Abstract
The siglec family of sialic acid-binding proteins are endocytic immune cell receptors that are recognized as potential targets for cell directed therapies. CD33 and CD22 are prototypical members and are validated candidates for targeting acute myeloid leukaemia and non-Hodgkin’s lymphomas due to their restricted expression on myeloid cells and B-cells, respectively. While nanoparticles decorated with high affinity siglec ligands represent an attractive platform for delivery of therapeutic agents to these cells, a lack of ligands with suitable affinity and/or selectivity has hampered progress. Herein we describe selective ligands for both of these siglecs, which when displayed on liposomal nanoparticles, can efficiently target the cells expressing them in peripheral human blood. Key to their identification was the development of a facile method for chemo-enzymatic synthesis of disubstituted sialic acid analogues, combined with iterative rounds of synthesis and rapid functional analysis using glycan microarrays.
Introduction
The majority of the siglec family of sialic acid-binding proteins exhibit restricted expression on subsets of white blood cells of the immune system, making them attractive targets for cell specific therapies.1–3 Because most siglecs are also endocytic receptors, they are ideal for a “Trojan Horse”-based strategy involving delivery of a therapeutic cargo into the cell when conjugated to antibodies or nanoparticles that target the desired siglec.4–6 Of particular interest in this regard are CD33 (Siglec-3) and CD22 (Siglec-2), which were identified in the mid-80’s as markers of primary acute myeloid leukaemia (AML) blasts and various non-Hodgkin’s lymphomas, respectively,7–11 leading to the development of anti-CD33 and anti-CD22 immunotoxins soon thereafter.12, 13 Gemtuzumab Ozagamicin, a calicheamicin-conjugated anti-CD33 antibody, was approved in 2000 for treatment of acute myeloid leukaemia after promising Phase I and Phase II data.14, 15 However, it was voluntarily withdrawn from the market in 2010 in the United States after disappointing Phase III results16 with evidence of increased treatment-related mortality.17 Despite this setback, new Phase III trials combining low dose Gemtuzumab Ozagamicin with chemotherapy look highly promising for providing benefit to patients with acute myeloid leukaemia.18 Similarly, in the last decade anti-CD22 based therapeutics including naked antibodies, immunotoxin conjugates, and radio-immunotherapeutic have also progressed through Phase I and Phase II clinical trials for treatment of B cell lymphomas/leukaemias with very encouraging results.19–24 In a very recent development, high expression of CD33 on brain microglial cells (macrophages) has emerged as a major risk factor for the development of late onset Alzheimer’s disease due to its ability to inhibit the uptake of neurofibrillary plaques.25–27 Thus, interest in CD33 and CD22 as clinical targets for cell directed therapies continues to grow.
Glycan ligand decorated nanoparticles represent a promising alternative to antibodies for in vivo targeting of siglec expressing cells. They are rapidly endocytosed and accumulate in intracellular organelles, delivering any payload they carry.3, 5, 24, 28–30 The major challenge, though, has been to identify ligands of sufficient avidity and selectivity to target cells expressing only the desired siglec. The most successful approach to date has been to use sialic acid as a privileged scaffold, with modifications made around the sugar ring, primarily at C9 and C5, to increase affinity and selectivity for the desired siglec.31–41
Despite significant progress in this arena, efforts have failed to identify ligands of CD22 and CD33 with sufficient avidity and selectivity needed for human clinical studies. For hCD33 in particular, there are no reports describing high affinity ligands of this siglec. In contrast, several groups have generated ligands of CD22 with 100-1000 fold higher affinity than the natural ligand, but the best of these have not demonstrated adequate selectivity.36, 38, 39, 41 For example, although we have shown that doxorubicin-loaded liposomes displaying a high affinity ligand of CD22 (Fig. 1, compound 4) are effective in prolonging life in a murine model of disseminated human B cell lymphoma, this ligand exhibits a major cross-reactivity with sialoadhesin (Siglec-1, mSn), expressed on macrophages, which mediate rapid clearance of the liposomes.28 Thus, a more selective ligand of hCD22 is needed for optimal targeting of B lymphoma cells.
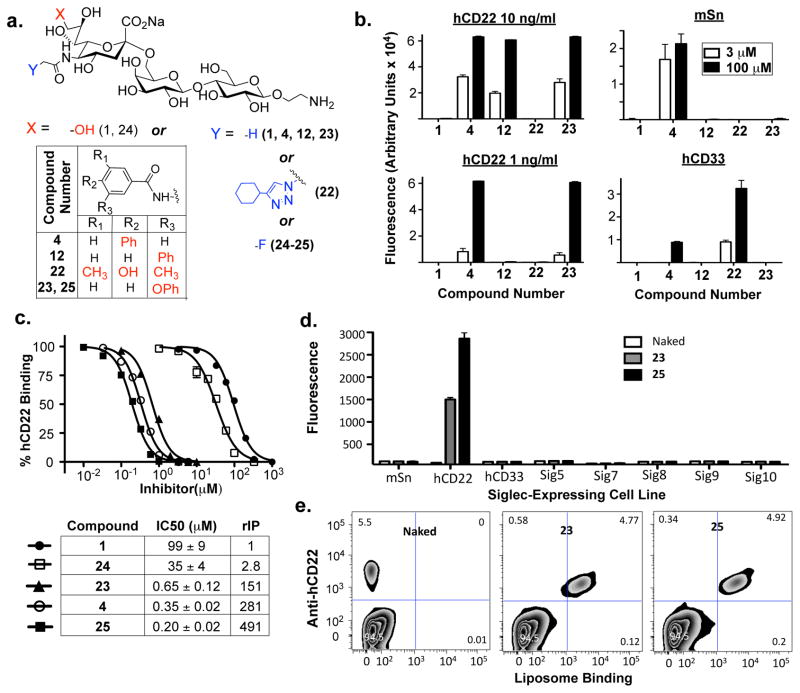
Fig. 1.
Iterative, hCD33-directed Library Synthesis and Screening for the Identification of Selective, High-affinity hCD33 Ligands. (a) Structures of the sialoside analogues that were constructed and screened. (b) Compiled results of the iterative microarray screening process for hCD33. In this process, screening with high and low concentrations of both hCD33 and the deposited sialoside allows for the identification of analogues that give increases in affinity. To aid the reader, key results, which were used to design the next library, are noted. (c) Counterscreening these libraries with related siglecs, hCD22 and mouse Siglec-1 (mSn), allows for the identification of ligands which also exhibit selectivity for hCD33. Shown are representative results from three independent experiments.
Here we report the development of high affinity ligands selective for hCD33 and hCD22. This was accomplished for hCD33 by carrying out iterative cycles of focused library synthesis followed by glycan microarray screening to assess relative avidity and specificity for selected siglecs. Ultimately this resulted in a ligand exhibiting ~350-fold increased affinity over a natural sialoside, and when displayed on liposomal nanoparticles exhibited high specificity for hCD33 over a panel of other human siglecs. During these screens we fortuitously identified a sialic acid analog showing increased affinity for hCD22 with no crossreactivity to Siglec-1 (mSn) or hCD33. Further optimization of this scaffold yielded a ligand with high affinity and selectivity for hCD22. Finally, we show that ligand-bearing liposomes displaying the ligands of hCD33 and hCD22 bind selectively to cells expressing their respective siglec in peripheral human blood.
Results and Discussion
Identification of high-affinity sialoside ligands selective for hCD33
We have previously shown that hCD33 binds to α2-6 linked sialoside analogues bearing unnatural hydrophobic substituents appended to C9 or C5.31 In this previous work, screening an extensive library of click-chemistry generated sialoside analogues identified compound 2, with a 4-cyclohexyl-1,2,3-triazole substituent at the C5 position, with a modestly increased affinity for hCD33 over the native scaffold (1), and without cross-reactivity to other siglecs in the screen (Fig. 1).31 Although triazole-containing substituents linked to the C9 position failed to yield affinity gains for hCD33, a previously identified high affinity hCD22/mSn ligand with a benzamide linkage (4) also exhibited an affinity gain for hCD33, albeit without selectivity (Fig. 1).31 These observations provided motivation to more exhaustively survey C9-substituted benzamide analogues as potential high-affinity CD33 ligands using iterative rounds of focused library synthesis coupled with glycan array screening to simultaneously address affinity and selectivity for this siglec. It was reasoned that an optimal C9 substituent combined with the 4-cyclohexyl-1,2,3-triazole at the C5 position could work synergistically to achieve high affinity and selectivity for hCD33.
As a first step towards this goal, an initial series of 9-benzamide substituents were synthesized and analysed by glycan array (Fig. 1, compounds 3-6). It was noted that replacing the biphenyl substituent with a single benzamido group (3) completely abolished binding to hCD33 (Fig. 1). Interestingly, however, addition of an acetylene moiety to the meta- (5) but not para- (6) position of the benzamide ring re-established this affinity gain and improved selectivity. Notably, click chemistry-derived products of (5) with a variety of azides completely abolished binding to hCD33 and suggested a potential steric clash of large moieties at this position (data not shown). Thus, we first sought to explore if other substituents at the meta position of the benzamide ring, particularly small ones, could yield further improvements over 5.
Accordingly, a small library of C9-analogues with meta-substituted benzamide rings were generated in the α2-6 linked scaffold (Fig. 1, compounds 7-12). This was accomplished through a simple synthetic strategy involving enzymatic transfer of a 9-amino sialic acid to an azide or Cbz-protected lactosyl-α-O-ethylamine scaffold (Scheme 1, A and B), followed by N-acylation of the C9 position of sialic acid, and deprotection of the linker to the free amine required for microcontact printing (Scheme 1).42 On a 5–10 mg scale, this procedure reproducibly offered compounds in excellent yield and purity. Utilizing this approach, analogues with both small (7-11) and large (12) substituents at the meta position of the benzamide ring were created. Upon glycan array analysis, compound 7, with a 3-methylbenzamido substituent, yielded the most promising increase in affinity and selectivity over 5 (Fig. 1b-c and Fig. S1, ESI). It should be noted that we routinely confirm that all compounds are equally printed using the α2-6-linkage specific plant lectin SNA, which is not affected by the presence of 9-substituents (Fig. S2, ESI).33, 43, 44
Scheme 1. Synthesis of Compounds 3-22.
Conditions and Reagents: i) CTP, N. Meningitidis CMP-NeuAc Synthetase, P. Damsella α2,6 Sialyltransferase. Yield: (C) = 98%, (D) = 90%. ii) For X = Cl, MeOH, CH2Cl2, NEt3. For X = N-hydroxysuccinimide, DMF, H2O, NaHCO3. iii) For R = NHCbz: Pd/C, H2, H2O. Yields: 77-92% over 2 steps. For R = N3: PMe3, THF, H2O. Yields 70-75% over 2 steps. iv) (A), Pyruvate, C. Perfringens NeuAc Aldolase, CTP, N. Meningitidis CMP-NeuAc Synthetase, P. Damsella α2,6 Sialyltransferase. Yield: 96%. v) H2O, MeOH, I2, pH 1.0. Yield: 75%. vi) DMF, NEt3. Yield: 86%. vii) PMe3, THF, H2O. Yield: 97%. viii) DMF, H2O, NaHCO3. Yield: 73%. ix) Pd/C, H2, H2O. Yield: 90%.
With a goal to improve upon compound 7, another library containing C9-appended, 3-methylbenzamide substituents, was designed with additional perturbations to the benzamide ring (Fig. 1, Compounds 13-16). From this library, 13, containing a 3,5-dimethylbenzamide substituent, gave a further improvement in affinity and selectivity for hCD33 (Fig. 1b–c and Fig. S1, ESI), while the 2,3-dimethyl isomer 14 abolished binding. Since the methyl group of the 3-methylbenzamide is important for binding to hCD33 (compare 3 and 7), the further increase in avidity for the 3,5-dimethylsubstituent may be an entropic effect due to the symmetry of the resulting ring.
It was notable that all substitutions at the 2 and 5-position of the benzamide ring abrogated binding to hCD33 (14 and 15), while modifications at the 4-positon were sometimes tolerated (4 and 16). To extend these observations, we constructed a panel of C9-substituted 3,5-dimethylbenzamide analogues with varying alterations at the 4-position (Fig. 1a, compounds 17-21). While all of these analogues improved affinity and retained or improved selectivity, compound 17 appeared to be the most promising ligand generated as shown by the fact that it is the only compound of this series detected at a printing concentration of 3 μM and a low hCD33 concentration (0.2 μg/ml, Fig. 1b bottom panel and Fig. S1, ESI). This was further supported by experiments where fluorescently labelled CHO cells expressing high levels of hCD33 cells (CHO-hCD33) were overlaid onto the array. In this case only 17 and 18 of this series can support binding of these cells, confirming that they exhibited highest avidity for CD33 (Fig. S3a, ESI).
Having optimized substituents at the 3, 4, and 5 positions on the C9-benzamide ring we next asked if the further addition of the previously identified C5 substituent, 4-cyclohexyl-1,2,3-triazole (compound 2), would provide further avidity.31 To accomplish the synthesis of a 9,5-disubstituted sialoside we employed a strategy involving chemo-enzymatic synthesis of a sialoside orthogonally protected at the two positions (Scheme 1), in addition to the aglycone. In this strategy we employ a three enzyme one-pot reaction45, 46 that converts a 6-azido-N-pentenoyl-mannosamine (E) into a 9-azido-5-N-pentenoyl sialic acid by condensation with pyruvate, which is then activated to the corresponding CMP-sialic acid followed by sialyltransferase-mediated α2-6 sialylation of the lactoside (A) to yield the trisaccharide precursor (F). Subsequent deprotection of the pentenoyl group afforded (G) to which the 4-cyclohexyl-1,2,3-triazole was installed using NHS chemistry. Reduction of the azide group at C9, followed by amine acylation, and hydrogenation of the Cbz group on the aglycone gave access to 22 in good overall yield. As exemplified by the synthesis of 22, we believe this approach represents a flexible strategy to synthesize 9,5-disubstitued sialosides.
Microarray analysis showed that 22 exhibited superior properties compared to the monosubstituted compounds, for hCD33. In particular, 22 exhibited higher avidity than both parent compounds, 17 and 2 (Fig. 1b bottom panel and Fig. S1, ESI), and showed increased selectivity for hCD33 over hCD22 and mSn (Fig. 1c). This increase in avidity was further supported by the fact that HL-60 cells, an AML cell line expressing intermediate levels of hCD33, bound only to compound 22, but not to any other analogue in our library (Fig. S3b, ESI).
Since glycan microarrays provide only qualitative measures of avidity and selectivity, we analysed the relative affinities of these compounds using solution-phase inhibition assays. Accordingly, IC50 values were determined using a flow cytometry assay, wherein compounds are evaluated for their ability to prevent the binding of fluorescently labelled hCD33 to ligand-coated beads, and these values were used to determine the relative inhibitory potency (rIP) for each compound compared to the native sialoside (rIP = 1). Encouragingly, the results of these assays were in remarkable agreement with the qualitative estimation of avidity gains obtained from our microarray studies (Fig. 2a). As expected the native sialoside (1) showed a relatively low affinity for hCD33 (IC50 = 3.78 mM).47 Relative to the native sialoside, the optimal 5-substituted analogue (2) gave only a 4-fold increase in affinity (IC50 = 997 μM, rIP = 3.9), and the 9-substituted, 3-methylbenzamide analogue (7) yielded a 20-fold increase (IC50 = 174 μM, rIP = 22). Each additional perturbation to the benzamide ring (compounds 13 and 17) added affinity gains of 2-3 fold. Gratifyingly, combining C5 and C9 substituents yielded a roughly additive increase in affinity, as exemplified by 22, with an IC50 of 11 μM. These results highlight the utility of microarrays for rapid qualitative analysis of avidity gains, enabling our iterative approach, and leading to the identification of compound (22) having a 350-fold increased affinity over the natural sialoside.
Fig. 2.
Validation of hCD33 Screening Hits Leads to Ligands with Suitable Affinity and Selectivity for Liposomal Targeting of hCD33-expressing Cells. (a) A solution phase inhibitor assay was used to validate and quantify the affinity gains observed for hCD33-hits identified via microarray screening and determine their relative inhibitory potency (rIP) compared to the native sialoside (rIP = 1) (b) hCD33-hits were coupled to lipids and formulated into fluorescent, ligand-displaying liposomes. Binding to the hCD33-expressing AML cell lines, U937 and HL-60, was assessed by flow cytometry. (c) A blocking antibody was used to demonstrate that binding of the best ligand-displaying liposomes, 17 and 22, to these cell lines was hCD33-dependent. (d) The selectivity of these ligand-displaying liposomes was then assessed against a panel of recombinant siglec-expressing cell lines. (e) Binding of these liposomes to white blood cells isolated from peripheral human blood further documents the selectivity of these liposomes for hCD33-expressing cells in a complex cellular mixture. Red arrow denotes monocytes and green arrow denotes neutrophils as determined by forward and side-scatter properties. In (a)–(e), representative results are shown for one of three independent experiments carried out in duplicate (a) or triplicate (b–e). The IC50 values in (a) are the average of three independent trials.
CD33 Targeted Nanoparticles
With a goal of targeting hCD33-expressing cells in complex biological systems, we initially assessed binding of ligand-bearing liposomes to two hCD33-expressing AML cell lines: HL-60 cells and U937 cells. For these experiments various sialoside analogues (2, 5, 7, 13, 17, and 22) were coupled to an NHS-activated PEGylated lipid and formulated into fluorescent, ~100 nm liposomal nanoparticles displaying a 5% molar amount of the various ligand-lipids or, as a control, 5% of a PEGylated lipid containing no ligand (‘Naked’). Liposome binding to both cell lines, as assessed by flow cytometry, was ligand-dependent and gave the expected trend wherein increased affinity correlated with increased binding (Fig. 2b). While this suggests that the binding is hCD33-dependent, this was further confirmed with an antibody that blocks the ligand-binding domain of hCD33 (Fig. 2c). In these experiments, the blocking antibody completely abrogated binding of the best hCD33-ligand bearing liposomes, 17- and 22-displaying liposomes, confirming that the interaction was specific and was mediated by hCD33 (Fig. 2c).
To determine the selectivity of the best ligand-bearing liposomes, we assessed binding to a panel of recombinant siglec-expressing cell lines. As shown in Fig. 2d, binding of 17- and 22-displaying liposomes was found only to cells expressing hCD33, but not any other siglec tested. These liposomes were then assessed for binding to CD33-expressing cells in peripheral human blood, reflecting a more physiologically relevant setting. As expected, binding was seen only to cell subsets, which express hCD33 (Fig. 2e). Notably, the binding intensity correlates with hCD33 expression as monocytes, with high hCD33 expression (red arrow), show a greater shift than neutrophils with an intermediate level of cell surface hCD33 (green arrow). These results further support the selectivity of our high affinity hCD33 ligands and demonstrate that targeting of primary hCD33-expressing cells is possible with the identified sialoside analogues.
CD22-Targeted Nanoparticles Selective for B cells
While the high-affinity hCD22 ligand (4) has been shown to be effective in targeting B-lymphoma cells in vivo, its crossreactivity with Siglec-1 limits its utility and potential for clinical application. Thus, during the course of our analysis of hCD33 ligands we were excited to note that a 3-biphenylcarboxamide analogue (12) showed selective binding to hCD22 without crossreactivity to other siglecs (Fig. 1). This finding, along with the fact that a 3-phenoxybenzamide analogue (23, Fig. 3) exhibited similar properties33, suggests that appending bulky substituents at the meta position of the C9-benzamide ring can increase affinity and selectivity for hCD22 over other siglecs.
Fig. 3.
CD22-Counterscreening Leads to Insight for Subsequent Generation of a Selective, High-Affinity hCD22 Ligand. (a) Structures of sialoside analogues synthesized and tested. (b) Microarray screening of a small library of sialosides, including 12 and 23, show that bulky substituents at the meta-position of the ring gives selective increases in affinity for hCD22 over other siglecs. (c) A solution-phase inhibitor assay was used to validate and quantify the affinity gains identified in (b), as well as for newly generated analogues 24 and 25 containing a 5-N-Fluoroacetamide substituent known previously to give selective increases in affinity for hCD22 over mSn. (d) The mono- and disubstituted CD22 ligands 23 and 25 were coupled to lipids and formulated into fluorescent ligand-displaying liposomes. These were then assessed for binding to a panel of recombinant siglec-expressing cell lines. (e) Binding of these liposomes to white blood cells isolated from peripheral human blood demonstrate the ability of 23 and 25 to selectively target hCD22-expressing cells in this complex cellular mixture. Results shown in all experiments are representative of three independent experiments carried out in (b) quintuplicate, (c) duplicate, or triplicate (d–e). The IC50 values in (c) are the average of these three independent trials.
To compare these analogues directly, a custom array containing 1, 4, 12, 22, and 23, printed at 100 μM and 3 μM printing concentration, was constructed. Using a sensitive 2-step detection approach (see Methods section) and evaluating binding at various concentrations of the hCD22-Fc, compound 4 showed a higher avidity than compound 12 (Fig. 3a–b and Fig. S4, ESI). However, the related analogue, 23, had comparable avidity to compound 4, and also exhibited excellent selectivity for hCD22 over other siglecs (Fig. 3b and Fig. S4, ESI).
To confirm these results, a solution-phase, competitive inhibition assay was used to determine IC50 values of compounds 1, 4, and 23 for hCD22. With this assay, the natural sialoside (1) yielded an IC50 value in the range of previous observations (IC50 = 99 μM).47–49 The 4-biphenyl derivative (4) had an IC50 of 0.35 μM, while compound 23 gave a roughly 2-fold higher value (IC50 = 0.65 μM). In order to boost the affinity of compound 23 yet retain selectivity for hCD22, we hypothesized that a N-fluoroacetamide group could be installed at the C5 position based on previous reports which documented that this modification yields a selective increase in affinity for hCD22 over Sn.36, 50 As such, both the mono- and disubstituted 5-N-fluoroacetamide containing compounds, 24 and 25, respectively, were synthesized (see ESI). As hoped, the 5-N-fluoroacetamide group gave an additive affinity increase (roughly 3-fold), with the most potent compound 25 yielding an IC50 of 0.2 μM.
Based on our previous results with compound (4)-displaying liposomes,28 we were confident that liposomes bearing 25 would bind avidly to CD22-expressing cells. It was uncertain, however, if the minor decrease in affinity of 23 would yield similar results. In testing these liposomes with the hCD22-expressing, non-Hodgkin’s lymphoma B-cell line, Ramos, both 23- and 25-displaying liposomes, at 4% molar ligand concentration, show excellent binding and, not surprisingly, the 25-bearing liposomes are superior (Fig. S5, ESI). Both of these ligand-bearing liposomes were then assessed for selectivity using our panel of siglec expressing cell lines (Fig. 3d). Notably, no binding was detected with mSn-expressing CHO cells or any other siglec in the series (Fig. 3d). Experiments with white blood cells isolated from peripheral human blood showed that only cells expressing CD22 are targeted, and moreover, the binding correlates with CD22 intensity (Fig. 3e). As expected due to the restricted expression of CD22 on B cells, this CD22+-liposome+ cell population consists entirely of CD19+ B cells (data not shown). In summary, we have developed high affinity hCD22-specific sialic analogues without cross-reactivity to other siglecs, opening the door for future studies aimed at targeting hCD22 for therapeutic gain.
Conclusions
Selective, high affinity ligands of siglecs have proven to have utility as novel chemical probes for elucidating the natural function of these receptors,30, 51, 52 and for targeting nanoparticles to siglec-expressing cells in vivo.28, 29 By loading these nanoparticles with various therapeutic payloads, siglec-targeted nanoparticles represent a versatile platform for cell-targeted therapies. In this regard, hCD22 and hCD33 have received considerable attention as pharmaceutical targets due to their restricted expression on primary AML cells7, 9, 17 and B-cell lymphomas,10, 12, 24 respectively, and more recently the finding that CD33 expression is notably upregulated on brain microglial cells in patients with Alzheimer’s disease.25–27
Here we use glycan microarrays and a versatile chemo-enzymatic strategy to rapidly synthesize and screen a wide variety of mono- and disubstituted sialic acid analogues allowing for rapid, simultaneous assessment of both affinity and selectivity. The strength of this approach is highlighted by the identification of compounds 22 and 25, which can selectively target hCD33 and hCD22, respectively, when conjugated to liposomal nanoparticles. This approach and synthetic methodology, should find utility in the identification of high affinity ligands for other siglecs, and potentially for other ligand-receptor systems.
With 22 and 25 in hand, the stage is set to assess their utility in in vitro and in vivo cancer models. Since a ligand-targeting approach has never been pursued before for hCD33, it will be important to document that these particles are efficiently endocytosed and can therefore deliver a chemotherapeutic drug to leukemic cells. For hCD22, on the other hand, progress has been hindered by the fact that our useful, yet promiscuous tool compound, (4), is crossreactive with Siglec-1 and thereby imposed significant experimental and therapeutic constraints.28 Since compound 25 has improved affinity and selectivity, further studies exploiting the ligand-binding domain of hCD22 for treating a variety of non-Hodgkin’s lymphomas, a broad and genetically diverse set of diseases, are currently underway.
Experimental Section
Compound Synthesis
Synthetic procedures and compound characterization can be found in the Supporting Information.
Glycan Array Printing and Screening
The noted compounds were spot-printed in five replicates at 100 μM or 3 μM printing concentration in 150 mM Phosphate Buffer, 0.005% Tween-20, pH 8.2, using previously established and reported techniques.31, 33, 42 Siglec-Fc chimeras were produced in-house using stable expression in CHO cells (hCD33 and mSn) or transient transfection into COS-cells as previously described.47 For binding studies shown in Fig. 1, hCD33-Fc was pre-complexed (10 μg/ml Fc-chimera) with an R-PE labelled anti-human IgG (5 μg/ml, Jackson Immunoresearch) and serially diluted onto the array. Analysis with hCD22-Fc and mSn-Fc was performed similarly. In Fig. 3, the same procedures were used for hCD33 and mSn; however, a more sensitive approach was used to better distinguish between high affinity hCD22 ligands. In this process, hCD22-Fc was applied to the array at various concentrations, the arrays were washed by dipping three times into a reservoir of PBS-Tween, followed by detection with the above R-PE labelled secondary antibody (10 μg/ml). Final washes in both procedures included dipping three times into reservoirs of PBS-Tween, PBS, and H2O, followed by centrifugation to dry. Slides were then scanned on a Perkin-Elmer ProScanArray Express and the images processed using IMAGENE. Data shown are the mean ± S.D. of the five printed spots.
Bead-Based Flow Cytometry Assays for Determining Compound IC50 Values
Streptavidin-coated magnetic beads (20 μl of 6.7×108 beads/ml, M-280 Dynabeads, Invitrogen Corp.) were washed with HBSS containing 0.5% BSA (HBSS/BSA), resuspended in 200 μl HBSS/BSA, and coated with the appropriate biotin-linked sialoside (see below) overnight at 4°C. After washing to remove unbound probe, they were taken up in 200 μl HBSS/BSA (6.7×107 beads/ml). To 96-well compatible flow cytometry tubes, siglec-Fc and inhibitors were added followed by addition of premixed siaoloside-beads (0.5 μl beads/tube) and FITC-anti-human IgG Fc (Jackson Immunoresearch). After shaking for 30 minutes at room temperature, samples were analysed by flow cytometry. We should note that, due to the low background of the assay, no wash step is necessary and for convenience samples were run without washing. No inhibitor treatment was used to set 100% binding, while no siglec-Fc was used to set 0% binding. For hCD33, 2 μg 6’BPCNeuAc-Biotin (see Supporting Information for structure and synthesis) was used to coat the bead and 0.03 μg hCD33-Fc and 0.13 μg anti-human IgG were added to each assay tube. For hCD22, 1 μg of a 1 MD NeuGcα2-6Galβ1-4GlcNAc-β-O-ethyl-PAA-biotin probe (Consortium for Functional Glycomics) was used to coat the bead and 0.025 μg hCD22-Fc and 0.31 μg anti-human IgG were added to each assay tube. Assays were carried out in duplicate and three independent measurements were performed. Data was analysed using Prism Graphpad Software. IC50 values and standard deviations are reported as the average of these three independent trials. The relative inhibitory potency (rIP) for each compound was determined by dividing the IC50 of the compound in question by the IC50 of the native sialoside.
Liposome Preparation and Cell-Binding Studies
Fluorescent, ~100 nm liposomes were prepared as previously described with the following composition: 0.1 mol % Alexa-Fluor 647 lipid: 5 mol % PEGylated lipid (= ‘naked’ lipid + siglec-ligand lipid): 57 mol % disteraoyl phosphatidylcholine, and 38 mol% cholesterol. For recombinant cell lines, cells (100 μl of 2 x 106 cells/ml) in HBSS/BSA were incubated with liposomes (5-50 μM) for 45 minutes at 37°C, followed by washing (2x with 200 μl HBSS/BSA), and flow cytometry analysis. For hCD33 experiments with HL-60 and U937 cell lines, 5% ligand-displaying liposomes were used. To conduct the antibody-blocking experiments the WM53 antibody or mouse IgG-κ isotype control (Biolegend, 10 μg/ml) was added to cells, allowed to incubate for 10 minutes at room temperature, liposomes were added, and binding was done as above. 100% binding was defined as cells with no pre-treatment condition, but incubated with fluorescent liposomes, while 0% binding was defined as completely untreated cells (i.e. cellular autofluorescence). For hCD33-ligand specificity analysis with overexpressing recombinant cell lines,28, 31, 32 1% ligand-displaying liposomes were used. This panel of cell lines consists of CHO cells expressing mSn, hSig3, hSig5, hSig8, hSig9, and hSig10, Jurkat cells expressing hSig7, and Ramos cells expressing hCD22 (other B-cell lines were found to express additional siglecs, data not shown). Notably, increased ligand percentages (up to 5%) does not alter selectivity (Fig. S6, ESI) of these liposomes in this experiment, but does increase binding to AML cell lines (Fig. S7, ESI). Similarly, for hCD22 ligand specificity studies, 4% ligand-bearing liposomes were used as these were found to be optimal for binding to peripheral blood B-cells (Fig. S7, ESI). For experiments with primary human cells, peripheral blood was obtained from the TSRI Normal Blood Donor Services and processed as previously described.31 For these experiments ~2 x 106 total cells were suspended in HBSS/BSA (100 μl) and 5-50 μM of the naked or targeted 5% (hCD33) or 4% (hCD22) ligand-displaying liposomes were added. Incubation was carried out at 37°C for 1 h, after which time Human Trustain FcX was added to block Fc receptors (Biolegend). After a 5-minute incubation at room temperature, cells were stained with anti-hCD33 R-PE (Biolegend) or anti-hCD22 R-PE (Biolegend) for 15-30 minutes at 37°C. Cells were washed 2 × with HBSS/BSA and then analysed by flow cytometry. Importantly, incubation of cells with liposomes followed by labelled antibody does not block binding of the liposomes, likely because they have been endocytosed in the initial incubation step. Finally, it should be noted that in all graphs of flow cytometry data, the fluorescence plotted is the mean fluorescence intensity (MFI).
Supplementary Material
Acknowledgments
This work was supported by the NIH (P01HL107151 to J.C.P., T32AI007606 to C.D.R., and GM087620 to V.V.F), a Human Frontiers Fellowship (M.S.M), a Schering-Plough Research Institute Postdoctoral Fellowship (to E.S.), and a Rubicon fellowship from the Netherlands Organization For Scientific Research (to E.S.).
Footnotes
Electronic supplementary information (ESI) available: All synthetic procedures and compound characterization, as well as supporting figures and schemes.
Notes and references
- 1.Kiwamoto T, Kawasaki N, Paulson JC, Bochner BS. Pharmacol Ther. 2012;135:327–336. doi: 10.1016/j.pharmthera.2012.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Crocker PR, Paulson JC, Varki A. Nat Rev Immunol. 2007;7:255–266. doi: 10.1038/nri2056. [DOI] [PubMed] [Google Scholar]
- 3.O’Reilly MK, Tian H, Paulson JC. J Immunol. 2011;186:1554–1563. doi: 10.4049/jimmunol.1003005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Jandus C, Simon HU, von Gunten S. Biochem Pharmacol. 2011;82:323–332. doi: 10.1016/j.bcp.2011.05.018. [DOI] [PubMed] [Google Scholar]
- 5.O’Reilly MK, Paulson JC. Trends Pharmacol Sci. 2009;30:240–248. doi: 10.1016/j.tips.2009.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.von Gunten S, Bochner BS. Ann N Y Acad Sci. 2008;1143:61–82. doi: 10.1196/annals.1443.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dinndorf PA, Andrews RG, Benjamin D, Ridgway D, Wolff L, Bernstein ID. Blood. 1986;67:1048–1053. [PubMed] [Google Scholar]
- 8.Griffin JD, Linch D, Sabbath K, Larcom P, Schlossman SF. Leuk Res. 1984;8:521–534. doi: 10.1016/0145-2126(84)90001-8. [DOI] [PubMed] [Google Scholar]
- 9.Drexler HG. Leukemia. 1987;1:697–705. [PubMed] [Google Scholar]
- 10.Mason DY, Stein H, Gerdes J, Pulford KA, Ralfkiaer E, Falini B, Erber WN, Micklem K, Gatter KC. Blood. 1987;69:836–840. [PubMed] [Google Scholar]
- 11.Dorken B, Moldenhauer G, Pezzutto A, Schwartz R, Feller A, Kiesel S, Nadler LM. J Immunol. 1986;136:4470–4479. [PubMed] [Google Scholar]
- 12.May RD, Vitetta ES, Moldenhauer G, Dörken B. Cancer Drug Deliv. 1986;3:261–272. doi: 10.1089/cdd.1986.3.261. [DOI] [PubMed] [Google Scholar]
- 13.Lambert JM, Goldmacher VS, Collinson AR, Nadler LM, Blattler WA. Cancer Res. 1991;51:6236–6242. [PubMed] [Google Scholar]
- 14.Sievers EL, Appelbaum FR, Spielberger RT, Forman SJ, Flowers D, Smith FO, Shannon-Dorcy K, Berger MS, Bernstein ID. Blood. 1999;93:3678–3684. [PubMed] [Google Scholar]
- 15.Sievers EL, Larson RA, Stadtmauer EA, Estey E, Löwenberg B, Dombret H, Karanes C, Theobald M, Bennett JM, Sherman ML, Berger MS, Eten CB, Loken MR, van Dongen JJ, Bernstein ID, Appelbaum FR, Group MS. J Clin Oncol. 2001;19:3244–3254. doi: 10.1200/JCO.2001.19.13.3244. [DOI] [PubMed] [Google Scholar]
- 16.Burnett AK, Russell NH, Hills RK, Kell J, Freeman S, Kjeldsen L, Hunter AE, Yin J, Craddock CF, Dufva IH, Wheatley K, Milligan D. J Clin Oncol. 2012;30:3924–3931. doi: 10.1200/JCO.2012.42.2964. [DOI] [PubMed] [Google Scholar]
- 17.Walter RB, Appelbaum FR, Estey EH, Bernstein ID. Blood. 2012;119:6198–6208. doi: 10.1182/blood-2011-11-325050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Castaigne S, Pautas C, Terre C, Raffoux E, Bordessoule D, Bastie JN, Legrand O, Thomas X, Turlure P, Reman O, de Revel T, Gastaud L, de Gunzburg N, Contentin N, Henry E, Marolleau JP, Aljijakli A, Rousselot P, Fenaux P, Preudhomme C, Chevret S, Dombret H. Lancet. 2012;379:1508–1516. doi: 10.1016/S0140-6736(12)60485-1. [DOI] [PubMed] [Google Scholar]
- 19.Kreitman RJ, Wilson WH, Bergeron K, Raggio M, Stetler-Stevenson M, FitzGerald DJ, Pastan I. N Engl J Med. 2001;345:241–247. doi: 10.1056/NEJM200107263450402. [DOI] [PubMed] [Google Scholar]
- 20.Kreitman RJ, Tallman MS, Robak T, Coutre S, Wilson WH, Stetler-Stevenson M, Fitzgerald DJ, Lechleider R, Pastan I. J Clin Oncol. 2012;30:1822–1828. doi: 10.1200/JCO.2011.38.1756. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kantarjian H, Thomas D, Jorgensen J, Jabbour E, Kebriaei P, Rytting M, York S, Ravandi F, Kwari M, Faderl S, Rios MB, Cortes J, Fayad L, Tarnai R, Wang SA, Champlin R, Advani A, O’Brien S. Lancet Oncol. 2012;13:403–411. doi: 10.1016/S1470-2045(11)70386-2. [DOI] [PubMed] [Google Scholar]
- 22.Advani A, Coiffier B, Czuczman MS, Dreyling M, Foran J, Gine E, Gisselbrecht C, Ketterer N, Nasta S, Rohatiner A, Schmidt-Wolf IG, Schuler M, Sierra J, Smith MR, Verhoef G, Winter JN, Boni J, Vandendries E, Shapiro M, Fayad L. J Clin Oncol. 2010;28:2085–2093. doi: 10.1200/JCO.2009.25.1900. [DOI] [PubMed] [Google Scholar]
- 23.Ogura M, Tobinai K, Hatake K, Uchida T, Kasai M, Oyama T, Suzuki T, Kobayashi Y, Watanabe T, Azuma T, Mori M, Terui Y, Yokoyama M, Mishima Y, Takahashi S, Ono C, Ohata J. Cancer Sci. 2010;101:1840–1845. doi: 10.1111/j.1349-7006.2010.01601.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Sullivan-Chang L, O’Donnell RT, Tuscano JM. BioDrugs. 2013;27:293–304. doi: 10.1007/s40259-013-0016-7. [DOI] [PubMed] [Google Scholar]
- 25.Naj AC, Jun G, Beecham GW, Wang LS, Vardarajan BN, Buros J, Gallins PJ, Buxbaum JD, Jarvik GP, Crane PK, Larson EB, Bird TD, Boeve BF, Graff-Radford NR, De Jager PL, Evans D, Schneider JA, Carrasquillo MM, Ertekin-Taner N, Younkin SG, Cruchaga C, Kauwe JS, Nowotny P, Kramer P, Hardy J, Huentelman MJ, Myers AJ, Barmada MM, Demirci FY, Baldwin CT, Green RC, Rogaeva E, St George-Hyslop P, Arnold SE, Barber R, Beach T, Bigio EH, Bowen JD, Boxer A, Burke JR, Cairns NJ, Carlson CS, Carney RM, Carroll SL, Chui HC, Clark DG, Corneveaux J, Cotman CW, Cummings JL, DeCarli C, DeKosky ST, Diaz-Arrastia R, Dick M, Dickson DW, Ellis WG, Faber KM, Fallon KB, Farlow MR, Ferris S, Frosch MP, Galasko DR, Ganguli M, Gearing M, Geschwind DH, Ghetti B, Gilbert JR, Gilman S, Giordani B, Glass JD, Growdon JH, Hamilton RL, Harrell LE, Head E, Honig LS, Hulette CM, Hyman BT, Jicha GA, Jin LW, Johnson N, Karlawish J, Karydas A, Kaye JA, Kim R, Koo EH, Kowall NW, Lah JJ, Levey AI, Lieberman AP, Lopez OL, Mack WJ, Marson DC, Martiniuk F, Mash DC, Masliah E, McCormick WC, McCurry SM, McDavid AN, McKee AC, Mesulam M, Miller BL, Miller CA, Miller JW, Parisi JE, Perl DP, Peskind E, Petersen RC, Poon WW, Quinn JF, Rajbhandary RA, Raskind M, Reisberg B, Ringman JM, Roberson ED, Rosenberg RN, Sano M, Schneider LS, Seeley W, Shelanski ML, Slifer MA, Smith CD, Sonnen JA, Spina S, Stern RA, Tanzi RE, Trojanowski JQ, Troncoso JC, Van Deerlin VM, Vinters HV, Vonsattel JP, Weintraub S, Welsh-Bohmer KA, Williamson J, Woltjer RL, Cantwell LB, Dombroski BA, Beekly D, Lunetta KL, Martin ER, Kamboh MI, Saykin AJ, Reiman EM, Bennett DA, Morris JC, Montine TJ, Goate AM, Blacker D, Tsuang DW, Hakonarson H, Kukull WA, Foroud TM, Haines JL, Mayeux R, Pericak-Vance MA, Farrer LA, Schellenberg GD. Nat Genet. 2011;43:436–441. doi: 10.1038/ng.801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bradshaw EM, Chibnik LB, Keenan BT, Ottoboni L, Raj T, Tang A, Rosenkrantz LL, Imboywa S, Lee M, Von Korff A, Morris MC, Evans DA, Johnson K, Sperling RA, Schneider JA, Bennett DA, De Jager PL. Nat Neurosci. 2013;16:848–850. doi: 10.1038/nn.3435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Griciuc A, Serrano-Pozo A, Parrado AR, Lesinski AN, Asselin CN, Mullin K, Hooli B, Choi SH, Hyman BT, Tanzi RE. Neuron. 2013;78:631–643. doi: 10.1016/j.neuron.2013.04.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chen WC, Completo GC, Sigal DS, Crocker PR, Saven A, Paulson JC. Blood. 2010;115:4778–4786. doi: 10.1182/blood-2009-12-257386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kawasaki N, Vela JL, Nycholat CM, Rademacher C, Khurana A, van Rooijen N, Crocker PR, Kronenberg M, Paulson JC. Proc Natl Acad Sci U S A. 2013;110:7826–7831. doi: 10.1073/pnas.1219888110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Macauley MS, Pfrengle F, Rademacher C, Nycholat CM, Gale AJ, von Drygalski A, Paulson JC. J Clin Invest. 2013;123:3074–3083. doi: 10.1172/JCI69187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Rillahan CD, Schwartz E, McBride R, Fokin VV, Paulson JC. Angew Chem Int Ed Engl. 2012;51:11014–11018. doi: 10.1002/anie.201205831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Rillahan CD, Schwartz E, Rademacher C, McBride R, Rangarajan J, Fokin VV, Paulson JC. ACS Chem Biol. 2013 doi: 10.1021/cb400125w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Blixt O, Han S, Liao L, Zeng Y, Hoffmann J, Futakawa S, Paulson J. J Am Chem Soc. 2008;130:6680–6681. doi: 10.1021/ja801052g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nycholat CM, Rademacher C, Kawasaki N, Paulson JC. J Am Chem Soc. 2012;134:15696–15699. doi: 10.1021/ja307501e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zeng Y, Rademacher C, Nycholat CM, Futakawa S, Lemme K, Ernst B, Paulson JC. Bioorg Med Chem Lett. 2011;21:5045–5049. doi: 10.1016/j.bmcl.2011.04.068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Mesch S, Lemme K, Wittwer M, Koliwer-Brandl H, Schwardt O, Kelm S, Ernst B. ChemMedChem. 2012;7:134–143. doi: 10.1002/cmdc.201100407. [DOI] [PubMed] [Google Scholar]
- 37.Mesch S, Moser D, Strasser DS, Kelm A, Cutting B, Rossato G, Vedani A, Koliwer-Brandl H, Wittwer M, Rabbani S, Schwardt O, Kelm S, Ernst B. J Med Chem. 2010;53:1597–1615. doi: 10.1021/jm901517k. [DOI] [PubMed] [Google Scholar]
- 38.Abdu-Allah HH, Tamanaka T, Yu J, Zhuoyuan L, Sadagopan M, Adachi T, Tsubata T, Kelm S, Ishida H, Kiso M. J Med Chem. 2008;51:6665–6681. doi: 10.1021/jm8000696. [DOI] [PubMed] [Google Scholar]
- 39.Abdu-Allah HH, Watanabe K, Completo GC, Sadagopan M, Hayashizaki K, Takaku C, Tamanaka T, Takematsu H, Kozutsumi Y, Paulson JC, Tsubata T, Ando H, Ishida H, Kiso M. Bioorg Med Chem. 2011;19:1966–1971. doi: 10.1016/j.bmc.2011.01.060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Abdu-Allah HH, Watanabe K, Hayashizaki K, Takaku C, Tamanaka T, Takematsu H, Kozutsumi Y, Tsubata T, Ishida H, Kiso M. Bioorg Med Chem Lett. 2009;19:5573–5575. doi: 10.1016/j.bmcl.2009.08.044. [DOI] [PubMed] [Google Scholar]
- 41.Kelm S, Madge P, Islam T, Bennett R, Koliwer-Brandl H, Waespy M, von Itzstein M, Haselhorst T. Angew Chem Int Ed Engl. 2013;52:3616–3620. doi: 10.1002/anie.201207267. [DOI] [PubMed] [Google Scholar]
- 42.Blixt O, Head S, Mondala T, Scanlan C, Huflejt ME, Alvarez R, Bryan MC, Fazio F, Calarese D, Stevens J, Razi N, Stevens DJ, Skehel JJ, van Die I, Burton DR, Wilson IA, Cummings R, Bovin N, Wong CH, Paulson JC. Proc Natl Acad Sci U S A. 2004;101:17033–17038. doi: 10.1073/pnas.0407902101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Song X, Yu H, Chen X, Lasanajak Y, Tappert MM, Air GM, Tiwari VK, Cao H, Chokhawala HA, Zheng H, Cummings RD, Smith DF. J Biol Chem. 2011;286:31610–31622. doi: 10.1074/jbc.M111.274217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Shibuya N, Goldstein IJ, Broekaert WF, Nsimba-Lubaki M, Peeters B, Peumans WJ. J Biol Chem. 1987;262:1596–1601. [PubMed] [Google Scholar]
- 45.Yu H, Chokhawala H, Huang S, Chen X. Nat Protoc. 2006;1:2485–2492. doi: 10.1038/nprot.2006.401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yu H, Huang S, Chokhawala H, Sun M, Zheng H, Chen X. Angew Chem Int Ed Engl. 2006;45:3938–3944. doi: 10.1002/anie.200600572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Blixt O, Collins BE, van den Nieuwenhof IM, Crocker PR, Paulson JC. J Biol Chem. 2003;278:31007–31019. doi: 10.1074/jbc.M304331200. [DOI] [PubMed] [Google Scholar]
- 48.Kelm S, Gerlach J, Brossmer R, Danzer CP, Nitschke L. J Exp Med. 2002;195:1207–1213. doi: 10.1084/jem.20011783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Collins BE, Blixt O, Han S, Duong B, Li H, Nathan JK, Bovin N, Paulson JC. J Immunol. 2006;177:2994–3003. doi: 10.4049/jimmunol.177.5.2994. [DOI] [PubMed] [Google Scholar]
- 50.Kelm S, Brossmer R, Isecke R, Gross HJ, Strenge K, Schauer R. Eur J Biochem. 1998;255:663–672. doi: 10.1046/j.1432-1327.1998.2550663.x. [DOI] [PubMed] [Google Scholar]
- 51.Pfrengle F, Macauley MS, Kawasaki N, Paulson JC. J Immunol. 2013;191:1724–1731. doi: 10.4049/jimmunol.1300921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Duong BH, Tian H, Ota T, Completo G, Han S, Vela JL, Ota M, Kubitz M, Bovin N, Paulson JC, Nemazee D. J Exp Med. 2010;207:173–187. doi: 10.1084/jem.20091873. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.