Abstract
Single Nucleotide Polymorphisms (SNPs) between microarray probes and RNA targets can affect the performance of expression array by weakening the hybridization. In this paper, we examined the effect of the SNPs on Affymetrix GeneChip probe set summaries and the expression quantitative trait loci (eQTL) mapping results in two eQTL datasets, one from mouse and one from human. We showed that removing SNP-containing probes significantly changed the probe set summaries and the more SNP-containing probes we removed the greater the change. Comparison of the eQTL mapping results between with and without SNP-containing probes showed that less than 70% of the significant eQTL peaks were concordant regardless of the significance threshold. These results indicate that SNPs do affect both probe set summaries and eQTLs (both cis and trans), thus SNP-containing probes should be filtered out to improve the performance of eQTL mapping.
Keywords: microarray, SNP, eQTL, cis-eQTL, trans-eQTL, mouse, human
Introduction
Microarray probes are designed to match the sequences of the target genes in the selected reference genome. However, due to individual and population variation, some probes may not exactly match the sequence of the RNA applied to the arrays. The polymorphisms that are most likely encountered in microarray experiments are single nucleotide polymorphisms (SNPs) because they are much more abundant than any other type of polymorphisms in most species including human and mouse [1]. Therefore, it is safe to speculate that a substantial number of probes on human and mouse microarrays overlap with SNPs in population studies, such as expression quantitative trait loci (eQTL) studies.
Sequence polymorphisms between a probe on the microarray and its target can affect the hybridization signal [2]. The effect of SNPs on probe-target hybridization has long been utilized in the Affymetrix short oligo microarrays. The Affymetrix expression GeneChips use a single base-pair mismatch in the middle of the probe sequence (mismatch probe) to estimate non-specific hybridization background of the corresponding perfect match probe [3]. This property is also fully utilized in the Affymetrix SNP Arrays, where each SNP is interrogated with approximately 40 different probes. The hybridization signal differences caused by a SNP are used to infer the genotype for an individual at a given locus[4].
The effect of SNPs on probe-target hybridization has also enabled the identification of novel sequence polymorphisms [5,6]. For example, Borevitz et. al. [6] used genomic DNA of Arobidopsis theliana to hybridize to RNA expression GeneChips. They were able to identify a large number of single feature polymorphisms (SFPs), which ranged from single nucleotide polymorphisms to large deletions. Comparing the SFPs and genomic sequences, Rostoks et al.,[5] showed that a large proportion of the identified SFPs with sequence information available contain SNPs. Rostoks et. al., [5] further confirmed that the SNPs located near the center of the probes are more likely to be identified as SFP.
Even though sequence differences can lead to different hybridization intensity, it is not clear how this affects the summary score of a probe set consisting of 11–16 probes on Affymetrix expression microarrays. If the existence of SNPs does affect the probe set summary, it is also likely to affect gene expression QTL (eQTL) studies for mapping the genetic factors controlling gene expression when a probe perfectly matches one allele but mismatches the other in the mapping population. In an eQTL study, the expression of each gene (the summary of each probe set for Affymetrix arrays) is considered as a quantitative trait for QTL study in a segregating population [7–11]. Most of eQTL studies identify both cis-eQTLs (eQTLs mapped to the same genomic location as the expressed gene) and trans-eQTLs (eQTL mapped to a different genomic location from the expressed gene) [12–14]. A few eQTL studies have briefly examined the effect of SNPs on eQTLs. One study indicated that SNPs are enriched in the cis-regulated eQTLs compared in the trans-regulated eQTLs [15]. Another study [14] made similar findings but stated that the net effect of the SNP on the eQTLs was likely small and “only a relatively small number of cis-acting eQTL can be attributed to probes overlapping SNPs”.
In this paper, we aim to evaluate the effect of SNPs on eQTL studies using a human dataset and a mouse dataset by comparing the results of eQTL studies with and without the probes that contain SNPs. Hereafter, we define a probe ‘SNP-containing probe’ if its target sequence contains one or more SNPs and ‘SNP-free probe’ otherwise. Similarly, a probe set is ‘SNP-containing probe set’ if any of its probes is a SNP-containing probe and ‘SNP-free probe set’ otherwise.
Results
Identification of SNPs in probe targeted sequences
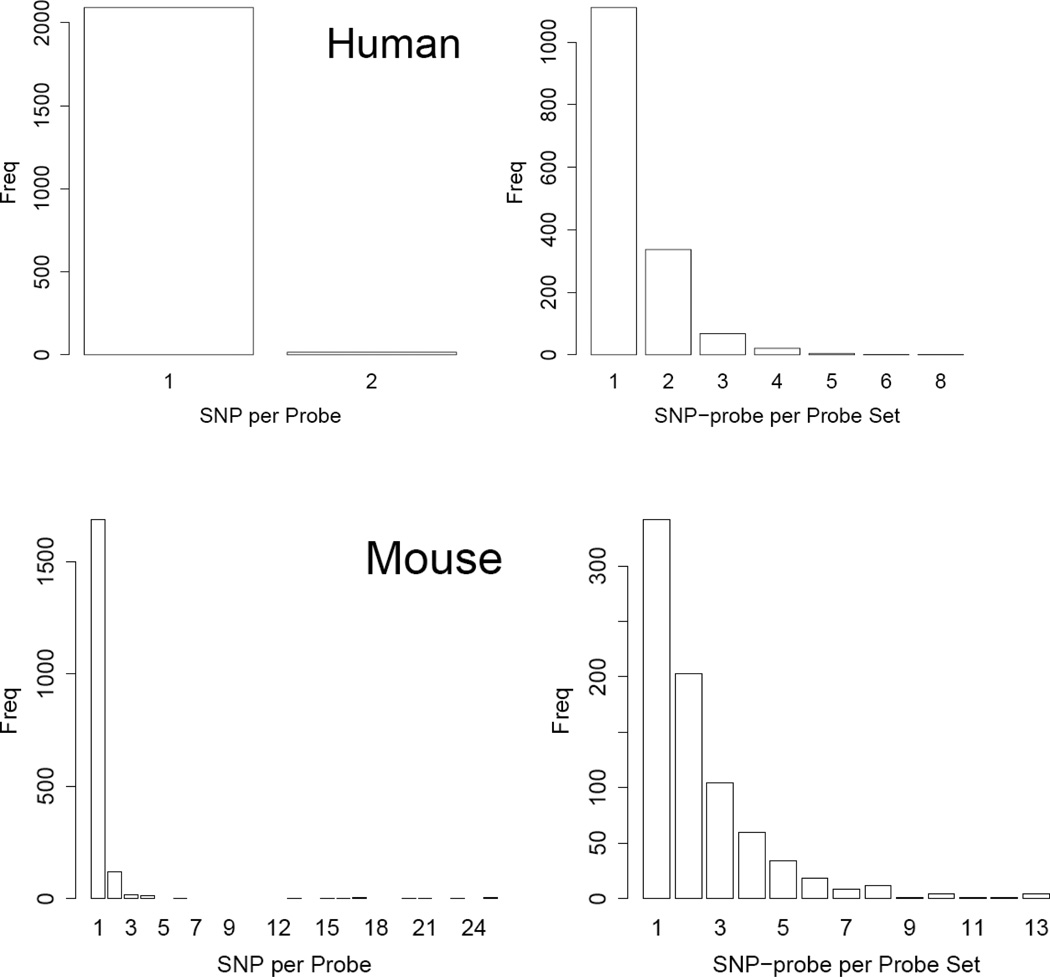
SNPs are the most common genetic variation (90%) in various genomes including human and mouse [1]. It is likely that a substantial number of probes on the microarrays used in the human and mouse eQTL studies overlap with genetic variation in populations. The Microarray lab of Molecular and Behavioral Neuroscience Institute at University of Michigan has compared human and mouse SNPs in the NCBI dbSNP database with the probe sequences on various Affymetrix GeneChips and identified the probes that overlap with any of the SNPs in the database (http://arrayanalysis.mbni.med.umich.edu/). We identified the SNPs in their list that were also found in the HapMap CEPH population and examined their overlap with probes on the Affymetrix Human Genome Focus Array, which was used in the human eQTL study using the CEPH population [8]. We found that 2107 (2.1%) of the total 98149 probes on the chip overlap with one or more SNPs present in the CEPH population. Most of these probes, 99.3%, overlap with just one SNP and only 0.7% of them overlap with two SNPs (Figure 1). At the probe set level, more than 17% (1543) of the 8793 probe sets on the chip overlap with one or more SNPs. Examining the 1543 SNP-containing probes sets showed that 71.6% of SNP-containing probe sets have just one SNP-containing probe (Figure 1) and 98.1% of the SNP-containing probe sets have one to three SNP-containing probes. Only a small number of probe sets (29) have more than three SNP-containing probes.
Figure 1.
Distributions of SNP-containing probes and SNP-containing probe sets on the Affymetrix arrays used in our study. The human data were generated using the Affymetrix Human Focus Arrays containing probes for 8,500 transcripts. The mouse data were generated using the Affymetrix Mouse U74Av2 microarrays, which contain 12488 probe sets. The SNP-containing probes of the human array in the studied population were established based on the HapMap Phase 3 data of CEPH population. The mouse SNP-containing probes were established based on the SNPs between the two parental inbred lines for the BXD RI population in the mouse SNP database. SNP-probe, SNP-containing probe
The same type of analysis was conducted for the BXD mouse data [16], where the Affymetrix U74Av2 chip was used. The U74Av2 chip contains 12488 probe sets, most of which (11820) have 16 probes. The BXD mouse eQTL population was originated from two inbred line (C57BL/6J and DBA/2J)[17], which have been sequenced. Comparing the SNPs between these two inbred lines and the probe sequences on the array revealed that 1854 probes harbor SNPs. As observed from the human dataset, most of these probes, 1689 (91.1%), contain just one SNP. A much smaller number, 120 (6.5%), have two SNPs and an even smaller number of probes have more than two SNPs. At the probe set level, 792 (6.3%), contain SNPs. Similarly to that in the human dataset, most of the SNP-containing probe sets have just one or two SNP-containing probes (Figure 1), 342 (43%) with one SNP-containing probe and 203 (26%) with two SNP-containing probes. A total of 104 and 143 probe sets have 3 and more than 3 SNP-containing probes, respectively.
SNP-containing probes affect the probe-level eQTL analysis
It has been shown that the SNP can affect the probe level analysis of gene expression and its effect is related to the position of the SNP on the probe [5,18]. However, it is not clear whether and how much it affects the probe level eQTL analysis. If the effect is large in probe level eQTL analysis, we would expect to see dramatic enrichment of cis-eQTL peaks from SNP-containing probes, especially from probes with SNPs near the center. We used the mouse data to test the effect of SNPs and SNP locations in probe level eQTL analysis for computational simplicity. For this analysis, we only considered the probes that contain one SNP for simplicity. If we find strong effects of one SNP, multiple SNP would likely to have even stronger effects. After RMA processing, the probe intensities (on log scale) were treated as quantitative traits and analyzed for QTL mapping as described in Materials and Methods. We considered a linkage peak as cis-eQTL if the peak falls within 10 Mb of the probes. The results showed that collectively only a very small proportions (2.85%) of the QTL peaks obtained from the SNP-free probes of the SNP-containing probe sets are cis eQTLs while a much larger proportion of the eQTL peaks (up to 30%) obtained from the SNP-containing probes in the same probe sets are cis-eQTL peaks, especially from the probes with SNPs in the center (Table 1). These results indicate that the SNP-containing probes tend to produce false positive cis-eQTLs in probe level mapping. Figure 2 shows an example of false positive cis-QTL as well as an example of false negative cis-QLT resulted from SNP-containing probes.
Table 1.
SNP position affects the number of cis-eQTL the probe level analysis.
| SNP position no SNP |
# of Probes 10805 |
Trans- QTLa 1637 |
Cis- eQTLs 48 |
Cis-eQTL Percentage 2.85% |
|---|---|---|---|---|
| 1 | 125 | 24 | 2 | 7.69% |
| 2 | 133 | 23 | 5 | 17.86% |
| 3 | 134 | 16 | 1 | 5.88% |
| 4 | 126 | 29 | 1 | 3.33% |
| 5 | 130 | 25 | 2 | 7.41% |
| 6 | 146 | 37 | 3 | 7.50% |
| 7 | 145 | 23 | 5 | 17.86% |
| 8 | 146 | 24 | 5 | 17.24% |
| 9 | 128 | 21 | 8 | 27.59% |
| 10 | 138 | 29 | 7 | 19.44% |
| 11 | 142 | 30 | 3 | 9.09% |
| 12 | 138 | 36 | 7 | 16.28% |
| 13 | 58 | 11 | 5 | 31.25% |
| Total | 1689 | 328 | 54 | 14.14% |
The mouse dataset was analyzed at probe level for the probe sets with only one SNP-containing probe. The SNP-free probes in these SNP-containing probe sets were used as negative control (no SNP row). The SNP position indicates the location of the SNP from the nearest end of the probe.
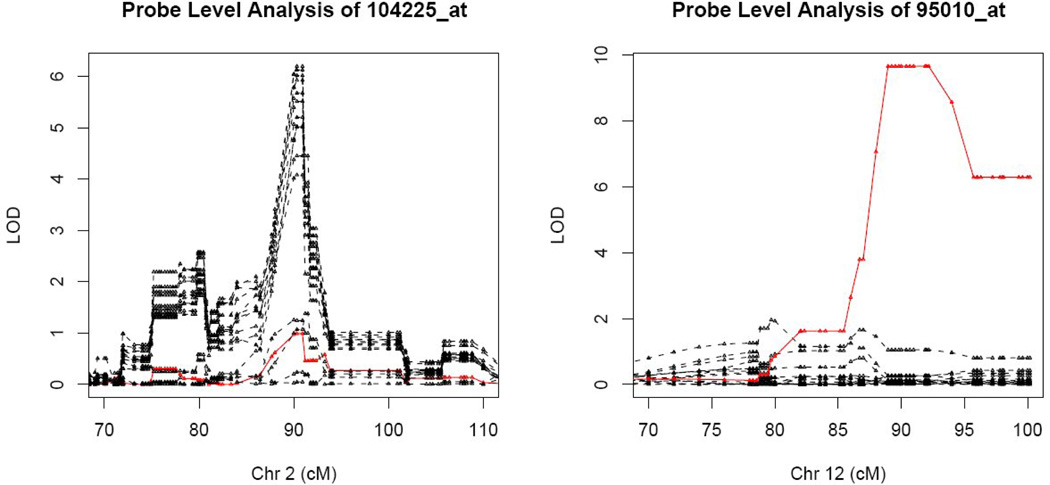
Figure 2.
Examples of false positive and false negative cis-QTLs caused by SNP-containing probes. The LOD scores are plotted at the regions surrounding the target gene locations in the genome for each probe in the probe set. The red solid and black dashed lines represent SNP-containing and SNP-free probes, respectively. LOD curves of probe set 104225_at show that 11 SNP-free probes have a cis-QTL at 90cM on chromosome 2 while the SNP-containing probe 104225_at1 does not have the QTL on that location, which illustrates the false negative caused by SNP-containing probe. LOD curves of probe set 95010_at show that SNP-containing probe 95010_at16 has a cis-QTL at 92cM on chromosome 12 while the SNP-free probes in the same probe set do not.
SNPs between probe and target sequences affect probe set summary scores
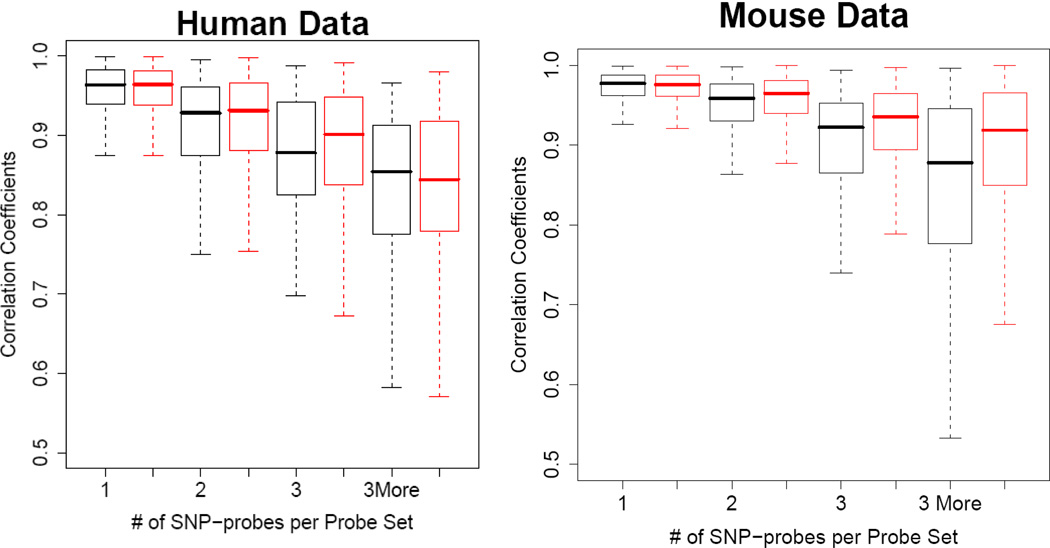
In almost all eQTL studies using Affymetrix GeneChips, the hybridization intensities of probes are first summarized for each probe set. The probe set summaries are then used for eQTL mapping. Therefore, it is important to evaluate the effect of SNPs on the probe set summaries. We examined the difference of the probe set summaries caused by filtering out the SNP-containing probes. After preprocessing using RMA based on filtered or unfiltered CDF files, we calculated the Pearson correlation coefficients of the probe set summaries for each probe set. Figure 3 illustrates the observed distributions for the correlation coefficients of the expression between the two types of probe set summaries. The results showed that the presence of a SNP in a probe set can change the expression summaries and the larger the number of SNP-containing probes, the greater the change. Due to the manner in which RMA normalizes and processes arrays the expression, values realized from the SNP-free probes could differ when realized with the filtered or unfiltered *.CDFs; however, the differences were minimal as expected. For the SNP-containing probe sets in the human data, the median correlation coefficients are around 0.96, 0.93, 0.88, and 0.85 for the probe sets with one, two, three, and more than 3 SNP-containing probes, respectively. For mouse, a similar reduction of coefficients was observed along with the increase of the number of SNP-containing probes. The median correlation coefficients are around 0.98, 0.96, 0.92, and 0.88 for the probe sets with one, two, three, and more than three SNP-containing probes, respectively.
Figure 3.
Box plots of Pearson correlation coefficients of probe set summaries. The coefficients were obtained from correlating the original probe set summaries with those after filtering out probes. The black plots are for correlations between the original summaries and those after filtering out the SNP-containing probes. The red plots are for correlations between the original summaries and those from random probe filtering. Correlation coefficients were calculated after normalizing all individuals in the population together.
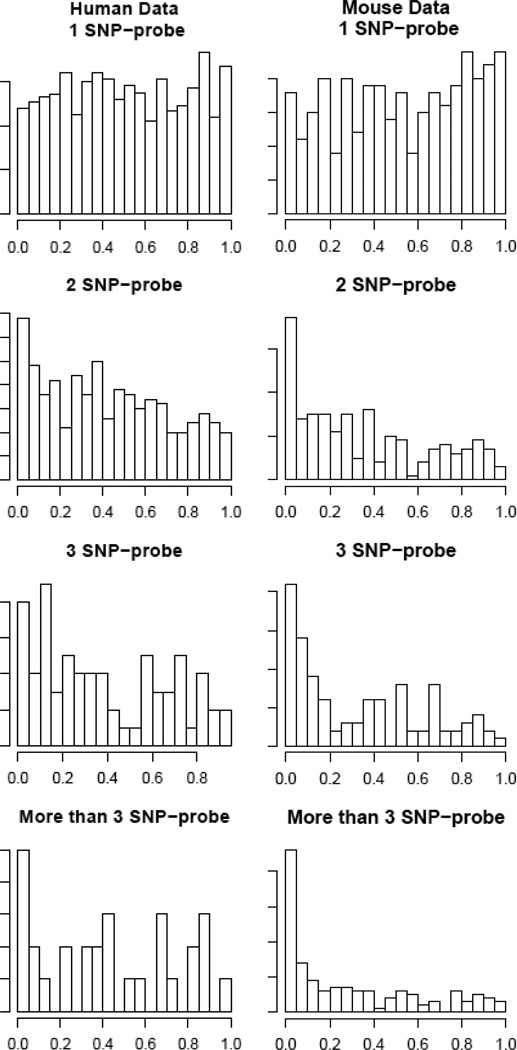
The reduction of the correlation coefficients could also come from the reduction of the number of probes in the probe sets. To evaluate the contribution of SNPs independent of the probe set size reduction, we randomly removed same number of probes from each SNP-containing probe set and calculated the correlation coefficients for 1000 times (red box plots in Figure 3). These coefficients are larger compared to the ones obtained from removing the SNP-containing probes (black box plots Figure 3) except the category of probe sets with more than three SNP-containing probes from human data. The lacking improvement of correlation in the random removing process could be due to the small number of probe sets in this category (29), which resulted in the unstable results. We also obtained p value for each SNP-containing probe removal based on the empirical distributions of the correlation coefficients obtained from randomly removing probes. If the SNPs have significant effect on the probe set summary, we would expect a left shifted distribution of the p-values, else a uniform one. We indeed observed left shifted distributions of p-values when there are more than one SNP-containing probes in a probe set although the proportion of excess small p values is relatively small (Figure 4). From these results we conclude that SNPs between probe and target sequences do affect probe set summaries.
Figure 4.
Histograms of p values obtained from testing the effect of removing SNP-containing probes on probe set summaries. The extra small p values compared with a uniform distribution indicate that in some probe sets correlation obtained from filtering out the SNP-containing probes is significantly weaker than that obtained from randomly filtering out equal number of probes. SNP-probe, SNP-containing probe.
SNPs between probe and target sequences affect probe set level eQTL analyses
Our goal in this study is to evaluate whether SNPs have an impact on the results of eQTL studies. If removing the SNP-containing probes had no effect, we would expect that the eQTLs identified in the filtered and unfiltered datasets largely agree with some allowance for the variability from the reduction of probe numbers and slightly different results from separate RMA preprocessing. The results from analyzing the human CEPH data showed that many of the eQTL peaks obtained after removing the SNP-containing probes agreed with those resulted from the original data for the SNP-containing probe sets (Table 2). In total, we found 145 eQTLs and 132 eQTLs for the 1543 SNP-containing probe sets before and after filtering, respectively. Among these eQTLs, 93 were common. When we examined the relationship of the agreement of results and the number of SNP-containing probes, we found that the discrepancy of linkage peaks increases as the number of SNP-containing probes increases. The probe sets with just one SNP-containing probe gave very similar linkage results (68.3% agreement) between filtered and unfiltered datasets (Table 2). For the group of probe sets with two and three SNP-containing probes, the overlap is reduced to 46.2% and 42.9% of those obtained without filtering, respectively. For the group of probe sets that have more than three SNP-containing probes, there is no overlap. A better measurement of agreement is positive agreement proportion (PAP) [19], which is defined as the proportion of agreed results among all the positives from both analyses. The PAP between eQTLs from the filtered and unfiltered data decreased from 70% to 46% as the number of the SNPs-containing probes increased from one to three (Table 2). These results indicate that the linkage mapping results are relatively robust against removing just one SNP-containing probe; however, when there is more than one SNP-containing probe, the linkage mapping result can change dramatically. Filtering out SNP-containing probes not only removed some of the eQTL peaks obtained from the unfiltered data but also produced some new eQTLs. For example, from the probe sets that contain just one SNP-containing probe in the human data, 39 original eQTLs were lost but 33 new eQTLs were gained after filtering out the SNP-containing probes (Table 2).
Table 2.
Comparisons between eQTL peaks identified before and after filtering out the SNP-containing probes.
| Human CEPH data | Mouse BXD data | |||||||
|---|---|---|---|---|---|---|---|---|
| # SNP-p per prs | 1 | 2 | 3 | >3 | 1 | 2 | 3 | >3 |
| # associated prs | 1105 | 339 | 70 | 29 | 342 | 203 | 104 | 143 |
| # overlapping QTLs | 84 | 6 | 3 | 0 | 59 | 27 | 6 | 13 |
| # QTLs w/o filtering (overlapping) | 123 (68.3%) | 13 (46.2%) | 7 (42.9%) | 2 (0.0%) | 94 (62.8%) | 50 (54%) | 20 (30%) | 43 (30.2%) |
| # QTLs w/filtering (overlapping) | 117 (71.8%) | 9 (66.7%) | 6 (50.0%) | 0 (NA) | 76 (77.6%) | 46 (58.7%) | 14 (42.9%) | 30 (43.3%) |
| PAP | 70.0% | 54.55% | 46.15% | NA | 69.4% | 56.3% | 35.3% | 35.6% |
The significance level for eQTLs is genome wide 0.05. The filtering was conducted before data preprocessing. All other steps for the filtered data were exactly the same as for the unfiltered data. Overlapping QTLs were defined as the QTLs located within 10cM. SNP-p, SNP-containing probe; prs, probe set. PAP, positive agreement proportion defined as the proportion of overlapping positives among all positives from both analyses.
Similar results were obtained from the mouse data. We identified 207 and 166 eQTL peaks from the 792 SNP-containing probe sets before and after filtering the SNP-probes, respectively. Among the eQTL peaks identified before filtering, 105 (50.7%) were also identified after filtering. The overlapping eQTL peaks also decrease as the number of the SNP-containing probes in the probe sets increases. For the probe sets with just one SNP-containing probe, the overlap of the eQTL is about 63% of those obtained without filtering. For the probe sets with two and three SNP-containing probes, the overlap is about 54% and 30%, respectively. These results are consistent for various significance levels (e.g. LOD 2, 4 etc.). The proportion of the positive agreement also decreased from 69% to 35% with the number of the SNP-containing probes in a probe set increased from one to three (Table 2). Notice that PAP of probe sets with three SNP-containing probes is slightly lower than that of probe sets with more than three SNP-containing probes. This could be due to the random fluctuation with small sample size (only 6 and 13 overlapping QTLs in these two categories).
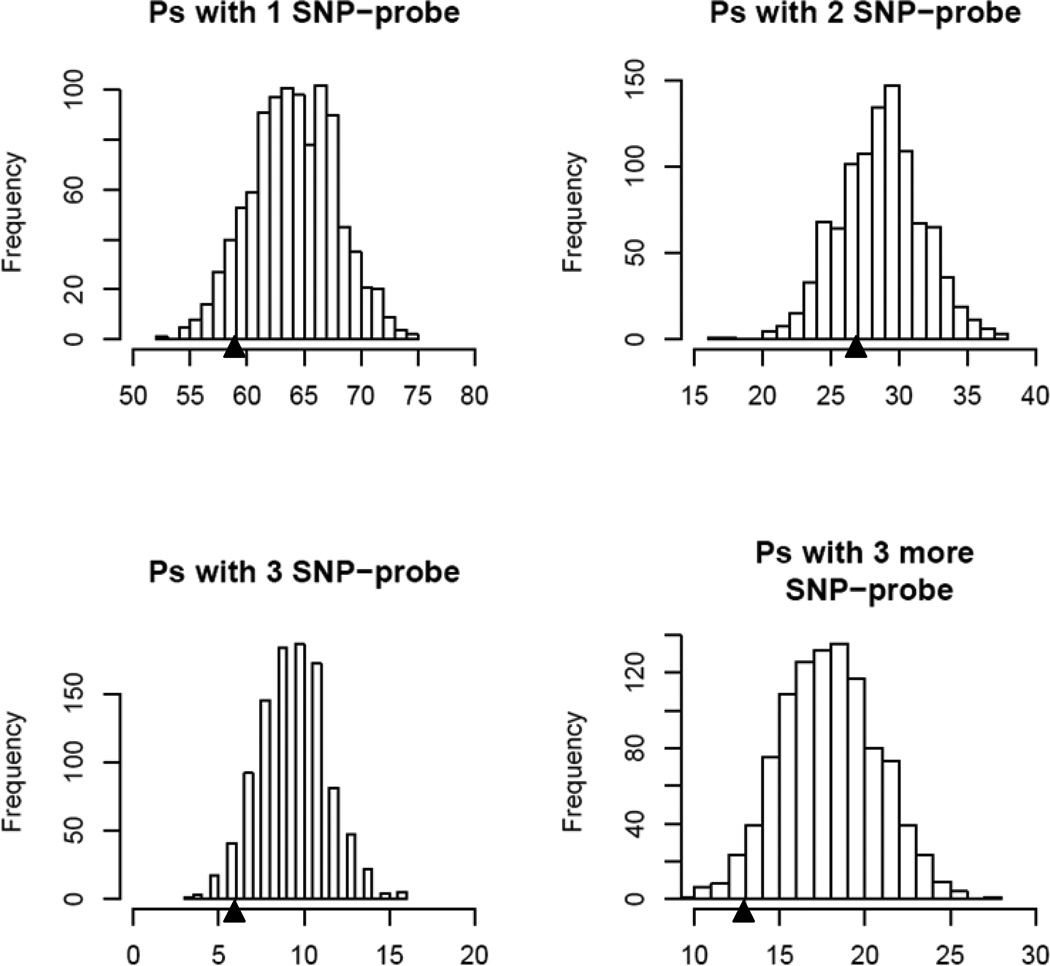
To examine whether the reduction of overlap is simply due to the reduction of probe set size we conducted a resampling study for the mouse data. Similar to what we did in assessing the effect of the SNP on probe set summaries, we randomly removed equal number of SNP-free probes from the SNP-containing probe sets 1000 times and examined the overlap of linkage peaks between with and without filtering probes each time. The distribution of the overlapping eQTL is shown in Figure 5. For the probe sets with just one SNP-containing probe, the number of overlapping eQTL peaks between with and without filtering is 59 (Table 2), which is considerably smaller than most of those from the resampling processes. This result indicates that removing the SNP-containing probes have substantial effect on the eQTL results beyond the effect of probe set size reduction. The empirical p-values are 0.095, 0.296, 0.062, 0.038 for one, two, three and more than three SNP-containing probes, respectively (Figure 5). To understand the large p value obtained from the probe sets with two SNP-containing probes, we randomly picked 16 probe sets from this category and compared the microarray probe sequences with the corresponding sequences of the two mouse strains. We found that only 7 probe sets with the two SNP-containing probes matching the same strain, while 5 probe sets with the two SNP-containing probes matching neither of the two strains, one probe set with the SNP-containing probes matching the two opposite strains, and 3 probe sets with one SNP-containing probe matching neither of the two strains. These findings indicate that only about half of the probe sets in this category have effects as expected for the SNP-containing probe sets with two SNP-containing probes.
Figure 5.
Histograms for the numbers of overlapping eQTLs between randomly removing probes and the original mouse data. Probes were randomly filtered out of the SNP-containing probe sets to generate new datasets. The same preprocess and QTL mapping methods were applied to the new datasets. The eQTLs from the new datasets were compared to the eQTLs obtained from the same probe sets in the original dataset. The triangles point to the number of overlapping eQTLs between the original dataset and the dataset with all the SNP-containing probes filtered out. Ps, probe sets; SNP-probe, SNP-containing probe
SNPs affect both cis- and trans-eQTLs
Most previous investigations on the effect of SNPs in eQTL studies have focused only on the cis-eQTL peaks [14,20]. Our results above included both cis- and trans-eQTLs. To examine the two types of eQTL peaks separately using the mouse data, we first established the baseline by examining the SNP-free probe sets for the similarity of cis- and trans-eQTL peaks before and after the filtering procedure. Although no probe was removed from these probe sets, the removing of the SNP-containing probes from other probe sets potentially affects the RMA preprocessing and normalization results; therefore, affects the SNP-free probe sets too. Our analysis generated 2346 tran-QTLs and 155 cis-QTLs in common between the two analyses (Table 3A). Only about 10 unique trans-eQTLs and one or two unique cis-eQTLs was found in each analysis. For the SNP-containing probes sets, the differences are much greater. For the cis-eQTLs, there are 15 common to both analyses but 3 and 4 unique to the without and with filtering, respectively. In addition, one cis-eQTL became trans-eQTL and one trans-eQTL became cis-eQTL (Table 3A). Overall, the gain and loss of cis-eQTLs is about the same. For the trans-eQTLs, the difference is even more dramatic. There are 84 trans-eQTLs in common between the two analyses; however, there are 50 and 86 trans-eQTLs unique to the without and with filtering, respectively. The overall gain of new trans-eQTLs after filtering is much greater than the loss. These results showed that filtering out SNP-containing probes not only affect the cis-eQTLs but also the trans-eQTL, which indicates that SNPs can cause both false positive cis- and trans-eQTLs. When we randomly removed same number of probes from SNP-containing probe sets and conducted eQTL-mapping, we found that a much larger proportion of trans-eQTL are in common with those obtained from the original data (Table 3B). This result indicates that the low overlap of trans-eQTLs is indeed partially due to the SNP-probes. For the cis-eQTL, we actually obtained a smaller overlap from the resampling procedure, the cause of which is not clear.
Table 3.
Comparing the cis- and trans-eQTLs between before and after filtering out the SNP-containing probes from the mouse data.
| A | ||||
|---|---|---|---|---|
| Trans-eQTLs w/o filter |
Cis-eQTLs w/o filter |
No eQTL | ||
| SNP-free probe sets | Trans-eQTLs w/filter | 2346 | 0 | 10 |
| Cis-eQTL w/filter | 0 | 155 | 2 | |
| No eQTL | 13 | 1 | 0 | |
| SNP-containing probe sets | Trans-eQTL w/filter | 84 | 1 | 86 |
| Cis-eQTL w/filter | 1 | 15 | 4 | |
| No eQTL | 50 | 3 | 0 | |
| B | ||||
|---|---|---|---|---|
| Trans-eQTLs w/o filter |
Cis-eQTLs w/o filter |
No eQTL | ||
| Resampling the SNP-containing probe sets | Trans-eQTL | 160.3 | 9.8 | 53.76 |
| Cis-eQTL | 8.883 | 10.2 | 1.9 | |
| No eQTL | 55.9 | 2.2 | 0 | |
A, comparisons when the SNP-containing probes were removed from the SNP-containing probe sets. B, comparisons when equal numbers of probes were randomly removed from the SNP-containing probe sets. The resampling process was repeated for 1000 times and the averages are shown in the table 3B. The significance level used for the eQTLs here is LOD 3. The common eQTL peaks are defined as within 10 cM.
Discussion
One important piece of evidence investigators have used in the past and continue to use to indicate the effect of SNP in eQTL studies is the enrichment of cis-acting eQTL peaks from probe sets with SNPs. Our mouse study showed that about 4% of the eQTLs obtained from the SNP-free probe sets were cis-eQTLs while 13% of the eQTLs obtained from the SNP-containing probe sets were cis-eQTLs. However, as Dross et. al., [14] pointed out, the excess of cis-eQTL from the SNP-containing probe sets could be due to the fact that SNP-containing probe sets tend to be localized in none identical by decent (IBD) regions, where sequence is highly polymorphic between the two mouse strains. The SNPs in the nearby region instead of the SNPs on the probes could be the cause of cis-acting peaks. To distinguish the effect of these two types of SNPs, we compared the cis-acting eQTLs before and after removing the SNP-probes. Our results showed that only 4 out of the 19 cis-QTLs disappeared after filtering SNP-containing probes. In addition, 5 new cis-eQTLs were gained after filtering in the mouse dataset. Albert et. al.,[21] flagged 25 of the 70 cisB6 eQTL (36%) as potentially false cis eQTLs, which is higher than our finding that 20% of the cis-QTLs resulted from the original data analysis without filtering were potential false cis-QTLs. However, we showed that false positive cis-eQTLs is just one aspect of the SNP effect. A substantial proportion of trans-eQTLs (38%) was also lost after filtering out the SNP-containing probes. On the other hand, both new cis-eQTL’s and trans-eQTLs were also gained after filtering. Interestingly, we observed even greater SNP effect on the trans-eQTLs. The positive agreement proportion for the trans-eQTLs is only 40% while that for the cis-eQTL is around 65%. In addition, a lot more new trans-eQTLs were gained after removing the SNP-containing probes compared with the ones lost. This could be explained by the inflated residual signal variation associated with SNP-containing probes in testing for the trans-eQTLs, which results in fewer significant trans-eQTL peaks. After removing the SNP-containing probes, the residual variance is reduced and more trans-eQTL peaks become significant.
Multiple factors contribute to the difficulty in comparing results regarding the effect of SNPs in eQTL mapping. One problem is that the definition of the cis-acting eQTL is inconsistent across studies. Investigators use Mb or cM to define a location for a cis-eQTL and the window size can range for 2–20 [8,11,14,22]. Depending on the sample size and power of the study, it might be more appropriate to chose different window size for the cis-acting eQTL, such as confident interval of the eQTL locations. In addition, eQTL studies are the high-dimensionality in nature. Both the number of the markers and the number of traits are large. Although most studies can establish genome-wide significant level for a single trait, it is hard to control the type I error rate for all traits without losing much power. The low power results in high false negative rate and thus we could easily underestimate the effects of SNPs. In our study, we did not adjust for multiple testing at the gene dimension to avoid extreme low power. We did try various significant levels and found that our results were relatively consistent.
Different microarray data preprocessing methods [23] can also cause difference in eQTL studies. Due to the robust nature against the outlier probes from probe sets, RMA likely minimizes the effect of SNP effects. Therefore, we choose the RMA preprocessing in this study. We also tried MAS 5.0 [3] for preprocessing. The results from MAS5 were similar (results not shown) to those obtained from RMA in respect to the decrease of overlapping QTLs with the increase of SNP-containing probes in a probe set. However, removing the SNP-containing probes resulted in much less overlapping eQTL peaks when MAS5.0 was used to preprocess data. This indicates that the choice of preprocessing methods affects the robustness of the analysis against SNP-containing probes. SNP probes can be considered as noise in the eQTL analysis; therefore, down-weighting or filtering out those probes provides more accurate measurement and improves the analysis.
Our probe-level eQTL analyses showed that SNP-probes cause a dramatic increase in cis-eQTLs compared with the SNP-free probes in the same probe set. These results indicate that the SNP-containing probes definitely need to be excluded if eQTL mapping is to be conducted at probe level. Affymetrix expression GeneChips use probe sets that contain more than 10 probes to interrogate the expression of one gene. As expected, the summary of a probe set is less sensitive to SNPs unless there are multiple SNP-probes in a probe set. However, we showed that the eQTL mapping results based on probe set summaries can be significantly affected by the presence of just one SNP-containing probe (Figure 5). Therefore, removing the SNP-containing probes is a worthwhile practice in eQTL studies using Affymetrix GeneChips. For microarray platforms with longer probes, the SNP effect on probe level eQTL analysis is likely less significant. It has been shown that the SNP effect on eQTL result is minor in a study using a 60-mer long oligo arrays [14]. It is not clear how many SNPs it takes to affect the performance of long oligo probes.
We showed that SNPs affect both probe set summaries and eQTL results based on the probe set summaries, especially when the SNP-containing probe sets have more than one SNP-containing probes. For both the human CEPH data and the mouse data, we only considered the SNPs found in the population or between the two parental lines, C57BL/6J (B) and DBA/2J (D) in the mouse case. However, there is still the possibility that the probe sequence does not match any of the alleles as we have shown using a handful of probe sets with two SNP-containing probes from the mouse data. In these cases, having the sequence polymorphism between probe and targets will not affect the eQTL results because the SNP affects the two alleles equally. In addition, it is also possible that some of the SNPs are fixed in the mouse RI lines due to the lack of representation of the second parent in some regions. Removing the probes that contain these SNPs are not necessary either. All these issues deserve further investigation to fully characterize the true effect of sequence polymorphism between probe and targets in eQTL studies. However, removing specific types of SNP-containing probes will also greatly complicate the procedure of probe-removing before eQTL mapping. Furthermore, SNP is only one type of sequence polymorphisms, other types of sequence polymorphisms are worth investigate too. Only after we remove the contributions of all these sources that affect probe hybridization, we can truly be confident that we are mapping the factors that affect the gene expression level.
Materials and Methods
Human eQTL dataset - CEPH Utah families data
The dataset consists of 14 three-generation Centre d’Etude du Polymorphisme Humain (CEPH) Utah pedigrees [24] with 194 individuals. The gene expression profile was measured on immortalized B cells using the Affymetrix Human Focus Arrays that contain probes for 8,500 transcripts. Only 82 individuals were profiled with two microarrays while the rest were profiled with one microarray each[8]. A total of 2,819 autosomal SNP markers were genotyped by The SNP Consortium (http://snp.cshl.org/linkage_maps). The average distance between two adjacent makers is 0.97 megabases (Mb) with median 0.06 Mb. Only 13 inter-marker distances were greater that 10 Mb.
Mouse eQTL dataset - mouse BXD bone marrow data
The mouse dataset consists of 22 BXD recombinant inbred lines [16]. Bone marrow cells were collected from the femurs and tibiae of three mice. The samples were then pooled within each inbred line and hybridized onto two Affymetrix Mouse U74Av2 microarrays, which contain 12488 probe sets consisting of 197993 probes. The marker genotypes are available for a total of 7636 informative markers that differ between the parental strains, C57BL/6J (B6) and DBA/2J (DBA). Marker genotypes were available at http://www.webQTL.com. A selected subset of 2325 markers that includes all markers with unique strain distribution patterns were used in the linkage analysis ftp://atlas.utmem.edu/public/BXD_WebQTL_Genotypes_June05.txt.
SNP filtering
We used Probefilter (version 1.4.0) (http://arrayanalysis.mbni.med.umich.edu/MBNIUM.html) to create customized Affymetrix probe set definitions (CDFs) with probes that contain SNPs removed. Probefilter provides the list of SNPs that overlaps the probe sequences on the Human Focus Array. Unfortunate, their list contains all SNPs existing in the dbSNP database (ftp://ftp.ncbi.nih.gov/snp/organisms/). For our purpose of removing only the SNPs existing in the human CEPH population, we identified the SNPs present in the HapMap Phase 3 data of CEPH population (NCBI build 36, dbSNP b126 at www.hapmap.org). All Probes on the array that overlap one or more SNPs in the HapMap Phase 3 data of CEPH population were identified as a SNP-containing probes. For the mouse data, the SNPs between the two parental inbred lines (B6 and DBA) of the BXD RI lines were used to identify SNP-containing probes. For both data sets, the SNP-containing probes were eliminated from the Affymetrix CDF files before data preprocessing. Probe sets containing less than three SNP-free probes were removed from the analysis due to unstable probe set summaries [25].
Microarray image processing
The Affymetrix CEL files were preprocessed using RMA method [26] as implemented in the affy package in Bioconductor (http://www.bioconductor.org) with default settings. The .cel files were processed separately with or without filtering the SNP-containing probes using different versions of CDF files.
Correlation coefficients analyses of probe set summary scores
For each probe set, the Pearson correlation coefficient (robs) was calculated between the probe set summary scores obtained before and after filtering the SNP-containing probes based on all individuals in the population. To establish an empirical null distribution for these correlation coefficients, we constructed a new filtered datasets by randomly removing the same number of probes from each SNP-containing probe set. The new filtered dataset was then processed using the RMA method with the same setting and a correlation coefficient (r*) was calculated between this new dataset and the original unfiltered dataset for each probe set. This procedure was repeated 1000 times to obtain an empirical distribution for correlation coefficients from each probe set. The observed correlation coefficients were compared to the empirical distributions from corresponding probe sets to obtain the p values (proportion of r*< robs).
Linkage analysis
For linkage analysis, the probe set summary scores from technical replicates were averaged for each individual after image processing. For the human data, we used MERLIN [27] to detect problematic genotypes. All genotype errors were set to missing. Linkage analyses were conducted using the variance components approach [28] and the summary score of each probe set was considered as a separate continuous phenotype. We skipped those markers that had Mendelian inconsistencies within a pedigree for calculation of the likelihood. LOD scores, Chi-square statistics, and their associated nominal p values were reported. Nominal p values less than or equal to 0.00005 were considered to be genome-wide significant at 0.05 according to the Ohrenstein-Uhlenbeck method [29].
For the mouse data, the average probe set intensities of the two technical replicates (two chips) after image processing were treated as quantitative trait. Haley-Knott regression implemented in R/qtl was used for the QTL mapping [30,31]. Pseudomarkers were generated every 2 centimorgan (cM) and genome-wide significance level was established using 1000 permutations.
We also performed probe level eQTL analysis for mouse data, the intensity of each probe was treated as a quantitative trait after background correction, normalization and log transformation. The same QTL mapping methods described above for mouse probe set level data were applied.
Comparison of linkage results
The linkage results were compared at the concordance and discordance of eQTL peaks at various significance levels. QTL peaks located within 10 Mb of each other were considered as the same peaks. This criterion was chosen based on what has been used in other studies. Mapping was conducted based on genetic distance cM. The genomic positions (Mb) of pseudo-markers were extrapolated using linear relation between cM and Mb between two adjacent markers. Extrapolation was based on the fact that the average genomic recombination rate is about 1.1cM per Mb in the human genome, although the rate can range from 0.01 cM to 130 cM per Mb [32]. For mouse, the average recombination rate per Mb was also based on the average genetic distance per Mb DNA calculated from the total mouse genome genetic distance, 1,355~1, 450 cM, and the total physical distance, 2,500 Mb [33–36].
Acknowledgement
We thank Guiming Gao and Nianjun Liu for fruitful discussions. The authors acknowledge the financial support from the National Institutes of Health U54CA100949, R01NS043530, and R01ES012933.
Reference List
- 1.Sachidanandam R, Weissman D, Schmidt SC, Kakol JM, Stein LD, Marth G, et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature. 2001;409:928–933. doi: 10.1038/35057149. [DOI] [PubMed] [Google Scholar]
- 2.Yamashita S, Nomoto T, Ohta T, Ohki M, Sugimura T, Ushijima T. Differential expression of genes related to levels of mucosal cell proliferation among multiple rat strains by using oligonucleotide microarrays. Mamm Genome. 2003;14:845–852. doi: 10.1007/s00335-003-2299-3. [DOI] [PubMed] [Google Scholar]
- 3.Affymetix. Statistical Algorithms Description Document. Technical Report. 2002 [Google Scholar]
- 4.Affymetrix. Single Nuceotide Polymorphism Marker Selection and Assay Validation. 2007 Affymetrix. Technical Notes. Internet Communication. [Google Scholar]
- 5.Rostoks N, Borevitz JO, Hedley PE, Russell J, Mudie S, Morris J, et al. Single-feature polymorphism discovery in the barley transcriptome. Genome Biology. 2005;6 doi: 10.1186/gb-2005-6-6-r54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Borevitz JO, Liang D, Plouffe D, Chang HS, Zhu T, Weigel D, et al. Large-scale identification of single-feature polymorphisms in complex genomes. Genome Res. 2003;13:513–523. doi: 10.1101/gr.541303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Monks SA, Leonardson A, Zhu H, Cundiff P, Pietrusiak P, Edwards S, et al. Genetic inheritance of gene expression in human cell lines. American Journal of Human Genetics. 2004;75:1094–1105. doi: 10.1086/426461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Morley M, Molony CM, Weber TM, Devlin JL, Ewens KG, Spielman RS, et al. Genetic analysis of genome-wide variation in human gene expression. Nature. 2004;430:743–747. doi: 10.1038/nature02797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Cheung VG, Spielman RS, Ewens KG, Weber TM, Morley M, Burdick JT. Mapping determinants of human gene expression by regional and genome-wide association. Nature. 2005;437:1365–1369. doi: 10.1038/nature04244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Greenwood TA, Cadman PE, Stridsberg M, Nguyen S, Taupenot L, Schork NJ, et al. Genome-wide linkage analysis of chromogranin B expression in the CEPH pedigrees: implications for exocytotic sympathochromaffin secretion in humans. Physiological Genomics. 2004;18:119–127. doi: 10.1152/physiolgenomics.00104.2003. [DOI] [PubMed] [Google Scholar]
- 11.Stranger BE, Forrest MS, Clark AG, Minichiello MJ, Deutsch S, Lyle R, et al. Genome-wide associations of gene expression variation in humans. PLoS Genet. 2005;1:e78. doi: 10.1371/journal.pgen.0010078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Deutsch S, Lyle R, Dermitzakis ET, Subrahmanyan L, Gehrig C, Parand L, et al. Gene expression variation and expression quantitative trait mapping of human chromosome 21 genes. Human Molecular Genetics. 2005;14:3741–3749. doi: 10.1093/hmg/ddi404. [DOI] [PubMed] [Google Scholar]
- 13.Hubner N, Yagil C, Yagil Y. Novel integrative approaches to the identification of candidate genes in hypertension. Hypertension. 2006;47:1–5. doi: 10.1161/01.HYP.0000197951.82190.c4. [DOI] [PubMed] [Google Scholar]
- 14.Doss S, Schadt EE, Drake TA, Lusis AJ. Cis-acting expression quantitative trait loci in mice. Genome Res. 2005;15:681–691. doi: 10.1101/gr.3216905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hubner N, Wallace CA, Zimdahl H, Petretto E, Schulz H, Maciver F, et al. Integrated transcriptional profiling and linkage analysis for identification of genes underlying disease. Nat Genet. 2005;37:243–253. doi: 10.1038/ng1522. [DOI] [PubMed] [Google Scholar]
- 16.Bystrykh L, Weersing E, Dontje B, Sutton S, Pletcher MT, Wiltshire T, et al. Uncovering regulatory pathways that affect hematopoietic stem cell function using 'genetical genomics'. Nat Genet. 2005;37:225–232. doi: 10.1038/ng1497. [DOI] [PubMed] [Google Scholar]
- 17.Potter M, Mock B. Mouse Genetics - Concepts and Applications - Silver,Lm. Science. 1995;270:1692–1693. [Google Scholar]
- 18.Gresham D, Ruderfer DM, Pratt SC, Schacherer J, Dunham MJ, Botstein D, et al. Genome-wide detection of polymorphisms at nucleotide resolution with a single DNA microarray. Science. 2006;311:1932–1936. doi: 10.1126/science.1123726. [DOI] [PubMed] [Google Scholar]
- 19.Spitzer RL, Fleiss JL. A re-analysis of the reliability of psychiatric diagnosis. The British journal of psychiatry : the journal of mental science. 1974;125:341–347. doi: 10.1192/bjp.125.4.341. [DOI] [PubMed] [Google Scholar]
- 20.Drake TA, Schadt EE, Lusis AJ. Integrating genetic and gene expression data: application to cardiovascular and metabolic traits in mice. Mammalian Genome. 2006;17:466–479. doi: 10.1007/s00335-005-0175-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Alberts R, Terpstra P, Li Y, Breitling R, Nap JP, Jansen RC. Sequence polymorphisms cause many false cis eQTLs. PLoS ONE. 2007;2:e622. doi: 10.1371/journal.pone.0000622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yvert G, Brem RB, Whittle J, Akey JM, Foss E, Smith EN, et al. Trans-acting regulatory variation in Saccharomyces cerevisiae and the role of transcription factors. Nature Genetics. 2003;35:57–64. doi: 10.1038/ng1222. [DOI] [PubMed] [Google Scholar]
- 23.Lam A, Schouten M, Aulchenko Y, Haley C, de Koning DJ. Rapid and robust association mapping of expression quantitative trait loci. BMC Proceedings. 2007;1:S144. doi: 10.1186/1753-6561-1-s1-s144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dausset J, Cann H, Cohen D, Lathrop M, Lalouel JM, White R. Centre d'etude du polymorphisme humain (CEPH): collaborative genetic mapping of the human genome. Genomics. 1990;6:575–577. doi: 10.1016/0888-7543(90)90491-c. [DOI] [PubMed] [Google Scholar]
- 25.Dai MH, Wang PL, Boyd AD, Kostov G, Athey B, Jones EG, et al. Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Research. 2005;33 doi: 10.1093/nar/gni179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003;4:249–264. doi: 10.1093/biostatistics/4.2.249. [DOI] [PubMed] [Google Scholar]
- 27.Abecasis GR, Cherny SS, Cookson WO, Cardon LR. Merlin--rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet. 2002;30:97–101. doi: 10.1038/ng786. [DOI] [PubMed] [Google Scholar]
- 28.Amos CI. Robust variance-components approach for assessing genetic linkage in pedigrees. Am J Hum Genet. 1994;54:535–543. [PMC free article] [PubMed] [Google Scholar]
- 29.Lander E, Kruglyak L. Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet. 1995;11:241–247. doi: 10.1038/ng1195-241. [DOI] [PubMed] [Google Scholar]
- 30.Haley CS, Knott SA. A Simple Regression Method for Mapping Quantitative Trait Loci in Line Crosses Using Flanking Markers. Heredity. 1992;69:315–324. doi: 10.1038/hdy.1992.131. [DOI] [PubMed] [Google Scholar]
- 31.Broman KW, Wu H, Sen S, Churchill GA. R/qtl: QTL mapping in experimental crosses. Bioinformatics. 2003;19:889–890. doi: 10.1093/bioinformatics/btg112. [DOI] [PubMed] [Google Scholar]
- 32.Deonier RC, Tavaré S, Waterman MS. Computational genome analysis : an introduction. New York: Springer; 2005. [Google Scholar]
- 33.Rowe LB, Barter ME, Kelmenson JA, Eppig JT. The comprehensive mouse radiation hybrid map densely cross-referenced to the recombination map: a tool to support the sequence assemblies. Genome Res. 2003;13:122–133. doi: 10.1101/gr.858103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Copeland NG, Jenkins NA, Gilbert DJ, Eppig JT, Maltais LJ, Miller JC, et al. A genetic linkage map of the mouse: current applications and future prospects. Science. 1993;262:57–66. doi: 10.1126/science.8211130. [DOI] [PubMed] [Google Scholar]
- 35.Dietrich WF, Miller J, Steen R, Merchant MA, Damron-Boles D, Husain Z, et al. A comprehensive genetic map of the mouse genome. Nature. 1996;380:149–152. doi: 10.1038/380149a0. [DOI] [PubMed] [Google Scholar]
- 36.Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, et al. Initial sequencing and comparative analysis of the mouse genome. Nature. 2002;420:520–562. doi: 10.1038/nature01262. [DOI] [PubMed] [Google Scholar]