Abstract
The composition of the typical commercial diet fed to zebrafish can dramatically vary. By utilizing defined diets we sought to answer two questions: 1) How does the embryonic zebrafish transcriptome change when the parental adults are fed a commercial lab diet compared with a sufficient, defined diet (E+)? 2) Does a vitamin Edeficient parental diet (E−) further change the embryonic transcriptome? We conducted a global gene expression study using embryos from zebrafish fed a commercial (Lab), an E+ or an E− diet. To capture differentially expressed transcripts prior to onset of overt malformations observed in E− embryos at 48 h post-fertilization (hpf), embryos were collected from each group at 36 hpf. Lab embryos differentially expressed (p < 0.01) 946 transcripts compared with the E+ embryos, and 2656 transcripts compared with the E− embryos. The differences in protein, fat and micronutrient intakes in zebrafish fed the Lab compared with the E + diet demonstrate that despite overt morphologic consistency, significant differences in gene expression occurred. Moreover, functional analysis of the significant transcripts in the E−embryos suggested perturbed energy metabolism, leading to overt malformations and mortality. Thus, these findings demonstrate that parental zebrafish diet has a direct impact on the embryonic transcriptome.
Keywords: Zebrafish, Defined diet, Vitamin E, Microarray, Transcriptome
1. Introduction
The zebrafish model is growing in popularity and is being used as a model to study human-related conditions, including: neurological disorders (Patton and Zon, 2001; Dambly-Chaudiere et al., 2003; Kokel et al., 2010; Rinkwitz et al., 2011; Becker and Rinkwitz, 2012), addiction (Petzold et al., 2009;Webb et al., 2009), disease treatment/drug development (Shin and Fishman, 2002; Lieschke and Currie, 2007; Mathew et al., 2007; Delvecchio et al., 2011; Truong et al., 2011) and gene–environment interactions (Hillwalker et al., 2010). The zebrafish model is becoming a prevalent model because the developmental processes are well conserved with other vertebrates, including humans (Gilbert, 2010). The embryos also develop rapidly and externally from the mother, and are optically transparent, allowing non-invasive whole organism modeling in vivo (Kimmel et al., 1995). Additionally, the embryonic zebrafish is genetically tractable and amenable to rapid-throughput analyses. However, unlike the rodent models, the diet of zebrafish has not been studied extensively or standardized (Lawrence, 2011). Commercial flake feeds are typically used in conjunction with various live feeds (e.g. Artemia nauplii, Paramecium and rotifers). These commercially prepared diets often include unknown or ill-defined ingredients (e.g. fish oil, or fish meal) and have variable compositions between batches. These variables could impact experimental results. In light of the increasing utilization of embryonic zebrafish as a research platform, it is important to begin to understand what impacts parental diets have on developing embryos.
The lack of consistent zebrafish laboratory diets is not a newconcern (Siccardi et al., 2009; Lawrence, 2011). We have previously reported outcomes fromz ebrafish fed a diet, which has fully defined components sufficient for zebrafish health (Lebold et al., 2011; Kirkwood et al., 2012; Miller et al., 2012). This diet has permitted the controlled removal of key nutrients from the parental diet, allowing the use of zebrafish to query nutrition-related questions. We previously demonstrated that the embryos from zebrafish fed a defined diet minus one nutrient, vitamin E (E−) were severely malformed and exhibited a heightened incidence of mortality compared to the complete diet (E+) (Miller et al., 2012).
Vitamin E was first discovered from its link to mammalian fertility (Evans and Bishop, 1922); its role in embryonic development, however, has remained largely unknown. The goal of this study was to identify developmental responses to different parental diets, and investigate the specific role of vitamin E. To understand the functional consequences of vitamin E deficiency, we employed a global transcriptome analysis of the embryos from adult zebrafish fed one of the three diets; Lab, E+ or E−.
2. Methods
2.1. Fish care and husbandry
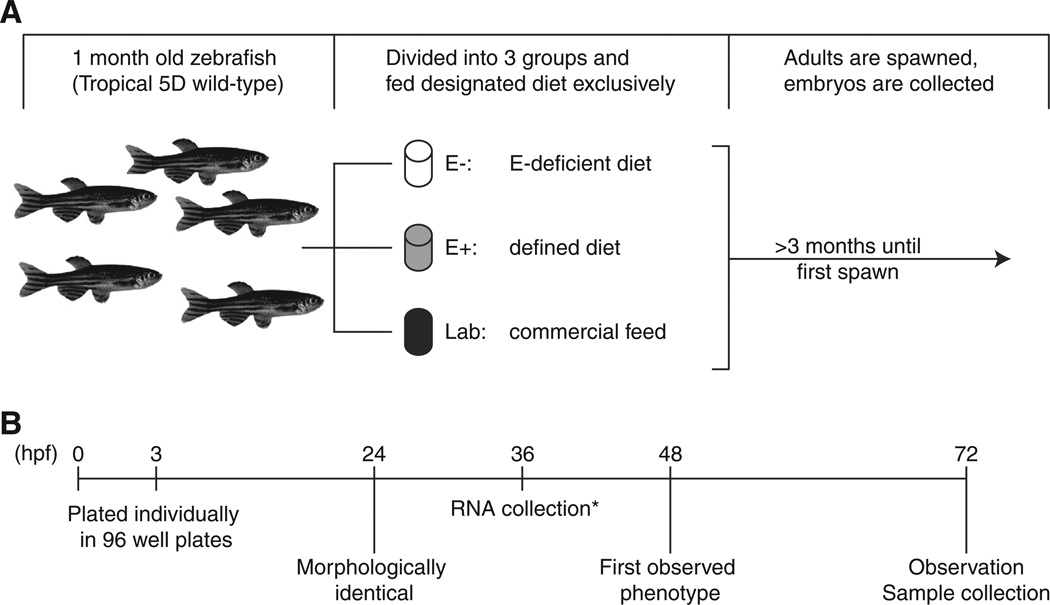
Tropical 5D (wild-type) zebrafish (Danio rerio) were reared and maintained at the Sinnhuber Aquatic Research Laboratory at Oregon State University, with protocols approved by the Oregon State University Institutional Animal Care and Use Committee (ACUP #3897). A schematic of the experimental design is shown in Fig. 1A. Briefly, juvenile zebrafish (~1 month of age)were divided into 3 groups and exclusively fed one of three designated diets: E−, E+ or Lab. Defined diets have been previously described (Lebold et al., 2011; Kirkwood et al., 2012; Miller et al., 2012), and similar diets were used (Table 1) with slightly lower ascorbic acid concentrations (350 mg/kg diet, added as Stay-C, Vitamin C-3, Argent Chemical Laboratories Inc., Redmond, WA, USA). The E+ diet contained 500 mg RRR-α-tocopherol/kg diet; the E− was made without added tocopherol. The standard laboratory diet is a mix of commercial feeds: Aquatox flake food (76.5% by weight, Ziegler Brothers, Gardener, CA, USA), Cyclopeeze (6.7% by weight, Argent Laboratories), and Golden Pearl 300–500 micron (16.8% by weight, Artemia International, Fairview, TX, USA). In addition to receiving this commercial flake mix, laboratory diet fish were also fed Artemia nauplii (GSL, Artemia International). Beginning at 3–4 months after diet initiation, adult zebrafish were group-spawned regularly and embryos collected and staged for subsequent experiments (Kimmel et al., 1995).
Fig. 1.
Experimental design and feeding scheme. A. Schematic of the experimental design and feeding regime. From left to right: 1-month old zebrafish were split into equal groups and exclusively fed one of three diets (E−, E+ or Lab diets). After 3 months of dietary initiation, zebrafish from each group were spawned regularly. B. Embryos fromeach diet were observed until 72 h post fertilization (hpf). Noted on the timeline are observations at 24, 48 and 72 hpf, samples were collected and euthanized at 72 hpf. *RNA samples were from one clutch only.
Table 1.
Defined diet composition.
| Ingredient | g/100 g diet |
|---|---|
| Wheat gluten (Dyets, Inc., Bethlehem, PA, USA) | 15 |
| Casein (Dyets, Inc.) | 30.5 |
| Egg whites (Dyets, Inc.) | 4 |
| Cellulose (Dyets, Inc.) | 3 |
| Vitamin mixa | 4 |
| Mineral mixb | 4 |
| Starch (National Starch Food Innovation, Bridgewater, NJ, USA) | 26.5 |
| Stripped soybean oil (Dyets, Inc.) | 7 |
| Lipoid PC 18:0/18:0 (Lipoid GmbH, Newark, NJ, USA) | 5 |
| Stay C (Argent Chemical Laboratories) | 3.5 |
| DL-α-tocopherol (where applicable; ADM, Decatur, IL, USA) | 0.05 |
| Ingredient | g/kg |
|---|---|
| Calcium carbonate | 19.23 |
| Calcium phosphate dibasic (2H2O) | 766.29 |
| Citric acid | 5.28 |
| Cupric carbonate | 0.36 |
| Ferric citrate | 2.99 |
| Magnesium oxide | 22.89 |
| Manganese carbonate | 5.65 |
| Sodium chloride | 28.02 |
| Disodium hydrogen phosphate | 11.89 |
| Zinc carbonate | 0.97 |
| Potassium phosphate dibasic | 74.16 |
| Potassium sulfate | 62.26 |
| Potassium iodide | 0.01 |
| Ingredient | g/kg |
|---|---|
| Vitamin A (500,000 IU/g) | 0.45 |
| Vitamin D3 (400,000 usp/µg) | 6.2445 |
| Vitamin K | 0.025 |
| Thiamine | 0.15 |
| Riboflavin | 0.25 |
| Vitamin B6 | 0.125 |
| Pantothenic acid | 0.75 |
| Niacin | 1.25 |
| Biotin | 0.005 |
| Folate | 0.05 |
| Vitamin B12 | 0.0005 |
| Myoinositol | 6.25 |
| PABA | 1 |
| Celufil (alpha cellulose) | 983.75 |
Mineral mix (Dyets Inc.):
Vitamin mix (Dyets Inc.):
2.2. Embryonic morphology
To characterize the morphologic effects, 3 hpf embryos from parents fed each of the three diets were placed into individual wells of a 96-well plate containing 100 μL of fish water (reverse osmosis filtered water containing 0.6% Instant Ocean salt solution) and observed daily for visible morphologic changes using a stereomicroscope at 24, 48 and 72 hpf (Fig. 1B). Results shown are from 3 separate spawns (combined totals: E−, n=358; E+, n=396; Lab, n= 296).α-Tocopherolwasmeasured in embryos from the abovementioned spawns (3 groups of 10 embryos per diet) at 24, 48 and 72 hpf, as previously described (Miller et al., 2012).
2.3. Microarray studies
Four biological replicates of 36 hpf embryos (n=35 per set), collected from adults fed one of the three diets (E−, E+ and Lab), were carefully age-matched and collected into RNALater (Life Technologies, Carlsbad, CA, USA). RNA was extracted with Trizol (Life Technologies) as per manufacturer's direction. RNA samples were further purified and characterized to specifications required for microarray analysis, previously described (Truong et al., 2013). Samples from the E− and Lab groups were kept as four independent biological replicates. Samples from the E+ diet were combined and split into four equal aliquots (technical replicates) to provide a common background comparison for the E− and Lab diet samples.
cDNA synthesis, labeling and array hybridization were performed by the University of Idaho Initiative for Bioinformatics and Evolutionary Studies (IBEST) core facility (Moscow, ID, USA) (Tal et al., 2012). The Nimble Gen Zebrafish 12 × 135 K Array platform (Roche NimbleGen, Madison, WI, USA) was used. The University of Idaho core facility normalized raw intensity scores and normalized values were used for subsequent analyses.
2.4. Bioinformatics data and pathway analyses
NimbleScan software (Roche NimbleGen) was used to extract raw and quantile normalized data. Gene call files were generated using Robust Multichip Average (RMA) algorithm. Statistical analysis of the normalized intensities was performed using GeneSpring GX 11 (Agilent Technologies, Santa Clara, CA, USA).
Functional and pathway analyses were performed on the output lists generated by GeneSpring (above). Entrez IDs were updated from the original NimbleGen annotation file (version 7 to version 9) prior to functional and pathway analyses. This updated list was generated by the Pacific Northwest National Laboratory (PNNL), and applied using the Bioinformatics Resource Manager (BRM Version 2.3) (Shah et al., 2007). Multi-Experiment Viewer (MeV) (Saeed et al., 2003) was used for heat map visualization, and analysis. The significant lists were subject to unsupervised bidirectional hierarchal clustering using Euclidean distance metric, and centroid linkage similarity clustering groups with similar expression patterns.
Functional enrichment statistics for the gene clusters were determined using the Database for Annotation, Visualization and Integrated Discovery (DAVID, http://david.abcc.ncifcrf.gov/, v6.7) (Huang da et al., 2009a,b,c). DAVID analyses were employed as previously described (Truong et al., 2013). Briefly, gene ontology category terms for significant genes were compared to a background containing the entire probe-set recognized by the NimbleGen Zebrafish 12 × 135 K Array platform. Functional annotation clustering with medium stringency was used to group similar annotations together into non-redundant functional groups. Functional Gene Ontology (GO) terms were deemed significant with a p < 0.05 as determined by DAVID using a modified Fisher's exact test (EASE score) (Huang da et al., 2009b). GO terms with the highest p-value from each cluster were compiled and used to describe the functional category of the clustered groups.
Ingenuity Pathway Analysis software (IPA, Ingenuity Systems, Redwood City, CA, USA) was used for further functional annotation of the gene sets. Mammalian Entrez gene identification numbers (obtained using BRM) and fold-change or p-values (obtained through GeneSpring analysis) were uploaded into the IPA database. The data sets were subject to IPA “Core analysis”, and the identified biological functions obtained were sorted by their z-score (putative activation/repression of biological function), number of affected molecules and p-value. Redundant functional groups were manually removed from the data set.
2.5. Statistical analyses
Initialmicroarray significance tests were performed using GeneSpring GX 11 software. Relative expression values were compared by unpaired Student's t-test with unequal variance; values were considered significant at p < 0.01.
Further statistical analyses were tabulated with Prism 5.0 (Graph Pad, La Jolla, CA, USA). Data are presented asmeans±SD, unless otherwise indicated. Comparisons across groups were by 2-tailed paired Student's t-test or 1-way ANOVA with Tukey's multiple-comparison post-hoc analysis; values of p < 0.05 were considered statistically significant.
3. Results
3.1. Embryo morphology and α-tocopherol concentrambryo morphology and α-tocophertions
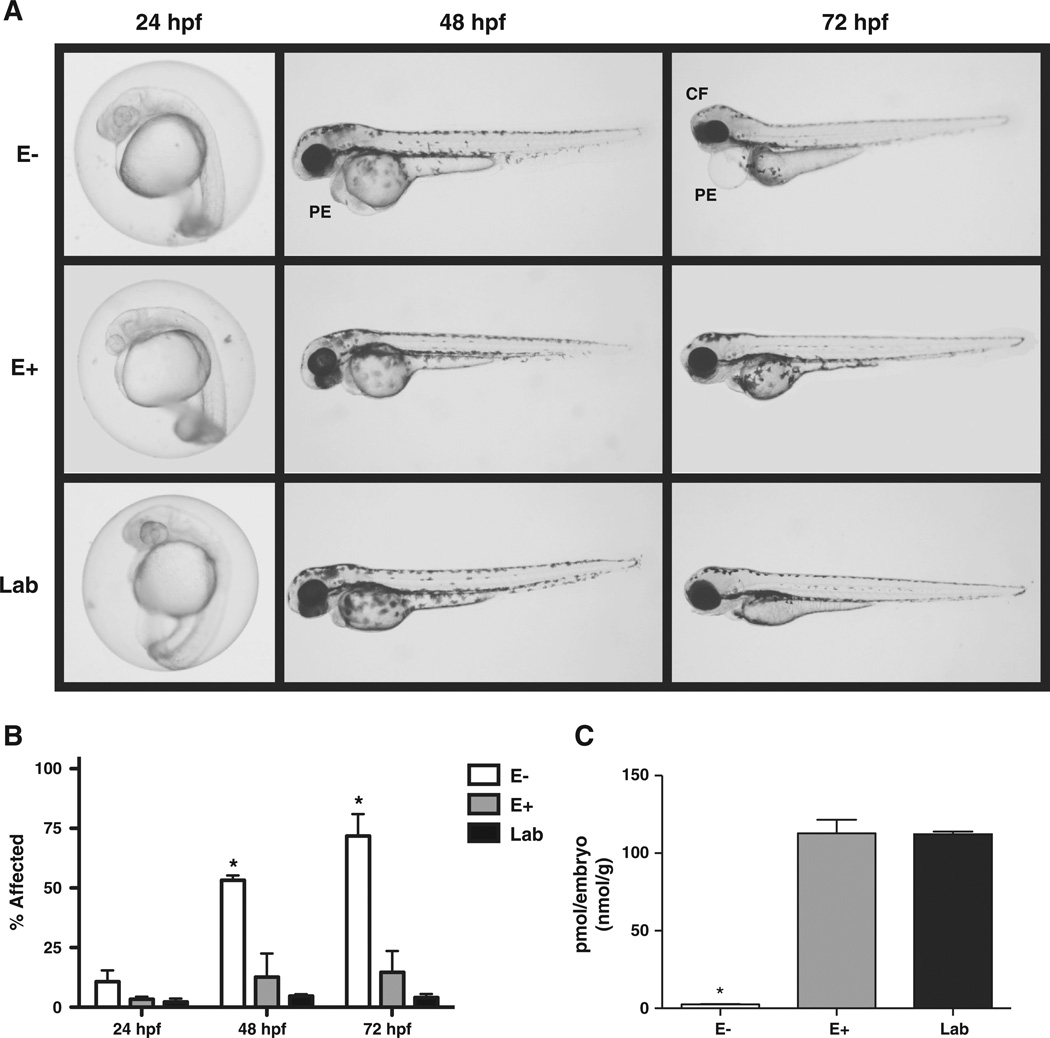
Embryos fromall diets developed properly when observed at 24 hpf (Fig. 2A and B). Those spawned from E+ and Lab diet fish continued to develop normally, showing no significant differences within the observation period (72 hpf). However, the E− embryos showed signs of abnormal development beginning at the 48 hpf (~50%). The first notable malformations in the E−embryos consisted of enlarged pericardial regions (pericardial edema, PE), slight craniofacial malformations (more noticeable at 72 hpf observation), and a discoloration of the yolk sack. These malformations increased in severity and affected ~70% of the embryos by 72 hpf (Fig. 2A and B).
Fig. 2.
Embryonic development differs with parental diet. A. Parental diet effects on embryomorphology. Representative images displaying embryomorphology at 3 time points for each diet. At 24 hpf, no adverse effects are noted across groups. At 48 hpf, the E− group begins to display pericardial defects (PE), which are dramatically apparent by 72 hpf in addition to cranial–facial malformations. The E+ and the Lab group embryos remain similar throughout. B. Embryos from each of the three groups are indistinguishable at 24 hpf, but at 48 hpf, >50% of embryos from the E− group show adverse outward signs of the vitamin E deficiency. At 72 hpf, nearly 70% of the E− embryos display abnormalities or mortalities. Embryos from the E+ and Lab diet groups remain similar throughout the experimental observations. C. Embryos spawned from E− diet-fed fish show deplete a-tocopherol at 72 hpf (n = 3 pools of 10 embryos for all diet groups). *Indicates p < 0.001 compared to E+ and Lab groups (one-way ANOVA, with Tukey's HSD post-test). Data are shown as mean ± SD.
Embryonic α-tocopherol status was confirmed in embryos from the three diet groups after the final morphological observation, 72 hpf. The E− embryos had nearly 40-fold lower α-tocopherol concentrations (2.6 ± 0.3 pmol/embryo) compared with either the E+ (112.8 ± 8.8 pmol/embryo) or the Lab diet embryos (112.2±1.8 pmol/embryo) (Fig. 2C).
3.2. Global transcriptional differences
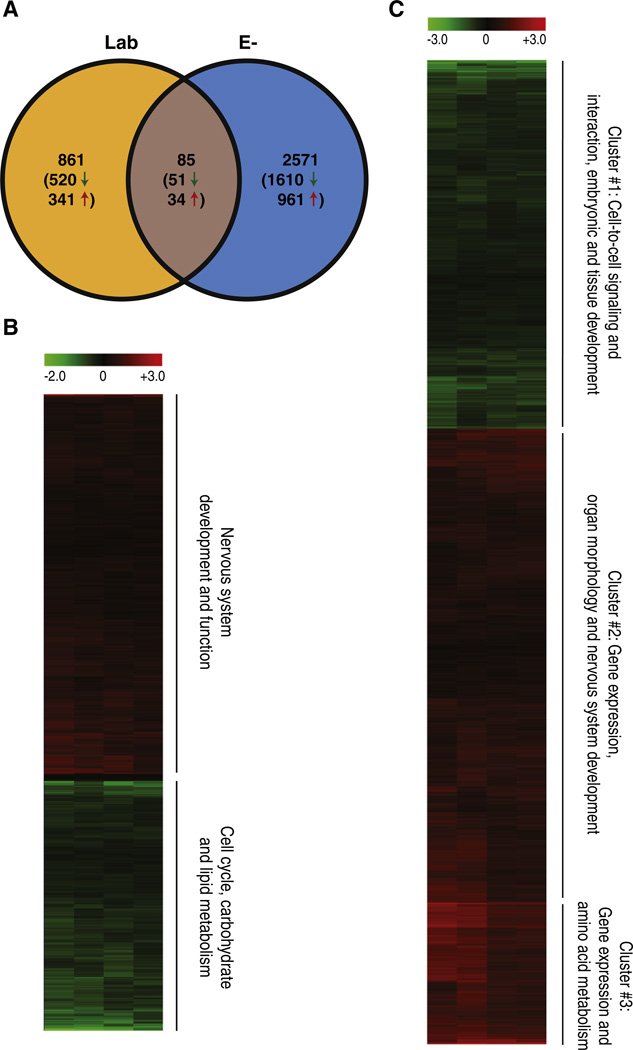
To elucidate the mechanism responsible for the abnormal development noted in the E− embryos, we collected RNA samples for global gene expression analysis at 36 hpf, a time prior to the appearance of any phenotypic malformations. At this time point, embryos spawned from all three groups were morphologically indistinguishable (data not shown). To normalize for the different diets fed to the adults, we used the E+ group as a control for the Lab and E−diets. Our first comparison was to determine if feeding zebrafish a defined diet in which every nutrient component is known, affected the transcriptome profile of the offspring. When doing this comparison (Lab vs E+ diet), we identified 946 statistically (p < 0.01) differentially regulated transcripts. The other comparison that was of interest was the significance and the perturbation that occurred at the transcript level when vitamin E was deficient in the adult, and passed onto their embryos. In this comparison (E− vs E+ diet), 2656 transcripts were differentially expressed. We found that 85 transcripts were common between the Lab and E− diets when compared to the E+ diet (Fig. 3A). Therefore, 861 transcripts were uniquely expressed in the Lab diet where 520 were repressed, and 341 were elevated, and in the E− diet, 2571 transcripts were unique with 1610 repressed and 961 elevated.
Fig. 3.
Global effects of parental diet on embryonic transcription. A. A Venn diagram displaying differentially represented transcripts. Both Lab and E− groups were first compared to the E+ defined diet control group; significant transcripts (p < 0.01) from each set were then compared as represented in the figure. B and C. Transcripts from Lab (B) and E−(C) that were significantly different from the E+ controls were grouped using bi-hierarchical clustering as displayed in the heat map. General annotations for each cluster are noted on the right side of the heat maps (annotation generated using IPA and DAVID).
3.3. Comparison of sufficient diet profiles (Lab vs E+)
Of the 946 differentially expressed transcripts between the Lab and the E+ diet embryos, 571 were decreased and 375 increased in the Lab diet (Fig. 3B). Using pathway analysis software, we first analyzed only the decreased transcripts and found that the genes were associated with metabolism and processing of carbohydrates and lipids, as well as basic cell cycle processes (Fig. 3B, bottom). Transcripts with increased expression were functionally clustered on nervous system development and function (Fig. 3B, top). Additionally, we analyzed the full set (946) and this revealed that the affected transcripts were linked to cellular growth, apoptosis, movement disorders, transcription, differentiation, and molecular transport in the Lab diet group (Table 2).
Table 2.
Biological process networks affected in Lab diet embryos.
| Biological process network | p-Value | # Molecules |
|---|---|---|
| Tissue development | 3.38E−07 | 123 |
| Cell death — apoptosis | 1.51E−02 | 92 |
| Organismal development — organogenesis | 1.55E−05 | 87 |
| Embryonic development — development of organ | 2.71E−05 | 85 |
| Cellular development — differentiation | 1.20E−02 | 74 |
| Cellular growth and proliferation — growth of cells | 3.38E−03 | 73 |
| Organismal survival — organismal death | 9.69E−03 | 62 |
| Gene expression — transcription of DNA | 4.36E−03 | 61 |
| Neurological disease — movement disorder | 1.99E−06 | 56 |
| Molecular transport — transport of molecule | 2.53E−04 | 56 |
3.4. Vitamin E deficiency induced transcriptional changes
The E− embryos had significant differential expression of 2656 transcripts compared to E+ diet embryos. Of these transcripts, 1661 were repressed and 995 were elevated in the E+ diet when compared to the E− diet embryo transcript levels (Fig. 3C). The elevated transcripts were implicated in gene expression, amino acid metabolism, organ morphology and nervous system development (clusters #2 and #3, Fig. 3C, Table 3). The repressed transcripts were associated with pathways involved in cell-to-cell signaling and interaction, embryonic and tissue development (cluster #1, Fig. 3C, and Table 3). Notably across all three clusters many differentially expressed transcripts were linked to control of cellular death, organogenesis, transcription of DNA and cellular growth and proliferation.
Table 3.
Biological process networks affected in E-embryos.
| Biological process network | p-Value | # Molecules |
|---|---|---|
| Cluster #1 | ||
| Cell death | 3.85E−03 | 142 |
| Cellular growth and proliferation — proliferation of cells | 3.17E−03 | 117 |
| Cell death — apoptosis | 1.26E−02 | 107 |
| Organismal survival — organismal death | 6.00E−04 | 78 |
| Organ morphology — morphology of organ | 2.43E−03 | 69 |
| Molecular transport — transport of molecule | 3.63E−04 | 63 |
| Cell cycle — cell cycle progression | 1.23E−03 | 53 |
| Cell death — cell survival | 1.48E−02 | 52 |
| Cellular function and maintenance — organization of cytoplasm | 1.53E−03 | 50 |
| Cell death — cell death of organ | 5.48E−03 | 48 |
| Cluster #2 | ||
| Gene expression — transcription of DNA | 1.19E−24 | 149 |
| Gene expression — activation of DNA endogenous promoter | 1.59E−21 | 111 |
| Tissue development — tissue development | 9.93E−16 | 198 |
| Cell death | 3.57E−07 | 196 |
| Cellular growth and proliferation — proliferation of cells | 4.36E−06 | 158 |
| Embryonic development — organogenesis | 2.66E−13 | 146 |
| Organismal development — development of organ | 1.72E−13 | 145 |
| Cellular development — differentiation | 3.00E−12 | 145 |
| Organismal survival — organismal death | 2.55E−09 | 115 |
| Organ morphology — morphology of organ | 2.62E−09 | 107 |
| Cluster #3 | ||
| Cell death | 2.70E−02 | 38 |
| Tissue development | 5.98E−03 | 35 |
| Gene expression — expression of RNA | 7.12E−05 | 34 |
| Cellular growth and proliferation — proliferation of cells | 3.55E−02 | 31 |
| Cellular development — differentiation | 5.70E−03 | 27 |
| Organismal survival — organismal death | 5.68E−03 | 23 |
| Cell cycle — cell cycle progression | 2.47E−04 | 20 |
| Neurological disease — encephalopathy | 4.29E−03 | 20 |
| Cellular function and maintenance — organization of cytoplasm | 2.24E−03 | 17 |
| Skeletal and muscular disorders — neuromuscular disease | 8.00E−03 | 16 |
The transcripts with altered expression levels also revealed numerous perturbed pathways. When these pathways were clustered together and analyzed utilizing bioinformatics programs (DAVID, and IPA), lipid peroxidation was predicted as a key affected process, as might be expected from vitamin E's antioxidant function. Evaluation of the data using the IPA core analysis function suggests that the lack of vitamin E perturbs energy metabolism. Transcripts in this analysis were significantly misex pressed in E− embryos: transcript levels of the amyloid precursor protein (APP) were decreased; apolipoprotein E (ApoE), nuclear receptor subfamily 4, group A, member 3 (NR4A3), cAMPresponse element-binding (CREB) binding protein (CREBBP), peroxi some proliferation-activated receptor gamma, co-activator 1-alpha (PGC1A) and peroxisome proliferation-activated receptor gamma, co-activator 1-beta (PGC1B) had increased transcript levels compared to E + control. These genes were misexpressed in E− embryos theoretically resulting in perturbed energy metabolism and mitochondrial function ultimately causing embryonic malformations and mortality.
4. Discussion
4.1. “Hidden” transcriptional differences
The differences in nutrient profiles between our E+ diet and the Lab diet are multifaceted (Lebold et al., 2011; Kirkwood et al., 2012; Miller et al., 2012), but adult zebrafish fed either diet produce viable, and apparently healthy embryos. Interestingly, without obvious phenotypic differences, our analysis revealed 946 differentially represented transcripts between embryos from the two diets. We anticipated many of these changes due to the different dietary compositions. The Lab diet has been formulated to provide more than adequate nutrition for the fish with an overabundance of many nutrients especially polyunsaturated fatty acids (PUFAs), while our defined diet is closely tailored to zebrafish nutritional requirements and to exacerbate the vitamin E deficiency by limiting the availability of both arachidonic and docosahexaenoic acids. The E+ diet, by design, contains only the essential fatty acids, linoleic and alpha-linolenic acids, but not longer chain, n−3 or n−6, fatty acids (Lebold et al., 2011). Additionally, the defined diets contain structured synthetic phosphatidyl choline with only stearic acids. The PUFA content and distribution are important for early central nervous system development (Guesnet and Alessandri, 2011). Fish fed the defined dietmust synthesize the additional required PUFAs from the dietary precursors (Lebold et al., 2011), thereby affecting the PUFA content of their progeny (data not shown). Alternatively, the Lab diet provides an excess of fish oil/meal, containing high levels of n−3 PUFAs (e.g. fish oils, DHA, EPA), and may contain oxidized fatty acids as well. These dietary differences likely drive the differences in genes responsible for lipid metabolism. Similarly, we attribute the differences in carbohydrate metabolism to the nutrient differences between the Lab and E+ diets; the Lab diet had a mixture of sources of carbohydrates, while the E+ diet only contained starch. In addition, there were major differences in various phytochemicals and dietary components (i.e. different levels of vitamins, minerals, phytochemicals, and toxicants).
The remaining transcriptional differences between Lab and E+ are less easily explained. There was increased transcription of genes associated with tissue development, organogenesis, cellular differentiation, transcription and molecular transport in the Lab embryos. Pathways involving apoptosis, cell growth, organismal death and neurological disease all had decreased transcript levels compared to the control expression pattern. These changes are due to the parental diet differences; thereby raising the concern that widespread use of a controlled and well-defined diet has not been properly addressed in this model (Hau and Schapiro, 2011; Lawrence, 2011). While the optimum nutrient levels are likely not met by either the E+ or the Lab diet, the effect of parental diet on embryonic transcription cannot be denied. More importantly, these concealed embryonic effects may have further reaching impact (e.g. behavior, fertility, life expectancy of the adult animal). The limitation in embryonic growth in the E+ diet animals once they are fed this diet also suggests that there are limiting components that have not yet been identified (unpublished observations, Tanguay laboratory).
The need to standardize zebrafish laboratory diet is recognized within the zebrafish community (DeKoven et al., 1992; Markovich et al., 2007; Siccardi et al., 2009; Lawrence, 2011). Current research diets include ambiguous ingredients, their content may vary between batches, and they may contain toxins, or toxic levels of some ingredients (e.g. mercury found in Artemia from the Great Salt Lake (Peterson and Gustin, 2008), and vitellogenin induction due to commercial diets in farmed tilapia (Davis et al., 2009)). Furthermore, these results beg the question of long-term effects on growth, chemical susceptibility, or behavior due to the parental diet-mediated developmental programming alterations. Previous groups have formulated partially defined diets for laboratory fish (DeKoven et al., 1992; Markovich et al., 2007; Siccardi et al., 2009), but the main source of fish nutritional research is found in large-scale aquaculture (National Research Council (U.S.). Committee on Animal Nutrition, 1993), which is focused on adult health.
4.2. Vitamin E deficiency
We previously reported that E-deficient embryos have a higher incidence of malformations and mortalities by 120 hpf (Miller et al., 2012). In the present study we characterized the onset of these malformations throughout early development to determine a morphologic anchor to study the affected mechanisms. The parental fish in the E− and E+ groups were fed identical, fully defined diets, differing only in the amount of vitamin E. The E-deficient state of the parents (data not shown, Miller et al., 2012) was passed on to their progeny (Fig. 2C). The lack of this single micronutrient in the adult fish resulted in developmental abnormalities of the offspring by 48 hpf (Fig. 2); by 72 hpf, >70% of E− embryos typically displayed gross malformations or mortality. The removal of vitamin E from the parental diet resulted in a higher number of significantly misregulated transcripts than using a defined diet. The transcriptional profile provides a range of possible mechanisms for a vitamin E requirement. Basic developmental functions were chief amongst affected transcripts: organ morphology was differentially regulated, appearing in all significant clusters of E− compared to E+ embryos; transcripts responsible for cell proliferation, differentiation and death were also decreased in the E−embryos. Although these changes preceded the development of a visible phenotype, many are indicative of a systemic problem, and as such are likely downstream of the initial E deficient effect. Similar transcriptional patterns have been seen in previous vitamin E deficiency studies. In cultured colon cancer cells, the vitamin E status influenced genes related to cell cycle (proliferation, differentiation and death), DNA damage repair, cell signaling and connective tissue (Lunec et al., 2004). Diet induced vitamin E deficiency in pregnant rats resulted in their offspring displaying dysregulation in similar pathways (cell cycle and signaling), as well as lipid transport and neuronal development (Roy et al., 2002). Similar effects have been noted usingα-tocopherol transfer protein knockout mice, a model commonly used to study vitamin E deficiency (Jishage et al., 2001; Gohil et al., 2003, 2004; Vasu et al., 2007). However, these studies are focused on specific tissues from adult animals making the comparison to the whole, developing zebrafish embryo difficult.
Vitamin E is known as a potent lipid soluble antioxidant (Burton and Ingold, 1981; Kamal-Eldin and Appelqvist, 1996; Traber and Atkinson, 2007), and while this action has been indirectly demonstrated in vivo (Mastaloudis et al., 2004; Bruno et al., 2005, 2006; Traber, 2013) the precise biological role of vitamin E is still unknown. Oxidative stress is produced through endogenous processes and is common throughout development (Jauniaux et al., 2004); vitamin E acts as a lipid-specific antioxidant to limit the lipid peroxidation chain reaction. The present analysis takes into account the entire developing embryo, thus allowing inferences to the systemic consequences of vitamin E deficiency. Analysis of the transcriptional changes suggested that energy metabolism is perturbed in developing animals due to the removal of vitamin E from the parental diet. In the presence of vitamin E, embryos remain in a basal, or ‘quiet’ (Leese, 2002; Leese et al., 2007), metabolic state; a matching basal transcription rate for metabolically active transcription factors such as PGC1A, PGC1B, CREBBP, or NR4A3. However, in the absence of vitamin E, lipid peroxidation would over whelm the antioxidant capacity of the embryo through rampant lipid oxidation (primarily in the highly-oxidizable PUFAs (Hashimoto et al., 2005; Spiteller, 2006; Andreo et al., 2011; Corsinovi et al., 2011)), perturbing the ‘quiet’ metabolic state in the embryo. Metabolic regulation would switch to an ‘active’ state in response to the increased stress from the lack of vitamin E. This is supported by the significantly increased expression of both PGC1A and PGC1B mRNA in the E− embryos. Both are nuclear receptor cofactors with global biologic functions, chiefly serving as mediators of energy metabolism (Handschin and Spiegelman, 2006; Liu and Lin, 2011). The system-wide importance of these cofactors explains the developmental defects, and increased mortalities caused by vitamin E deficiency. PGC1A and PGC1B are integral to zebrafish health and development specifically, they are necessary for oxytocin production in the zebrafish brain (Blechman et al., 2011), and their expression is colocalized with many known zebrafish nuclear receptors (Bertrand et al., 2007). Coupled with the inherent advantages and tractability of the vitamin E deficient zebrafish model, this hypothesized mechanism illuminates a plausible role for vitamin E in development.
In summary, the diet fed to adult zebrafish should not be overlooked when studying embryonic zebrafish. As our results illustrate, the parental diet impacts the basal embryonic transcriptome. These changes in gene expression could lead to long-term physiological or functional effects, and could also alter susceptibility to other stressors. Collectively, all these data emphasize that a defined laboratory diet is needed to ensure greater intra-laboratory consistency. Efforts to develop a standardize laboratory diet will further advance the zebrafish as a preeminent biomedical model.
Acknowledgments
These studies were partially funded by NIH grants RO1 HD062109 and P30 ES000210. The authors would like to thank Scott Leonard and Charlotte Wright for assistance with vitamin E measurements.
References
- Andreo U, Elkind J, Blachford C, Cederbaum AI, Fisher EA. Role of superoxide radical anion in the mechanism of apoB100 degradation induced by DHA in hepatic cells. FASEB J. 2011;25:3554–3560. doi: 10.1096/fj.11-182725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becker TS, Rinkwitz S. Zebrafish as a genomics model for human neurological and polygenic disorders. Dev. Neurobiol. 2012;72:415–428. doi: 10.1002/dneu.20888. [DOI] [PubMed] [Google Scholar]
- Bertrand S, Thisse B, Tavares R, Sachs L, Chaumot A, Bardet PL, Escriva H, Duffraisse M, Marchand O, Safi R, Thisse C, Laudet V. Unexpected novel relational links uncovered by extensive developmental profiling of nuclear receptor expression. PLoS Genet. 2007;3:e188. doi: 10.1371/journal.pgen.0030188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blechman J, Amir-Zilberstein L, Gutnick A, Ben-Dor S, Levkowitz G. The metabolic regulator PGC-1 alpha directly controls the expression of the hypothalamic neuropeptide oxytocin. J. Neurosci. 2011;31:14835–14840. doi: 10.1523/JNEUROSCI.1798-11.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruno RS, Ramakrishnan R, Montine TJ, Bray TM, Traber MG. α-Tocopherol disappearance is faster in cigarette smokers and is inversely related to their ascorbic acid status. Am. J. clin. Nutr. 2005;81:95–103. doi: 10.1093/ajcn/81.1.95. [DOI] [PubMed] [Google Scholar]
- Bruno RS, Leonard SW, Atkinson J, Montine TJ, Ramakrishnan R, Bray TM, Traber MG. Faster plasma vitamin E disappearance in smokers is normalized by vitamin C supplementation. Free. Radic. Biol. Med. 2006;40:689–697. doi: 10.1016/j.freeradbiomed.2005.10.051. [DOI] [PubMed] [Google Scholar]
- Burton GW, Ingold KU. Autoxidation of biological molecules. 1. The antioxidant activity of vitamin-E and related chain-breaking phenolic antioxidants in vitro. J. Am. Chem. Soc. 1981;103:6472–6477. [Google Scholar]
- Corsinovi L, Biasi F, Poli G, Leonarduzzi G, Isaia G. Dietary lipids and their oxidized products in Alzheimer'sdisease. Mol. Nutr. Food. Res. 2011;55(Suppl. 2):S161–S172. doi: 10.1002/mnfr.201100208. [DOI] [PubMed] [Google Scholar]
- Dambly-Chaudiere C, Sapede D, Soubiran F, Decorde K, Gompel N, Ghysen A. The lateral line of zebrafish: a model system for the analysis of morphogenesis and neural development in vertebrates. Biol. Cell. 2003;95:579–587. doi: 10.1016/j.biolcel.2003.10.005. [DOI] [PubMed] [Google Scholar]
- Davis LK, Fox BK, Lim C, Hiramatsu N, Sullivan CV, Hirano T, Grau EG. Induction of vitellogenin production in male tilapia (Oreochromismos sambicus) by commercial fish diets. Comp. Bio. chem. Physiol. A. 2009;154:249–254. doi: 10.1016/j.cbpa.2009.06.009. [DOI] [PubMed] [Google Scholar]
- DeKoven DL, Nunez JM, Lester SM, Conklin DE, Marty GD, Parker LM, Hinton DE. A purified diet for medaka (Oryzias latipes): refining a fish model for toxicological research. Lab. Anim. Sci. 1992;42:180–189. [PubMed] [Google Scholar]
- Delvecchio C, Tiefenbach J, Krause HM. The zebrafish: a powerful platform for in vivo, HTS drug discovery. Assay Drug Dev. Technol. 2011;9:354–361. doi: 10.1089/adt.2010.0346. [DOI] [PubMed] [Google Scholar]
- Evans HM, Bishop KS. On the existence of a hitherto unrecognized dietary factor essential for reproduction. Science. 1922;56:650–651. doi: 10.1126/science.56.1458.650. [DOI] [PubMed] [Google Scholar]
- Gilbert SF. Developmental Biology. 9th ed. Sunderland, Mass: Sinauer Associates; 2010. [Google Scholar]
- Gohil K, Schock BC, Chakraborty AA, Terasawa Y, Raber J, Farese RV, Jr, Packer L, Cross CE, Traber MG. Gene expression profile of oxidant stress and neurode-generation in transgenic mice deficient in alpha-tocopherol transfer protein. Free Radic. Biol. Med. 2003;35:1343–1354. doi: 10.1016/s0891-5849(03)00509-4. [DOI] [PubMed] [Google Scholar]
- Gohil K, Godzdanker R, O'Roark E, Schock BC, Kaini RR, Packer L, Cross CE, Traber MG. Alpha-tocopherol transfer protein deficiency in mice causes multi-organ deregulation of gene networks and behavioral deficits with age. Ann. N. Y. Acad. Sci. 2004;1031:109–126. doi: 10.1196/annals.1331.012. [DOI] [PubMed] [Google Scholar]
- Guesnet P, Alessandri JM. Docosahexaenoic acid (DHA) and the developing central nervous system (CNS) — implications for dietary recommendations. Biochimie. 2011;93:7–12. doi: 10.1016/j.biochi.2010.05.005. [DOI] [PubMed] [Google Scholar]
- Handschin C, Spiegelman BM. Peroxisome proliferator-activated receptor gamma coactivator 1 coactivators, energy homeostasis, and metabolism. Endocr. Rev. 2006;27:728–735. doi: 10.1210/er.2006-0037. [DOI] [PubMed] [Google Scholar]
- Hashimoto M, Tanabe Y, Fujii Y, Kikuta T, Shibata H, Shido O. Chronic administration of docosahexaenoic acid ameliorates the impairment of spatial cognition learning ability in amyloid beta-infused rats. J. Nutr. 2005;135:549–555. doi: 10.1093/jn/135.3.549. [DOI] [PubMed] [Google Scholar]
- Hau J, Schapiro SJ. Handbook of Laboratory Animal Science: Essential Principles and Practices. 3rd. ed. Boca Raton, FL: CRC Press/Taylor & Francis Group; 2011. [Google Scholar]
- Hillwalker WE, Allan SE, Tanguay RL, Anderson KA. Exploiting lipid-free tubing passive samplers and embryonic zebrafish to link site specific contaminant mixtures to biological responses. Chemosphere. 2010;79:1–7. doi: 10.1016/j.chemosphere.2010.02.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009a;37:1–13. doi: 10.1093/nar/gkn923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009b;4:44–57. doi: 10.1038/nprot.2008.211. [DOI] [PubMed] [Google Scholar]
- Huang da W, Sherman BT, Zheng X, Yang J, Imamichi T, Stephens R, Lempicki RA. Extracting biological meaning from large gene lists with DAVID. Current protocols in bioinformatics/editoral board, Andreas D. Baxevanis … [et al.] 2009c;Chapter13:13.11.11–13.11.13. doi: 10.1002/0471250953.bi1311s27. [DOI] [PubMed] [Google Scholar]
- Jauniaux E, Cindrova-Davies T, Johns J, Dunster C, Hempstock J, Kelly FJ, Burton GJ. Distribution and transfer pathways of antioxidant molecules inside the first trimester human gestational sac. J. Clin. Endocrinol. Metab. 2004;89:1452–1458. doi: 10.1210/jc.2003-031332. [DOI] [PubMed] [Google Scholar]
- Jishage K, Arita M, Igarashi K, Iwata T, Watanabe M, Ogawa M, Ueda O, Kamada N, Inoue K, Arai H, Suzuki H. Alpha-tocopherol transfer protein is important for the normal development of placental labyrinthine trophoblasts in mice. J. Biol. Chem. 2001;276:1669–1672. doi: 10.1074/jbc.C000676200. [DOI] [PubMed] [Google Scholar]
- Kamal-Eldin A, Appelqvist LA. The chemistry and antioxidant properties of tocopherols and tocotrienols. Lipids. 1996;31:671–701. doi: 10.1007/BF02522884. [DOI] [PubMed] [Google Scholar]
- Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF. Stages of embryonic development of the zebrafish. Dev. Dyn. 1995;203:253–310. doi: 10.1002/aja.1002030302. [DOI] [PubMed] [Google Scholar]
- Kirkwood JS, Lebold KM, Miranda CL, Wright CL, Miller GW, Tanguay RL, Barton CL, Traber MG, Stevens JF. Vitamin C deficiency activates the purine nucleotide cycle in zebrafish. J. Biol. Chem. 2012;287:3833–3841. doi: 10.1074/jbc.M111.316018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kokel D, Bryan J, Laggner C, White R, Cheung CY, Mateus R, Healey D, Kim S, Werdich AA, Haggarty SJ, Macrae CA, Shoichet B, Peterson RT. Rapid behavior-based identification of neuroactive small molecules in the zebrafish. Nat. Chem. Biol. 2010;6:231–237. doi: 10.1038/nchembio.307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrence C. Advances in zebrafish husbandry and management. Methods. Cell. Biol. 2011;104:429–451. doi: 10.1016/B978-0-12-374814-0.00023-9. [DOI] [PubMed] [Google Scholar]
- Lebold KM, Jump DB, Miller GW, Wright CL, Labut EM, Barton CL, Tanguay RL, Traber MG. Vitamin E deficiency decreases long-chain PUFA in zebrafish (Danio rerio) J. Nutr. 2011;141:2113–2118. doi: 10.3945/jn.111.144279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leese HJ. Quiet please, do not disturb: a hypothesis of embryo metabolism and viability. BioEssays. 2002;24:845–849. doi: 10.1002/bies.10137. [DOI] [PubMed] [Google Scholar]
- Leese HJ, Sturmey RG, Baumann CG, McEvoy TG. Embryo viability and metabolism: obeying the quiet rules. Hum. Reprod. 2007;22:3047–3050. doi: 10.1093/humrep/dem253. [DOI] [PubMed] [Google Scholar]
- Lieschke GJ, Currie PD. Animal models of human disease: zebrafish swim into view. Nat. Rev. Genet. 2007;8:353–367. doi: 10.1038/nrg2091. [DOI] [PubMed] [Google Scholar]
- Liu C, Lin JD. PGC-1 coactivators in the control of energy metabolism. Acta Biochim. Biophys. Sin. 2011;43:248–257. doi: 10.1093/abbs/gmr007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lunec J, Halligan E, Mistry N, Karakoula K. Effect of vitamin E on gene expression changes in diet-related carcinogenesis. Ann. N. Y. Acad. Sci. 2004;1031:169–183. doi: 10.1196/annals.1331.016. [DOI] [PubMed] [Google Scholar]
- Markovich ML, Rizzuto NV, Brown PB. Diet affects spawning in zebrafish. Zebrafish. 2007;4:69–74. doi: 10.1089/zeb.2006.9993. [DOI] [PubMed] [Google Scholar]
- Mastaloudis A, Morrow JD, Hopkins DW, Devaraj S, Traber MG. Antioxidant supplementation prevents exercise-induced lipid peroxidation, but not inflammation, in ultramarathon runners. Free Radic. Biol. Med. 2004;36:1329–1341. doi: 10.1016/j.freeradbiomed.2004.02.069. [DOI] [PubMed] [Google Scholar]
- Mathew LK, Sengupta S, Kawakami A, Andreasen EA, Lohr CV, Loynes CA, Renshaw SA, Peterson RT, Tanguay RL. Unraveling tissue regeneration pathways using chemical genetics. J. Biol. Chem. 2007;282:35202–35210. doi: 10.1074/jbc.M706640200. [DOI] [PubMed] [Google Scholar]
- Miller GW, Labut EM, Lebold KM, Floeter A, Tanguay RL, Traber MG. Zebrafish (Dani orerio) fed vitamin E-deficient diets produce embryos with increased morphologic abnormalities and mortality. J. Nutr. Biochem. 2012;23:478–486. doi: 10.1016/j.jnutbio.2011.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- National Research Council (U.S.) Nutrient Requirements of Fish. Washington, D.C.; National Academy Press; 1993. Committee on Animal Nutrition. [Google Scholar]
- Patton EE, Zon LI. The art and design of genetic screens: zebrafish. Nat. Rev. Genet. 2001;2:956–966. doi: 10.1038/35103567. [DOI] [PubMed] [Google Scholar]
- Peterson C, Gustin M. Mercury in the a, water and biota at the Great Salt Lake (Utah, USA) Sci. Total Environ. 2008;405:255–268. doi: 10.1016/j.scitotenv.2008.06.046. [DOI] [PubMed] [Google Scholar]
- Petzold AM, Balciunas D, Sivasubbu S, Clark KJ, Bedell VM, Westcot SE, Myers SR, Moulder GL, Thomas MJ, Ekker SC. Nicotine response genetics in the zebrafish. Proc. Natl. Acad. Sci. U. S. A. 2009;106:18662–18667. doi: 10.1073/pnas.0908247106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinkwitz S, Mourrain P, Becker TS. Zebrafish: an integrative system for neurogenomics and neurosciences. Prog. Neurobiol. 2011;93:231–243. doi: 10.1016/j.pneurobio.2010.11.003. [DOI] [PubMed] [Google Scholar]
- Roy S, Lado BH, Khanna S, Sen CK. Vitamin E sensitive genes in the developing rat fetal brain: a high-density oligonucleotide microarray analysis. FEBS Lett. 2002;530:17–23. doi: 10.1016/s0014-5793(02)03309-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, Braisted J, Klapa M, Currier T, Thiagarajan M, Sturn A, Snuffin M, Rezantsev A, Popov D, Ryltsov A, Kostukovich E, Borisovsky I, Liu Z, Vinsavich A, Trush V, Quackenbush J. TM4: a free, open-source system for microarray data management and analysis. BioTechniques. 2003;34:374–378. doi: 10.2144/03342mt01. [DOI] [PubMed] [Google Scholar]
- Shah AR, Singhal M, Klicker KR, Stephan EG, Wiley HS, Waters KM. Enabling high-throughput data management for systems biology: the Bioinformatics Resource Manager. Bioinformatics. 2007;23:906–909. doi: 10.1093/bioinformatics/btm031. [DOI] [PubMed] [Google Scholar]
- Shin JT, Fishman MC. From Zebrafish to human: modular medical models. Annu. Rev. Genomics Hum. Genet. 2002;3:311–340. doi: 10.1146/annurev.genom.3.031402.131506. [DOI] [PubMed] [Google Scholar]
- Siccardi AJ, III, Garris HW, Jones WT, Moseley DB, D'Abramo LR, Watts SA. Growth and survival of zebrafish (Danio rerio) fed different commercial and laboratory diets. Zebrafish. 2009;6:275–280. doi: 10.1089/zeb.2008.0553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spiteller G. Peroxyl radicals: inductors of neurodegenerative and other inflammatory diseases. Their origin and how they transform cholesterol, phospholipids, plasmalogens, polyunsaturated fatty acids, sugars, and proteins into deleterious products. Free Radic. Biol. Med. 2006;41:362–387. doi: 10.1016/j.freeradbiomed.2006.03.013. [DOI] [PubMed] [Google Scholar]
- Tal TL, Franzosa JA, Tilton SC, Philbrick KA, Iwaniec UT, Turner RT, Waters KM, Tanguay RL. MicroRNAs control neurobehavioral development and function in zebrafish. FASEB. J. 2012;26:1452–1461. doi: 10.1096/fj.11-194464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Traber MG. Mechanisms for the prevention of vitamin E excess. J. Lipid Res. 2013;54:2295–2306. doi: 10.1194/jlr.R032946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Traber MG, Atkinson J. Vitamin E, antioxidant and nothing more. Free Radic. Biol. Med. 2007;43:4–15. doi: 10.1016/j.freeradbiomed.2007.03.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Truong L, Harper SL, Tanguay RL. Evaluation of embryotoxicity using the zebrafish model. Methods Mol Biol. 2011;691:271–279. doi: 10.1007/978-1-60761-849-2_16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Truong L, Tilton SC, Zaikova T, Richman E, Waters KM, Hutchison JE, Tanguay RL. Surface functionalities of gold nanoparticles impact embryonic gene expression responses. Nanotoxicology. 2013;7:192–201. doi: 10.3109/17435390.2011.648225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasu VT, Hobson B, Gohil K, Cross CE. Genome-wide screening of alpha tocopherol sensitive genes in heart tissue from alpha-tocopherol transfer protein null mice (ATTP(−/−)) FEBS Lett. 2007;581:1572–1578. doi: 10.1016/j.febslet.2007.03.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Webb KJ, Norton WH, Trumbach D, Meijer AH, Ninkovic J, Topp S, Heck D, Marr C, Wurst W, Theis FJ, Spaink HP, Bally-Cuif L. Zebrafish reward mutants reveal novel transcripts mediating the behavioral effects of amphetamine. Genome Biol. 2009;10:R81. doi: 10.1186/gb-2009-10-7-r81. [DOI] [PMC free article] [PubMed] [Google Scholar]