Abstract
Molecular methods for the rapid identification of methicillin-resistant Staphylococcus aureus (MRSA) are generally based on the detection of an S. aureus-specific gene target and the mecA gene. However, such methods cannot be applied for the direct detection of MRSA from nonsterile specimens such as nasal samples without the previous isolation, capture, or enrichment of MRSA because these samples often contain both coagulase-negative staphylococci (CoNS) and S. aureus, either of which can carry mecA. In this study, we describe a real-time multiplex PCR assay which allows the detection of MRSA directly from clinical specimens containing a mixture of staphylococci in <1 h. Five primers specific to the different staphylococcal cassette chromosome mec (SCCmec) right extremity sequences, including three new sequences, were used in combination with a primer and three molecular beacon probes specific to the S. aureus chromosomal orfX gene sequences located to the right of the SCCmec integration site. Of the 1,657 MRSA isolates tested, 1,636 (98.7%) were detected with the PCR assay, whereas 26 of 569 (4.6%) methicillin-susceptible S. aureus (MSSA) strains were misidentified as MRSA. None of the 62 nonstaphylococcal bacterial species or the 212 methicillin-resistant or 74 methicillin-susceptible CoNS strains (MRCoNS and MSCoNS, respectively) were detected by the assay. The amplification of MRSA was not inhibited in the presence of high copy numbers of MSSA, MRCoNS, or MSCoNS. The analytical sensitivity of the PCR assay, as evaluated with MRSA-negative nasal specimens containing a mixture of MSSA, MRCoNS, and MSCoNS spiked with MRSA, was ∼25 CFU per nasal sample. This real-time PCR assay represents a rapid and powerful method which can be used for the detection of MRSA directly from specimens containing a mixture of staphylococci.
Staphylococcus aureus is a major pathogen that causes a wide spectrum of clinical manifestations, such as wound infections, pneumonia, septicemia, and endocarditis. Beta-lactam antimicrobial agents are the preferred drugs for serious S. aureus infections. However, since the introduction of methicillin into clinical use, methicillin-resistant S. aureus (MRSA) strains have emerged worldwide as important nosocomial pathogens, and the prevalence of these strains in the community is now increasing substantially (6, 10, 18).
Methicillin resistance in S. aureus is caused by the acquisition of an exogenous gene, mecA, that encodes an additional β-lactam-resistant penicillin-binding protein (PBP), termed PBP 2a (or PBP2′) (15). The mecA gene is carried by a mobile genetic element, designated staphylococcal cassette chromosome mec (SCCmec), inserted near the chromosomal origin of replication (32). SCCmec is characterized by the presence of terminal inverted and direct repeats, a set of site-specific recombinase genes (ccrA and ccrB), and the mec gene complex (16). The SCCmec DNAs are integrated at a specific site (attBscc) in the methicillin-susceptible S. aureus (MSSA) chromosome which is located at the 3′ end of an open reading frame (ORF), orfX, of unknown function (16).
The rapid and accurate identification of MRSA in clinical specimens has important implications for the therapy and management of both colonized and infected patients. Numerous molecular approaches that reduce the time for identification of MRSA have been described (3, 12, 13, 26, 30, 35, 42, 46, 48, 49). However, the molecular tests developed to date for the detection of MRSA are based on the detection of an S. aureus-specific gene and/or mecA. Thus, they cannot be applied for the direct detection of MRSA from nonsterile specimens such as nasal samples without the previous isolation, capture, or enrichment of MRSA because these samples often contain both coagulase-negative staphylococci (CoNS) and S. aureus, either of which can carry mecA (1).
In this study, we describe a real-time multiplex PCR assay that is useful for the detection of MRSA directly from specimens containing a mixture of staphylococci. This assay comprises five primers specific to the various SCCmec right extremity (SRE) sequences, including three new sequences, in combination with a primer and three molecular beacon probes (MBPs) specific to the S. aureus chromosomal orfX gene located to the right of the SCCmec integration site. We have validated this real-time PCR assay by using a variety of gram-negative and gram-positive bacterial species as well as strains of MSSA, MRSA, methicillin-susceptible CoNS (MSCoNS), and methicillin-resistant CoNS (MRCoNS) from various countries. The assay was also used to detect MRSA directly from nasal specimens.
MATERIALS AND METHODS
Bacterial strains.
The reference strains used for this study are listed in Tables 1 and 2. The staphylococcal clinical isolates used for this study are part of the SENTRY program collection and several supplier's collections. The S. aureus clinical isolates originated from Africa (n = 15) as well as the following 29 countries: Albania (n = 2), Argentina (n = 50), Australia (n = 71), Austria (n = 2), Belgium (n = 3), Brazil (n = 78), Canada (n = 601), Chile (n = 42), China (n = 70), Denmark (n = 33), Egypt (n = 1), France (n = 36), Germany (n = 27), Greece (n = 4), Ireland (n = 5), Israel (n = 19), Italy (n = 61), Japan (n = 62), Mexico (n = 1), The Netherlands (n = 179), Poland (n = 33), Portugal (n = 76), Singapore (n = 20), Slovenia (n = 1), Spain (n = 30), Switzerland (n = 13), Turkey (n = 28), United Kingdom (n = 10), and United States (n = 535). The CoNS clinical isolates originated from the following eight countries: Argentina (n = 13), Canada (n = 95), China (n = 7), Denmark (n = 21), France (n = 4), Japan (n = 28), United Kingdom (n = 44), and United States (n = 37). Confirmation of the identification of the staphylococcal strains was done by use of the MicroScan WalkAway Panel Type Positive Breakpoint Combo 13 system when required (Dade Behring Canada Inc., Mississauga, Ontario, Canada).
TABLE 1.
Reference S. aureus strains used for this studya
| Source or strain no. |
|---|
| HARMONY collection of European epidemic MRSA strains (type designation)b |
| 96158 (B) |
| 97117 (A) |
| 97118 (A) |
| 97120 (B) |
| 97151 (B) |
| 97392 (B) |
| 97393 (A) |
| BM10827 (C) |
| 162 (A) |
| 920 (B) |
| 95035 (A) |
| 97121 (B) |
| BM10828 (C) |
| BM10882 (C) |
| 97S97 (Belgian epidemic clone 1a) |
| 98S46 (Belgian epidemic clone 3b) |
| 97S96 (Belgian epidemic clone 1a) |
| 97S98 (Belgian epidemic clone 1b) |
| 97S99 (Belgian epidemic clone 2a) |
| 97S100 (Belgian epidemic clone 2b) |
| 97S101 (Belgian epidemic clone 3a) |
| 359/96 (Berlin epidemic EMRSA IVc) |
| 792/96 (Berlin epidemic EMRSA IVd) |
| 844/96 (Berlin epidemic EMRSA IVb) |
| 1966/97 (Hannover area EMRSA IIIc) |
| 2594-2/97 (S. German EMRSA IIb) |
| 131/98 (S. German EMRSA II d2) |
| 406/98 (N. German EMRSA I c1) |
| 408/98 (N. German EMRSA I c2) |
| 61974 (Helsinki I E1) |
| 62176 (Kotka E10) |
| 62305 (mecA- Tampere I E12) |
| 62396 (Helsinki II E2) |
| 75541 (Tampere II E13) |
| 75916 (Helsinki V E5) |
| 76167 (Kemi E17) |
| 98442 (Helsinki VI E19) |
| 98514 (Helsinki VII E20) |
| 98541 (Lohja E24) |
| 3717 (EMRSA-GR1b) |
| 3680 (EMRSA-GR1) |
| 3713 (EMRSA-GR1a) |
| HS 2 (I) |
| AO 17934/97 (II) |
| N8-890/99 (Sa 543 VI) |
| N8-3756/90 (Sa544 I) |
| 9805-01937 (V) |
| AK 541 (IV) |
| ON 408/99 (VII) |
| AO 9973/97 (III) |
| 98/10618 (EMRSA-15/b2) |
| 98/26821 (EMRSA-15/b3) |
| 98/24344 (EMRSA-15/b7) |
| 99/1139 (EMRSA-16/a2) |
| 99/159 (EMRSA-16/a14) |
| M307 (EMRSA-3) |
| 90/10685 (EMRSA-15) |
| 98/14719 (EMRSA-15/b4) |
| 872/98 (Hannover area EMRSA IIIb) |
| 1155-1/98 (S. German EMRSA II c) |
| 1163/98 (S. German EMRSA II d1) |
| 1869/98 (N. German EMRSA I d) |
| 134/93 (N. German EMRSA I) |
| 1000/93 (Hannover area EMRSA III) |
| 1450/94 (N. German EMRSA I a) |
| 825/96 (Berlin epidemic EMRSA IV) |
| 842/96 (Berlin epidemic EMRSA IV a) |
| 2594-1/97 (S. German EMRSA II a) |
| 1155-2/98 (S. German EMRSA II) |
| 1442/98 (Hannover area EMRSA III a) |
| 234/95 (N. German EMRSA I b) |
| 37481 (Seinajoki E 14) |
| 54511 (Turku I E6) |
| 54518 (Turku II E7) |
| 96/32010 (EMRSA-16) |
| 99/579 (EMRSA-16/a3) |
| 5 (E1) |
| 6 (D) |
| 13 (A′) |
| 14 (A′) |
| 18 (A) |
| 20 (E) |
| 25 (FI) |
| 30 (G) |
| 33 (F) |
| 54 (B) |
| 60 (A̋) |
| 80 (E) |
| 98 (C) |
| Public collections (type designation) |
| ATCC 6538c |
| ATCC 13301c |
| ATCC 23235c |
| ATCC 25923c |
| ATCC 27660c |
| ATCC 29737c |
| ATCC 29213c |
| ATCC 29247c |
| ATCC 33591 |
| ATCC 33592 |
| ATCC 33593 |
| ATCC 43300 |
| ATCC BAA-38 (Archaic clone of MRSA)d |
| ATCC BAA-39 (Hungarian clone of MRSA)d |
| ATCC BAA-40 (Portuguese clone of MRSA)d |
| ATCC BAA-41 (New York clone of MRSA)d |
| ATCC BAA-42 (Pediatric clone of MRSA)d |
| ATCC BAA-43 (Brazilian clone of MRSA)d |
| ATCC BAA-44 (Iberian clone of MRSA)d |
| CCUG 41787 (Sa 501 V)b |
| CCUG 38266 (II)b |
| NCTC 8325c |
| NCTC 11939 (EMRSA-1)b |
| Canadian epidemic MRSA strainse |
| CMRSA-1 |
| CMRSA-2 |
| CMRSA-3 |
| CMRSA-4 |
| CMRSA-5 |
| CMRSA-6 |
All S. aureus strains are resistant to oxacillin except where otherwise indicated.
Information on these strains and type designations based on pulsed-field gel electrophoresis are from http://www.phls.co.uk/inter/harmony/menu.htm.
Susceptible to oxacillin (MSSA).
Information on these strains and type designations based on pulsed-field gel electrophoresis are from reference 7.
Information on these strains and type designations based on pulsed-field gel electrophoresis are from reference 47.
TABLE 2.
Reference strains other than S. aureus used for this study
| Bacterial group or strain |
|---|
| CoNS |
| S. arlettae ATCC 43957 (MSCoNS) |
| S. auricularis ATCC 33753 (MSCoNS) |
| S. capitis subsp. capitis ATCC 27840 (MSCoNS) |
| S. capitis subsp. ureolyticus ATCC 49326 (MSCoNS) |
| S. caprae ATCC 35538 (MSCoNS) |
| S. carnosus subsp. carnosus ATCC 51365 (MSCoNS) |
| S. chromogenes ATCC 43764 (MSCoNS) |
| S. cohnii subsp. cohnii ATCC 29974 (MSCoNS) |
| S. delphini ATCC 49171 (MSCoNS) |
| S. epidermidis ATCC 12228 (MSCoNS) |
| S. epidermidis ATCC 14990 (MSCoNS) |
| S. epidermidis ATCC 35983 (MRCoNS) |
| S. epidermidis ATCC 35984 (MRCoNS) |
| S. equorum ATCC 43958 (MSCoNS) |
| S. felis ATCC 49168 (MSCoNS) |
| S. gallinarum ATCC 35539 (MSCoNS) |
| S. haemolyticus ATCC 29970 (MSCoNS) |
| S. hominis subsp. hominis ATCC 27844 (MSCoNS) |
| S. hominis subsp. hominis ATCC 35982 (MSCoNS) |
| S. hyicus ATCC 11249 (MSCoNS) |
| S. intermedius ATCC 29663 (MSCoNS) |
| S. kloosii ATCC 43959 (MSCoNS) |
| S. lentus ATCC 29070 (MSCoNS) |
| S. lugdunensis ATCC 43809 (MSCoNS) |
| S. pasteuri ATCC 51129 (MSCoNS) |
| S. pulvereri ATCC 51698 (MSCoNS) |
| S. saccharolyticus ATCC 14953 (MSCoNS) |
| S. saprophyticus ATCC 43867 (MSCoNS) |
| S. saprophyticus ATCC 35552 (MSCoNS) |
| S. saprophyticus ATCC 15305 (MSCoNS) |
| S. schleiferi subsp. coagulans ATCC 49545 (MSCoNS) |
| S. schleiferi subsp. schleiferi ATCC 43808 (MSCoNS) |
| S. sciuri subsp. sciuri ATCC 29060 (MRCoNS) |
| S. sciuri subsp. sciuri ATCC 29062 (MSCoNS) |
| S. simulans ATCC 27848 (MSCoNS) |
| S. warneri ATCC 27836 (MSCoNS) |
| S. xylosus ATCC 29971 (MRCoNS) |
| Nonstaphylococcal gram-positive bacteria |
| Abiotrophia defectiva ATCC 49176 |
| Bacillus cereus ATCC 13472 |
| Corynebacterium bovis ATCC 7715 |
| Corynebacterium cervicis NCTC 10604 |
| Corynebacterium flavescens ATCC 10340 |
| Corynebacterium genitalium ATCC 33031 |
| Enterococcus avium ATCC 14025 |
| Enterococcus casseliflavus ATCC 25788 |
| Enterococcus cecorum ATCC 43198 |
| Enterococcus dispar ATCC 51266 |
| Enterococcus durans ATCC 19432 |
| Enterococcus faecalis ATCC 19433 |
| Enterococcus faecium ATCC 19434 |
| Enterococcus flavescens ATCC 49996 |
| Enterococcus gallinarum ATCC 49573 |
| Enterococcus hirae ATCC 8043 |
| Enterococcus mundtii ATCC 43186 |
| Enterococcus pseudoavium ATCC 49372 |
| Enterococcus raffinosus ATCC 49427 |
| Enterococcus saccharolyticus ATCC 43076 |
| Enterococcus solitarius ATCC 49428 |
| Gemella haemolysans ATCC 10379 |
| Kocuria rhizophila ATCC 9341 |
| Lactobacillus crispatus ATCC 33820 |
| Leifsonia aquatica ATCC 14665 |
| Listeria monocytogenes ATCC 15313 |
| Streptococcus agalactiae ATCC 13813 |
| Streptococcus anginosus ATCC 33397 |
| Streptococcus bovis ATCC 33317 |
| Streptococcus constellatus subsp. constellatus ATCC 27823 |
| Streptococcus cristatus ATCC 51100 |
| Streptococcus intermedius ATCC 27335 |
| Streptococcus mitis ATCC 49456 |
| Streptococcus mutans ATCC 25175 |
| Streptococcus parasanguinis ATCC 15912 |
| Streptococcus pneumoniae ATCC 6303 |
| Streptococcus pyogenes ATCC 19615 |
| Streptococcus salivarius ATCC 7073 |
| Streptococcus sanguinis ATCC 10556 |
| Streptococcus suis ATCC 43765 |
| Gram-negative bacteria |
| Acinetobacter baumannii ATCC 19606 |
| Bordetella pertussis ATCC 9797 |
| Burkholderia cepacia ATCC 25416 |
| Citrobacter freundii ATCC 8090 |
| Enterobacter cloacae ATCC 13047 |
| Escherichia coli ATCC 25922 |
| Haemophilus influenzae ATCC 9007 |
| Hafnia alvei ATCC 13337 |
| Klebsiella pneumoniae ATCC 13883 |
| Moraxella catarrhalis ATCC 43628 |
| Neisseria gonorrhoeae ATCC 35201 |
| Neisseria meningitidis ATCC 13077 |
| Pasteurella aerogenes ATCC 27883 |
| Proteus mirabilis ATCC 25933 |
| Providencia stuartii ATCC 33672 |
| Pseudomonas aeruginosa ATCC 35554 |
| Pseudomonas fluorescens ATCC 13525 |
| Salmonella choleraesuis subsp. choleraesuis ATCC 14028 |
| Serratia marcescens ATCC 13880 |
| Shigella sonnei ATCC 29930 |
| Stenotrophomonas maltophilia ATCC 13637 |
| Yersinia enterocolitica ATCC 9610 |
MIC determination.
The MICs for oxacillin were either provided by the strain suppliers or determined in our laboratory by the Etest method (AB Biodisk, Solna, Sweden) according to the manufacturer's instructions. The results were interpreted according to the standards of the NCCLS (37). S. aureus strains ATCC 29213 and ATCC 43300 were used as quality controls for antimicrobial susceptibility testing.
DNA sequencing.
The primers used to amplify and sequence the SRE junctions (MREJs) of various MRSA strains are listed in Table 3. The MREJ comprises the right extremity of SCCmec, the SCCmec integration site, and the orfX gene.
TABLE 3.
Oligonucleotides used for this study
| Oligonucleotide | Oligonucleotide sequence (5′ → 3′) | Nucleotide position | Amplification product size (bp) |
|---|---|---|---|
| S. aureus orfX-specific primer | |||
| Xsau325 | GGATCAAACGGCCTGCACA | 325a | |
| SCCmec-specific primers | |||
| mecii574 | GTCAAAAATCATGAACCTCATTACTTATG | 472,b 574c | 176d or 278e |
| meciii519 | ATTTCATATATGTAATTCCTCCACATCTC | 519f | 223g |
| meciv511 | CAAATATTATCTCGTAATTTACCTTGTTC | 511h | 215i |
| mecv492 | CTCTGCTTTATATTATAAAATTACGGCTG | 492j | 196k |
| mecvii512 | CACTTTTTATTCTTCAAAGATTTGAGC | 512l | 214m |
| S. aureus orfX-specific probes | |||
| XsauB5-FAMn | CCCGCGCGTAGTTACTGCGTTGTAAGACGTCCGCGGGo | 346a | |
| XsauB8-FAMn | CCCGCGCATAGTTACTGCGTTGTAAGACGTCCGCGGGo | 346p | |
| XsauB9-FAMn | CCCGCGCGTAGTTACTACGTTGTAAGACGTCCGCGGGo | 346q | |
| Internal control probe | |||
| PSARM-TETr | CCGGCGATGCCTCTTCACATTGCTCCACCTTTCCTCGCCGGo | 2,038s | |
| Sequencing primers | |||
| IS431626 | TCTACGGATTTTCGCCATGC | 626t | 1,200u |
| Xsau401 | ATCAAATGATGCGGGTTGTGT | 401a | |
| mecA1059 | AACAGGTGAATTATTAGCACTTGTAAG | 1,059v | >8,000w |
| SA0022Sau2673 | GAGGACCAAACGACATGAAAATC | 2,673x | |
| Internal sequencing primers | |||
| meciv1013 | CAATCGGTATCTGTAAATATCAAAT | 1,013h | |
| meciv1411 | TCGCATACCTGTTTATCTTCTACT | 1,411h | |
| meciv916 | TTGGTTCCATCTGAACTTTGAG | 916h | |
| mecvii1126 | ACTAGAATCTCCAAATGAATCCAGT | 1,126l | |
| mecv1029 | TTTAAATTCAGCTATATGGGGAGA | 1,029j | |
| mecv968 | TTCCGTTTTGCTATTCCATAAT | 968j | |
| tetK1169 | CCTCTGATAAAAAACTTGTGAAAT | 1,169y | |
| tetK136 | ACTACTCCTGGAATTACAAACTGG | 136y | |
| Xsau193 | GCCAAAATTAAACCACAATCCAC | 193a | |
| Xsau367 | CATTTTGCTGAATGATAGTGCGTA | 367a |
Nucleotide position in orfX (start codon = nt 1 to 3) from GenBank accession no. AP003129.
Nucleotide position in SCCmec from the orfX gene (start codon = nt 1 to 3) from GenBank accession no. AB033763.
Nucleotide position in SCCmec from the orfX gene (start codon = nt 1 to 3) from GenBank accession no. AP003129.
Amplification product size generated with primer Xsau325 using DNA from MRSA with MREJ type i.
Amplification product size generated with primer Xsau325 using DNA from MRSA with MREJ type ii.
Nucleotide position in SCCmec from the orfX gene (start codon = nt 1 to 3) from GenBank accession no. AB037671.
Amplification product size generated with primer Xsau325 using DNA from MRSA with MREJ type iii.
Nucleotide position in the SCCmec from the orfX gene (start codon = nt 1 to 3) from GenBank accession no. AY267374.
Amplification product size generated with primer Xsau325 using DNA from MRSA with MREJ type iv.
Nucleotide position in the SCCmec from the orfX gene (start codon = nt 1 to 3) from GenBank accession no. AY267381.
Amplification product size generated with primer Xsau325 using DNA from MRSA with MREJ type v.
Nucleotide position in the SCCmec from the orfX gene (start codon = nt 1 to 3) from GenBank accession no. AY267375.
Amplification product size generated with primer Xsau325 using DNA from MRSA with MREJ type vii.
FAM, 6-carboxylfluorescein.
The underlined sequences constitute the stem of each molecular beacon.
Nucleotide position in orfX (start codon = nt 1 to 3) from GenBank accession no. AB037671.
Nucleotide position in orfX (start codon = nt 1 to 3) from GenBank accession no. AP004822.
TET, tetrachloro-6-carboxylfluorescein.
Nucleotide position from GenBank accession no. D83207.
Nucleotide position in the IS431 transposase gene (start codon = nt 1 to 3) from GenBank accession no. AB037671.
Amplification product size generated with primer Xsau401.
Amplification product size generated with primer Xsau401 or SA0022Sau2673.
Nucleotide position from the SA0022 orf (start codon = nt 1 to 3) from GenBank accession no. AP003129.
Nucleotide position in tetK (start codon = nt 1 to 3) from GenBank accession no. S67449.
The MREJ fragments to be sequenced were amplified on a PTC-200 thermocycler (MJ Research Inc., Watertown, Mass.), using purified genomic DNA. Genomic DNAs were purified by use of a Gnome kit (Qbiogene Inc., Carlsbad, Calif.) according to the manufacturer's instructions. Depending on the MRSA strain, amplification products of 1.2 or >8 kb were obtained. Amplification products of 1.2 kb were amplified as previously described (29), whereas those of >8 kb were amplified with Herculase Enhanced DNA polymerase (Stratagene, La Jolla, Calif.). Purification of the amplification products and sequencing reactions were performed as previously described (29).
Primers and probes.
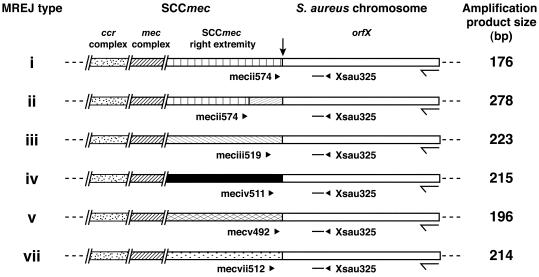
A multiple nucleotide sequence alignment of the MREJ sequences of several staphylococcal species obtained for this study or available from public databases was performed with the Pileup program from the GCG package (version 10; Accelrys, San Diego, Calif.). This alignment allowed the design of a set of PCR primers specific to the various SRE sequences (mecii574, meciii519, meciv511, mecv492, and mecvii512) as well as a primer specific to S. aureus orfX (Xsau325). This set of primers was used in combination with three MBPs (XsauB5-FAM, XsauB8-FAM, and XsauB9-FAM) targeting orfX sequences within the MREJ amplification products, allowing the detection of all S. aureus orfX variants. A fourth MBP (PSARM-TET) was used to detect pSARM internal control amplification products. MBPs were designed as previously described (4). Primers were designed with the help of Oligo Primer Analysis software, version 6.22 (Molecular Biology Insights, Cascade, Colo.). Primers were synthesized with a DNA synthesizer (model 391; PE Applied Biosystems, Foster City, Calif.) and MBPs were obtained from Biosearch Technologies (Novato, Calif.). The PCR primers and probes used for this study are listed in Table 3 and are shown in Fig. 1.
FIG. 1.
Schematic representation of the MREJs of MRSA strains with MREJ types i to vii. Each MREJ type exhibits a distinct SRE sequence, with the exception of MREJ types i and ii, which differ by a 102-bp insertion. The positions of forward (▸) and reverse (◂) primers and of internal probes (bars) included in the MRSA assay are shown. Three MBPs, XsauB5, XsauB8, and XsauB9, were used to detect the MRSA-specific amplification products. The locations within SCCmec of the crr complex and the mec complex containing mecA are shown, but these genetic elements are not drawn to scale. The vertical arrow at the top indicates the SCCmec integration site within the chromosomal S. aureus orfX gene. The half- arrows show the positions of the orfX start codon and the direction of transcription. More details regarding primers and probes are found in Table 3.
Construction of internal control.
An internal control was constructed as described previously (28). A 324-bp DNA fragment consisting of a 276-bp sequence not found in MRSA flanked by the sequences of the meciv511 and mecv492 primers (Table 3) was used as a template for the internal control. This fragment was cloned into the pCR2.1 vector (Invitrogen, Burlington, Ontario, Canada) and isolated as previously described (28). The purified recombinant plasmid, named pSARM, was linearized with BamHI (New England Biolabs, Mississauga, Ontario, Canada) and serially diluted. The concentration of the linearized plasmid was optimized to permit the amplification of the 324-bp internal control product without a significant detrimental effect on MRSA-specific amplification.
PCR assay.
For all bacterial strains, amplification was performed with either purified genomic DNAs or crude DNA extracts prepared from bacterial suspensions whose turbidity was adjusted to that of a 0.5 McFarland standard, which corresponds to approximately 1.5 × 108 bacteria per ml. Crude DNA extracts were prepared for PCRs by a rapid DNA extraction method (Infectio Diagnostic Inc., Ste-Foy, Canada) (28). One microliter of purified genomic DNA or of a crude DNA extract was transferred directly to a 24-μl PCR mixture containing oligonucleotide primers and probes (Table 3), four deoxyribonucleoside triphosphates (Pharmacia Biotech, Baie d'Urfé, Québec, Canada), a 10 mM Tris-HCl buffer (pH 9.0), 4.75 mM MgCl2, 2.5 mg of bovine serum albumin per ml, and 1.25 U of Taq DNA polymerase (Roche, Mississauga, Ontario, Canada) combined with a TaqStart antibody (Clontech Laboratories, Palo Alto, Calif.). The internal control (pSARM) was included in the PCR assay to ensure that there was no significant PCR inhibition by the test sample. The thermal cycling protocol was as follows: 3 min at 95°C for initial denaturation followed by 48 cycles of three steps consisting of 5 s at 95°C for denaturation, 15 s at 60°C for annealing, and 20 s at 72°C for extension. Real-time detection of the PCR products was performed on a Smart Cycler (Cepheid, Sunnyvale, Calif.) by measuring the fluorescence signal emitted by the MBPs hybridized to their targets at the end of each annealing step. The specificity and ubiquity (i.e., the ability to detect all or most MRSA strains) (5) of the PCR assay were verified by use of a panel of 22 gram-negative bacterial species, 40 gram-positive nonstaphylococcal bacterial species, 14 species (212 strains) of MRCoNS, 27 species (74 strains) of MSCoNS, 569 MSSA isolates, and 1,657 MRSA strains from various geographic areas. For determination of the analytical sensitivity of the PCR assay, 1-μl samples of serial twofold dilutions of purified genomic DNA (ranging from 64 to 1 genome copy per μl) from MRSA strains of different MREJ types were used to determine the minimal number of genomes which were detected. To verify that MRSA amplification was not inhibited in the presence of DNAs from MSSA, MRCoNS, and MSCoNS, we performed the following experiment. The equivalent of 10 genome copies of DNA from MRSA strain ATCC 43300 (carrying MREJ type ii) per PCR was amplified by the real-time PCR assay in the presence of 0, 10, 102, 103, and 104 genome copies of DNA (per PCR) from either (i) MSSA strain ATCC 29213 (MREJ negative), (ii) methicillin-resistant Staphylococcus epidermidis strain ATCC 35983 (carrying MREJ type ii), or (iii) methicillin-susceptible S. epidermidis strain ATCC 14990 (MREJ negative).
MREJ and SCCmec typing.
The MREJ types of the MRSA strains described in this study were determined by sequence analysis or by examining the PCR amplification products generated by the multiplex PCR assay described above by standard agarose gel electrophoresis (44). The SCCmec types and subtypes of the MRSA strains described in this study were identified by using a previously described typing method (38). Primers mecIVc70 (5′-TGGGGTATTTTTATCTTCAACTC-3′) and mecIVc1079 (5′-TGGGATTTTAAAGCAGAATATCA-3′) were designed to identify SCCmec type IVc based on the SCCmec sequence of MRSA strain MR108 (25). Primers mecIVd26 (5′-ACGGGAGATTAGGAGATGTTAT-3′) and mecIVd307 (5′-CAGCCATCAATTTTGTTTCACC-3′) were designed to identify SCCmec type IVd based on the SCCmec sequence of MRSA strain JCSC 4469 (GenBank accession number AB097677).
Evaluation of the sensitivity of the PCR assay by using nasal samples.
The minimal number of CFU that can be detected in nasal specimens was evaluated by using 18 nasal swabs obtained from nine volunteers (2 nasal swabs/volunteer) who were not colonized by MRSA. Nasal specimens were collected with a collection and transport system for aerobes (Venturi Transystem; Copan Canada, Richmond Hill, Ontario, Canada). A swab was carefully inserted a short distance into each nostril and gently rotated for 5 s. Swabs were inserted into the transport medium immediately after samples were obtained. Each swab was streaked directly on mannitol salt agar to determine the presence of MSSA, MRSA, MRCoNS, or MSCoNS. The agar medium was examined after 1 and 2 days of incubation for typical staphylococcal colonies. The identification of suspicious staphylococcal colonies was based on catalase production, slide agglutination (Staphaurex; Murex Biotech Limited), and the tube coagulase test. Identification of and oxacillin MIC determinations for the staphylococcal isolates were performed as described above. None of the specimens contained MRSA isolates, whereas 10 specimens contained MSCoNS isolates, 4 specimens contained MRCoNS isolates, and 4 specimens contained both MSSA and MSCoNS isolates. Nasal specimens from the 18 swabs were resuspended in buffer and pooled. The pooled MRSA-negative specimens, which contained a mixture of MSSA, MRCoNS, and MSCoNS, were divided into 18 aliquots. The samples were spiked with serial 10-fold dilutions of MRSA strain ATCC 43300 during the logarithmic phase of growth (optical density at 600 nm of ∼0.6) in phosphate-buffered saline. Each 10-fold dilution was added directly to the clinical specimens and processed prior to PCR amplification by a rapid DNA extraction method (28). The number of CFU was estimated by standard plating procedures.
Nucleotide sequence accession numbers.
GenBank accession numbers for the S. aureus MREJ sequences are as follows: for strain ATCC 33592, AY267373; for strain ATCC BAA-40, AY267374; for strain CMRSA-1, AY267375; for strain CCRI-1263, AY267376; for strain CCRI-1311, AY267377; for strain CCRI-1331, AY267379; for strain CCRI-1377, AY267380; for strain CCRI-2025, AY267381; for strain CCRI-8895, AY267382; for strain CCRI-8903, AY267383; and for strain CCRI-9583, AY267384.
RESULTS
New SRE sequences of MRSA.
A molecular method named mec right extremity polymorphism (MREP) typing, which takes advantage of the polymorphism among SRE sequences, was previously developed (17, 23). This method combines primers specific to each distinct SRE sequence with a primer specific to the S. aureus chromosome located to the right of the SCCmec integration site to detect the different SCCmec types encountered. MREP types i, ii, and iii were defined according to the respective SCCmec types I, II, and III (17, 23). By using this strategy, we have developed a new set of primers for the specific identification of MRSA by using the known SRE sequences of SCCmec types I, II, III, IVa, IVb, and IVc (2, 23-25, 32, 34). The first primer, mecii574, recognizes the right extremity sequences of SCCmec types I, II, IVa, IVb, and IVc. The right extremities of SCCmec types II, IVa, IVb, and IVc are identical but differ from the right extremity of SCCmec type I by a 102-bp insertion (2, 23-25, 32, 34). The second primer, meciii519, was specific to the right extremity sequence of SCCmec type III. These two primers were used in combination with a primer specific to the S. aureus orfX gene, located to the right of the SCCmec integration site (Xsau325). We used this set of primers to amplify the MREJs of a variety of MRSA strains. We found that 15 of 206 MRSA strains tested could not be amplified with this assay (data not shown). The MREJs of 11 of these 15 strains were sequenced in order to establish the new sequences at the right extremity of SCCmec to allow the development of a PCR assay to detect more MRSA strains. The MREJ sequences obtained for these 11 MRSA strains were compared to sequences available from public databases.
Portions of the MREJ sequence corresponding to the orfX gene for these 11 MRSA strains had identities ranging from 97 to 100% with the publicly available orfX sequences of S. aureus and from 79.1 to 82.5% with those of other staphylococcal species (S. epidermidis, S. hominis, and S. haemolyticus). The characteristic right chromosome-SCCmec junction attL sequence, which comprises the inverted repeat IRscc-R and the direct repeat DRscc (23), was present within the 3′ end of the orfX gene for all 11 strains. However, the DNA sequence within the right extremity of SCCmec was shown to be very different from those of types I, II, III, IVa, IVb, and IVc which were previously described (2, 23-25, 32, 34). Three novel SRE sequences were characterized. These sequences were designated MREJ types iv, v, and vii. The MREJs comprising distinct MREP types were named according to the previously described MREP numbering scheme (17, 23). Hence, MREP type i comprises MREJ type i, MREP type ii comprises MREJ type ii, and so on, up to MREP type vii.
The SRE sequences obtained for strains ATCC 33592, ATCC BAA-40, CCRI-1331, CCRI-8895, and CCRI-8903 were nearly identical to each other and exhibited 100% identity with a DNA fragment found within SCCmec type III of MRSA strain 85/2082, which includes the sequence located between the second and third IS431 copies, the inverted repeat sequence of IS431, and part of the IS431 transposase gene sequence (23). However, our sequence data revealed the location of this fragment at the right extremity of SCCmec to be adjacent to the DRscc. These new sequences were designated MREJ type iv because the SRE sequences for these five MRSA strains were different from those of SCCmec type I from MRSA strain NCTC 10442, SCCmec type II from MRSA strain N315, SCCmec type III from MRSA strain 85/2082, SCCmec type IVa from MRSA strain CA05, SCCmec type IVb from MRSA strain 8/6P, and SCCmec type IVc from MRSA strain MR108 (23, 25, 34).
The SRE sequences obtained for strains CCRI-1263, CCRI-1311, CCRI-1377, and CCRI-2025 were nearly identical to each other, were different from those of all four SCCmec types and MREJ type iv, and consequently, were designated MREJ type v. The SRE sequence of MREJ type v did not show any significant homology with any published sequences when compared with publicly available sequences by use of BLAST.
The SRE sequences obtained for strains CCRI-9583 and CMRSA-1 were also different from those of all four SCCmec types and MREJ types iv and v, and consequently, were designated MREJ type vii. Upon a BLAST search, the SRE sequence of MREJ type vii was also shown to be unique, exhibiting no significant homology to any published sequences.
Four of the 15 MRSA strains that were not detected by the PCR assay were not sequenced, as they were shown to carry either MREJ type v or vii based on agarose gel electrophoresis analysis of the amplification products.
Comparison of MREJ and SCCmec types of MRSA strains.
The MREJ types defined for several MRSA strains in this study were compared with the SCCmec types established for these strains by using a previously described SCCmec typing method (38). As shown in Table 4, there was no correlation between the MREJ and SCCmec types. The MRSA strain with MREJ type i had SCCmec type I and those with MREJ type iii all had SCCmec type III. It was not possible to identify the SCCmec type of the epidemic strain ATCC BAA-42, which carries MREJ type ii. This strain was shown to carry a class B mec gene complex. However, primers targeting the ccrA and ccrB genes used for this typing method (38) did not generate any amplification products. All other MRSA strains with MREJ type ii carried SCCmec type I, II, or IVd. The two MRSA strains with the new MREJ type vii defined in this study carried SCCmec type II, whereas MRSA strains with the new MREJ type v had either SCCmec type IVa or IVc. MRSA strains with the new MREJ type iv carried SCCmec type III or IVa.
TABLE 4.
Comparison of MREJ and SCCmec typing for the MRSA strains described in this study, including well-known epidemic clones
| MRSA strain number | MREJ type | SCCmec type |
|---|---|---|
| ATCC BAA-38 | i | I |
| CMRSA-2 | ii | II |
| CMRSA-4 | ii | II |
| CMRSA-5 | ii | IVda |
| ATCC BAA-41 | ii | II |
| ATCC BAA-42 | ii | NDb |
| ATCC BAA-44 | ii | I |
| ATCC BAA-39 | iii | III |
| ATCC BAA-43 | iii | III |
| CMRSA-3 | iii | III |
| ATCC 33592 | iv | III |
| ATCC BAA-40 | iv | III |
| CCRI-1331 | iv | IVa |
| CCRI-8895 | iv | III |
| CCRI-8903 | iv | III |
| CCRI-1263 | v | IVcc |
| CCRI-1311 | v | IVa |
| CCRI-1377 | v | IVa |
| CCRI-2025 | v | IVa |
| CMRSA-1 | vii | II |
| CCRI-9583 | vii | II |
SCCmec subtype IVd was defined by using primers mecIVd26 and mecIVd307 as described in Materials and Methods.
It was not possible to define the SCCmec type of this strain by using the previously described ccrA- and ccrB-specific primers (39). This strain carries novel ccrA and ccrB gene sequences (39).
SCCmec subtype IVc was defined by using primers mecIVc70 and mecIVc1079 as described in Materials and Methods.
Analytical sensitivity and specificity of real-time multiplex PCR assay.
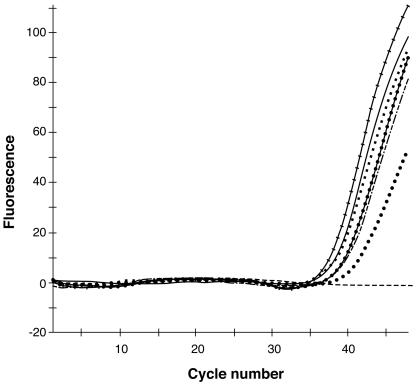
By using the three new SRE sequences for MRSA, we developed a real-time multiplex PCR assay containing six primers and four probes. The detection limit of the assay was determined by using genomic DNAs purified from 24 MRSA strains of different MREJ types and was found to be two to eight genome copies per PCR (Fig. 2). The specificity of the PCR assay was tested by using 40 nonstaphylococcal gram-positive bacterial species, 22 gram-negative bacterial species, 212 MRCoNS isolates, 74 MSCoNS isolates, and 569 MSSA strains from various geographic areas (Tables 1, 2, and 5). None of the nonstaphylococcal MRCoNS and MSCoNS strains tested cross-reacted with the PCR assay. Among the 569 MSSA strains tested, 26 (4.6%) were misidentified as MRSA based on the PCR assay. The ubiquity of the PCR assay was tested with a variety of MRSA strains originating from various geographic areas, including well-known epidemic clones (Tables 1 and 5). Of the 1,657 MRSA strains tested, 1,636 (98.7%) were specifically detected by the PCR assay. Only 1.3% of these MRSA strains, representing a broad variety of origins, were not detected by the assay. An agarose gel analysis of the PCR amplification products revealed that there was no amplification of the genomic DNAs from these 21 strains (data not shown). The amplification of MRSA (∼10 genome copies per PCR) was not inhibited in the presence of increasing concentrations (up to 104 genome copies per PCR) of MSSA, MRCoNS, or MSCoNS (data not shown).
FIG. 2.
Example showing the FAM fluorescence detection of MRSA, using 10 copies of genomic DNAs purified from MRSA strains with MREJ types i (solid line), ii (dashed line with dots), iii (circles), iv (solid line with circles), v (solid line with hash marks), and vii (squares). Dashed line, negative control.
TABLE 5.
Evaluation of MRSA PCR assay using DNAs from a variety of methicillin-susceptible and methicillin-resistant staphylococcal strains
| Staphylococcal strain type (no. of strains)a | No. (%) with PCR result
|
|
|---|---|---|
| Positive | Negative | |
| MRSA (1,657) | 1,636 (98.7) | 21 (1.3) |
| MSSA (569) | 26 (4.6) | 543 (95.4) |
| MRCoNS (212) | 0 | 212 (100) |
| MSCoNS (74) | 0 | 74 (100) |
Detection of MRSA directly from nasal specimens.
The detection limit of the PCR assay with nasal specimens was evaluated by using 18 MRSA-negative nasal specimens obtained from volunteers. The MRSA-negative specimens, which contained a mixture of MSSA, MRCoNS, and MRCoNS, were spiked with serial 10-fold dilutions of MRSA in the logarithmic phase of growth. The detection limit of the PCR assay was ∼25 CFU of MRSA per nasal swab. There was no PCR inhibition for the 18 nasal samples tested based on the amplification efficiency of the internal control (data not shown).
DISCUSSION
The emergence of methicillin resistance in S. aureus is of great concern, as MRSA strains are often multidrug resistant (6). Infections with MRSA are known to be associated with considerable morbidity and mortality (8). Many studies have shown that effective control measures, including the systematic screening of persons exposed to MRSA, can confine or even eliminate the nosocomial spread of MRSA (31, 36, 41, 43, 50). However, standard culture methods for the identification of S. aureus and the determination of oxacillin susceptibility are time-consuming, usually requiring 2 to 4 days. For these reasons, it has become important to develop rapid diagnostic tests for the detection of MRSA.
The molecular detection of MRSA directly from clinical specimens containing a mixture of staphylococci, such as screening swabs from anterior nares, represents an important challenge for the rapid detection of MRSA carriers (26). To overcome this challenge, we have developed a multiplex PCR assay which provides a link between mecA and the S. aureus chromosome. This assay is based on the integration site of the SCCmec DNA containing mecA in the S. aureus chromosome and uses a previously described strategy (17, 23). By using this PCR strategy, we found that 15 of 206 MRSA strains were not detected, suggesting that new SCCmec or new SRE sequences were present in these strains. Nucleotide sequencing of the MREJs found in these MRSA strains revealed three new SRE sequences, designated MREJ types iv, v, and vii. The SRE sequences with MREJ types v and vii did not show any significant homology with sequences in databases, whereas those with MREJ type iv exhibited nearly 100% identity with a portion of the SCCmec type III carried by MRSA strain 85/2082 (23). This SRE sequence was also recently found at the right extremity of the SCCmec of MRSA strain HDG2 (39).
It was possible to assign an SCCmec type for most of the MRSA strains with the new MREJ types described in this study, thereby showing that strains with new SRE sequences do not carry a new SCCmec but rather have structural variations at the SCCmec right extremity. We found that MRSA strains with the new MREJ types v and vii carried SCCmec types IV (IVa or IVc) and II, respectively, whereas MRSA strains with the new MREJ type iv had SCCmec type III or IVa. It has been shown that SCCmec type III of MRSA strain 85/2082 comprises two separate SCCmec or SCC elements that were sequentially integrated in the chromosome. This was assessed by the presence of a 15-bp direct repeat sequence between the second and third copies of IS431 within the SCCmec of this strain (23). A region identical to the one found between the second and third copies of IS431 of SCCmec type III of MRSA strain 85/2082 is present at the right extremity of SCCmec of the MRSA strains with MREJ type iv carrying SCCmec type III described in this study. This suggests that this genetic arrangement may be the result of an excision of the second SCC element in MRSA strains carrying MREJ type iv or that MRSA strains with MREJ type iv may be the ancestral recipients of the second SCC element. SCCmec types I, II, and III have been typically associated with MREJ types i, ii, and iii, respectively (17, 23). Surprisingly, the prototype of the MRSA Iberian clone of MRSA ATCC BAA-44, which was shown to have SCCmec type I, carries MREJ type ii. Another exception is MRSA strain 93/H44, which was shown to carry SCCmec type III and MREJ type i (23). It was not possible to define the SCCmec type for the prototype of the pediatric clone of MRSA ATCC BAA-42 carrying MREJ type ii. This strain was shown to carry an SCCmec containing a class B mec gene complex, but no amplification product was detected with the ccr-specific primers described by Okuma et al. (38). The following three types of ccr gene complexes have been described: (i) type 1, containing the ccrA1 and ccrB1 genes; (ii) type 2, containing the ccrA2 and ccrB2 genes; and (iii) type 3, containing the ccrA3 and ccrB3 genes (38). It was recently shown that MRSA strain ATCC BAA-42 carries new ccrA and ccrB gene sequences (39). The ccrA gene of this strain exhibits 58 to 61% identity with the ccrA1, ccrA2, and ccrA3 genes, whereas the ccrB gene of this strain exhibits 70.2 to 73% identity with the ccrB1, ccrB2, and ccrB3 genes (23, 34). Therefore, according to the typing nomenclature defined by Okuma et al. (38), this strain carries a new ccr gene complex, and consequently, a new SCCmec type.
The sequence of the complete SCCmec was not characterized for the strains described in this study, but our findings suggest that more polymorphisms are found at the right extremity of SCCmec than was originally suggested (17, 23).
After the discovery of new SRE sequences in MRSA strains, we developed a multiplex PCR assay which combines a primer specific to S. aureus orfX with a set of primers specific to each SRE sequence and with four MBPs for real-time fluorescence detection of MRSA. None of the gram-negative or gram-positive bacterial species tested other than S. aureus, including a variety of MRCoNS and MSCoNS strains, were detected with the PCR assay, showing that it was specific and did not cross-react with bacterial species other than S. aureus. We then compared conventional culture methods and PCR for the identification of S. aureus and the determination of susceptibility to oxacillin for 2,226 S. aureus isolates from various geographic areas. Whereas 1,636 (98.7%) of the 1,657 MRSA strains were correctly identified by the PCR assay, 26 (4.6%) of the 569 MSSA strains tested were misidentified as MRSA. The absence of the mecA gene in these 26 MSSA strains was confirmed by use of a previously described mecA-specific assay (35; also data not shown). This incorrect identification with the PCR assay could be explained by the presence of a residual SCCmec right extremity fragment following the deletion of a chromosomal region containing mecA or the presence of an SCC which does not contain mecA (23, 25). An analysis by agarose gel electrophoresis of the amplification products generated with DNAs from these MSSA strains showed that they carry SCC fragments with different MREJ types (i, ii, iii, or vii). The spontaneous loss of the mecA region in vitro (i) during the storage or long-term cultivation of MRSA strains in antibiotic-free medium or (ii) from cultures which were starved, grown at an elevated temperature, or given small doses of UV radiation has already been described (14, 19, 20, 22, 40). The deletion of a large chromosomal region has been identified for some strains with a deletion of mecA. The deletion was shown to start precisely from the left boundary of IS431mec and to extend leftwards for various distances beyond the mecA gene (51), suggesting that a residual SCCmec right extremity fragment would still be present in these deletants.
The deletion of the mec region from MRSA strains has also been observed in vivo, but the deleted mec fragments have not been characterized in most cases (9, 11, 21, 33). On the other hand, it was recently shown that two MSSA strains, which were originally shown to be MRSA based on spa sequence typing, had a deletion of the complete SCCmec (45). For these MRSA strains, the SCCmec would have been completely excised from the chromosome by the Ccr recombinases, which are specifically involved in the integration and excision of SCCmec in the S. aureus chromosome (27). In this study, the PCR detection of 26 mecA-negative S. aureus strains suggests that an SCC fragment is present in the chromosome. Some of these MSSA strains were isolated concurrently with an MRSA strain from the same patient's specimen. If a deletion event has occurred in these strains, it would be interesting to know if it was produced in vivo or in vitro during cultivation. Work is in progress to further characterize the deletion fragment of these MSSA strains and to verify if the MSSA and MRSA strains which occur together in the same patient are of the same genetic background.
Twenty-one (1.3%) of the MRSA strains tested in this study were not detected by the PCR assay. However, for all of these MRSA strains except one, it was possible to assign an SCCmec type (data not shown). The lack of amplification of the MREJ fragments of these strains may be attributable to the presence of polymorphisms in the region recognized by the amplification primers or to new SRE sequences. It should be noted that these strains do not seem to represent epidemic clones in the respective geographic areas from which they originated. Sequencing of the MREJs of these strains is in progress.
To further evaluate the usefulness of the real-time PCR assay for the detection of MRSA directly from nonsterile specimens containing a mixture of staphylococci, we performed amplification of a low genome copy number of MRSA (i.e., ∼10) in the presence of up to 104 genome copies of MSSA, MSCoNS, or MRCoNS. The data showed that the amplification of MRSA was not inhibited in the presence of these potentially competing DNAs, suggesting that the assay could be applied for the detection of MRSA directly from clinical samples containing mixed populations of staphylococci. We have used this assay to detect MRSA directly from nasal specimens. The PCR assay was performed directly from nasal material prepared with a rapid specimen preparation protocol and allowed the detection of MRSA in <1 h. The detection limit of the PCR assay was found to be ∼25 CFU per nasal swab. A clinical study with >100 nasal swabs obtained during an MRSA surveillance program showed that the sensitivity of the PCR assay (>97%) compared to that of the standard culture method was sufficient to detect MRSA directly from nasal specimens (A. Huletsky, P. Lebel, M. Gagnon, M. Bernier, K. Truchon, V. Rossbach, F. Gagnon, N. Boucher, L. Guay, F. J. Picard, and M. G. Bergeron, Abstr. 42nd Intersci. Conf. Antimicrob. Agents Chemother., abstr. D-2006, 2002).
With this study, we have developed a powerful real-time multiplex PCR assay which can be used to detect MRSA directly from nonsterile clinical specimens containing a mixture of staphylococci in <1 h. As opposed to all other published molecular methods for the detection of MRSA, this assay does not require any previous isolation, capture, or enrichment of the bacteria, thus reducing the number of sample preparation steps and the time to results. The use of this rapid PCR assay should help to reduce the workload associated with MRSA surveillance programs and the spread of MRSA in clinical settings.
Acknowledgments
This study was supported by Infectio Diagnostic Inc. (Sainte-Foy, Québec, Canada) and by grant PA-15586 from the Canadian Institutes of Health Research (CIHR). M.O. is the holder of a Canada Research Chair in Antimicrobial Resistance.
We thank W. van Leeuwen (Erasmus MC) for his help in the validation of the assay. We also thank L. Jetté (Laboratoire de Santé Publique du Québec, Sainte-Anne-de-Bellevue, Québec, Canada), R. N. Jones (The Jones Group/JMI Laboratories, North Liberty, Iowa), J. M. Bell (Women's and Children's Hospital, North Adelaide, Australia), A. C. Fluit (University Medical Center Utrecht, Utrecht, The Netherlands), A. McGeer and D. E. Low (Mount Sinai Hospital, Toronto, Ontario, Canada), P.-Y. Donnio (Centre Hospitalier Universitaire, Rennes, France), W. Fu (HuaShan Hospital, Shanghai, China), N. El Solh and J.-F. Vieu (Institut Pasteur, Paris France), F. C. Tenover (CDC, Atlanta, Ga.), T. Takenouchi (Sankyo Co., Ltd., Tokyo, Japan), J. Sutcliffe (Pfizer Inc., Groton, N.Y.), A. D. Larsen and L. Pallesen (Statens Serum Institut, Copenhagen, Denmark), Y. Nakajima (Hokkaido College of Pharmacy, Hokkaido, Japan), D. Centron (Universidad de Buenos Aires, Buenos Aires, Argentina), P. Lebel (Hôpital Général de Montréal, Montréal, Québec, Canada), T. Weller (City Hospital NHS Trust, Birmingham, United Kingdom), Z. Hussain (Health Sciences Center, London, Ontario, Canada), K. Hiramatsu (Research Institute International Medical Center of Japan, Tokyo, Japan), and G. F. Brooks (University of California, San Francisco, Calif.) for providing staphylococcal isolates.
REFERENCES
- 1.Aires De Sousa, M., I. Santos Sanches, M. L. Ferro, and H. De Lencastre. 2000. Epidemiological study of staphylococcal colonization and cross-infection in two West African Hospitals. Microb. Drug Resist. 6:133-141. [DOI] [PubMed] [Google Scholar]
- 2.Baba, T., F. Takeuchi, M. Kuroda, H. Yuzawa, K. Aoki, A. Oguchi, Y. Nagai, N. Iwama, K. Asano, T. Naimi, H. Kuroda, L. Cui, K. Yamamoto, and K. Hiramatsu. 2002. Genome and virulence determinants of high virulence community-acquired MRSA. Lancet 359:1819-1827. [DOI] [PubMed] [Google Scholar]
- 3.Baron, E. J. 1995. Genetic aspects of methicillin resistance in Staphylococcus aureus and methods used for its detection in clinical laboratories in the United States. J. Chemother. 7(Suppl. 3):87-92. [PubMed] [Google Scholar]
- 4.Bélanger, S. D., M. Boissinot, N. Clairoux, F. J. Picard, and M. G. Bergeron. 2003. Rapid detection of Clostridium difficile in feces by real-time PCR. J. Clin. Microbiol. 41:730-734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Boissinot, M., and M. G. Bergeron. 2002. Toward rapid real-time molecular diagnostic to guide smart use of antimicrobials. Curr. Opin. Microbiol. 5:478-482. [DOI] [PubMed] [Google Scholar]
- 6.Chambers, H. F. 2001. The changing epidemiology of Staphylococcus aureus? Emerg. Infect. Dis. 7:178-182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chung, M., H. de Lencastre, P. Matthews, A. Tomasz, I. Adamsson, M. Aries de Sousa, T. Camou, C. Cocuzza, A. Corso, I. Couto, A. Dominguez, M. Gniadkowski, R. Goering, A. Gomes, K. Kikuchi, A. Marchese, R. Mato, O. Melter, D. Oliveira, R. Palacio, R. Sa-Leao, I. Santos Sanches, J. H. Song, P. T. Tassios, and P. Villari. 2000. Molecular typing of methicillin-resistant Staphylococcus aureus by pulsed-field gel electrophoresis: comparison of results obtained in a multilaboratory effort using identical protocols and MRSA strains. Microb. Drug Resist. 6:189-198. [DOI] [PubMed] [Google Scholar]
- 8.Cosgrove, S. E., G. Sakoulas, E. N. Perencevich, M. J. Schwaber, A. W. Karchmer, and Y. Carmeli. 2003. Comparison of mortality associated with methicillin-resistant and methicillin-susceptible Staphylococcus aureus bacteremia: a meta-analysis. Clin. Infect. Dis. 36:53-59. [DOI] [PubMed] [Google Scholar]
- 9.Deplano, A., P. T. Tassios, Y. Glupczynski, E. Godfroid, and M. J. Struelens. 2000. In vivo deletion of the methicillin resistance mec region from the chromosome of Staphylococcus aureus strains. J. Antimicrob. Chemother. 46:617-620. [DOI] [PubMed] [Google Scholar]
- 10.Diekema, D. J., M. A. Pfaller, F. J. Schmitz, J. Smayevsky, J. Bell, R. N. Jones, and M. Beach. 2001. Survey of infections due to Staphylococcus species: frequency of occurrence and antimicrobial susceptibility of isolates collected in the United States, Canada, Latin America, Europe, and the Western Pacific region for the SENTRY Antimicrobial Surveillance Program, 1997-1999. Clin. Infect. Dis. 32(Suppl. 2):S114-S132. [DOI] [PubMed] [Google Scholar]
- 11.Dominguez, M. A., H. de Lencastre, J. Linares, and A. Tomasz. 1994. Spread and maintenance of a dominant methicillin-resistant Staphylococcus aureus (MRSA) clone during an outbreak of MRSA disease in a Spanish hospital. J. Clin. Microbiol. 32:2081-2087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.François, P., D. Pittet, M. Bento, B. Pepey, P. Vaudaux, D. Lew, and J. Schrenzel. 2003. Rapid detection of methicillin-resistant Staphylococcus aureus directly from sterile or nonsterile clinical samples by a new molecular assay. J. Clin. Microbiol. 41:254-260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Grisold, A. J., E. Leitner, G. Muhlbauer, E. Marth, and H. H. Kessler. 2002. Detection of methicillin-resistant Staphylococcus aureus and simultaneous confirmation by automated nucleic acid extraction and real-time PCR. J. Clin. Microbiol. 40:2392-2397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Grubb, W. B., and D. I. Annear. 1972. Spontaneous loss of methicillin resistance in Staphylococcus aureus at room-temperature. Lancet ii:1257. [DOI] [PubMed] [Google Scholar]
- 15.Hiramatsu, K., L. Cui, M. Kuroda, and T. Ito. 2001. The emergence and evolution of methicillin-resistant Staphylococcus aureus. Trends Microbiol. 9:486-493. [DOI] [PubMed] [Google Scholar]
- 16.Hiramatsu, K., Y. Katayama, H. Yuzawa, and T. Ito. 2002. Molecular genetics of methicillin-resistant Staphylococcus aureus. Int. J. Med. Microbiol. 292:67-74. [DOI] [PubMed] [Google Scholar]
- 17.Hiramatsu, K., N. Kondo, and T. Ito. 1996. Genetic basis for molecular epidemiology of MRSA. J. Infect. Chemother. 2:117-129. [DOI] [PubMed] [Google Scholar]
- 18.Hiramatsu, K., K. Okuma, X. X. Ma, M. Yamamoto, S. Hori, and M. Kapi. 2002. New trends in Staphylococcus aureus infections: glycopeptide resistance in hospital and methicillin resistance in the community. Curr. Opin. Infect. Dis. 15:407-413. [DOI] [PubMed] [Google Scholar]
- 19.Hiramatsu, K., E. Suzuki, H. Takayama, Y. Katayama, and T. Yokota. 1990. Role of penicillinase plasmids in the stability of the mecA gene in methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 34:600-604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hurlimann-Dalel, R. L., C. Ryffel, F. H. Kayser, and B. Berger-Bachi. 1992. Survey of the methicillin resistance-associated genes mecA, mecR1-mecI, and femA-femB in clinical isolates of methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 36:2617-2621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Inglis, B., W. el-Adhami, and P. R. Stewart. 1993. Methicillin-sensitive and -resistant homologues of Staphylococcus aureus occur together among clinical isolates. J. Infect. Dis. 167:323-328. [DOI] [PubMed] [Google Scholar]
- 22.Inglis, B., P. R. Matthews, and P. R. Stewart. 1990. Induced deletions within a cluster of resistance genes in the mec region of the chromosome of Staphylococcus aureus. J. Gen. Microbiol. 136:2231-2239. [DOI] [PubMed] [Google Scholar]
- 23.Ito, T., Y. Katayama, K. Asada, N. Mori, K. Tsutsumimoto, C. Tiensasitorn, and K. Hiramatsu. 2001. Structural comparison of three types of staphylococcal cassette chromosome mec integrated in the chromosome in methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 45:1323-1336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ito, T., Y. Katayama, and K. Hiramatsu. 1999. Cloning and nucleotide sequence determination of the entire mec DNA of pre-methicillin-resistant Staphylococcus aureus N315. Antimicrob. Agents Chemother. 43:1449-1458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ito, T., K. Okuma, X. X. Ma, H. Yuzawa, and K. Hiramatsu. 2003. Insights on antibiotic resistance of Staphylococcus aureus from its whole genome: genomic island SCC. Drug Resist. Update 6:41-52. [DOI] [PubMed] [Google Scholar]
- 26.Jonas, D., M. Speck, F. D. Daschner, and H. Grundmann. 2002. Rapid PCR-based identification of methicillin-resistant Staphylococcus aureus from screening swabs. J. Clin. Microbiol. 40:1821-1823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Katayama, Y., T. Ito, and K. Hiramatsu. 2000. A new class of genetic element, staphylococcus cassette chromosome mec, encodes methicillin resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 44:1549-1555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ke, D., C. Ménard, F. J. Picard, M. Boissinot, M. Ouellette, P. H. Roy, and M. G. Bergeron. 2000. Development of conventional and real-time PCR assays for the rapid detection of group B streptococci. Clin. Chem. 46:324-331. [PubMed] [Google Scholar]
- 29.Ke, D., F. J. Picard, F. Martineau, C. Ménard, P. H. Roy, M. Ouellette, and M. G. Bergeron. 1999. Development of a PCR assay for rapid detection of enterococci. J. Clin. Microbiol. 37:3497-3503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kearns, A. M., P. R. Seiders, J. Wheeler, R. Freeman, and M. Steward. 1999. Rapid detection of methicillin-resistant staphylococci by multiplex PCR. J. Hosp. Infect. 43:33-37. [DOI] [PubMed] [Google Scholar]
- 31.Kotilainen, P., M. Routamaa, R. Peltonen, J. Oksi, E. Rintala, O. Meurman, O. P. Lehtonen, E. Eerola, S. Salmenlinna, J. Vuopio-Varkila, and T. Rossi. 2003. Elimination of epidemic methicillin-resistant Staphylococcus aureus from a university hospital and district institutions, Finland. Emerg. Infect. Dis. 9:169-175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kuroda, M., T. Ohta, I. Uchiyama, T. Baba, H. Yuzawa, I. Kobayashi, L. Cui, A. Oguchi, K. Aoki, Y. Nagai, J. Lian, T. Ito, M. Kanamori, H. Matsumaru, A. Maruyama, H. Murakami, A. Hosoyama, Y. Mizutani-Ui, N. K. Takahashi, T. Sawano, R. Inoue, C. Kaito, K. Sekimizu, H. Hirakawa, S. Kuhara, S. Goto, J. Yabuzaki, M. Kanehisa, A. Yamashita, K. Oshima, K. Furuya, C. Yoshino, T. Shiba, M. Hattori, N. Ogasawara, H. Hayashi, and K. Hiramatsu. 2001. Whole genome sequencing of methicillin-resistant Staphylococcus aureus. Lancet 357:1225-1240. [DOI] [PubMed] [Google Scholar]
- 33.Lawrence, C., M. Cosseron, P. Durand, Y. Costa, and R. Leclercq. 1996. Consecutive isolation of homologous strains of methicillin-resistant and methicillin-susceptible Staphylococcus aureus from a hospitalized child. J. Hosp. Infect. 33:49-53. [DOI] [PubMed] [Google Scholar]
- 34.Ma, X. X., T. Ito, C. Tiensasitorn, M. Jamklang, P. Chongtrakool, S. Boyle-Vavra, R. S. Daum, and K. Hiramatsu. 2002. Novel type of staphylococcal cassette chromosome mec identified in community-acquired methicillin-resistant Staphylococcus aureus strains. Antimicrob. Agents Chemother. 46:1147-1152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Martineau, F., F. J. Picard, N. Lansac, C. Ménard, P. H. Roy, M. Ouellette, and M. G. Bergeron. 2000. Correlation between the resistance genotype determined by multiplex PCR assays and the antibiotic susceptibility patterns of Staphylococcus aureus and Staphylococcus epidermidis. Antimicrob. Agents Chemother. 44:231-238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Muto, C. A., J. A. Jernigan, B. E. Ostrowsky, H. M. Richet, W. R. Jarvis, J. M. Boyce, and B. M. Farr. 2003. SHEA guideline for preventing nosocomial transmission of multidrug-resistant strains of Staphylococcus aureus and enterococcus. Infect. Control Hosp. Epidemiol. 24:362-386. [DOI] [PubMed] [Google Scholar]
- 37.National Committee for Clinical Laboratory Standards. 2000. Performing standards for antimicrobial susceptibility testing. Eleventh informational supplement, vol. 21. National Committee for Clinical Laboratory Standards, Wayne, Pa.
- 38.Okuma, K., K. Iwakawa, J. D. Turnidge, W. B. Grubb, J. M. Bell, F. G. O'Brien, G. W. Coombs, J. W. Pearman, F. C. Tenover, M. Kapi, C. Tiensasitorn, T. Ito, and K. Hiramatsu. 2002. Dissemination of new methicillin-resistant Staphylococcus aureus clones in the community. J. Clin. Microbiol. 40:4289-4294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Oliveira, D. C., A. Tomasz, and H. de Lencastre. 2001. The evolution of pandemic clones of methicillin-resistant Staphylococcus aureus: identification of two ancestral genetic backgrounds and the associated mec elements. Microb. Drug Resist. 7:349-361. [DOI] [PubMed] [Google Scholar]
- 40.Poston, S. M., and F. L. Li Saw Hee. 1991. Genetic characterisation of resistance to metal ions in methicillin-resistant Staphylococcus aureus: elimination of resistance to cadmium, mercury and tetracycline with loss of methicillin resistance. J. Med. Microbiol. 34:193-201. [DOI] [PubMed] [Google Scholar]
- 41.Que, T. L., P. L. Ho, K. T. Yip, H. L. Ng, F. Y. Leung, K. K. Lai, and K. Y. Yuen. 2003. Three-year study of targeted screening for methicillin-resistant Staphylococcus aureus at hospital admission. Eur. J. Clin. Microbiol. 22:268-270. [DOI] [PubMed] [Google Scholar]
- 42.Reischl, U., H. J. Linde, M. Metz, B. Leppmeier, and N. Lehn. 2000. Rapid identification of methicillin-resistant Staphylococcus aureus and simultaneous species confirmation using real-time fluorescence PCR. J. Clin. Microbiol. 38:2429-2433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Rubinovitch, B., and D. Pittet. 2001. Screening for methicillin-resistant Staphylococcus aureus in the endemic hospital: what have we learned? J. Hosp. Infect. 47:9-18. [DOI] [PubMed] [Google Scholar]
- 44.Sambrook, J., and D. W. Russell. 2001. Molecular cloning: a laboratory manual, 3rd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 45.Shopsin, B., B. Mathema, X. Zhao, J. Martinez, J. Kornblum, and B. N. Kreiswirth. 2000. Resistance rather than virulence selects for the clonal spread of methicillin-resistant Staphylococcus aureus: implications for MRSA transmission. Microb. Drug Resist. 6:239-244. [DOI] [PubMed] [Google Scholar]
- 46.Shrestha, N. K., M. J. Tuohy, G. S. Hall, C. M. Isada, and G. W. Procop. 2002. Rapid identification of Staphylococcus aureus and the mecA gene from BacT/ALERT blood culture bottles by using the LightCycler system. J. Clin. Microbiol. 40:2659-2661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Simor, A., M. Ofner-Agostini, E. Bryce, A. McGeer, S. Paton, M. R. Mulvey, and Canadian Hospital Epidemiology Committee and Canadian Nosocomial Infection Surveillance Program, Health Canada. 2002. Laboratory characterization of methicillin-resistant Staphylococcus aureus in Canadian hospital: results of 5 years of national surveillance, 1995-1999. J. Infect. Dis. 186:652-660. [DOI] [PubMed] [Google Scholar]
- 48.Tan, T. Y., S. Corden, R. Barnes, and B. Cookson. 2001. Rapid identification of methicillin-resistant Staphylococcus aureus from positive blood cultures by real-time fluorescence PCR. J. Clin. Microbiol. 39:4529-4531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Vannuffel, P., J. Gigi, H. Ezzedine, B. Vandercam, M. Delmee, G. Wauters, and J. L. Gala. 1995. Specific detection of methicillin-resistant Staphylococcus species by multiplex PCR. J. Clin. Microbiol. 33:2864-2867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Verhoef, J., D. Beaujean, H. Blok, A. Baars, A. Meyler, C. van der Werken, and A. Weersink. 1999. A Dutch approach to methicillin-resistant Staphylococcus aureus. Eur. J. Clin. Microbiol. Infect. Dis. 18:461-466. [DOI] [PubMed] [Google Scholar]
- 51.Wada, A., Y. Katayama, K. Hiramatsu, and T. Yokota. 1991. Southern hybridization analysis of the mecA deletion from methicillin-resistant Staphylococcus aureus. Biochem. Biophys. Res. Commun. 176:1319-1325. [DOI] [PubMed] [Google Scholar]