red, top row) and distribution (white, bottom row) in cells using the indicated concentrations of 20(S)-yne for labeling and Alexa 594-azide for the click reaction. The low magnification images (top row, scale bar, 50 μm) were taken with the same settings to directly compare intensities, and the high magnification images (bottom row, scale bar, 10 μm) were taken with different settings to optimally show the subcellular distribution of 20(S)-yne. C, quantification and statistical analysis of total cellular 20(S)-yne fluorescence from multiple images of the type shown in B. Each point denotes the total fluorescence in a single cell. D, Alexa 594-azide fluorescence (red) in cells incubated with 20(S)-yne, 20(S)-OHC, or no sterol. Scale bar, 10 μm. E, no sterol and no catalyst controls for 20(S)-yne staining. 20(S)-yne staining of the Golgi apparatus is fully dependent on the presence of 20(S)-yne and the cycloaddition reaction; removing either the catalyst (middle panel) or 20(S)-yne (right panel) eliminated staining. Scale bar, 10 μm. F, 20(S)-yne uptake in cells incubated with 20(S)-yne alone (left panel) or 20(S)-yne in the presence of a 10-fold excess of 20(S)-OHC (middle panel) or ent-20(S)-OHC (right panel). Scale bar, 50 μm. G, Alexa 594 staining in a peri-nuclear compartment was seen even after incubation with low levels of 20(S)-yne. C, red bars show the mean fluorescence (±95% confidence interval). All populations are significantly different from each other (**, p < 0.01, and ****, p < 0.0001, using 1-way ANOVA plus Bonferroni multiple comparison test, n >45 for each condition). In all images, DAPI (blue) is used to stain nuclei, and arrowheads point to an asymmetric perinuclear focus of 20(S)-yne accumulation. AU, arbitrary units.
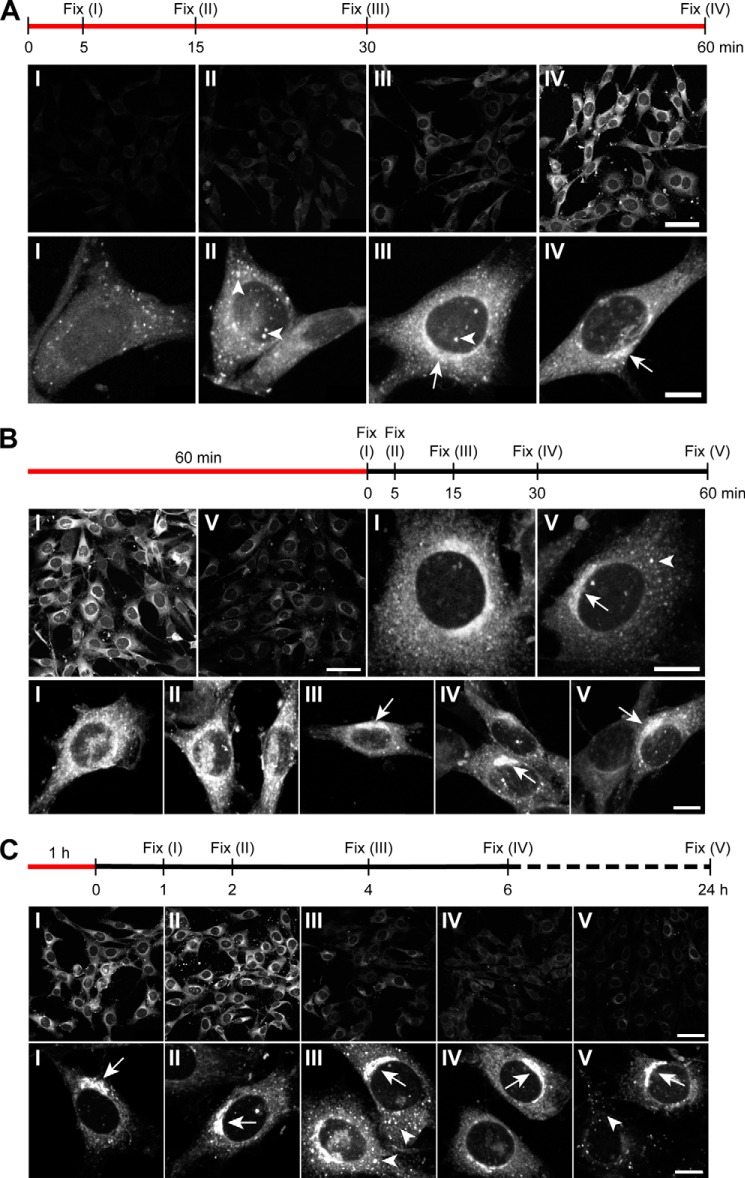
FIGURE 5.
Kinetics of 20(S)-yne accumulation and retention in the Golgi. The sequence of metabolic labeling prior to fixation is indicated above each panel, with red lines denoting the duration of the 20(S)-yne pulse and the black line denoting the duration of the sterol-free chase. The images at low magnification (top rows of each panel, scale bar, 50 μm) were taken at identical settings to compare intensities at different time points, whereas the high magnification images (bottom row of each panel, scale bar, 10 μm) were taken with different settings to optimally show the subcellular pattern of staining. Arrowheads and arrows indicate punctate and asymmetric perinuclear staining, respectively, and the roman numerals on the images correspond to the point on the timeline shown above each panel. A, 20(S)-yne signal intensity (top row) and distribution (bottom row) after various times of incubation. A representative image depicting the most prevalent staining pattern for each time point is shown in the bottom row. B and C show 20(S)-yne staining after a short and long chase, respectively.
Measurement of Sterols by Mass Spectrometry
Cell pellets were harvested and resuspended in phosphate-buffered saline (PBS). An aliquot was used to measure DNA content, which was used to normalize measured oxysterol levels. Deuterated 27-hydroxycholesterol (OHC)-d5 was added to the second aliquot as an internal standard for quantification of the oxysterols. The oxysterol extraction from cell pellets was performed by Bligh-Dyer extraction method, followed by N,N-dimethylglycine derivatization (40) of sterols for improvement of MS sensitivity of sterol detection.
The N,N-dimethylglycine derivatized oxysterols were quantified using the multiple reaction monitoring method from positive ion electrospray ionization using an API-4000 quadrupole mass spectrometer interfaced to a Leap PAL autosampler, Shimadzu 20 AD HPLC system, and Agilent 1100 HPLC. The quantification was performed by Analyst software (Applied Biosystems) against calibration curves of oxysterols (20(S)-OHC and 20(S)-yne) with the deuterated 27-OHC internal standard.
Quantitative RT-PCR
3T3 cells (3.5 × 105) were seeded in a 24-well plate and cultured until they reached confluency. Cells were treated for 16 h in 0.5% FBS DMEM with DMSO (0.05% v/v), 5 μm 20(S)-OHC, or 5 μm 20(S)-yne. TRIzol reagent (Invitrogen) was used to extract RNA following the manufacturer's instructions. The following primers were used: Gapdh for, ggccttccgtgtttc, and rev, tgtcatcatacttg; Gli1 for, ccaagccaaacttta, and rev, agcccgcttctttg; Ldlr for, ccaaatggcatcacactagatctt, and rev, cgattgcccccattgaca; Srebp2 for, gaccagcacccatactcag, and rev, acaccatttaccagccacag; Hmgcr for, tgtggtttgtgaagccgtcat, and rev, tcaaccatagcttccgtagttgtc; Hmgcs1 for, gggccaaacgctcctctaat, and rev, agtcataggcatgctgcatgtg; and Abca1 for, ggtttggagatggttatacaatagtt, and rev, ttcccggaaacgcaagtc.
Fluorescence Dequenching Assays
Liposomes containing carboxyfluorescein were prepared using a reverse phase vaporization technique. Briefly, 10 mg of 1,2-dioleoyl-sn-glycero-3-phosphocholine (Avanti Polar Lipids) were dissolved in chloroform and dried to a film using a rotary evaporator (Buchi Rotavapor R-210). The lipid film was placed under vacuum at 50 °C overnight to remove residual chloroform. Afterward, 500 μl of diethyl ether and 500 μl of 20 mm carboxyfluorescein in 115 mm NaCl, 10 mm Tris, pH 7.0, buffer were added to the film, and the mixture was sonicated three times for 20 s each using a probe sonicator (Branson Sonifier 250). The ether was removed by rotary evaporation, and the resulting vesicles were sized by extrusion through a 200-nm polycarbonate filter (Nucleopore). Free carboxyfluorescein was then removed by size exclusion chromatography using Sephadex G-25 beads (Sigma). The measured liposome size was 188.2 nm, determined by dynamic light scattering using a Zetaplus analyzer (Brookhaven Instruments Corp.). For fluorescence dequenching assays, a 3.3 μg/ml preparation of liposomes in 115 mm NaCl, 10 mm Tris, pH 7.0, buffer was used, and baseline fluorescence was observed for 2 min using a Cary Eclipse fluorescence spectrophotometer (Varian). A 5 mm stock solution of 20(S)-OHC or 20(S)s-yne in isopropyl alcohol was then added to achieve the appropriate sterol concentration, and fluorescence was observed for 8 min. To calculate fractional dequenching, average baseline fluorescence was subtracted from the observed fluorescence signal, and the signal was normalized to lysis by 1% Triton X-100.
RESULTS
20(S)-yne Labels Specific Membranes in NIH/3T3 Cells
Although 20(S)-OHC has been previously detected in tissues (7, 41), very little is known about its biosynthetic pathways, distribution, or trafficking. In our previous studies of Hh signaling, we synthesized 20(S)-yne, an alkynyl derivative of 20(S)-OHC, that retained activity as a direct SMO agonist (Fig. 1A) (18). In that study, the alkynyl group was used to immobilize 20(S)-yne on a solid support using click cycloaddition for affinity chromatography. Given the widespread availability of azide-linked fluorophores, we asked if this same bio-orthogonal reaction could be exploited to reveal the subcellular location of 20(S)-yne.
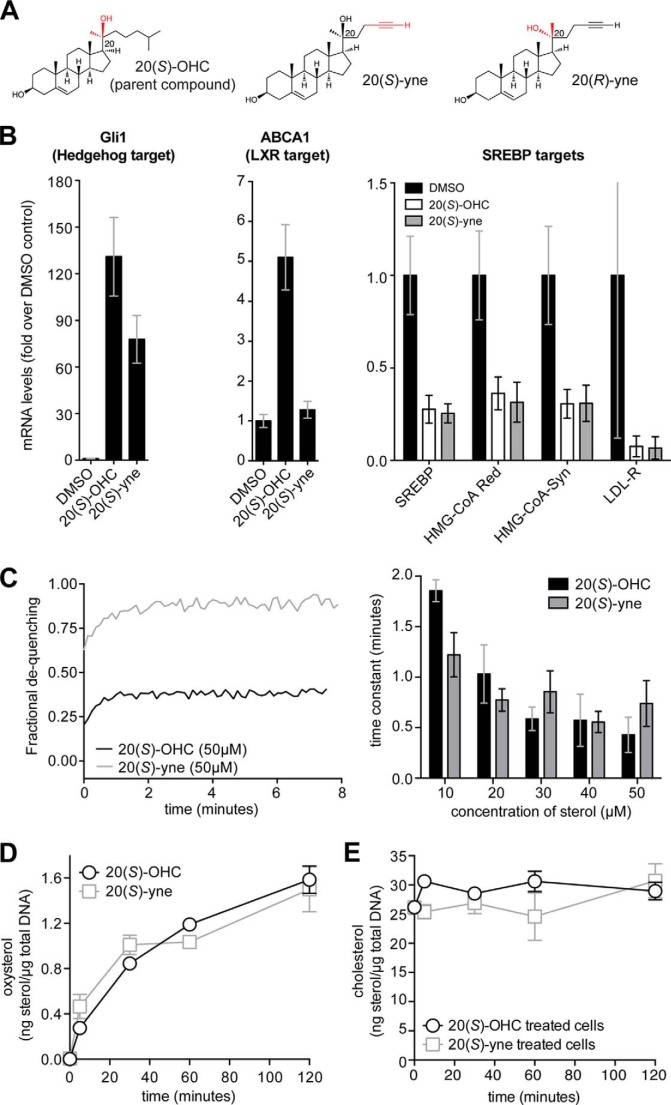
FIGURE 1.
Comparison of 20(S)-yne and 20(S)-OHC. A, 20(S)-yne is a clickable analog of 20(S)-OHC in which an alkyne group replaces the iso-butyl group at carbon 22. In 20(R)-yne, the chirality at carbon 20 is changed from the S to the R configuration. B, to measure the transcriptional response to oxysterols, NIH/3T3 cells were treated with vehicle control (DMSO), 20(S)-OHC, or 20(S)-yne at 5 μm for 16 h. The levels of the target genes Gli1, ABCA1, SREBP, HMG-CoA reductase, HMG-CoA synthase, and LDL-R were measured using quantitative RT-PCR. C, carboxyfluorescein-loaded vesicles were incubated with the indicated concentrations of 20(S)-OHC or 20(S)-yne, and vesicle expansion was followed by the increase in fluorescence caused by dequenching (left panel). Time constants for the de-quenching reaction, which indicate the rate at which the oxysterols accumulated in the vesicles, are shown on the right panel. The temporal course of oxysterol uptake (D) and cholesterol levels (E) in NIH/3T3 cells were determined by mass spectrometry.
We reasoned that the unobtrusive alkyne group in 20(S)-yne (Fig. 1A) would make it a good imaging probe because the structure and polarity are only moderately altered relative to the parent compound. We used signaling assays, cellular uptake assays, and in vitro membrane expansion assays to compare 20(S)-yne to its parent 20(S)-OHC. Cells were treated with these sterols, and quantitative RT-PCR was used to measure the levels of target genes for three distinct oxysterol-responsive signaling pathways as follows: the Hh pathway, the SREBP pathway, and the LXR pathway (Fig. 1B). Both 20(S)-OHC and 20(S)-yne had very similar effects on Hh and SREBP target genes; however, only 20(S)-OHC could activate an LXR target gene, suggesting that 20(S)-yne cannot bind or activate LXR proteins.
Because oxysterols are thought to mediate a subset of their effects through alterations in the biophysical properties of membranes, we used a fluorescence de-quenching assay to compare the membrane-expanding effects of 20(S)-OHC and 20(S)-yne on unilamellar vesicles composed of a synthetic dioleoylphosphatidic acid bilayer and loaded with carboxyfluorescein (Fig. 1C). Both oxysterols caused significant membrane expansion of the bilayer, but the final extent of expansion was larger for 20(S)-yne, perhaps because it was more disruptive to the packing of phospholipids. However, both oxysterols accumulated in the vesicles at similar rates across a range of concentrations, as measured by time constants for the dequenching reaction (Fig. 1C). When added to live cells, both 20(S)-OHC and 20(S)-yne were taken up with similar kinetics and to a similar final extent were measured using mass spectrometry (MS) (Fig. 1D). Neither compound had significant effects on total cellular levels of cholesterol over the duration of uptake assays described in this study (Fig. 1E). Taken together, these results suggest that there is significant, but not complete, overlap between the properties of 20(S)-OHC and 20(S)-yne and that both oxysterols may follow similar uptake and transport pathways in cells.
Considering that 20(S)-yne appeared to be a reasonable surrogate for 20(S)-OHC, we used its alkyne moiety to image its distribution in cells. NIH/3T3 cells were metabolically labeled with a pulse of 20(S)-yne, fixed with paraformaldehyde, and subjected to a click reaction on coverslips with an Alexa 594-azide fluorophore (Fig. 2A). Similar methods have been used to image other biomolecules, including sugars (42), nucleic acids (43, 44), proteins (45), phospholipids (46), and most recently cholesterol itself (47). We established the specificity of the staining protocol in a variety of ways. Incubation of cells with increasing concentrations of 20(S)-yne prior to fixation and click labeling with a constant concentration of Alexa 594-azide led to an increase in the intensity of the Alexa 594 fluorescence seen within cells (Fig. 2, B and C). These concentrations of 20(S)-yne can fully activate Hh signaling without compromising cell viability even at long labeling times (18). Alexa 594-azide staining of cells depended on the cycloaddition reaction; Alexa 594 fluorescence was not seen if the cells were not incubated with 20(S)-yne or were incubated with 20(S)-OHC instead (Fig. 2D) or if the copper and ascorbic acid catalysts were omitted (Fig. 2E). These controls exclude the possibility that Alexa 594-azide passively accumulates in cells during the click staining procedure. Importantly, the uptake of 20(S)-yne occurred through a saturable pathway, because the staining intensity could be reduced by including an excess of “cold” competitor, the nonclickable parent 20(S)-OHC, during labeling (Fig. 2F).
Given the purported rapid rate of oxysterol transfer between membranes, we were surprised to find that 20(S)-yne accumulated in a distinct, asymmetric perinuclear structure at a range of concentrations (Fig. 2, B, D, and G, arrowheads). The distribution of 20(S)-yne was quite different from the uniform labeling of all cellular membranes reported with clickable choline phospholipids (46). Bright staining was also apparent in punctate cytoplasmic structures, whereas other cellular endomembranes demonstrated lower levels of staining (Fig. 2, B and D). 20(S)-yne did not accumulate in the nucleus or at the plasma membrane.
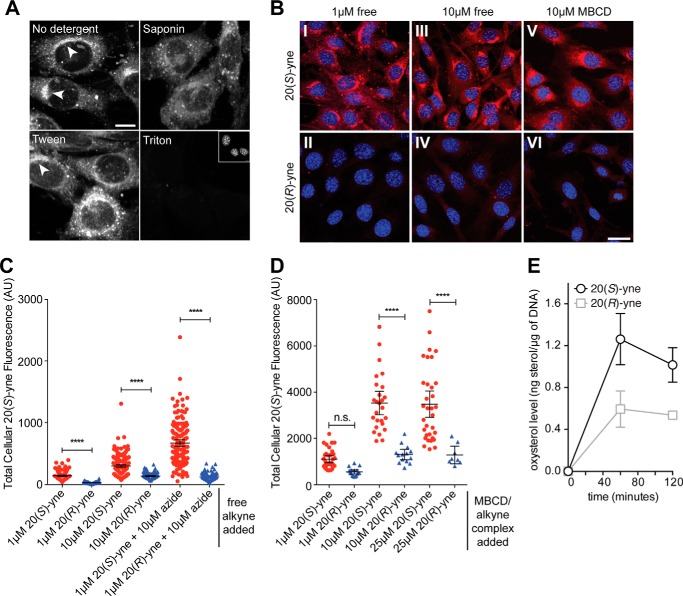
20(S)-yne was likely incorporated into cellular membranes because treatment of cells after the click reaction with 0.1% v/v Triton X-100, a detergent commonly used to solubilize membranes, completely eliminated staining (Fig. 3A). In contrast, treatment with 0.1% v/v Tween 20 or saponin, two milder detergents, allowed partial retention of the 20(S)-yne staining intensity but altered the morphology of the stained structures. To permeabilize cells for immunofluorescence after fixation, we found that only low concentrations of digitonin (0.003%) or saponin (0.03%) maintained both the intensity and distribution of 20(S)-yne staining. This detergent sensitivity profile suggests that 20(S)-yne is associated with membranes rather than being in a soluble pool.
FIGURE 3.
Detergent sensitivity and stereoselectivity of 20(S)-yne accumulation in cells. A, residual Alexa 594 staining in cells that were permeabilized with 0.1% v/v concentrations of the indicated detergents after fixation and click labeling. Scale bar, 10 μm. B, cellular uptake (red) of 20(S)-yne or its epimer 20(R)-yne added to media as the free sterol (panels I–IV) or added as a conjugate with MBCD (panels V and VI). Images within each column were taken with the same settings and are directly comparable. Scale bar, 25 μm. Quantitative analysis of total cellular fluorescence from multiple images from the experiments shown in B is shown in C and D for the free alkyne and MBCD conjugates, respectively. Bars indicate mean cellular fluorescence (±95% confidence interval), and each point denotes a single cell. Comparisons were made using 1-way ANOVA plus Bonferroni multiple comparison test (****, p < 0.0001, n.s., not significant). E, bulk uptake of 20(S)-yne was compared with that of 20(R)-yne in NIH/3T3 cells by mass spectrometry. AU, arbitrary units.
To determine whether the stereochemistry of the hydroxyl group at carbon 20 was important for cellular localization, we used 20(R)-yne (Fig. 1A) (48). When cells were labeled with 20(R)-yne, either alone or as MBCD conjugate to control for solubility differences, the Alexa 594-azide staining intensity was markedly reduced compared with the 20(S)-yne (Fig. 3, B, C, and D). Bulk uptake of 20(S)-yne, which includes both specific uptake and passive partitioning into membranes, was 2-fold higher than that of 20(R)-yne when measured by mass spectrometry (Fig. 3E). The significantly larger difference in Golgi staining seen in the microscopy-based assay may reflect that fact that 20(S)-yne is retained at the Golgi by protein interactions, making its localization more durable during the fixation and wash steps performed prior to the cycloaddition reaction. Thus, the uptake of 20(S)-yne is unlikely to be the result of nonspecific accumulation in membranes; instead, the stereoselectivity suggests a pathway that involves oxysterol-protein interactions that are sensitive to the chirality of the 20-hydroxyl group.
20(S)-yne Accumulates in the Golgi Apparatus
We identified the prominent perinuclear structure stained by 20(S)-yne as the Golgi because the staining co-localized with two markers of the Golgi, giantin and IGnT (Fig. 4, A–F′). Giantin and IGnT are integral Golgi membrane proteins with structural and enzymatic roles, respectively. Thus, oxysterols can accumulate in specific organelle membranes with striking selectivity, much like proteins and other lipids such as phosphoinositides.
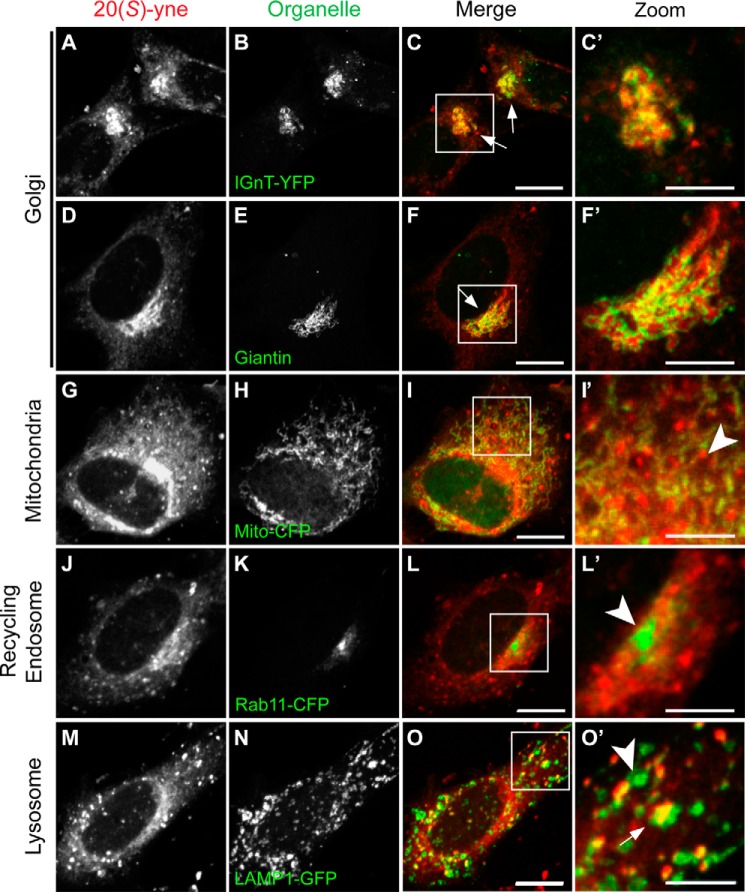
FIGURE 4.
20(S)-yne accumulates in the Golgi. The localization of 20(S)-yne (red) was visualized in cells transiently expressing FP-tagged markers (green) of the Golgi (IGnT-YFP, A–C′), mitochondria (Mito-CFP, G–I′), recycling endosomes (Rab11-CFP, J–L′), and lysosomes (Lamp1-GFP, M–O′). Golgi localization was confirmed by looking at the localization of 20(S)-yne relative to the immunostaining pattern obtained with an antibody against the cis- and medial-Golgi marker giantin, D–F′). Arrows and arrowheads denote, respectively, regions where the organelle marker does or does not overlap with the 20(S)-yne staining. Zoomed-in areas shown in the 4th column are indicated by solid white boxes in the 3rd column. Scale bar, 10 μm in the 3rd and 5 μm in the 4th column.
We also tested the co-localization of 20(S)-yne with markers of mitochondria, recycling endosomes, and lysosomes. Both the perinuclear and punctate staining patterns failed to significantly co-localize with either Mito-CFP or Rab11-CFP (Fig. 4, G–L′). 20(S)-yne staining encircled the pericentriolar recycling endosome compartment marked by RAB11, consistent with Golgi localization (49).
Although we could not identify all membrane compartments represented by the punctate 20(S)-yne structures, co-localization studies with the lysosome marker LAMP1 showed partial overlap between the 20(S)-yne puncta and lysosomes (Fig. 4, M–O′). Many of the 20(S)-yne puncta were either adjacent to or overlapping with LAMP1-positive structures; however, not all LAMP1-positive structures contained 20(S)-yne.
Although 20(S)-yne can induce SMO accumulation into the primary cilium (18), it did not show accumulation in the membrane of the primary cilium (data not shown). However, the high cell confluence required to promote ciliation in NIH/3T3 cells reduced overall uptake of 20(S)-yne, possibly reducing cilia staining below the limits of detection.
On a technical note, the use of organelle markers for 20(S)-yne co-localization studies was challenging for two reasons. First, many of the commonly used antibodies against organelle proteins did not detect their antigens in the mild saponin permeabilization conditions required to maintain the Alexa 594-azide staining (Fig. 3A and associated discussion). Second, the fluorescence intensity of FP-tagged organelle markers was substantially diminished after the copper-catalyzed click reaction (50).
Kinetics of 20(S)-yne Uptake and Release
To follow the route taken by 20(S)-yne from the culture medium into the cell and to the Golgi, we varied either the time of labeling (Fig. 5A) or the time of the label-free chase period (Fig. 5, B and C). Increasing the time of labeling prior to fixation led to progressively stronger Alexa 594-azide staining (Fig. 5A, top row). Interestingly, the subcellular distribution of labeling changed over time, short incubation times produced labeling in a largely punctate pattern (Fig. 5A, bottom row, arrowheads), which subsequently evolved over time into perinuclear Golgi staining (Fig. 5A, bottom row, arrows). At higher doses of 20(S)-yne, we observed an increase in low level membranous staining distributed throughout the cytoplasm, possibly reflecting the nonspecific partitioning of amphipathic 20(S)-yne into various organelle membranes (Figs. 4G and 5A, panel III). These results suggest that 20(S)-yne passed through the punctate structures en route to the Golgi.
To examine 20(S)-yne release from cells, we performed chase experiments for time periods ranging from 5 min to 24 h after a 1-h labeling period (Fig. 5, B and C). Overall staining intensity decreased significantly even after a 1-h chase; however, high magnification views demonstrated that the signal in the Golgi was significantly more persistent compared with the diffuse membrane signal (Fig. 5B, panel V). In fact, even after 24 h, the Alexa 594-azide staining was clearly detectable in the Golgi (Fig. 5C, panel V). We conclude that the 20(S)-yne pool that partitions into all cellular membranes is rapidly lost by diffusion, but a distinct pool of this oxysterol is retained against diffusive loss in the Golgi, perhaps by interaction with a protein or by an active transport pathway. We note that this finding critically depended on our ability to visualize the localization of 20(S)-yne within cells with high resolution microscopy; techniques using radiolabeled sterols that measure total cellular uptake would not have detected these differences.
20(S)-yne Uptake Is Sensitive to ATP Levels and Temperature
Current models propose that oxysterols can rapidly transfer between membranes through a nonvesicular transport pathway, perhaps chaperoned by the ORP proteins (31). In contrast, cholesterol is internalized through a vesicle-dependent endocytic pathway (51). To distinguish between a passive transport process and an active, energy-dependent one such as vesicle trafficking or motor-driven movement, we explored the dependence of the uptake process on ATP and its sensitivity to temperature.
A brief depletion of cellular ATP levels through sodium azide and deoxyglucose treatment significantly inhibited 20(S)-yne uptake, favoring an energy-dependent transport process (Fig. 6, A and B). The total cellular uptake of 20(S)-yne and its accumulation in the Golgi were strongly dependent on temperature, with both processes reduced at 22 °C and nearly blocked at 15 °C (Fig. 6, C and D). Different steps in the endocytic pathway show different sensitivity to temperature; the rate of internalization from the plasma membrane drops continuously with temperatures between 37 and 10 °C, but transport from endosomes to lysosomes stops abruptly between 18 and 20 °C (52, 53). These results suggest that an active transport pathway, possibly vesicle trafficking through late endosomes and lysosomes, controls 20(S)-yne uptake and distribution. We find the data to be inconsistent with passive or facilitated accumulation of 20(S)-yne in cellular membranes.
20(S)-yne Uptake Is Dependent on Lysosome Function
Given the overlap between 20(S)-yne puncta and lysosomes, we tested if lysosome function was important for 20(S)-yne trafficking. Treatment of cells with either NH4Cl or chloroquine, two structurally distinct lysosomotropic weak bases, inhibited the cellular uptake and Golgi targeting of 20(S)-yne (Fig. 7, A and B). These weak bases accumulate in their charged, protonated forms in the acidic milieu of lysosomes, inhibiting lysosome proteolysis by causing a rise in the pH of the lumen and leading to the structural distortion and vacuolation of lysosomes (54). Importantly, these agents often do not impair the initial cellular uptake of many protein cargoes that use the classical endocytic pathway (55).
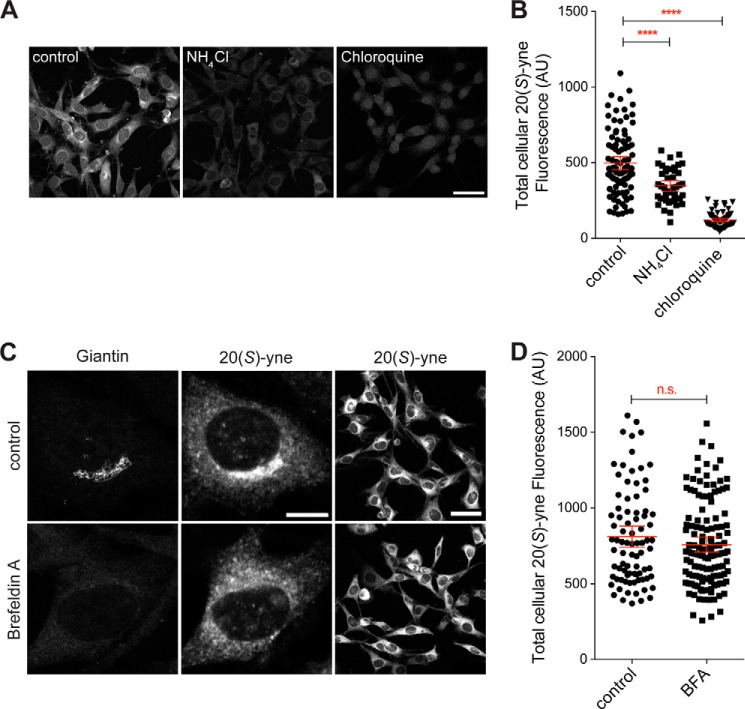
FIGURE 7.
Lysosomotropic weak bases inhibit the cellular uptake of 20(S)-yne. A and B, 20(S)-yne uptake in cells pretreated with either NH4Cl (10 mm) or chloroquine (100 μm). C and D, giantin staining (left column) and 20(S)-yne staining (middle and right columns) in cells treated with brefeldin A or left untreated. B and D, the total cellular 20(S)-yne fluorescence of multiple individual cells is plotted, and the bars denote the mean fluorescence (±95% confidence interval). B, differences between the conditions are evaluated 1-way ANOVA plus Bonferroni multiple comparison test (****, p < 0.0001, n >40 per condition). D, lack of a statistically significant difference was observed by an unpaired t test (p = 0.21, n >80 per condition, n.s., not significant). Scale bars in the high magnification images are 10 μm and in the low magnification images are 50 μm. AU, arbitrary units.
Because the ultimate destination of 20(S)-yne is the Golgi, we perturbed Golgi function with brefeldin A and examined the uptake of 20(S)-yne. Brefeldin A inhibits the activation of several ARF family GTPases, leading to a block in anterograde ER to Golgi transport and the tubulation of many organelles. Because retrograde Golgi→ER transport continues, Golgi stacks completely disassemble in brefeldin A-treated cells, and many Golgi proteins are resorbed into the ER (56). This can be seen by the loss of giantin, a peripheral membrane protein that connects Golgi stacks, in brefeldin A-treated cells (Fig. 7C, left column). Brefeldin A induced the dispersal of the perinuclear 20(S)-yne staining, confirming that this structure is indeed the Golgi (Fig. 7C, middle column). Yet, disassembly of the Golgi did not diminish the total cellular uptake of 20(S)-yne (Fig. 7D), which still accumulated in the tubulated intracellular membranes.
In summary, 20(S)-yne is taken up into cells and retained in the Golgi by a pathway that depends on the proper functioning of lysosomes but that does not depend on trafficking steps regulated by brefeldin A-sensitive ARF family GTPases.
20(S)-yne Accumulates in the Lysosomes of Cells Lacking Niemann-Pick C1 Protein
Niemann-Pick C1 (NPC1) protein is a 13-pass transmembrane protein that binds to both oxysterols and cholesterol and has been found to be defective in >90% of children with Niemann-Pick Type C disease (57). NPC1 is required for the efficient export of lipoprotein-derived cholesterol from the late endosome and lysosome (LE/L) compartments (58–60). In addition to accumulating cholesterol to high levels in LE/L, cells lacking NPC1 function have a variety of defects in the trafficking of other lipids, such as glycosphingolipids, and proteins through the endocytic pathway (61, 62).
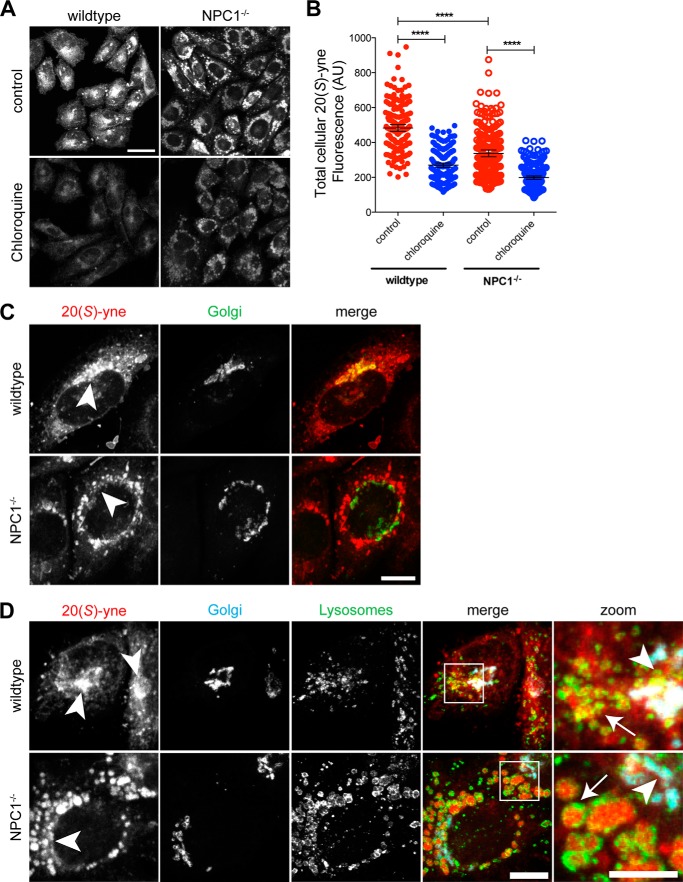
We compared 20(S)-yne accumulation in an NPC1 null cell line (NPC1−/−), CHO-M12, and a wild-type line, CHO-K1 (63). NPC1−/− cells accumulated ∼20% less total 20(S)-yne compared with wild-type cells (Fig. 8, A and B). However, there was a dramatic difference in the intracellular distribution of this internalized oxysterol between the two cell types. In wild-type CHO cells, the highest level of 20(S)-yne was observed in the Golgi, consistent with its localization in NIH/3T3 cells (Fig. 8C, top row). However, in NPC1−/− cells, 20(S)-yne was found at the highest levels in large vesicular structures that did not overlap with Golgi markers (Fig. 8C, bottom row). This staining pattern was reminiscent of the dilated LE/L laden with trapped cholesterol and other lipids that are characteristic of NPC1−/− cells (63). To define the localization of 20(S)-yne in these cells, we transiently transfected an FP-tagged Golgi marker and permeabilized the cells with 0.03% saponin, a mild procedure that maintained the distribution of 20(S)-yne staining while allowing immunostaining of LE/L with an antibody against the membrane glycoprotein LGP95. Indeed, in NPC1−/− cells, the majority of the 20(S)-yne was in the lumen of LE/L, surrounded by a rim of LGP95 staining; the signal in the Golgi was significantly lower (Fig. 8D). Thus, the trafficking defects caused by the loss of NPC1 seem to inhibit the targeting of 20(S)-yne to its ultimate Golgi destination, leading instead to its accumulation in LE/L.
FIGURE 8.
20(S)-yne accumulates in lysosomes of CHO cells lacking NPC1 function. A and B, accumulation of 20(S)-yne in wild-type and NPC1−/− CHO cells, either untreated or treated with chloroquine (100 μm). Total cellular fluorescence is plotted in B for multiple individual cells, along with bars that show the mean fluorescence (±95% confidence interval). 1-way ANOVA plus Bonferroni multiple comparison test shows significant differences between the conditions (****, p < 0.0001, n>180 per condition). Scale bar, 25 μm. C, 20(S)-yne localization (red) relative to the Golgi (Golgi-CFP, green, arrowheads) in wild-type or NPC1−/− CHO cells. Scale bar, 10 μm. D, 20(S)-yne localization (red) in wild-type or NPC1−/− CHO cells relative to the Golgi (Golgi-CFP marker, cyan) and the lysosomes (anti-Lgp95, green). Areas of overlap between the Golgi and the 20(S)-yne staining appear white in the merge panels. Areas shown in rightmost “zoomed” column are indicated by white box in merge column. Scale bar, 10 μm for the merge column and 5 μm for the zoom column. AU, arbitrary units.
The block in 20(S)-yne uptake seen in NPC1−/− cells appears to be at a more downstream step in the pathway compared with the blocks seen with either chloroquine treatment or low temperature, both of which seemed to prevent overall uptake rather than causing LE/L accumulation. This is consistent with the observation that chloroquine also significantly diminished overall cellular uptake in NPC1−/− cells (Fig. 8B). Taken together, these results suggest that the late LE/L vesicle transport pathway plays an important role in the cellular uptake and distribution of 20(S)-yne.
DISCUSSION
We have used bioorthogonal click chemistry to provide high resolution images of the subcellular distribution of the 20(S)-yne, an analog of the endogenous oxysterol, 20(S)-OHC, which can activate Hh signaling, regulate ciliary protein trafficking, and drive osteogenesis in vivo (20–22, 64). The unexpected result that emerges from these images is the preferential accumulation and subsequent retention of 20(S)-yne in the Golgi. These findings have significant implications for the regulatory functions of oxysterols in cells, suggesting roles in organelle identity and organelle-specific regulation of membrane properties.
Cholesterol itself is found in a gradient in cellular membranes, with highest levels in the plasma membrane, followed in order of decreasing concentration by the endosome recycling compartment, the Golgi, and the ER (65). However, it was previously unknown whether organelle membranes could vary in their composition of oxysterols, which incur a smaller energetic penalty for membrane desorption and thus show much higher rates of inter-membrane transfer. Given that diverse proteins can bind oxysterols, differences in membrane oxysterol composition could potentially be exploited to recruit or retain proteins to specific organelles or membrane domains.
Interestingly, side chain oxysterols such as 20(S)-OHC, which carry the second hydroxyl on the iso-octyl chain (Fig. 1A), can dramatically alter the biophysical properties of membranes, increasing lipid disorder as well as membrane expansion and membrane fluidity (35). These effects oppose the membrane condensing and ordering properties of cholesterol, especially in environments with a high proportion of sphingolipids or phospholipids with saturated acyl chains. Thus, differences in oxysterol composition of organelle membranes can allow differential tuning of membrane biophysical properties and consequently influence the activities of embedded protein and lipids. For instance, oxysterols are thought to increase the amount of accessible or “active” cholesterol by weakening cholesterol-phospholipid interactions (34, 35). Although the total cellular concentrations of oxysterols are much lower than cholesterol, our results show that the local concentrations of oxysterols can be significantly higher in specific membranes.
How is 20(S)-yne concentrated and retained in the Golgi? In the simplest model, oxysterols would passively transfer between cellular membranes along a concentration gradient and then be retained in the Golgi due to interactions with a protein or a lipid. However, a model based only on lipid interactions is inconsistent with our findings that the structurally similar stereoisomer, 20(R)-yne, exhibits significantly less intracellular localization; this stereoselectivity suggests that a protein is involved in either the trafficking or retention steps. Also, a purely passive diffusion-trap model is inconsistent with the energy requirement we observed for this process. Instead, sensitivity to genetic or pharmacological perturbation of the LE/L system suggests an important role for vesicular pathways.
A recent study used an approach similar to ours to visualize the subcellular distribution of an alkyne cholesterol analog (47). This molecule showed significant accumulation in the Golgi, similar to 20(S)-yne, but in addition showed much higher levels of labeling in the ER, the mitochondria, and the plasma membrane. The Golgi staining pattern seen in 20(S)-yne and alkyne cholesterol-stained cells suggests that common mechanisms may mediate the uptake of both 20(S)-yne and cholesterol. Interestingly, a vesicular pathway has been described for transport of LDL-derived cholesterol from the compartment containing NPC1 to the trans-Golgi network (66). However, these pathways must diverge at some point because the highest levels of cholesterol are found in the plasma membrane and the endocytic recycling compartment (67), both regions where we find very little 20(S)-yne staining. The pattern of 20(S)-yne uptake and concentration in the Golgi also resembles that of glycosphingolipids, which are endocytosed from the plasma membrane and pass through a LE/L compartment en route to the Golgi (68). The methods described here will be useful in elucidating the molecular details of the trafficking pathways driving the accumulation of 20(S)-yne in Golgi.
Moving forward, it will be interesting to see if click-based approaches can be used to image the subcellular distribution of other classes of oxysterols in cells or tissues (46), especially because oxysterols have been suggested to form chemotactic gradients (19). This will depend on whether other oxysterols can be modified with chemical groups (such as alkyne or azide moieties) compatible with bio-orthogonal chemistry without destroying their biological activities. Another future challenge will be to adapt this methodology to image the dynamics of oxysterols in live cells. This has been possible for cholesterol using the naturally fluorescent sterol dehydroergosterol or cholesterol analogs carrying fluorophores (69, 70). An azide-modified oxysterol could be detected in live cells using a strained cyclooctyne fluorophore without the use of the cytotoxic copper catalyst (71). Alkyne moieties can also be detected in live cells directly by Coherent Anti-Stokes Raman Scattering microscopy (72). In summary, we expect that the click chemistry toolkit will provide a new lens through which to explore sterol cell biology.
Acknowledgments
We thank Maika Deffieu and Suzanne Pfeffer for CHO-K1 cells, LAMP1-GFP and IGNT-YFP constructs, and valuable advice; Takanari Inoue for various CFP organelle markers, and Andres Lebensohn and Ganesh Pusapati for comments on the manuscript.
This work was supported, in whole or in part, by National Institutes of Health Grants GM47969 and HL67773 (to D. F. C.), HL67773 (to D. S. O.), and Training Grant 5 T32 HL007275 (to L. K. M.). This work was also supported by a Stand Up to Cancer Innovation Award from the American Association for Cancer Research (to R. R.) and by a National Science Foundation Graduate Research Fellowship (to S. N.).
- 20(S)-OHC
- 20(S)-hydroxycholesterol
- Hh
- Hedgehog
- Smo
- Smoothened
- DOPC
- dioleoylphosphatidic acid
- ER
- endoplasmic reticulum
- NPC1
- Niemann-Pick C1
- LE/L
- late endosome and lysosome
- for
- forward
- rev
- reverse
- ANOVA
- analysis of variance
- CFP
- cyan fluorescent protein
- FP
- fluorescent protein
- MBCD
- methyl-β-cyclodextrin
- SREBP
- sterol regulatory element-binding protein
- LXR
- liver X receptor.
REFERENCES
- 1. Chang P. V., Bertozzi C. R. (2012) Imaging beyond the proteome. Chem. Commun. 48, 8864–8879 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Zaccolo M., De Giorgi F., Cho C. Y., Feng L., Knapp T., Negulescu P. A., Taylor S. S., Tsien R. Y., Pozzan T. (2000) A genetically encoded, fluorescent indicator for cyclic AMP in living cells. Nat. Cell Biol. 2, 25–29 [DOI] [PubMed] [Google Scholar]
- 3. Grynkiewicz G., Poenie M., Tsien R. Y. (1985) A new generation of Ca2+ indicators with greatly improved fluorescence properties. J. Biol. Chem. 260, 3440–3450 [PubMed] [Google Scholar]
- 4. Várnai P., Balla T. (2006) Live cell imaging of phosphoinositide dynamics with fluorescent protein domains. Biochim. Biophys. Acta 1761, 957–967 [DOI] [PubMed] [Google Scholar]
- 5. Russell D. W. (2000) Oxysterol biosynthetic enzymes. Biochim. Biophys. Acta 1529, 126–135 [DOI] [PubMed] [Google Scholar]
- 6. Lütjohann D., Breuer O., Ahlborg G., Nennesmo I., Sidén A., Diczfalusy U., Björkhem I. (1996) Cholesterol homeostasis in human brain: Evidence for an age-dependent flux of 24S-hydroxycholesterol from the brain into the circulation. Proc. Natl. Acad. Sci. U.S.A. 93, 9799–9804 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Roberts K. D., Bandy L., Lieberman S. (1969) The occurrence and metabolism of 20α-hydroxycholesterol in bovine adrenal preparations. Biochemistry 8, 1259–1270 [DOI] [PubMed] [Google Scholar]
- 8. Lange Y., Ye J., Strebel F. (1995) Movement of 25-hydroxycholesterol from the plasma membrane to the rough endoplasmic reticulum in cultured hepatoma cells. J. Lipid Res. 36, 1092–1097 [PubMed] [Google Scholar]
- 9. Meaney S., Bodin K., Diczfalusy U., Björkhem I. (2002) On the rate of translocation in vitro and kinetics in vivo of the major oxysterols in human circulation: Critical importance of the position of the oxygen function. J. Lipid Res. 43, 2130–2135 [DOI] [PubMed] [Google Scholar]
- 10. Theunissen J. J., Jackson R. L., Kempen H. J., Demel R. A. (1986) Membrane properties of oxysterols. Interfacial orientation, influence on membrane permeability and redistribution between membranes. Biochim. Biophys. Acta 860, 66–74 [DOI] [PubMed] [Google Scholar]
- 11. Björkhem I. (2002) Do oxysterols control cholesterol homeostasis? J. Clin. Invest. 110, 725–730 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Radhakrishnan A., Ikeda Y., Kwon H. J., Brown M. S., Goldstein J. L. (2007) Sterol-regulated transport of SREBPs from endoplasmic reticulum to Golgi: Oxysterols block transport by binding to Insig. Proc. Natl. Acad. Sci. U.S.A. 104, 6511–6518 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Chen W., Chen G., Head D. L., Mangelsdorf D. J., Russell D. W. (2007) Enzymatic reduction of oxysterols impairs LXR signaling in cultured cells and the livers of mice. Cell Metab. 5, 73–79 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Janowski B. A., Willy P. J., Devi T. R., Falck J. R., Mangelsdorf D. J. (1996) An oxysterol signalling pathway mediated by the nuclear receptor LXRα. Nature 383, 728–731 [DOI] [PubMed] [Google Scholar]
- 15. Sacchetti P., Sousa K. M., Hall A. C., Liste I., Steffensen K. R., Theofilopoulos S., Parish C. L., Hazenberg C., Richter L. A., Hovatta O., Gustafsson J. A., Arenas E. (2009) Liver X receptors and oxysterols promote ventral midbrain neurogenesis in vivo and in human embryonic stem cells. Cell Stem Cell 5, 409–419 [DOI] [PubMed] [Google Scholar]
- 16. Hannedouche S., Zhang J., Yi T., Shen W., Nguyen D., Pereira J. P., Guerini D., Baumgarten B. U., Roggo S., Wen B., Knochenmuss R., Noël S., Gessier F., Kelly L. M., Vanek M., Laurent S., Preuss I., Miault C., Christen I., Karuna R., Li W., Koo D. I., Suply T., Schmedt C., Peters E. C., Falchetto R., Katopodis A., Spanka C., Roy M. O., Detheux M., Chen Y. A., Schultz P. G., Cho C. Y., Seuwen K., Cyster J. G., Sailer A. W. (2011) Oxysterols direct immune cell migration via EBI2. Nature 475, 524–527 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Liu C., Yang X. V., Wu J., Kuei C., Mani N. S., Zhang L., Yu J., Sutton S. W., Qin N., Banie H., Karlsson L., Sun S., Lovenberg T. W. (2011) Oxysterols direct B-cell migration through EBI2. Nature 475, 519–523 [DOI] [PubMed] [Google Scholar]
- 18. Nachtergaele S., Mydock L. K., Krishnan K., Rammohan J., Schlesinger P. H., Covey D. F., Rohatgi R. (2012) Oxysterols are allosteric activators of the oncoprotein Smoothened. Nat. Chem. Biol. 8, 211–220 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Yi T., Wang X., Kelly L. M., An J., Xu Y., Sailer A. W., Gustafsson J. A., Russell D. W., Cyster J. G. (2012) Oxysterol gradient generation by lymphoid stromal cells guides activated B cell movement during humoral responses. Immunity 37, 535–548 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Corcoran R. B., Scott M. P. (2006) Oxysterols stimulate Sonic hedgehog signal transduction and proliferation of medulloblastoma cells. Proc. Natl. Acad. Sci. U.S.A. 103, 8408–8413 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Dwyer J. R., Sever N., Carlson M., Nelson S. F., Beachy P. A., Parhami F. (2007) Oxysterols are novel activators of the hedgehog signaling pathway in pluripotent mesenchymal cells. J. Biol. Chem. 282, 8959–8968 [DOI] [PubMed] [Google Scholar]
- 22. Rohatgi R., Milenkovic L., Scott M. P. (2007) Patched1 regulates hedgehog signaling at the primary cilium. Science 317, 372–376 [DOI] [PubMed] [Google Scholar]
- 23. LeBlanc M. A., McMaster C. R. (2010) Lipid binding requirements for oxysterol-binding protein Kes1 inhibition of autophagy and endosome-trans-Golgi trafficking pathways. J. Biol. Chem. 285, 33875–33884 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Raychaudhuri S., Prinz W. A. (2010) The diverse functions of oxysterol-binding proteins. Annu. Rev. Cell Dev. Biol. 26, 157–177 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Burgett A. W., Poulsen T. B., Wangkanont K., Anderson D. R., Kikuchi C., Shimada K., Okubo S., Fortner K. C., Mimaki Y., Kuroda M., Murphy J. P., Schwalb D. J., Petrella E. C., Cornella-Taracido I., Schirle M., Tallarico J. A., Shair M. D. (2011) Natural products reveal cancer cell dependence on oxysterol-binding proteins. Nat. Chem. Biol. 7, 639–647 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. de Saint-Jean M., Delfosse V., Douguet D., Chicanne G., Payrastre B., Bourguet W., Antonny B., Drin G. (2011) Osh4p exchanges sterols for phosphatidylinositol 4-phosphate between lipid bilayers. J. Cell Biol. 195, 965–978 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Du X., Kumar J., Ferguson C., Schulz T. A., Ong Y. S., Hong W., Prinz W. A., Parton R. G., Brown A. J., Yang H. (2011) A role for oxysterol-binding protein-related protein 5 in endosomal cholesterol trafficking. J. Cell Biol. 192, 121–135 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Blanc M., Hsieh W. Y., Robertson K. A., Kropp K. A., Forster T., Shui G., Lacaze P., Watterson S., Griffiths S. J., Spann N. J., Meljon A., Talbot S., Krishnan K., Covey D. F., Wenk M. R., Craigon M., Ruzsics Z., Haas J., Angulo A., Griffiths W. J., Glass C. K., Wang Y., Ghazal P. (2013) The transcription factor STAT-1 couples macrophage synthesis of 25-hydroxycholesterol to the interferon antiviral response. Immunity 38, 106–118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Liu S. Y., Aliyari R., Chikere K., Li G., Marsden M. D., Smith J. K., Pernet O., Guo H., Nusbaum R., Zack J. A., Freiberg A. N., Su L., Lee B., Cheng G. (2013) Interferon-inducible cholesterol-25-hydroxylase broadly inhibits viral entry by production of 25-hydroxycholesterol. Immunity 38, 92–105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Massey J. B., Pownall H. J. (2006) Structures of biologically active oxysterols determine their differential effects on phospholipid membranes. Biochemistry 45, 10747–10758 [DOI] [PubMed] [Google Scholar]
- 31. Olkkonen V. M., Hynynen R. (2009) Interactions of oxysterols with membranes and proteins. Mol. Aspects Med. 30, 123–133 [DOI] [PubMed] [Google Scholar]
- 32. Olsen B. N., Schlesinger P. H., Baker N. A. (2009) Perturbations of membrane structure by cholesterol and cholesterol derivatives are determined by sterol orientation. J. Am. Chem. Soc. 131, 4854–4865 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. McIntosh T. J., Simon S. A. (2006) Roles of bilayer material properties in function and distribution of membrane proteins. Annu. Rev. Biophys. Biomol. Struct. 35, 177–198 [DOI] [PubMed] [Google Scholar]
- 34. Steck T. L., Lange Y. (2010) Cell cholesterol homeostasis: Mediation by active cholesterol. Trends Cell Biol. 20, 680–687 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Bielska A. A., Schlesinger P., Covey D. F., Ory D. S. (2012) Oxysterols as non-genomic regulators of cholesterol homeostasis. Trends Endocrinol. Metab. 23, 99–106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Gale S. E., Westover E. J., Dudley N., Krishnan K., Merlin S., Scherrer D. E., Han X., Zhai X., Brockman H. L., Brown R. E., Covey D. F., Schaffer J. E., Schlesinger P., Ory D. S. (2009) Side chain oxygenated cholesterol regulates cellular cholesterol homeostasis through direct sterol-membrane interactions. J. Biol. Chem. 284, 1755–1764 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Mesmin B., Maxfield F. R. (2009) Intracellular sterol dynamics. Biochim. Biophys. Acta 1791, 636–645 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Christian A. E., Haynes M. P., Phillips M. C., Rothblat G. H. (1997) Use of cyclodextrins for manipulating cellular cholesterol content. J. Lipid Res. 38, 2264–2272 [PubMed] [Google Scholar]
- 39. Carpenter A. E., Jones T. R., Lamprecht M. R., Clarke C., Kang I. H., Friman O., Guertin D. A., Chang J. H., Lindquist R. A., Moffat J., Golland P., Sabatini D. M. (2006) CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Jiang X., Sidhu R., Porter F. D., Yanjanin N. M., Speak A. O., te Vruchte D. T., Platt F. M., Fujiwara H., Scherrer D. E., Zhang J., Dietzen D. J., Schaffer J. E., Ory D. S. (2011) A sensitive and specific LC-MS/MS method for rapid diagnosis of Niemann-Pick C1 disease from human plasma. J. Lipid Res. 52, 1435–1445 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Lin Y. Y., Welch M., Lieberman S. (2003) The detection of 20S-hydroxycholesterol in extracts of rat brains and human placenta by a gas chromatograph/mass spectrometry technique. J. Steroid Biochem. Mol. Biol. 85, 57–61 [DOI] [PubMed] [Google Scholar]
- 42. Laughlin S. T., Bertozzi C. R. (2009) Imaging the glycome. Proc. Natl. Acad. Sci. U.S.A. 106, 12–17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Jao C. Y., Salic A. (2008) Exploring RNA transcription and turnover in vivo by using click chemistry. Proc. Natl. Acad. Sci. U.S.A. 105, 15779–15784 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Salic A., Mitchison T. J. (2008) A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc. Natl. Acad. Sci. U.S.A. 105, 2415–2420 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Liu J., Xu Y., Stoleru D., Salic A. (2012) Imaging protein synthesis in cells and tissues with an alkyne analog of puromycin. Proc. Natl. Acad. Sci. U.S.A. 109, 413–418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Jao C. Y., Roth M., Welti R., Salic A. (2009) Metabolic labeling and direct imaging of choline phospholipids in vivo. Proc. Natl. Acad. Sci. U.S.A. 106, 15332–15337 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Hofmann K., Thiele C., Schött H. F., Gaebler A., Schoene M., Kiver Y., Friedrichs S., Lütjohann D., Kuerschner L. (2014) A novel alkyne cholesterol to trace cellular cholesterol metabolism and localization. J. Lipid Res. 55, 583–591 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Nachtergaele S., Whalen D. M., Mydock L. K., Zhao Z., Malinauskas T., Krishnan K., Ingham P. W., Covey D. F., Siebold C., Rohatgi R. (2013) Structure and function of the Smoothened extracellular domain in vertebrate Hedgehog signaling. Elife 2, e01340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Uchida Y., Hasegawa J., Chinnapen D., Inoue T., Okazaki S., Kato R., Wakatsuki S., Misaki R., Koike M., Uchiyama Y., Iemura S., Natsume T., Kuwahara R., Nakagawa T., Nishikawa K., Mukai K., Miyoshi E., Taniguchi N., Sheff D., Lencer W. I., Taguchi T., Arai H. (2011) Intracellular phosphatidylserine is essential for retrograde membrane traffic through endosomes. Proc. Natl. Acad. Sci. U.S.A. 108, 15846–15851 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Bernardin A., Cazet A., Guyon L., Delannoy P., Vinet F., Bonnaffé D., Texier I. (2010) Copper-free click chemistry for highly luminescent quantum dot conjugates: application to in vivo metabolic imaging. Bioconjug. Chem. 21, 583–588 [DOI] [PubMed] [Google Scholar]
- 51. Schattner A. (1991) Colchicine–Expanding horizons. Postgrad. Med. J. 67, 223–226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Dunn W. A., Hubbard A. L., Aronson N. N., Jr. (1980) Low temperature selectively inhibits fusion between pinocytic vesicles and lysosomes during heterophagy of 125I-asialofetuin by the perfused rat liver. J. Biol. Chem. 255, 5971–5978 [PubMed] [Google Scholar]
- 53. Marsh M., Bolzau E., Helenius A. (1983) Penetration of Semliki forest virus from acidic prelysosomal vacuoles. Cell 32, 931–940 [DOI] [PubMed] [Google Scholar]
- 54. de Duve C., de Barsy T., Poole B., Trouet A., Tulkens P., Van Hoof F. (1974) Commentary. Lysosomotropic agents. Biochem. Pharmacol. 23, 2495–2531 [DOI] [PubMed] [Google Scholar]
- 55. Huotari J., Helenius A. (2011) Endosome maturation. EMBO J. 30, 3481–3500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Klausner R. D., Donaldson J. G., Lippincott-Schwartz J. (1992) Brefeldin A: Insights into the control of membrane traffic and organelle structure. J. Cell Biol. 116, 1071–1080 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Carstea E. D., Morris J. A., Coleman K. G., Loftus S. K., Zhang D., Cummings C., Gu J., Rosenfeld M. A., Pavan W. J., Krizman D. B., Nagle J., Polymeropoulos M. H., Sturley S. L., Ioannou Y. A., Higgins M. E., Comly M., Cooney A., Brown A., Kaneski C. R., Blanchette-Mackie E. J., Dwyer N. K., Neufeld E. B., Chang T. Y., Liscum L., Strauss J. F., 3rd, Ohno K., Zeigler M., Carmi R., Sokol J., Markie D., O'Neill R. R., van Diggelen O. P., Elleder M., Patterson M. C., Brady R. O., Vanier M. T., Pentchev P. G., Tagle D. A. (1997) Niemann-Pick C1 disease gene: homology to mediators of cholesterol homeostasis. Science 277, 228–231 [DOI] [PubMed] [Google Scholar]
- 58. Abi-Mosleh L., Infante R. E., Radhakrishnan A., Goldstein J. L., Brown M. S. (2009) Cyclodextrin overcomes deficient lysosome-to-endoplasmic reticulum transport of cholesterol in Niemann-Pick type C cells. Proc. Natl. Acad. Sci. U.S.A. 106, 19316–19321 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Liscum L., Faust J. R. (1987) Low density lipoprotein (LDL)-mediated suppression of cholesterol synthesis and LDL uptake is defective in Niemann-Pick type C fibroblasts. J. Biol. Chem. 262, 17002–17008 [PubMed] [Google Scholar]
- 60. Pentchev P. G., Comly M. E., Kruth H. S., Patel S., Proestel M., Weintroub H. (1986) The cholesterol storage disorder of the mutant BALB/c mouse. A primary genetic lesion closely linked to defective esterification of exogenously derived cholesterol and its relationship to human type C Niemann-Pick disease. J. Biol. Chem. 261, 2772–2777 [PubMed] [Google Scholar]
- 61. Choudhury A., Sharma D. K., Marks D. L., Pagano R. E. (2004) Elevated endosomal cholesterol levels in Niemann-Pick cells inhibit rab4 and perturb membrane recycling. Mol. Biol. Cell 15, 4500–4511 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Pipalia N. H., Hao M., Mukherjee S., Maxfield F. R. (2007) Sterol, protein and lipid trafficking in Chinese hamster ovary cells with Niemann-Pick type C1 defect. Traffic 8, 130–141 [DOI] [PubMed] [Google Scholar]
- 63. Millard E. E., Srivastava K., Traub L. M., Schaffer J. E., Ory D. S. (2000) Niemann-pick type C1 (NPC1) overexpression alters cellular cholesterol homeostasis. J. Biol. Chem. 275, 38445–38451 [DOI] [PubMed] [Google Scholar]
- 64. Kim W. K., Meliton V., Tetradis S., Weinmaster G., Hahn T. J., Carlson M., Nelson S. F., Parhami F. (2010) Osteogenic oxysterol, 20(S)-hydroxycholesterol, induces notch target gene expression in bone marrow stromal cells. J. Bone Miner. Res. 25, 782–795 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Maxfield F. R., van Meer G. (2010) Cholesterol, the central lipid of mammalian cells. Curr. Opin. Cell Biol. 22, 422–429 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Urano Y., Watanabe H., Murphy S. R., Shibuya Y., Geng Y., Peden A. A., Chang C. C., Chang T. Y. (2008) Transport of LDL-derived cholesterol from the NPC1 compartment to the ER involves the trans-Golgi network and the SNARE protein complex. Proc. Natl. Acad. Sci. U.S.A. 105, 16513–16518 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Hao M., Lin S. X., Karylowski O. J., Wüstner D., McGraw T. E., Maxfield F. R. (2002) Vesicular and non-vesicular sterol transport in living cells. The endocytic recycling compartment is a major sterol storage organelle. J. Biol. Chem. 277, 609–617 [DOI] [PubMed] [Google Scholar]
- 68. Marks D. L., Pagano R. E. (2002) Endocytosis and sorting of glycosphingolipids in sphingolipid storage disease. Trends Cell Biol. 12, 605–613 [DOI] [PubMed] [Google Scholar]
- 69. Hölttä-Vuori M., Uronen R. L., Repakova J., Salonen E., Vattulainen I., Panula P., Li Z., Bittman R., Ikonen E. (2008) BODIPY-cholesterol: a new tool to visualize sterol trafficking in living cells and organisms. Traffic 9, 1839–1849 [DOI] [PubMed] [Google Scholar]
- 70. Mukherjee S., Zha X., Tabas I., Maxfield F. R. (1998) Cholesterol distribution in living cells: Fluorescence imaging using dehydroergosterol as a fluorescent cholesterol analog. Biophys. J. 75, 1915–1925 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Baskin J. M., Prescher J. A., Laughlin S. T., Agard N. J., Chang P. V., Miller I. A., Lo A., Codelli J. A., Bertozzi C. R. (2007) Copper-free click chemistry for dynamic in vivo imaging. Proc. Natl. Acad. Sci. U.S.A. 104, 16793–16797 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Yamakoshi H., Dodo K., Palonpon A., Ando J., Fujita K., Kawata S., Sodeoka M. (2012) Alkyne-tag Raman imaging for visualization of mobile small molecules in live cells. J. Am. Chem. Soc. 134, 20681–20689 [DOI] [PubMed] [Google Scholar]