Abstract
The mouths of three human infants were examined from birth to age 2 years to detect colonization of Actinomyces naeslundii genospecies 1 and 2. These bacteria did not colonize until after tooth eruption. The diversity of posteruption isolates was determined by ribotyping. Using immunoblotting and enzyme-linked immunosorbent assay, we determined the reactivity of secretory immunoglobulin A (SIgA) antibodies in saliva samples collected from each infant before and after colonization against cell wall proteins from their own A. naeslundii strains and carbohydrates from standard A. naeslundii genospecies 1 and 2 strains. A. naeslundii genospecies 1 and 2 carbohydrate-reactive SIgA antibodies were not detected in any saliva sample. However, SIgA antibodies reactive with cell wall proteins were present in saliva before these bacteria colonized the mouth. These antibodies could be almost completely removed by absorption with A. odontolyticus, a species known to colonize the human mouth shortly after birth. However, after colonization by A. naeslundii genospecies 1 and 2, specific antibodies were induced that could not be removed by absorption with A. odontolyticus. Cluster analysis of the patterns of reactivity of postcolonization salivary antibodies from each infant with antigens from their own strains showed that not only could these antibodies discriminate among strains but antibodies in saliva samples collected at different times showed different reactivity patterns. Overall, these data suggest that, although much of the salivary SIgA antibodies reactive with A. naeslundii genospecies 1 and 2 are directed against genus-specific or more broadly cross-reactive antigens, species, genospecies, and possibly strain-specific antibodies are induced in response to colonization.
Secretory immunoglobulin A (SIgA) is considered to be the principal mediator of host defense at mucosal surfaces. Consistent with the large surface area of mucosae requiring protection and the fact that mucosal surfaces are open systems, more IgA is synthesized than all of the other immunoglobulin isotypes combined (22). The stimulus for IgA synthesis at mucosal surfaces appears to be colonization of these surfaces by commensal bacteria bearing polyclonal mitogens such as lipopolysaccharide (23) and perhaps lipoteichoic acid, since germfree mammals have underdeveloped mucosa-associated lymphoid tissues and lack SIgA in their secretions (9). Although SIgA is thought to act to exclude extrinsic pathogenic microorganisms, it appears to be without effect on commensal bacteria, since these microbes colonize and persist on mucosal and tooth surfaces despite its presence (6). The reasons for this persistence are unknown, although immune tolerance and antigenic variation have been proposed.
The response of the host to bacterial colonization of the mouth may provide a degree of host immune protection if these autochthonous organisms carry antigens that cross-react with antigens of pathogens that are important in virulence. Such antigens could be proteins involved in the adherence of pathogens to tissues (20). Conversely, tolerance of the host to such antigens that contribute to the virulence of pathogens could compromise immune protection. In addition, commensal gram-positive bacteria, in particular lactobacilli and oral streptococci, have been proposed as viable vectors of protective antigens in vaccines (1, 20, 30). In order for such vaccines to be effective, it is important that the organisms colonize the host for a sufficient period of time and stimulate a protective immune response. At present, testing the effectiveness of these vaccine strains has been limited to animals; however, it is proposed that such vaccines could be valuable in providing protection in humans. If this is the case, understanding the development of infants' salivary SIgA responses to bacteria colonizing the mouth could assist in determining the optimal time for oral immunization to promote both the persistence of vaccine strains and also a significant immune response. The human oral cavity with easy access, convenient surfaces, and saliva containing SIgA provides an ideal habitat to study the nature of mucosal immune responses.
However, in common with other areas supporting a commensal microbiota, the study of the generation and specificity of the immune response in the mouth is not without its difficulties. Among the most significant of these is the variation of strains of species of commensal bacteria during colonization, introducing the possibility of antigenic variation or drift of the colonizing species over time. We (13) and others (17, 18) have confirmed that strains of commensal bacteria colonizing oral mucosal surfaces demonstrate extensive diversity and, in particular, the streptococci show clonal replacement during colonization (13, 17). In addition, in studies in early infancy the volumes of saliva that can be obtained are small, and sensitive methods are necessary to demonstrate the amount and specificity of SIgA antibody.
The problem of potential variation of strains during colonization cannot be controlled but genetic typing of the predominant strains can give information on their stability during longitudinal analysis of the immune response. Moreover, storage of these isolates provides strains from individual infants to measure both the magnitude and the specificity of the response of individual infants to strains colonizing their mouths.
In longitudinal studies of human infants with standard strains of bacteria as antigen, we have shown that the salivary SIgA immune response to Actinomyces naeslundii genospecies 1 and 2 (7) and to selected species of viridans streptococci (8) is limited. Also, these commensal bacteria induce an SIgA antibody response in saliva with changing specificity that peaks at 6 months of age and wanes thereafter. Similarly, in the mouse, colonization by commensal enteric bacteria induces a self-limiting mucosal immune response and a state of chronic hyporesponsiveness (28). Consistent with the high degree of diversity among strains of Actinomyces and streptococci, Western blots of cell wall antigens of these bacteria probed with infants' saliva have shown extensive variability (7, 8). We have proposed, based on these data, that the induction of a limited immune response, perhaps as a result of antigenic variation, may be among the mechanisms used by commensal bacteria to avoid immune elimination and persist in the oral cavity and at other mucosal surfaces.
In our previous studies (7, 8), we analyzed the SIgA antibody response in saliva to selected commensal oral bacteria by enzyme-linked immunosorbent assay (ELISA) and Western immunoblotting with whole cells or cell wall extracts of standard and type strains. In our studies of Actinomyces spp. (7), cluster analysis was used to analyze Western blots of cell wall antigens of two standard strains of A. naeslundii genospecies 1 and 2 reacted with infants' saliva samples taken during colonization of their mouths. The clusters formed had high internal similarities of 90 to 96% and included saliva samples from different infants at different collection times. This result showed that the SIgA antibodies in the saliva from different infants reacted equivalently with the standard antigens, suggesting that all infants produced closely similar responses that did not allow discrimination within or among them.
We considered that one reason for this observation could be that standard laboratory strains of A. naeslundii genospecies 1 and 2 were used as antigen for all of the saliva samples. However, it is possible that cluster analysis of the patterns of salivary SIgA antibodies reactive with antigens from strains that colonized each infant, rather than standard strains might provide a more accurate picture of the specificity of an infant antibody response.
Furthermore, consideration of the identities of organisms that initiate the SIgA response prior to colonization by A. naeslundii raised the possibility that such responses may be the result of stimulation by antigens common to strains of closely related genera or species. Thus, the response to A. naeslundii could result, in part, from stimulation by strains of A. odontolyticus, because this species has been shown to colonize infants' mouths as early as 2 months postpartum (27). This suggestion is further strengthened because A. naeslundii and A. odontolyticus have a common peptidoglycan structure, the walls of both include rhamnose and 6-deoxytalose, and there is evidence of serological cross-reaction between strains of the species (4). The removal of SIgA antibody activity to selected species of oral streptococci by absorption of saliva with cells of Enterococcus faecalis (8) provided further support for this assertion.
Consequently, in an attempt to determine the specificity of the SIgA antibody response in individual infants, we examined SIgA antibodies in the saliva of three infants to their own genetically typed strains (ribotypes) of A. naeslundii genospecies 1 and 2. In addition, we measured the effect of absorption of saliva from six other infants with cells of A. odontolyticus on their reactivity with cells of A. naeslundii genospecies 1 and 2 to confirm the presence of cross-reacting antigens.
MATERIALS AND METHODS
Study population.
Nine healthy, full-term, breast-fed infants—referred to here as infants 1, 7, 8, 12, 14, 18, 20, 21, and 24—were selected for this study. They comprised five males and four females. Six were white, two were black, and one was Hispanic. The details of the study population have been described previously (7, 8, 13, 14). The experimental procedures used were approved by the Institutional Review Board of Georgetown University Medical Center.
Sample collection and processing. (i) Whole-mouth saliva.
Whole saliva was collected at 1 to 3 days, at 2 and 4 weeks, and at 2, 4, 6, 8 10, 12, and 15 months postpartum corresponding to visits 1 to 10. Saliva was collected by using sterile 3-ml plastic transfer pipettes. Immediately, EDTA was added to a final concentration of 5 mM to prevent formation of heterotypic calcium ion-dependent immunoglobulin-mucin complexes and to inhibit the IgA1 protease activity in the saliva (14). The saliva samples were held at −80°C until assay. Saliva collected from infant 18 at 2, 4, and 10 months (visits 4, 5, and 8), from infant 20 at 4, 10, and 15 months (visits 5, 8, and 10), from infant 21 at 2, 4, 8, 10, and 15 months (visits 4, 5, 7, 8, and 10), and from infants 1, 7, 8, 12, 14, and 24 at 4 and 15 months (visits 5 and 10) were used in the current study. The volumes of saliva that could be collected, particularly at the earlier visits, were low, such that only a limited number of assays could be carried out on each sample. Consequently, although saliva samples from infants 18, 20, and 21 were used for Western blotting with their homologous strains, it was necessary to use saliva samples from infants 1, 7, 8, 12, 14, and 24 for absorption. In addition, Western blots of cell wall proteins were limited to the same infant and it was not possible to perform analyses among the babies.
(ii) Oral swabs.
In parallel, and after each collection of whole saliva, the mucosal surfaces of the cheeks, buccal sulci, edentulous ridges, tongue and hard palate and teeth, when erupted, were swabbed by using the swab from a Vacutainer anaerobic specimen collector (Becton Dickinson Microbiology Systems, Cockeysville, Md.). The swab was then returned to the sealed tube of the collector and transported under anaerobic conditions to the laboratory within 1 h of collection. After the swab was placed in 2 ml of reduced transport fluid (29), bacteria were released from the swab and dispersed by ultrasound at 80 W for 10 s with a Branson Sonifier 250 (Branson Ultrasonics Corp., Danbury, Conn.) equipped with a microprobe. The dispersed samples were serially diluted in reduced transport fluid to 10−5 and plated onto various media.
Recovery and identification of Actinomyces species.
Trypticase soy agar containing 5% sheep blood (TSASB); Columbia agar containing 5% sheep blood, cysteine HCl, palladium chloride, dithiothreitol, and hemin (CASB); and CFAT agar (32) plates (all from Remel, Lenexa, Kans.) were inoculated by using a spiral plater (Spiral Systems, Cincinnati, Ohio). TSASB plates were incubated at 37°C for 48 h in air. CASB plates were incubated at 37°C for 3 to 5 days in an anaerobic chamber containing an atmosphere of 80% N2, 10% CO2, and 10% H2, and CFAT plates were incubated at 37°C for 48 h in 5% CO2 in air. After incubation, total counts were made of each colony morphotype on plates of the nonselective and selective media that contained between 30 and 300 CFU. Representatives of each morphotype were selected under a dissecting microscope at ×20 magnification and subcultured to purity on TSASB. The purified isolates were stained by Gram's method and tested for the production of catalase. Gram-positive pleomorphic rods were identified as Actinomyces species by slide agglutination with a panel of specific rabbit antisera (5, 11, 24).
Ribotyping of A. naeslundii genospecies 1 and 2 isolates.
Ribotyping of the A. naeslundii genospecies 1 and 2 isolates obtained from the mouths of infants 18, 20, and 21 was performed as described in detail previously (2, 3).
Preparation of wall extracts of A. naeslundii genospecies 1 and 2 isolates.
Cell wall extracts of the A. naeslundii genospecies 1 and 2 isolates were prepared as described previously (7). Briefly, cells of each isolate were suspended in ice-cold 10 mM HEPES (pH 7.4) and subjected to four 1-min cycles of ultrasound. The bacteria were then removed by centrifugation and stained by Gram's method. Microscopic examination confirmed that the bacterial cells remained intact, and there was no evidence of cellular debris. The supernatants were stored at −80°C until use.
Analysis of wall extracts of A. naeslundii genospecies 1 and 2 isolates.
The wall extracts and molecular weight standards (Bio-Rad, Hercules, Calif.) were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis with the Mini Protean II system (Bio-Rad). The separating gel was 11%, and the stacking gel was 5%. Three sets of gels were run. One set of gels was stained to detect proteins by using Coomassie brilliant blue R-250. The separated wall extracts on the other two sets of gels were transferred to polyvinylidene difluoride membranes (Immobilon-P; Millipore) by using a Trans-Blot SD system (Bio-Rad). One set of blots was processed for the simultaneous detection of carbohydrate and protein by using the DIG Glycan/Protein double labeling kit (Roche Molecular Biochemicals, Indianapolis, Ind.) according to the manufacturers instructions. The second set of blots was blocked for 1 h in 4% bovine serum albumin (BSA) in Tris-buffered saline (TBS; pH 8.3) containing 0.02% NaN3 in preparation for immunoblotting.
Extraction of A. naeslundii genospecies 1 and 2 cell wall carbohydrate.
Cell wall carbohydrate from A. naeslundii genospecies 1 (ATCC 12104) and 2 (WVU 627) was obtained by extraction from isolated cell walls with trichloroacetic acid as described previously (5). Briefly, cell walls were produced after cell disruption by a Mickle tissue disintegrator (Mickle Laboratory Engineering Co. Gomshall, Surrey, United Kingdom), washed thrice in distilled water, and lyophilized (model EF 03; Edwards High Vacuum, Crawley, Sussex, United Kingdom). Protein was removed by treatment with protease from Streptomyces griseus (Sigma-Aldrich, St. Louis, Mo.). A 50-mg portion of cell wall was incubated for 16 h at 37°C in 30 ml of 0.1 M phosphate buffer (pH 8.0) containing 0.2 mg of protease/ml. Protease-treated walls were washed thrice with distilled water and lyophilized. Lyophilized walls were extracted in 30 ml of 5% aqueous trichloroacetic acid (Fisher Scientific, Pittsburgh, Pa.) for 16 h at 37°C. The wall suspension was centrifuged, and 5 volumes of acetone (Fisher) was added to the separated supernatant. The precipitated carbohydrate from the supernatant was allowed to sediment for 16 h at 4°C, dissolved in distilled water, dialyzed against distilled water, and lyophilized. The carbohydrate was free of protein as determined by the Bradford assay (Bio-Rad).
Detection of SIgA antibodies in saliva reactive with A. naeslundii genospecies 1 and 2 cell wall carbohydrate.
The ELISA used in the present study has been described in detail elsewhere (7, 8). Briefly, Immulon-2 plates (Dynatech, Chantilly, Va.) were coated overnight with cell wall carbohydrate from A. naeslundii genospecies 1 or 2 at a concentration of 5 μg/ml in 0.05 M carbonate buffer (pH 9.6). Unbound carbohydrate was washed from the wells that were then blocked with phosphate-buffered saline (PBS; pH 8.0) containing 0.1% BSA. After the wells were washed, dilutions of saliva samples from each infant were added in duplicate and incubated overnight with shaking. After the diluted saliva was washed out, a biotinylated affinity-purified polyclonal goat anti-human α chain (BGAHα) antibody (Jackson ImmunoResearch, West Grove, Pa.) at 2.0 μg/ml was used to detect bound SIgA antibodies. After being washed, streptavidin conjugated with horseradish peroxidase (SA-HRP; BioSource International, Camarillo, Calif.) at 0.1 μg/ml was added to the wells to detect the biotinylated antibodies. The wells were washed again, and o-phenylenediamine (1 mg/ml) in citrate-phosphate buffer (pH 4.5) containing 0.012% hydrogen peroxide was added to each well. The optical density at 450 nm was measured with a Spectra-Rainbow automated microplate reader (Tecan U.S., Inc., Research Triangle Park, N.C.). As positive controls, rabbit antibodies reactive in precipitin tests with the cell wall carbohydrates and adult human saliva samples (from J. L. Kirchherr) were run in the ELISA. Bound rabbit IgG antibodies were detected by using an affinity-purified goat anti-rabbit IgG antibody conjugated with horseradish peroxidase (Jackson ImmunoResearch) and detected as described above.
Fine specificity of SIgA anti-A. naeslundii genospecies 1 and 2 antibodies in saliva.
Individual lanes were cut from the polyvinylidene difluoride membranes so that the wall extract obtained from each of the isolates could be reacted with all of the saliva samples obtained from each infant. After a blocking step in 4% BSA in 10 mM TBS (pH 7.4), the strips were incubated with the saliva samples overnight at 4°C. They were then subjected to three 10-min washes in TBS containing 0.1% Tween and 0.5 M NaCl. Bound SIgA antibodies were detected with BGAHα and SA-HRP as described above for ELISA except that 0.5 mg of diaminobenzidine/ml in 0.01 M PBS (pH 7.4) containing 0.01% H2O2 was used as substrate. Digital images of all lanes, including those containing molecular weight standards, were obtained with an AlphaImager (Alpha Innotech Corp., San Leandro, Calif.). The image files were imported into an analytic software program (GelCompar 4.0; Applied Maths, Kortrijk, Belgium) for analysis.
Cluster analysis of Western blot band patterns.
Cluster analysis was performed within GelCompar 4.0 by the method of Ward (31).
Absorption of saliva.
The specificity of A. naeslundii genospecies 1- and 2-reactive SIgA antibodies in saliva was tested by absorption. Selected pairs of saliva samples were obtained from three male and three female infants, i.e., infants 1, 7, 8, 12, 14, and 24. Saliva samples selected were those obtained at 4 months of age (visit 5) before A. naeslundii genospecies 1 and 2 were isolated from their mouths (termed “early samples”) and at 15 months of age (visit 10) after A. naeslundii genospecies 1 and 2 were isolated from their mouths (termed “late samples”). Since A. odontolyticus has been shown to be an early colonizer of the mouth (27), we elected to determine whether A. naeslundii-reactive antibodies could have been induced by colonization by A. odontolyticus. Briefly, A. odontolyticus NCTC 9935 (serotype 1), A. odontolyticus WVU 482 (serotype 2), and A. odontoloyticus MCB 120, a serotype 2 isolate obtained from an infant in our study, were grown in Actinomyces Broth (Becton Dickinson Microbiology Systems, Sparks, Md.) at 37°C in 5% CO2 for 5 days. Aliquots of the saliva samples were serially diluted in PBS-Tween and mixed with an equal volume of packed, washed cells of the Actinomyces strains. The suspensions were incubated at 37°C for 2 h and then at 4°C overnight. After absorption, the bacteria were removed by centrifugation, and an ELISA was used to measure the antibacterial antibodies in absorbed and unabsorbed dilutions of the saliva samples as described above. The percentages of residual antibody activity against A. naeslundii genospecies 1 and 2 after independent absorption with the A. odontolyticus strains compared to the unabsorbed control were determined in microtiter wells coated with A. naeslundii genospecies 1 and 2. The titration curves of unabsorbed and absorbed serial dilutions of the saliva were approximately parallel. The effect of absorption with A. odontolyticus on A. naeslundii reactive SIgA antibodies in the saliva samples was calculated from the linear portion of the titration curve by dividing the optical density at 450 nm of each dilution of the absorbed saliva samples by that of the corresponding dilution of unabsorbed saliva and multiplying by 100. The effect of absorption was expressed as: (i) the number of absorbed samples that retained activity against A. naeslundii genospecies 1 and 2 and (ii) the medians of the percentages of residual antibody in the absorbed samples. The variances in the data were expressed as the 25th and 75th percentiles.
RESULTS
Ribotypes.
As we have reported previously (7) A. naeslundii genospecies 1 and 2 do not colonize the human oral cavity until approximately 4 months after the eruption of teeth. In our study population primary teeth began to erupt at 6 months of age and all infants had erupted teeth by 12 months (7). A. naeslundii genospecies 1 and 2 first were detected in infants 18, 20, and 21 at 10 months of age (visit 8). Twenty-two isolates of A. naeslundii obtained from the three infants were examined. The 6 isolates from infant 18 and the 3 isolates from infant 21 were identified as A. naeslundii genospecies 1. A. naeslundii genospecies 1 (3 isolates) and A. naeslundii genospecies 2 (10 isolates) were obtained from the third infant (infant 20).
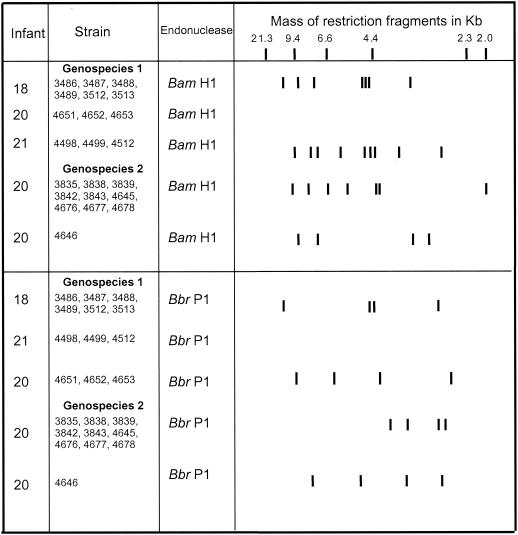
A schematic of the ribotype patterns of isolates of A. naeslundii from the infants is shown in Fig. 1. The ribotypes produced by separate digestion of the genomic DNA from A. naeslundii genospecies 1 isolates with endonucleases BamHI and BbrPI showed that each was colonized by a single unique clone. However, although use of these endonucleases allowed differentiation between the A. naeslundii clones colonizing the infants the clones shared common patterns. A. naeslundii genospecies 1 isolates from infants 18 and 20 shared a common ribotype pattern when genomic DNA was digested with BamHI and isolates from infants 18 and 21 shared a common BbrPI ribotype. Thus, overall, the A. naeslundii genospecies 1 isolates from infants 18, 20, and 21 were closely related. In contrast, the two clones of A. naeslundii genospecies 2 obtained from infant 20 were distinct. Nine isolates of A. naeslundii genospecies 2 grouped into one clone with identical BamHI and BbrPI ribotype patterns, whereas the BamHI and BbrPI ribotype patterns of the remaining isolate were different from those of the other nine isolates.
FIG. 1.
Ribotype patterns of the strains of A. naeslundii genospecies 1 and 2 isolated from the infants. The ribotypes were produced by separate digestion of genomic DNA with the restriction endonucleases BamHI and BbrPI.
Analysis of protein profiles of A. naeslundii genospecies 1 and 2.
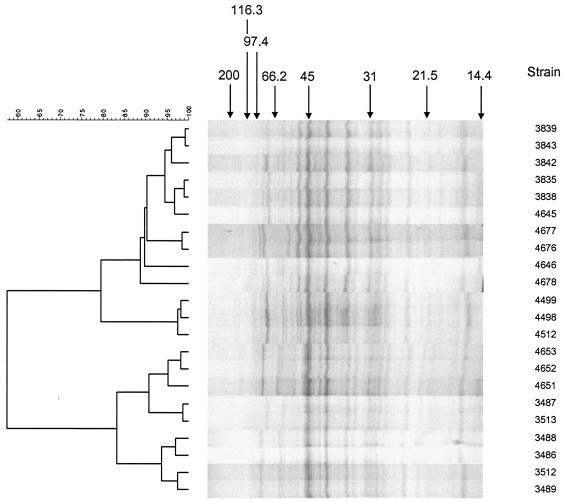
The cell wall extracts of A. naeslundii genospecies 1 and 2 were resolved into approximately 40 bands by sodium dodecyl sulfate-polyacrylamide gel electrophoresis with 11% running gels (Fig. 2). Overall, the A. naeslundii protein profiles were 57.5% similar and were readily separated into genospecies by using cluster analysis. In the genospecies 1 cluster, the protein profiles of the six isolates obtained from infant 18 (isolates 3486, 3487, 3488, 3489, 3512, and 3513) that comprised a single clone were 92% similar. The protein profiles of the three isolates from infant 20 (isolates 4651, 4652, and 4653) that comprised a second clone were 94% similar, and the protein profiles of the three isolates from infant 21 (isolates 4498, 4499, and 4512) that comprised a third clone were 97% similar.
FIG. 2.
Cluster analysis of protein profiles of isolates of A. naeslundii genospecies 1 and 2 obtained from infants 18, 20, and 21. The genospecies are separated at 57.5% similarity. Genospecies 2 isolates form the upper cluster and genospecies 1 isolates form the lower cluster.
The genospecies 2 isolates from infant 20 comprised two clones: one with nine isolates and the second with a single isolate. The protein profiles of the 10 genospecies 2 isolates (isolates 3835, 3838, 3839, 3842, 3843, 4645, 4646, 4676, 4677, and 4678) were 90.5% similar, and the protein profiles of the nine isolates comprising the larger of the clones (all of the above accept 4646) were 91.5% similar. These results indicate that the predominant strains of A. naeslundii in each infant were relatively stable over time compared to Streptococcus mitis and, unlike the streptococci, these strains did not show evidence of clonal replacement (17, 18).
Effect of absorption of the infant saliva with A. odontolyticus cells on A. naeslundii-reactive SIgA antibodies.
The results of absorption of the saliva samples are shown in Table 1. The saliva samples were absorbed with cells of the type strain of A. odontolyticus (NCTC 9935) (serotype 1), strain WVU (West Virginia University) 482 (serotype 2), and (MCB 120) (serotype 2). Only serotype 2 A. odontolyticus strains were recovered from the infants in our study, and MCB 120 was one of these. Absorbed saliva was tested by ELISA with cells of these strains and also A. naeslundii ATCC 12104 (genospecies 1) and A. naeslundii W1053 (genospecies 2) as antigen. Examination of the data for the early saliva samples shows clearly that absorption with strains of A. odontolyticus reduced the SIgA antibody activity against both genospecies of A. naeslundii either to background (0% median residual antibody activity for genospecies 1) or to nearly background levels (0 to 2.6% median residual antibody activity for genospecies 2). However, in contrast to the early saliva samples, cells of the A. odontolyticus strains were much less effective at absorbing A. naeslundii genospecies 1-reactive SIgA antibodies (14.6 to 30.3% median residual antibody activity) and genospecies 2-reactive SIgA antibodies (9.0 to 21.0% median residual antibody activity) in the late saliva samples.
TABLE 1.
Residual A. naeslundii genospecies 1 and 2 SIgA antibody activity in infant saliva after absorption with A. odontolyticus serotypes 1 and 2a
| Test strain (serotype) and parameter | Early saliva samples
|
Late saliva samples
|
||||
|---|---|---|---|---|---|---|
| Absorbed with A. odontolyticus strain (serotype):
|
Absorbed with A. odontolyticus strain (serotype):
|
|||||
| NCTC 9935 (1) | WVU 482 (2) | MCB 120 (2) | NCTC 9935 (1) | WVU 482 (2) | MCB 120 (2) | |
| A. odontolyticus | ||||||
| NCTC 9935 (1) | ||||||
| No. positive | 0 | 9 | 8 | 0 | 12 | 11 |
| Median (%) | 0 | 3.9 | 2.1 | 0 | 19.9 | 15.3 |
| 25th percentile | 0 | 0.7 | 0 | 0 | 18.2 | 4.1 |
| 75th percentile | 0 | 21.7 | 7.4 | 0 | 23.3 | 27.8 |
| WVU 483 (2) | ||||||
| No. positive | 3 | 0 | 3 | 12 | 0 | 6 |
| Median (%) | 0 | 0 | 0 | 18.1 | 0 | 1.1 |
| 25th percentile | 0 | 0 | 0 | 7.2 | 0 | 0 |
| 75th percentile | 1.1 | 0 | 0.3 | 27.2 | 0 | 21.6 |
| MCB 120 (2) | ||||||
| No. positive | 8 | 9 | 0 | 12 | 11 | 0 |
| Median (%) | 2.5 | 6.8 | 0 | 14.0 | 10.6 | 0 |
| 25th percentile | 0 | 0 | 0 | 6.9 | 3.1 | 0 |
| 75th percentile | 8.0 | 15.7 | 0 | 21.0 | 20.4 | 0 |
| A. naeslundii | ||||||
| ATCC 12104 (1) | ||||||
| No. positive | 3 | 5 | 5 | 11 | 12 | 11 |
| Median (%) | 0 | 0 | 0 | 19.9 | 30.3 | 14.6 |
| 25th percentile | 0 | 0 | 0 | 12.7 | 17.4 | 3.5 |
| 75th percentile | 0.1 | 13.3 | 3.3 | 38.1 | 37.1 | 52.5 |
| W1053 (2) | ||||||
| No. positive | 1 | 6 | 7 | 9 | 11 | 6 |
| Median (%) | 0 | 1.4 | 2.6 | 11.0 | 21.0 | 9.0 |
| 25th percentile | 0 | 0 | 0 | 1.3 | 5.6 | 0 |
| 75th percentile | 0 | 10.9 | 5.9 | 25.0 | 32.2 | 28.2 |
Whole saliva was collected from six infants before (early samples) and after (late samples), they were colonized with A. naeslundii genospecies 1 and 2. These saliva samples were absorbed with A. odontolyticus serotypes 1 and 2 whole cells. After absorption, an ELISA was used to measure A. naeslundii genospecies 1 and 2 SIgA antibodies in absorbed and unabsorbed dilutions of the saliva samples. The percentages of residual antibody activity against A. naeslundii genospecies 1 and 2 after independent absorption with the A. odontolyticus strains were determined by dividing the optical density at 450 nm of each absorbed saliva by that of the corresponding unabsorbed saliva at the same dilution and multiplying that value by 100. The results are expressed as the number of samples that retained activity against A. naeslundii genospecies 1 and 2 after absorption with A. odontolyticus, the medians of the percentages of residual antibody in the absorbed samples and the 25th and 75th percentiles. Duplicate samples of saliva from six infants were assayed on two separate occasions. Serotypes are indicated in parentheses.
Reactivity of salivary SIgA with wall carbohydrate from A. naeslundii genospecies 1 and 2.
After Western transfer of the wall extracts one of each of three replicate strips of each isolate was developed by using the DIG Glycan/Protein double labeling kit (Roche Molecular Biochemicals) to detect glycoconjugates, and several bands staining for both protein and sugars were detected. These bands reacted with the infants salivary SIgA antibody (data not shown). However, when purified wall carbohydrates from the A. naeslundii genospecies 1 and 2 reference strains were used to coat the wells of microtiter trays, no carbohydrate-reactive SIgA antibodies were detected in saliva from the infants by using a sensitive ELISA. In contrast, the control rabbit antisera and adult saliva gave strong positive reactions with the purified wall carbohydrate.
Reactivity of salivary SIgA antibodies with wall extracts.
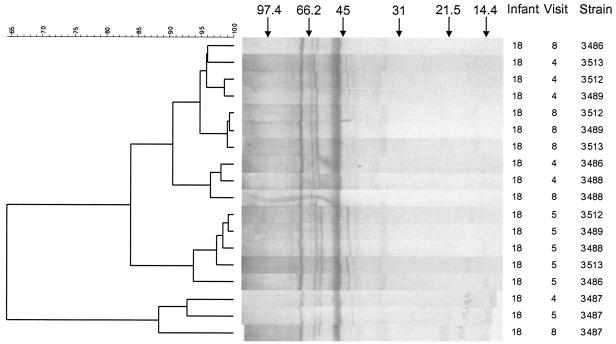
Immunoblot patterns were analyzed by using the Ward method (31). An example of the dendrogram from cluster analysis of the immunoblots from infant 18 is shown in Fig. 3. The majority of the A. naeslundii genospecies 1 and 2 bands recognized by salivary SIgA antibodies had apparent molecular masses of between 75 and 31 kDa. However, in some lanes salivary SIgA antibodies recognized between one and three bands with molecular masses of >75 kDa and several lower molecular mass bands that ran between the 31- and 14.4-kDa standards. SIgA antibodies in all but one saliva sample recognized two bands with apparent molecular masses of approximately 75 and 53 kDa that were common to both of the A. naeslundii genospecies 1 and 2 wall extracts. Computer analysis identified between 7 and 17 bands (mean, 11 bands) of the A. naeslundii genospecies 1 wall extracts recognized by salivary SIgA antibodies and between 6 and 17 bands (mean, 12 bands) of A. naeslundii genospecies 2 wall extracts recognized by salivary SIgA antibodies. Comparisons were made between the patterns of the blots of the wall extract of each isolate reacted with saliva from each visit within each infant, and details of the strains within clusters and the saliva samples are shown in Table 2. Comparisons of the clusters formed showed that they could be divided into three types termed A, B, and C based on the isolates and the visit at which the saliva samples were obtained (Table 2). Type A clusters were those formed by antibody reactivity patterns of saliva samples from more than one visit reacted with extracts of a single isolate. Type B clusters included antibody reactivity patterns of a saliva sample from a single visit reacted with several different isolates. However, these divisions were not absolute, and some clusters included antibody reactivity patterns of saliva from more than one visit reacted with several isolates. These were termed type C clusters. In order to simplify the presentation of the data, the points of separation and the internal similarities of clusters for each infant are shown as schematics in Fig. 4 and 5. As an example of the manner in which the simplified data are presented, consider the type A cluster of infant 18 based on strain 3487 (Table 2), which is shown in the dendrogram in Fig. 3 and the schematic in Fig. 4. In the Fig. 4 schematic for infant 18, saliva samples from visit 4 (2 months), visit 5 (4 months), and visit 8 (10 months) form a type A cluster at 88% similarity, and this cluster connects to the B2, C2, B1, and C1 clusters at 64% similarity. Similarly, in the same infant, cluster B2 (based on saliva from visit 5 [4 months] and strains 3486, 3488, 3489, 3512, and 3513) is formed at 94% similarity and joins the other clusters at 83% similarity.
FIG. 3.
Cluster analysis of immunoblots from infant 18. Wall extracts of autochthonous strains of A. naeslundii genospecies 1 were incubated with saliva collected at 2, 4, and 10 months (visits 4, 5, and 8) and probed to detect SIgA antibodies.
TABLE 2.
The clusters formed from analysis of Western blot patterns of SIgA antibodies in infants' saliva reacted with cell wall proteins from their autochthonous strains of A. naeslundii genospecies 1 and 2
| Infant no. | Genospeciesa | Overall pattern similarityb (%) | No. of clusters | Cluster typesc | Strain no. | Visit no. | Infant age(s) (mo) |
|---|---|---|---|---|---|---|---|
| 18 | 1 | 64 | 1 | A | 3487 | 4, 5, 8 | 2, 4, 10 |
| 2 | B1 | 3489, 3512, 3513 | 8 | 10 | |||
| B2 | 3486, 3488, 3489, 3512, 3513 | 5 | 4 | ||||
| 2 | C1 | 3486, 3488 | 4, 8 | 2, 10 | |||
| C2 | 3486, 3489, 3512, 3513 | 4, 8 | 2, 10 | ||||
| 20 | 1 | 42 | 3 | B1 | 4651, 4562, 4653 | 5 | 4 |
| B2 | 4651, 4562, 4653 | 10 | 15 | ||||
| B3 | 4651, 4562, 4653 | 8 | 10 | ||||
| 2 | 15 | 1 | A | 4676 | 5, 8, 10 | 4, 10, 15 | |
| 5 | B1 | 3835, 3838, 4645, 4646 | 5 | 4 | |||
| B2 | 3835, 3839, 3842, 3843, 4646 | 10 | 15 | ||||
| B3 | 4645, 4677, 4678 | 10 | 15 | ||||
| B4 | 3839, 3842, 3843 | 8 | 10 | ||||
| B5 | 3839, 3842, 3843 | 5 | 4 | ||||
| 2 | C1 | 4645, 4677, 4678 | 5, 8 | 4, 10 | |||
| C2 | 3835, 3836, 4646, 4677 | 5, 8 | 4, 10 | ||||
| 21 | 1 | 74 | 1 | A | 4512 | 8, 10 | 10, 15 |
| 4 | C1 | 4499, 4512 | 5, 7 | 4, 8 | |||
| C2 | 4498, 4512 | 4, 5 | 2, 4 | ||||
| C3 | 4498, 4499 | 7, 10 | 8, 15 | ||||
| C4 | 4498, 4499 | 4, 8, 10 | 2, 10, 15 |
A. naeslundii genospecies 1 or 2.
The level of similarity that includes all of the patterns.
Type A clusters are based on a single strain, type B clusters are based on a single saliva sample, and type C clusters contain mixed strains and saliva samples.
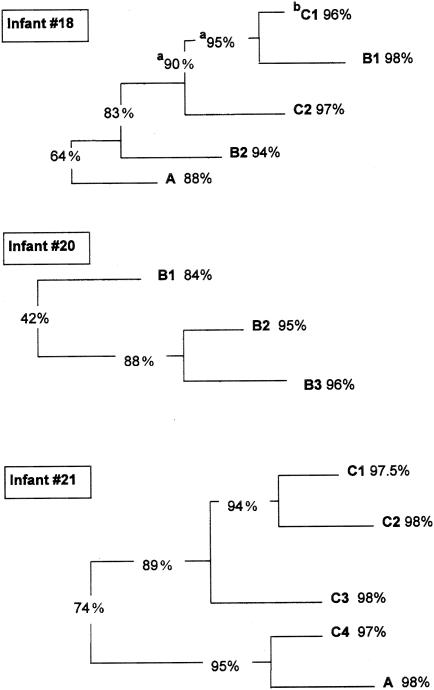
FIG. 4.
Separation and internal similarities among Western blot patterns produced by reaction of SIgA antibodies in saliva from infants 18, 20, and 21 with cell wall antigens of autochthonous strains of A. naeslundii genospecies 1. Superscripts: a, percentage similarity separating the cluster; b, internal similarity of the cluster.
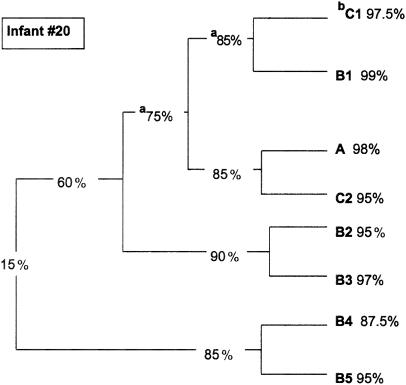
FIG. 5.
Separation and internal similarities among Western blot patterns produced by reaction of SIgA antibodies in saliva from infant 20, with cell wall antigens of autochthonous strains of A. naeslundii genospecies 2. Superscripts: a, percent similarity separating the cluster; b, internal similarity of the cluster.
The overall similarity among the band patterns recognized by the SIgA antibodies in the saliva samples collected from the same infant at different visits varied from 74% for genospecies 1 strains from infant 21 (Fig. 4) to 15% for genospecies 2 strains from infant 20 (Fig. 5).
The results of cluster analysis of A. naeslundii genospecies 1 isolates are shown in Table 2 and Fig. 4. Despite the overall separation at high similarities, some clusters were defined at lower similarities of between 42 and 83%. For infant 18, cluster A (strain 3487) and cluster B2 (saliva visit 5 [4 months]) separated at 64 and 83%, respectively. For infant 20, cluster B1 (saliva visit 5 [4 months]) separated from clusters B2 and B3 at 42%, making it quite distinct. For infant 21, cluster A (strain 4512) and cluster C4 were separated at 74% from clusters C1, C2, and C3.
In all, eight clusters were defined among the strains of A. naeslundii genospecies 2 from infant 20 (Table 2 and Fig. 5). Of particular note, were clusters B4 and B5 (both from saliva collected at visits 5 [4 months] and 8 [10 months]) that were separated from the other clusters at 15% similarity. Also, clusters B2 and B3 based on saliva from visit 10 (15 months) were separated at 60%. The single type A cluster (strain 4676) among the A. naeslundii genospecies 2 isolates separated at 75% from clusters B2 to B5.
DISCUSSION
The objective of the present study was to examine the specificity of the SIgA antibody responses in individual infants to the strains of A. naeslundii that colonized their mouths. The isolates were ribotyped in order to reveal the diversity of strains within an infant and to detect any obvious clonal replacement. Insufficient numbers of strains were isolated from the infants to detect genetic variants that could be present in low numbers. However, we believe that our results reveal the predominant A. naeslundii ribotype in these infants and confirm that the ribotypes isolated persisted in their mouths over time, as has been shown to be the case in adults (J. Johnson, M. Kinard, G. Bowden, and C. Schachtele, abstract from the 72nd General Session of the International Association for Dental Research, J. Dent. Res. 73:346, 1994). Each of the three infants (infants 18, 20, and 21) in our study was colonized by a unique ribotype of A. naeslundii genospecies 1 from age 10 months (visit 8) when the bacteria were first detected in their mouths until the last sampling at age 15 months (visit 10). Infant 21 was also colonized by A. naeslundii genospecies 2, and the isolates were predominantly one ribotype, reflecting the findings for A. naeslundii genospecies 1. However, one isolate recovered at age 15 months (visit 10) was a different ribotype, suggesting that infants, like adults (M. Kinard, J. Johnson, G. Bowden, and C. Schachtele, abstract from the 72nd General Session of the International Association for Dental Research, J. Dent. Res. 73:248, 1994) can be colonized by more than one ribotype of A. naeslundii genospecies 2.
The limited clonal diversity and stability of A. naeslundii is in marked contrast to S. mitis that exhibits extensive clonal diversity and replacement in infants (13, 17, 18). The finding of limited diversity among isolates of A. naeslundii colonizing these infants could be considered to be an advantage when comparisons are drawn among antibody responses because it could be argued that, because clonal diversity is minimal, the antigenic stimulus to the infants' immune system would be relatively stable. Against this it must be recognized that ribotyping may not be sufficiently discriminating to reveal antigenic drift or variation among these strains. Indeed, the persistence of a single ribotype may reflect a balance that has been established between the immune response of the host and that unique ribotype.
Establishing the specificity of the infants' salivary SIgA antibody responses to A. naeslundii was first approached by absorbing saliva samples from six infants with strains of A. odontolyticus, a species known to colonize the mucous membranes of infants prior to tooth eruption (27). Absorption of the early saliva samples (Table 1) with the A. odontolyticus strains removed at least 97.4% of A. naeslundii genospecies 1- and 2-reactive antibodies. This finding provides support for the suggestion that, in large part, the antibodies were either stimulated by this species or were directed to antigens common among the genus Actinomyces (4) or common among other gram-positive bacteria inhabiting the mouth and/or gastrointestinal tract of these infants. For example, it has been reported that some species of viridans streptococci, lactobacilli, and actinomyces share extracellular and cell wall antigens (27a). Such common antigens of gram-positive bacteria may include glucans and teichoic acid-associated phosphorylcholine (15). Alternatively, it is possible that the binding of SIgA1 antibodies in saliva to the Actinomyces species was nonimmune and mediated by interaction of bacterial lectins with O-linked oligosaccharides at the hinge region of the α1 heavy chain (26). However, the fact that salivary SIgA antibodies bound multiple bands on the Western blots of the cell wall proteins argues against a nonimmune interaction. Another possibility is that A. naeslundii-reactive SIgA antibodies present in saliva prior to colonization of this bacterium are examples of polyreactive antibody (25). Polyreactive SIgA antibodies, some with high functional affinity, have been detected in human saliva (25). Although, if this were the case one might have expected reactivity with the Actinomyces wall polysaccharide, since polyreactive antibodies commonly bind carbohydrates (25). However, the infants saliva samples were uniformly negative when tested with cell wall carbohydrate.
After the establishment of A. naeslundii in the mouth the salivary SIgA antibody response appeared to become more specific for this bacterium. The amounts of residual A. naeslundii-genospecies reactive antibodies remaining after absorption with the A. odontolyticus strains increased. Also, the immunoblot patterns of antibody reactivity of late saliva samples were generally more complex than those of early samples.
In a previous study (7) we examined the fine specificity of the salivary SIgA antibody response in infants' saliva by using two standard strains of A. naeslundii. The majority of the clusters observed were those that contained saliva from more than one visit that reacted with several isolates (type C clusters), suggesting that there was no individual specificity of the SIgA antibodies in the saliva of a given infant. However, we reasoned that this apparent lack of specificity might be attributed to the use of standard strains as common antigens to test the saliva samples. Therefore, we decided to use strains isolated from individual infants as antigens to test the reactivity of their saliva. The results of cluster analysis of the reactivity of SIgA antibody in saliva samples from individual infants and their homologous strains generated some type C clusters. However, unlike our previous results (7), two other types of clusters, types A and B, were observed. The presence of these clusters adds support for the proposal that the SIgA response to A. naeslundii in infants shows specificity in recognizing strains and also exhibits maturation with increased specificity. Type A clusters were less common than type B clusters, but one example was detected in each infant (Table 2). Type A clusters were based on a single isolate, and these clusters indicate that more than one saliva sample from an infant recognized this strain as unique and separate from other isolates. Therefore, it is most likely that these strains carried unique epitopes recognized by infants' salivary SIgA, supporting the concept that SIgA antibodies in saliva can separate and interact with individual strains of A. naeslundii.
Type B clusters were more common than type A clusters ranging from two in infant 18 for A. naeslundii genospecies 1 (Table 2 and Fig. 4) to five among the A. naeslundii genospecies 2 isolates from infant 20 (Table 2 and Fig. 5). However, no type B clusters were formed from the A. naeslundii genospecies 1 strains of infant 21 (Table 2 and Fig. 4). The presence of type B clusters indicates differences among the reactivity of SIgA antibodies in saliva collected at different visits. This is particularly evident with cluster B1 of A. naeslundii genospecies 1 in infant 20 that separates at 42% similarity (Fig. 4) and with clusters B4 and B5 of A. naeslundii genospecies 2 in the same infant that separate from the other clusters at 15% (Fig. 5). Also, the extent of differentiation among the clusters could be related to maturation of the immune response if, as the child's immune system developed, differences could be shown between the reaction of the earliest and the latest saliva samples to antigens from the bacteria. However, such observations require at least two type B clusters for an infant. This is the case in infant 20 for both A. naeslundii genospecies 1 and 2. In infant 20, A. naeslundii genospecies 1 wall proteins reacted with their homologous saliva only gave three clusters, all of type B (Table 2 and Fig. 4). Cluster B1 was based on visit 5 (4 months) and separated from the later visits, 8 and 10 (10 and 15 months), and at 42% similarity compared to B2 and B3 that separated at 88%, suggesting a distinction between the antibody activity in saliva from visit five compared to the later visits. Thus, in infant 20 the reactivity pattern of the earliest saliva sample was relatively distinct from those of later samples. This pattern was even more evident when the infant 20 A. naeslundii genospecies 2 clusters were considered. Clusters B4 (visit 8; 10 months) and B5 (visit 5; 4 months) separated from clusters B1 (visit 5; 4 months) and B2 and B3 (visit 10; 15 months) at 15% similarity. Moreover, clusters B2 and B3 separated from B1 at 60%, clearly distinguishing the later visit 10 (15 months) saliva from those of the earlier visits. The separation of the saliva from the visit 5 (B5) (4 months) and visit 10 (B3) (15 months) clusters could be based on the isolates involved, which were different for these clusters. If so, this would lend support to the concept that the strains of A. naeslundii isolated, despite being very similar genetically, had variations in their antigenic composition that were recognized by the SIgA antibodies in saliva.
The results of cross-absorption support the proposal that the SIgA antibodies in saliva samples collected after colonization of the mouths of the infants by A. naeslundii show more specificity than those collected prior to colonization by this species. When these data are considered together with those for the type B clusters, the suggestion of maturation and increased specificity of the salivary SIgA response is strengthened. Furthermore, the presence of type A clusters and the division of type B clusters such as B1, B2, and B3 from infant 20 (Fig. 4) indicate that the SIgA antibody response can recognize antigenic variations among isolates of the same ribotypes of A. naeslundii.
In the mouse, the lamina propria of the gut is populated by two types of IgA-secreting cells that have different origins (19). Conventional or B-2 B cells are derived from the bone marrow, predominate in the Peyer's patches and require cognate help from CD4+ T cells. As a result of affinity maturation (16), IgA antibodies produced by these cells are specific and of high affinity. In contrast, B-1 cells are derived from the peritoneal and pleural cavities early in ontogeny, do not require cognate help from CD4+ T cells, and are self-replenishing (19). As a consequence their receptor repertoire, established during the neonatal period, is limited, but it appears to be selected by colonization of the neonatal mouse by its commensal bacteria. It has been suggested that IgA antibodies produced by B-1 B cells are designed for regulation of indigenous bacteria, whereas IgA antibody produced by conventional B cells is targeted at extrinsic pathogenic bacteria (19). It is intriguing to speculate that the SIgA A. naeslundii genospecies 1- and 2-reactive antibodies present in the saliva samples collected from the infants prior to the establishment of this species in their mouths may reflect, at least in part, a B-1 cell-mediated plurispecific mucosal immune response (25). This B-1 response may then undergo transition into a B-2 response after colonization by these organisms.
Many studies exploring the potential utility of commensal bacteria as vaccine vectors have used the human commensal viridans streptococcus, S. gordonii, in rodent models (12, 21). Although this bacterium is autochthonous in the human oral cavity it is allochthonous in rodents. Thus, the ecological relationship between the bacterium and its host is different in rodents than in humans. Therefore, data obtained from rodent experiments concerning the immune response to the vector organism and the vaccine antigens it expresses and, indeed, the ease with which the organism can be stably introduced into the climax community, may not be readily extrapolated to man. In this context species belonging to the genus Actinomyces that are autochthonous to the oral cavity of humans and many other mammalian species (10) may provide a more suitable vector. Be that as it may, a fuller understanding of the mucosal immune response to commensal bacteria, such as has been begun in the present study, is needed before commensal bacteria can be successfully exploited for vaccine delivery in human beings.
Acknowledgments
This study was supported by Public Health Service grant DE08178 from the National Institute of Dental and Craniofacial Research. G.H.W.B. is supported by the Canadian Institutes for Health Research (Canadian Medical Research Council grant MT 7611).
REFERENCES
- 1.Bolken, T. C., C. A. Franze, K. F. Jones, R. H. Bell, R. M. Swanson, D. S. King, V. A. Fischetti, and D. E. Hruby. 2002. Analysis of factors affecting surface expression and immunogenicity of recombinant proteins expressed by gram-positive commensal vectors. Infect. Immun. 70:2487-2491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bowden, G., J. Johnson, and C. Schachtele. 1993. Characterization of Actinomyces with genomic DNA fingerprints and rRNA gene probes. J. Dent. Res. 72:1171-1179. [DOI] [PubMed] [Google Scholar]
- 3.Bowden, G. H., N. Nolette, H. Ryding, and B. M. Cleghorn. 1999. The diversity and distribution of the predominant ribotypes of Actinomyces naeslundii genospecies 1 and 2 in samples from enamel and from healthy and carious root surfaces of teeth. J. Dent. Res. 78:1800-1809. [DOI] [PubMed] [Google Scholar]
- 4.Bowden, G. H. W. 1998. Actinomyces, p. 445-462. In A. Balows and B. I. Duerden (ed.), Topley and Wilson's microbiology and microbial infections, 9th ed. Arnold, London, England.
- 5.Bowden, G. H. W., J. M. Hardie, and E. D. Fillery. 1976. Antigens from Actinomyces species and their value in identification. J. Dent. Res. 55A:192-204. [DOI] [PubMed] [Google Scholar]
- 6.Cole, M. F. 1985. Influence of secretory immunoglobulin A on ecology of oral bacteria, p. 131-135. In S. E. Merganhagen and B. Rosan (ed.), Molecular basis of oral microbial adhesion. American Society for Microbiology, Washington, D.C.
- 7.Cole, M. F., S. Bryan, M. K. Evans, C. L. Pearce, M. J. Sheridan, P. A. Sura, R. Wientzen, and G. H. W. Bowden. 1998. Humoral immunity of commensal oral bacteria in human infants: salivary antibodies reactive with Actinomyces naeslundii genospecies 1 and 2 during colonization. Infect. Immun. 66:4283-4289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cole, M. F., S. Bryan, M. K. Evans, C. L. Pearce, M. J. Sheridan, P. A. Sura, R. Wientzen, and G. H. W. Bowden. 1999. Humoral immunity to commensal oral bacteria in human infants: salivary secretory immunoglobulin A antibodies reactive with Streptococcus mitis biovar 1, Streptococcus oralis, Streptococcus mutans, and Enterococcus faecalis during the first two years of life. Infect. Immun. 67:1878-1886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Crabbe, P. A., D. R. Nash, H. Bazin, H. Eyssen, and J. F. Heremans. 1970. Immunohistochemical observations on lymphoid tissues from conventional and germ-free mice. Lab. Investig. 22:448-457. [PubMed] [Google Scholar]
- 10.Dent, V. E. 1979. The bacteriology of dental plaque from a variety of zoo-maintained mammalian species. Archs. Oral Biol. 24:277-282. [DOI] [PubMed] [Google Scholar]
- 11.Fillery, E. D., G. H. W. Bowden, and J. M. Hardie. 1978. A comparison of strains of bacteria designated Actinomyces viscosus and Actinomyces naeslundii. Caries Res. 12:299-312. [DOI] [PubMed] [Google Scholar]
- 12.Fischetti, V. A., D. Medaglini, and G. Pozzi. 1996. gram-positive commensal bacteria for mucosal vaccine delivery. Curr. Opin. Biotechnol. 7:659-666. [DOI] [PubMed] [Google Scholar]
- 13.Fitzsimmons, S., M. Evans, C. Pearce, M. J. Sheridan, R. Wientzen, G. Bowden, and M. F. Cole. 1996. Clonal diversity of Streptococcus mitis biovar 1 isolates from the oral cavity of human neonates. Clin. Diagn. Lab. Immunol. 3:517-522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Fitzsimmons, S. P., M. K. Evans, C. L. Pearce, M. J. Sheridan, R. Weintzen, and M. F. Cole. 1994. Immunoglobulin A subclasses in infants' saliva and in saliva and milk from their mothers. J. Pediatr. 124:566-573. [DOI] [PubMed] [Google Scholar]
- 15.Gmur, R., T. Thurnheer, and B. Guggenheim. 1999. Dominant cross-reactive antibodies generated during the response to a variety of oral bacterial species detect phosphorylcholine. J. Dent. Res. 78:77-85. [DOI] [PubMed] [Google Scholar]
- 16.Gonzalez-Fernandez, A., and C. Milstein. 1993. Analysis of somatic hypermutation in mouse Peyer's patches using immunoglobulin kappa light-chain transgenes. Proc. Natl. Acad. Sci. USA 90:9862-9866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hohwy, J., and M. Kilian. 1995. Clonal diversity of the Streptococcus mitis biovar 1 population in the human oral cavity and pharynx. Oral Microbiol. Immunol. 10:19-25. [DOI] [PubMed] [Google Scholar]
- 18.Hohwy, J., J. Reinholdt, and M. Kilian. 2001. Population dynamics of Streptococcus mitis in its natural habitat. Infect. Immun. 69:6055-6063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kroese, F. G., R. de Waard, and N. A. Bos. 1996. B-1 cells and their reactivity with the murine intestinal microflora. Semin. Immunol. 8:11-18. [DOI] [PubMed] [Google Scholar]
- 20.Lee, S. F. 2003. Oral colonization and immune responses to Streptococcus gordonii: potential use as a vector to induce antibodies against respiratory pathogens. Curr. Opin. Infect. Dis. 16:231-235. [DOI] [PubMed] [Google Scholar]
- 21.Medaglini, D., G. Pozzi, T. P. King, and V. A. Fischetti. 1995. Mucosal and systemic immune responses to a recombinant protein expressed on the surface of the oral commensal bacterium Streptococcus gordonii after oral colonization. Proc. Natl. Acad. Sci. USA 92:6868-6872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Mestecky, J., and J. R. McGhee. 1987. Immunoglobulin A (IgA): molecular and cellular interactions involved in IgA synthesis and immune response. Adv. Immunol. 40:153-245. [DOI] [PubMed] [Google Scholar]
- 23.Murakami, M., T. Tsubata, R. Shinkura, S. Nisitani, M. Okamoto, H. Yoshioka, T. Usui, S. Miyawaki, and T. Honjo. 1994. Oral administration of lipopolysaccharides activates B-1 cells in the peritoneal cavity and lamina propria of the gut and induces autoimmune symptoms in an autoantibody transgenic mouse. J. Exp. Med. 180:111-121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Putnins, E. E., and G. H. Bowden. 1993. Antigenic relationships among oral Actinomyces isolates, Actinomyces naeslundii genospecies 1 and 2, Actinomyces howellii, Actinomyces denticolens, and Actinomyces slackii. J. Dent. Res. 72:1374-1385. [DOI] [PubMed] [Google Scholar]
- 25.Quan, C. P., A. Berneman, R. Pires, S. Avrameas, and J.-P. Bouvet. 1997. Natural polyreactive secretory immunoglobulin A autoantibodies as a possible barrier to infection in humans. Infect. Immun. 65:3997-4004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ruhl, S., A. L. Sandberg, M. F. Cole, and J. O. Cisar. 1996. Recognition of immunoglobulin A1 by oral actinomyces and streptococcal lectins. Infect. Immun. 64:5421-5424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sarkonen, N., E. Kononen, P. Summanen, A. Kanervo, A. Takala, and H. Jousimies-Somer. 2000. Oral colonization with Actinomyces species in infants by two years of age. J. Dent. Res. 79:864-867. [DOI] [PubMed] [Google Scholar]
- 27a.Scholler, M., J. P. Klein, and R. M. Frank. 1981. Common antigens of streptococcal and nonstreptococcal oral bacteria: immunochemical studies of extracellular and cell wall-associated antigens from Streptococcus sanguis, Streptococcus mutans, Lactobacillus salivarius, and Actinomyces viscosus. Infect. Immun. 31:52-60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Shroff, K. E., K. Meslin, and J. J. Cebra. 1995. Commensal enteric bacteria engender a self-limiting humoral mucosal immune response while permanently colonizing the gut. Infect. Immun. 63:3904-3913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Syed, S. A., and W. J. Loesche. 1972. Survival of human dental plaque bacteria in various transport media. Appl. Microbiol. 24:638-644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Thole, J. E. R., P. J. van Dalen, C. E. G. Havenith, P. H. Pouwels, J. F. M. L. Seegers, F. D. Tielen, M. D. van der Zee, N. D. Zegers, and M. Shaw. 2000. Live bacterial delivery systems for development of mucosal vaccines. Curr. Opin. Mol. Ther. 2:94-99. [PubMed] [Google Scholar]
- 31.Ward, J. H. 1963. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 58:236-344. [Google Scholar]
- 32.Zylber, L., and H. V. Jordan. 1982. Development of a selective medium for detection and enumeration of Actinomyces viscosus and Actinomyces naeslundii in dental plaque. J. Clin. Microbiol. 15:253-259. [DOI] [PMC free article] [PubMed] [Google Scholar]