Abstract
Superoxide dismutase type 1 (SOD1) mutations cause protein aggregation and decrease protein stability, which are linked to amyotrophic lateral sclerosis (ALS) disease. This research utilizes the world's largest traditional Chinese medicine (TCM) database to search novel inhibitors of mutant SOD1, and molecular dynamics (MD) simulations were used to analyze the stability of protein that interacted with docked ligands. Docking results show that hesperidin and 2,3,5,4′-tetrahydroxystilbene-2-O-β-D-glucoside (THSG) have high affinity to mutant SOD1 and then dopamine. For MD simulation analysis, hesperidin and THSG displayed similar value of RMSD with dopamine, and the migration analysis reveals stable fluctuation at the end of MD simulation time. Interestingly, distance between the protein and ligand has distinct difference, and hesperidin changes the position from initial binding site to the other place. In flexibility of residues analysis, the secondary structure among all complexes does not change, indicating that the structure are not affect ligand binding. The binding poses of hesperidin and THSG are similar to dopamine after molecular simulation. Our result indicated that hesperidin and THSG might be potential lead compound to design inhibitors of mutant SOD1 for ALS therapy.
1. Introduction
Mutations in Cu/Zn-binding superoxide dismutase type 1 (SOD1) could decrease protein stability and increase aggregation; SOD1 variants are associated with amyotrophic lateral sclerosis (ALS) [1–4]. ALS belongs to motor neuron degenerative disease in the cortex, brainstem, and spinal cord [5, 6], which is similar to Alzheimer's disease, Parkinson's disease, and Huntington's disease, and the symptoms of ALS include degenerative disorder of upper/lower motor neurons and denervation of muscle fibres, leading to loss of motor neuron, progressive muscular paralysis, and muscle atrophy [7, 8], resulting in weakness of voluntary muscles till death because of respiratory failure [9]. A recent study indicated that mutation of SOD1 gene has genetic linkage in ALS disease, ALS-associated SOD 1 mutations including alanine 4 by valine (A4V) [10–13], histidine 46 by arginine (H46R) [14–16], and I113T mutation [17–19]. The most popular ALS-causing mutation is A4V mutant type in United States; near 50% of SOD1-ALS patients are associated with the A4V mutation [20]. H46R has been identified in Japanese, about 80% Japanese familial ALS in the affected members [21]. I113T type mutation is another one of the common SOD1 mutations of FALS [22], and many cases of clinical manifestations are linked to the I113T SOD1 mutation [23]. They proposed a toxic gain of function caused by mutant SOD due to the aggregation [24, 25]; hence, designing novel drugs for inhibition of SOD1 aggregation and stabilization have been used in ALS treatments [26].
The aim of this study is to focus on drug design of mutant SOD1. The complex of mutant SOD1 and dopamine was used to investigate the novel inhibitors of mutant SOD1 for inhibiting aggregation. Computer-aided drug design (CADD) is rapid approach for drug discovery [27–30], which is based on risk factor studies [31–36], theory [37], and web server [38] for developing novel leading compounds. CADD has been wildly used to design new drugs in many cases, such as virus [39, 40], inflammation [41], cancer [42–45], insomnia [46], weight loss [47], erectile dysfunction [48–50], nerve system [51–53], and diabetes [54]. Traditional Chinese medicine (TCM) has been used over two thousand years in clinical therapy, and many studies utilized TCM to investigate new therapies [55–58]. In our study, small molecules from the world TCM database [59] was used to screen for searching potential compounds with high affinity in mutant SOD1 active site, and we further utilized molecular dynamics (MD) simulation to verify the stability between protein and ligands for binding assay. Synthetic drug often has side effect in clinical treatments, and our results provided nature product as drug candidate, which is safer and reduce adverse reactions.
2. Materials and Methods
2.1. Database Screening
The crystal structure of mutant SOD1 was obtained from PDB database (PDB code: 4A7V) [60]. We employed Prepare Protein module of Accelrys Discovery Studio 2.5.5.9350 (DS 2.5) software [61] to clean up mistakes of each residue on mutant SOD structure, such as deleting alternate conformations, modeling missing loops, and removing water molecules. This module also predicts titration site pKs for each amino acid, and the pH value of 7.4 was used to protonate all residues. PONDR-FIT [62] was used to predict the order/disorder in mutant SOD1 structure. The 61000 TCM compounds were downloaded from the TCM Database@Taiwan [59] for database screening, we also employed TCM compounds from Chang' lab for binding assay [63], and MM2 force field [64] of ChemBioOffice 2010 software was carried out to optimize and calculate the 3D conformation of TCM compounds. All compounds generate different conformations by Monte-Carlo techniques under LigandFit module [65] of DS 2.5, which were docked into mutations SOD1 binding site for protein-ligand interaction analysis. Minimization of all docking poses was based on CHARMm force field [66], and we used Smart minimizer algorithm as minimization algorithm for ligands minimization [67, 68], which contains steepest descent and conjugate gradient. The steepest descent performed 1,000 steps and followed by conjugate gradient minimization.
2.2. Molecular Dynamics (MD) Simulation
Protein-ligand structures were obtained from results of docking study, and the starting conformation of protein-ligand complex was performed using GROMACS 4.5.5 package [69] for molecular dynamic simulation, using charmm27 force field. The protein structure was placed in cubic box containing TIP3P water molecules. The distance between protein and box was set to 1.2 nm, and the van der Waals cutoff to 1.4 nm. Particle mesh Ewald (PME) method is regard as coulomb type for calculating electrostatic interaction, and LINCS algorithm was used to restrain the lengths of all bonds among all simulations. For obtaining topology file and parameters of small compounds, we employed SwissParam to generate these data and compatible with the CHARMM all atoms force field for GROMACS simulation. In system neutralization, we added Na and Cl ions to randomly replace solvent molecules in simulation systems, and the concentration of NaCl model was set as 0.145 M. The time step was set to 0.002 ps for MD simulation. Steepest descent algorithm was applied to energy minimization for 5,000 cycle steps. The following procedure is equilibration, which was performed under position restraints for 100 ps to relax solvent in protein structure under constant temperature dynamics (NVT) condition. Production simulations perform 5000 ps at final step for all simulation systems under constant pressure and temperature (NPT) dynamics. Temperature of all simulation systems was set to 310 K. All MD frames were saved every 20 ps for trajectory analysis.
2.3. Analysis of MD Simulation
Trajectory analysis of MD conformations was calculated by GROMACS 4.5.5 [69], including root mean square deviation (RMSD), root mean square fluctuation (RMSF), and mean square displacement (MSD). The secondary structures analysis was performed by DSSP program under GROMACS 4.5.5. Linkage clustering algorithm was used to identify the most populated structural representations of conformation during MD simulations. The RMSD cutoff for cluster analysis was set as 0.13.
3. Results and Discussion
3.1. Docking Results of Database Screening
To analysis disorder region, we employed PONDR-FIT [62] to predict the order/disorder in mutant SOD1 structure. The sequence number from 21 to 32 and from 98 to 100 are binding site of mutant SOD1 (Figure 1). The disorder disposition values among this range are below 0.5, which indicates that the binding site is folded orderly and that the protein structure may not affect ligand binding [70, 71]. For docking analysis, we based on -PLP1, -PLP2, -PMF, and Dock Score to evaluate the docking pose of traditional Chinese medicine (TCM) compounds. From scoring analysis, dopamine was regarded as control for comparing with TCM compounds. The score values from docking poses of TCM compounds are shown in Table 1. All docked ligands are ranked by Dock Score, and we found that hesperidin and 2,3,5,4′-tetrahydroxystilbene-2-O-β-D-glucoside (THSG) [63] with Dock Score (including score values of -PLP1, -PLP2, and -PMF) are higher than dopamine. We selected hesperidin, THSG and dopamine for further studies, and chemical scaffolds of these small molecular are shown in Figure 2. Docking poses of dopamine, which displayed H bond with Glu100, the surrounding residues include Lys30, Lys23, Glu21, Pro28, and Gln22 (Figure 3(a)). For hesperidin, there are three amino acids (Glu21 and Glu100) generated H-bond interaction, and the surrounding residues are Trp32, Pro28, and Lys30 (Figure 3(b)). THSG has two amino acids generated H-bond for ligand binding, which are Glu21, Lys30 and Glu100, and amino acids that include Lys23, Pro28, and Trp32 are near the docked ligand (Figure 3(c)). It is worth to know that Glu100 is the common residue for each ligand binding, and the Lys30 can be found in all binding residues of mutant SOD1. In further study, we utilized molecular dynamics simulation to analyze variation of each ligand in protein structure.
Figure 1.
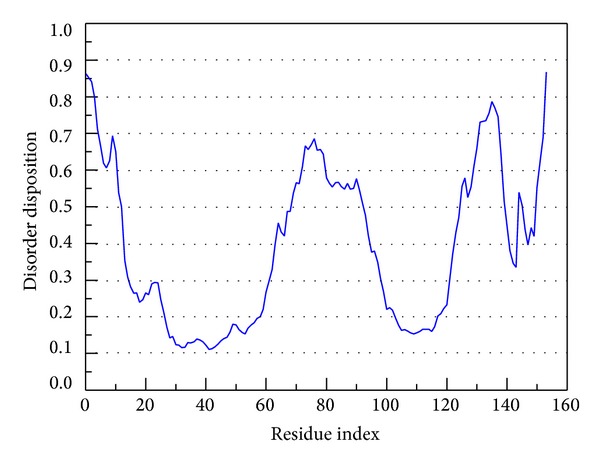
Disorder analysis of sequence of mutant SOD1 from result of PONDR-FIT prediction. The value of disorder disposition above 0.5 in disorder disposition is indicated as disorder residues.
Table 1.
Results of TCM compounds interacted with mutant SOD1 structure by LigandFit docking analysis.
| Name | -PLP1 | -PLP2 | -PMF | DOCK SCORE |
|---|---|---|---|---|
| Hesperidin | 33.72 | 39.61 | 95.25 | 91.91 |
| THSG | 37.36 | 41.61 | 101.23 | 88.16 |
| Hyperoside | 32.16 | 41.58 | 79.96 | 79.85 |
| Dopamine* | 26.79 | 30.68 | 37.71 | 51.94 |
| Nobiletin | 44.73 | 44.01 | 109.57 | 41.14 |
| Ursolic acid | 36.61 | 36.50 | 114.59 | 34.69 |
| Tangeretin | 29.24 | 32.45 | 73.20 | 33.28 |
| Nobiletin | 59.41 | 46.54 | 117.14 | 29.82 |
| Lupeol | 42.26 | 42.03 | 115.03 | 23.97 |
| Emodin | 30.29 | 32.13 | 84.07 | 22.02 |
| Physcion | 39.79 | 37.57 | 98.34 | 19.94 |
*Control.
Figure 2.
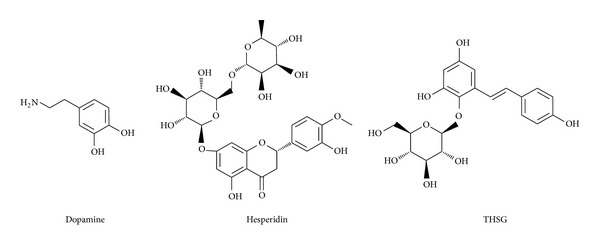
Chemical scaffold of dopamine, hesperidin, and THSG.
Figure 3.
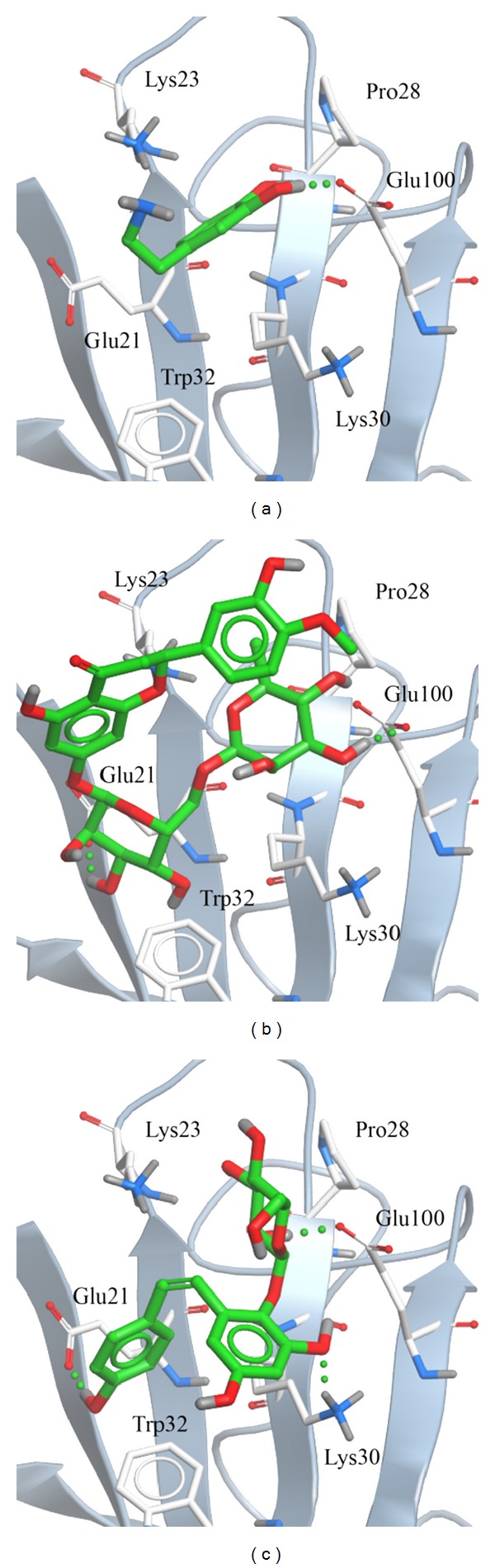
Binding poses of docking result: (a) dopamine, (b) hesperidin, and (c) THSG.
3.2. Stability Analysis of Molecular Dynamics Simulation
To determine stability of conformations among MD simulations, we utilized root mean square deviation (RMSD), radius of protein gyration, and total energy to analyze deviation of all complexes with docked ligand. The RMSD values of protein structure were used to verify stability among MD simulations. Figure 4 shows all values within the range from 0.16 to 0.24 nm, which indicate that all protein structures from protein-ligand complexes are stable during 5000 ps simulation, and the 5000 ps simulation time is enough for decreasing the fluctuation of all complexes. For ligand RMSD (Figure 5), it is obvious that the conformation of dopamine displayed high degree of difference during dynamic simulation, and the value of ligand RMAD increased to 0.24 nm from 3000 ps to the end. Ligand RMSD of TCM candidates shows slightly deviation, and the values of hesperidin and THSG are in average of 0.15 and 0.10, respectively.
Figure 4.
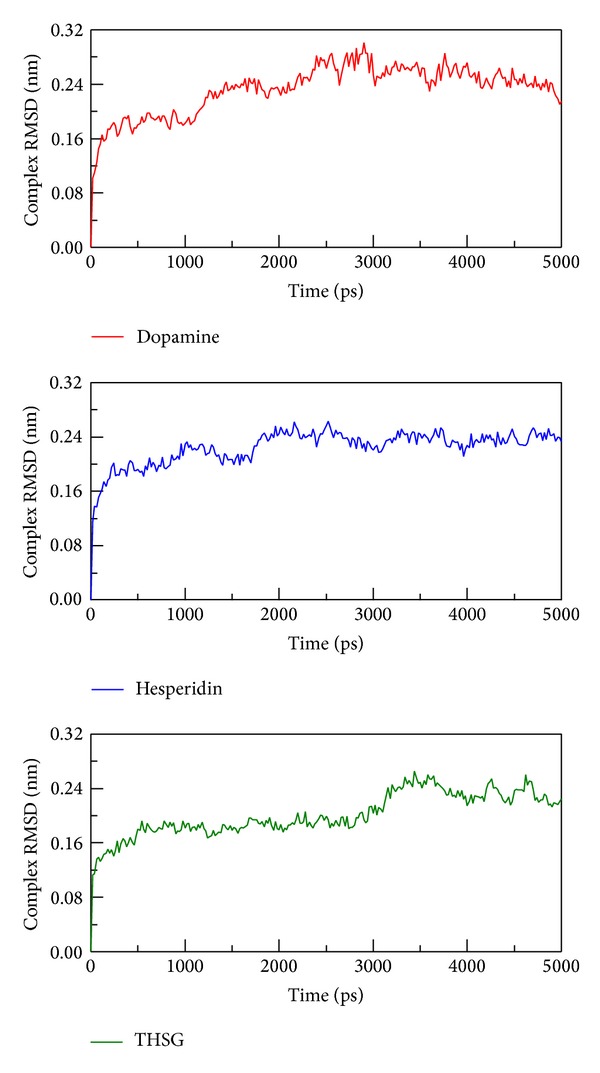
RMSD values of complex structures with docked ligand: dopamine, hesperidin, and THSG among 5000 ps simulation.
Figure 5.
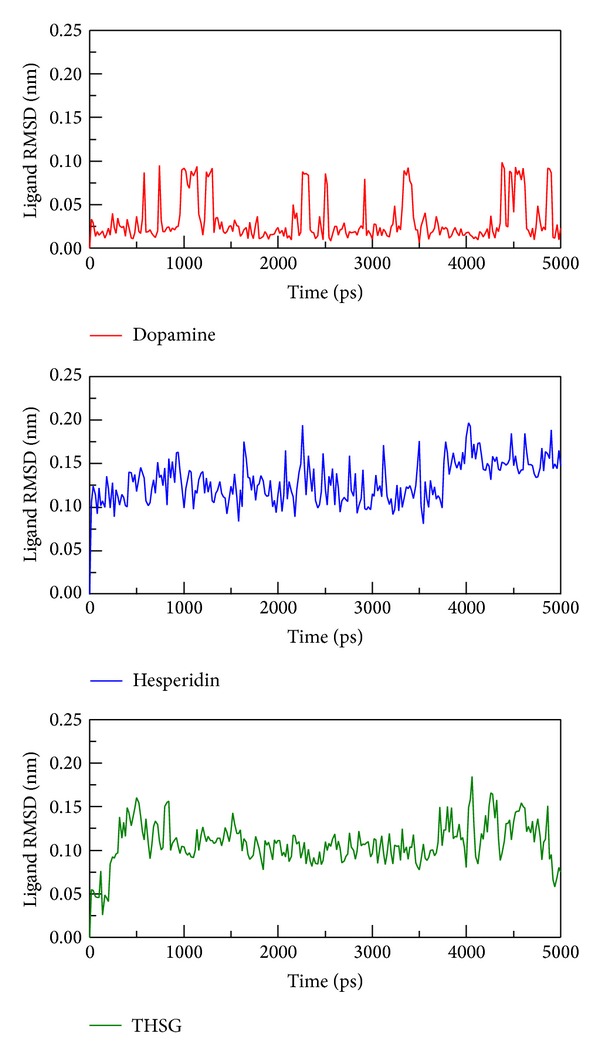
RMSD values of three compounds: dopamine, hesperidin, and THSG in protein complex during 5000 ps simulation time.
3.3. Migration Analysis of Molecules
In order to assess the variation of each ligand after being docked into protein binding site, MSD analysis was used to measure the migration of docked ligand during MD simulation. MSD value of hesperidin is the most distinct from the other TCM candidates, which displayed a rapid increase during initial simulation to the end of 5000 ps (Figure 6). In further study, we measured the distance between centers mass of protein and each ligand among all simulation times to understand movement of docked compounds. Interestingly, hesperidin shows a long distance with 2.6 nm (Figure 7) and turned to 2.0 nm after 2000 ps. Indicating that hesperidin moved away from the initial binding position and transferred to another site of the protein structure. The results suggest that each ligand could bind with mutant SOD1 during 5000 ps.
Figure 6.
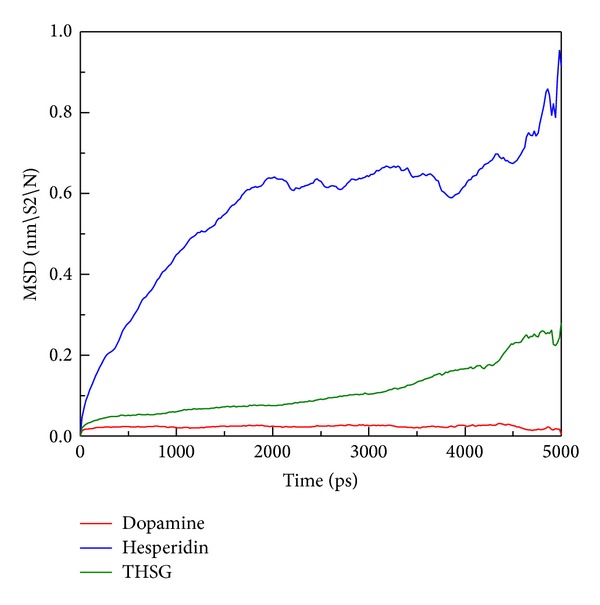
MSD values of three compounds: dopamine, hesperidin, and THSG in protein complex during 5000 ps simulation time.
Figure 7.
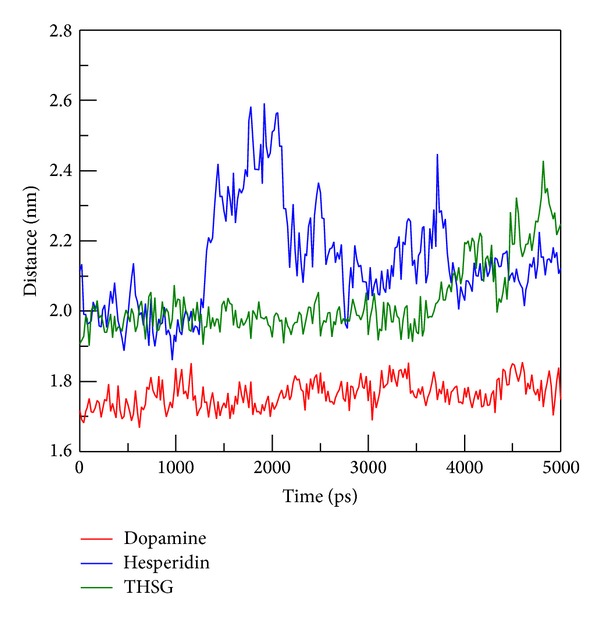
Distance between the centers of mass of mutant SOD1 and ligands.
3.4. Flexibility of Residues Analysis
We calculated root mean squared fluctuation (RMSF) to analyze the flexibility of residues on protein structure, and the binding region (from 21 to 32 and from 98 to 100 residues) shows no significant increment on structure of mutant SOD1 with all ligands (Figure 8). From DSSP analysis, all helices and beta-sheets of secondary structure for all complexes continued to exist during 5000 ps simulation times, and the number of residues is not variable among all conformations (Figure 9). The results suggest that the protein structure in each complex remained stable after MD simulations.
Figure 8.
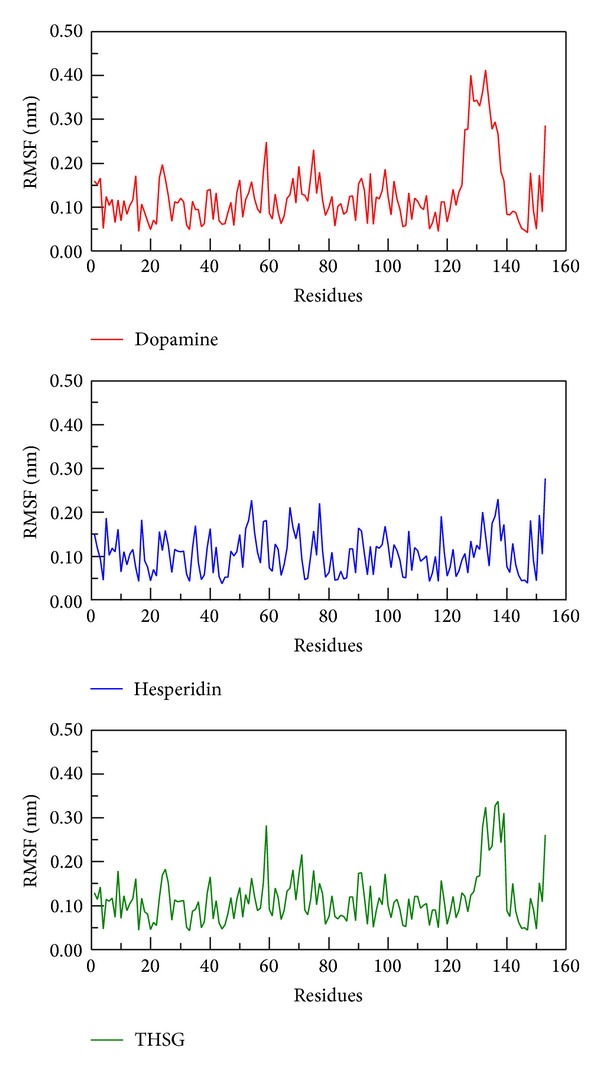
RMSF values of protein residues with docked ligand: dopamine, hesperidin, and THSG among 5000 ps simulation.
Figure 9.
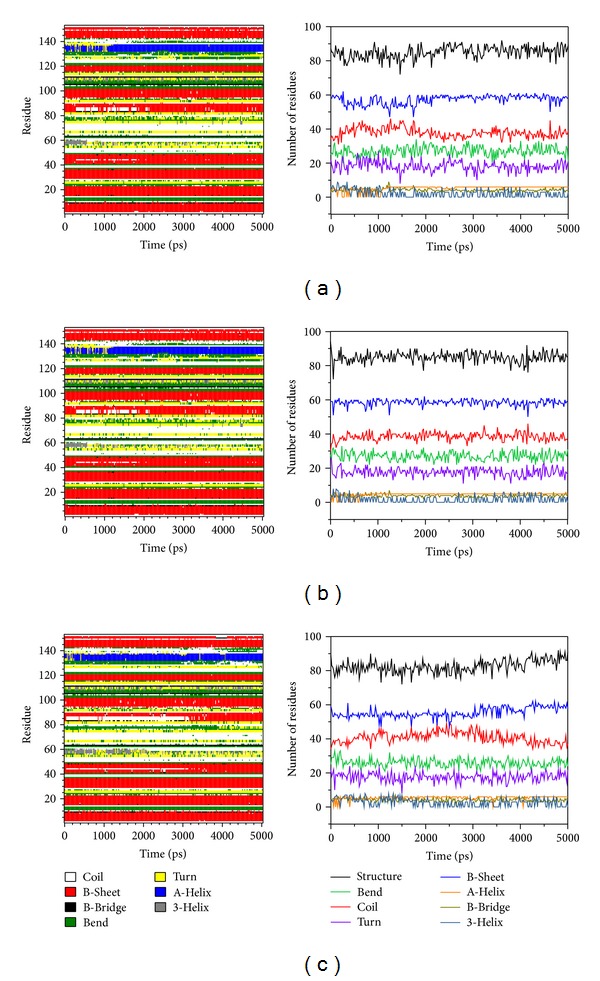
DSSP analysis of complexes with ligands: (a) dopamine (b) hesperidin, and (c) THSG. The “Structure” is summarized by residue number of A-Helix, B-Sheet, B-Bridge, and Turn.
3.5. Snapshots Analysis
In order to identify the most stable structure during the entire MD simulation for understanding the interaction of docked ligands, all conformations from MD simulation were clustered into seventeen or eighteen groups (Figure 10). We pick up the middle conformation from final groups of clusters as represented structure, and each middle frame is listed in Table 2. In the next study, we analyze protein-ligand interactions of represented structure (Figure 11). Dopamine has H-bond with Glu100 and Glu21, and the pi interaction is formed in Lys23 (Figure 11(a)). Hesperidin generate two H bonds with Lys23 and Glu21; besides, there are two pi interactions displayed on Lys30 and Trp32, respectively (Figure 11(b)). For THSG binding interaction, H bond was found on Glu24 and pi interaction was formed on Lys30 (Figure 11(c)). This result shows that hesperidin and THSG have similar binding residues to dopamine, suggesting that the binding conformations of two candidates are not significantly variable after MD simulations.
Figure 10.
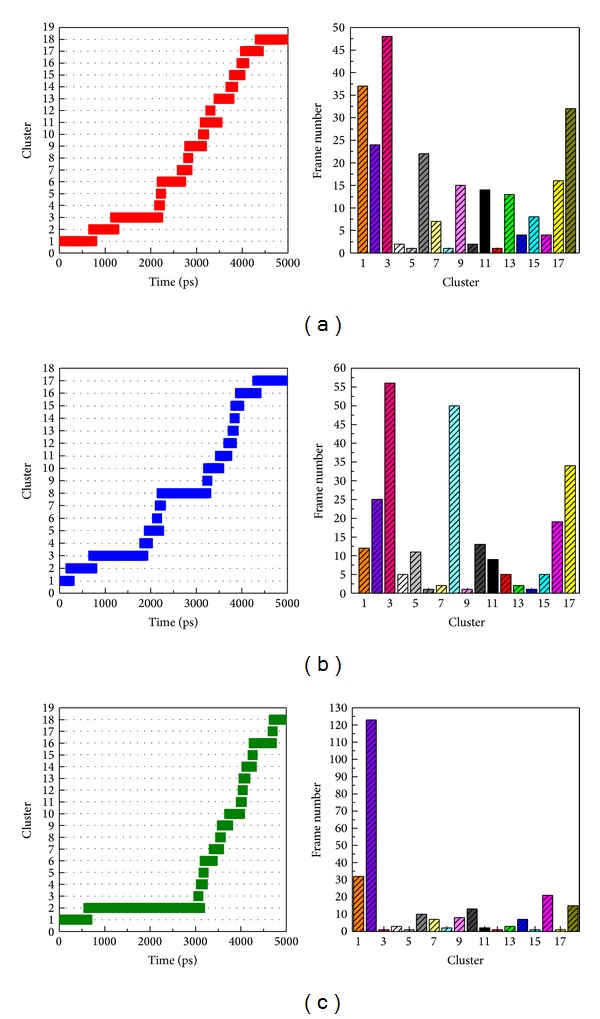
Clustering analyses of all conformations of mutant SOD1 complexes among 5000 ps simulation times.
Table 2.
Time of middle structure in each cluster among MD simulation times.
| Cluster | Time of middle frame (ps) | ||
|---|---|---|---|
| Dopamine | Hesperidin | THSG | |
| 1 | 460 | 160 | 300 |
| 2 | 940 | 360 | 2080 |
| 3 | 1520 | 1480 | 3060 |
| 4 | 2180 | 1880 | 3140 |
| 5 | 2220 | 2040 | 3180 |
| 6 | 2500 | 2140 | 3280 |
| 7 | 2740 | 2200 | 3460 |
| 8 | 2820 | 2920 | 3540 |
| 9 | 2960 | 3240 | 3660 |
| 10 | 3140 | 3380 | 3840 |
| 11 | 3380 | 3580 | 4000 |
| 12 | 3300 | 3740 | 4040 |
| 13 | 3560 | 3800 | 4080 |
| 14 | 3780 | 3840 | 4160 |
| 15 | 3900 | 3900 | 4260 |
| 16 | 4020 | 4180 | 4580 |
| 17 | 4280 | 4620 | 4700 |
| 18 | 4740 | — | 4860 |
Figure 11.
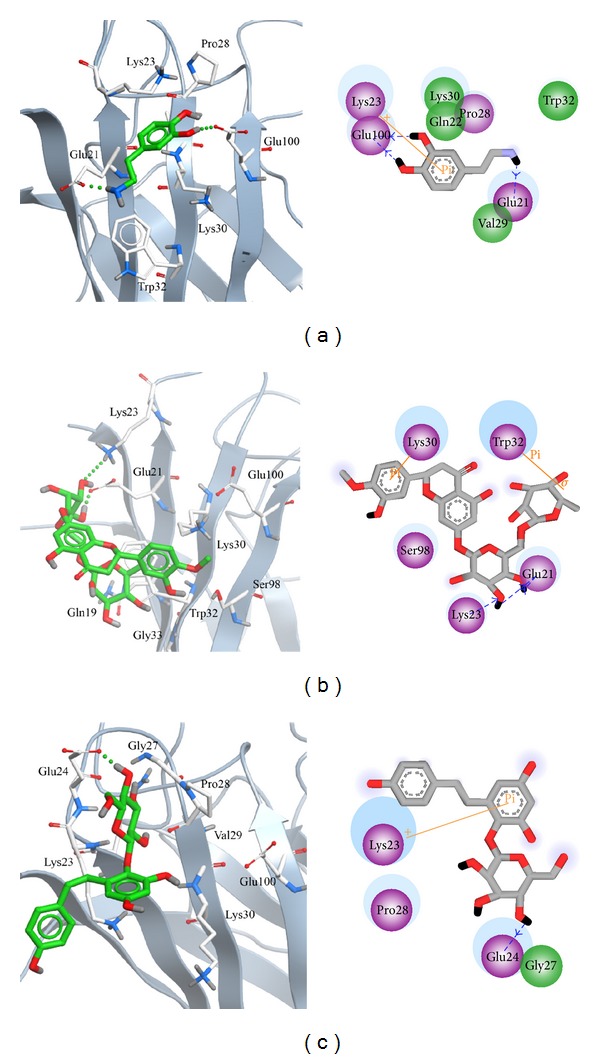
Binding poses of each representative conformation of mutant SOD1 from cluster analyses in 2D and 3D diagrams. Pi interactions (orange), polar residues (purple circle), and van der Waals residues (green circle) are displaced in 2D diagram.
4. Conclusion
For docking analysis, both Hesperidin and THSG have higher Dock Score than dopamine, and they displayed stable movements for mutant SOD1 from MSD and center mass distance analysis, which was correlated with the low affinity in docking results. For RMSF and DSSP assay, the secondary structure of mutant SOD1 did not change significantly during MD simulation, suggesting that the docked ligands are not affected by protein structure. Hesperidin and THSG have high affinity with SOD1 and the binding interactions are similar to dopamine among all molecular simulations. Our result indicated that hesperidin and THSG might be potential lead compound to design inhibitors of mutant SOD1 for ALS therapy.
Acknowledgments
The research was supported by grants from the National Science Council of Taiwan (NSC102-2325-B039-001, NSC102-2221-E-468-027-), Asia University (ASIA100-CMU-2, ASIA101-CMU-2, 102-ASIA-07), China Medical University (CMU99-TC-25), and China Medical University Hospital (DMR-103-058, DMR-103-001, and DMR-103-096). This study is also supported in part by Taiwan Department of Health Clinical Trial and Research Center of Excellence (DOH102-TD-B-111-004), Taiwan Department of Health Cancer Research Center of Excellence (MOHW103-TD-B-111-03), and CMU under the AIM for Top University Plan of the Ministry of Education, Taiwan.
Conflict of Interests
The authors reaffirm that they have no conflict of interests to declare.
Authors' Contribution
Hung-Jin Huang, Tung-Ti Chang, and Hsin-Yi Chen contributed equally to this paper.
References
- 1.Brown RH., Jr. SOD1 aggregates in ALS: cause, correlate or consequence? Nature Medicine. 1998;4(12):1362–1364. doi: 10.1038/3945. [DOI] [PubMed] [Google Scholar]
- 2.Wang L, Popko B, Roos RP. An enhanced integrated stress response ameliorates mutant SOD1-induced ALS. Human Molecular Genetics. 2014;23(10):2629–2638. doi: 10.1093/hmg/ddt658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Brasil AA, Belati A, Mannarino SC, Panek AD, Eleutherio ECA, Pereira MD. The involvement of GSH in the activation of human Sod1 linked to FALS in chronologically aged yeast cells. FEMS Yeast Research. 2013;13(5):433–440. doi: 10.1111/1567-1364.12045. [DOI] [PubMed] [Google Scholar]
- 4.Rosen DR, Siddique T, Patterson D, et al. Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature. 1993;362(6415):59–62. doi: 10.1038/362059a0. [DOI] [PubMed] [Google Scholar]
- 5.Tandan R, Bradley WG. Amyotrophic lateral sclerosis: part I. Clinical features, pathology, and ethical issues in management. Annals of Neurology. 1985;18(3):271–280. doi: 10.1002/ana.410180302. [DOI] [PubMed] [Google Scholar]
- 6.Tandan R, Bradley WG. Amyotrophic lateral sclerosis: part 2. Etiopathogenesis. Annals of Neurology. 1985;18(4):419–431. doi: 10.1002/ana.410180402. [DOI] [PubMed] [Google Scholar]
- 7.Staats KA, van Helleputte L, Jones AR, et al. Genetic ablation of phospholipase C delta 1 increases survival in SOD1(G93A) mice. Neurobiology of Disease. 2013;60:11–17. doi: 10.1016/j.nbd.2013.08.006. [DOI] [PubMed] [Google Scholar]
- 8.Shinde S, Arora N, Bhadra U. A complex network of micrornas expressed in brain and genes associated with amyotrophic lateral sclerosis. International Journal of Genomics. 2013;2013:12 pages. doi: 10.1155/2013/383024.383024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Al-Chalabi A, Hardiman O. The epidemiology of ALS: a conspiracy of genes, environment and time. Nature Reviews Neurology. 2013;9(11):617–628. doi: 10.1038/nrneurol.2013.203. [DOI] [PubMed] [Google Scholar]
- 10.Valentine JS, Hart PJ. Misfolded CuZnSOD and amyotrophic lateral sclerosis. Proceedings of the National Academy of Sciences of the United States of America. 2003;100(7):3617–3622. doi: 10.1073/pnas.0730423100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ip P, Mulligan VK, Chakrabartty A. ALS-causing SOD1 mutations promote production of copper-deficient misfolded species. Journal of Molecular Biology. 2011;409(5):839–852. doi: 10.1016/j.jmb.2011.04.027. [DOI] [PubMed] [Google Scholar]
- 12.Shi P, Ström A-L, Gal J, Zhu H. Effects of ALS-related SOD1 mutants on dynein- and KIF5-mediated retrograde and anterograde axonal transport. Biochimica et Biophysica Acta: Molecular Basis of Disease. 2010;1802(9):707–716. doi: 10.1016/j.bbadis.2010.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zinman L, Liu HN, Sato C, et al. A mechanism for low penetrance in an ALS family with a novel SOD1 deletion. Neurology. 2009;72(13):1153–1159. doi: 10.1212/01.wnl.0000345363.65799.35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Muller FL, Liu Y, Jernigan A, Borchelt D, Richardson A, van Remmen H. MnSOD deficiency has a differential effect on disease progression in two different ALS mutant mouse models. Muscle and Nerve. 2008;38(3):1173–1183. doi: 10.1002/mus.21049. [DOI] [PubMed] [Google Scholar]
- 15.Yoshii Y, Otomo A, Pan L, Ohtsuka M, Hadano S. Loss of glial fibrillary acidic protein marginally accelerates disease progression in a SOD1H46R transgenic mouse model of ALS. Neuroscience Research. 2011;70(3):321–329. doi: 10.1016/j.neures.2011.03.006. [DOI] [PubMed] [Google Scholar]
- 16.Winkler DD, Schuermann JP, Cao X, et al. Structural and biophysical properties of the pathogenic SOD1 variant H46R/H48Q. Biochemistry. 2009;48(15):3436–3447. doi: 10.1021/bi8021735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kokubo Y, Kuzuhara S, Narita Y, et al. Accumulation of neurofilaments and SOD1-immunoreactive products in a patient with familial amyotrophic lateral sclerosis with I113T SOD1 mutation. Archives of Neurology. 1999;56(12):1506–1508. doi: 10.1001/archneur.56.12.1506. [DOI] [PubMed] [Google Scholar]
- 18.Ince PG, Shaw PJ, Slade JY, Jones C, Hudgson P. Familial amyotrophic lateral sclerosis with a mutation in exon 4 of the Cu/Zn superoxide dismutase gene: pathological and immunocytochemical changes. Acta Neuropathologica. 1996;92(4):395–403. doi: 10.1007/s004010050535. [DOI] [PubMed] [Google Scholar]
- 19.Honjo Y, Kaneko S, Ito H, et al. Protein disulfide isomerase-immunopositive inclusions in patients with amyotrophic lateral sclerosis. Amyotrophic Lateral Sclerosis. 2011;12(6):444–450. doi: 10.3109/17482968.2011.594055. [DOI] [PubMed] [Google Scholar]
- 20.Cudkowicz ME, McKenna-Yasek D, Sapp PE, et al. Epidemiology of mutations in superoxide dismutase in amyotrophic lateral sclerosis. Annals of Neurology. 1997;41(2):210–221. doi: 10.1002/ana.410410212. [DOI] [PubMed] [Google Scholar]
- 21.Aoki M, Ogasawara M, Matsubara Y, et al. Familial amytrophic lateral sclerosis (ALS) in Japan associated with H46R mutation in Cu/Zn superoxide dismutase gene: a possible new subtype of familial ALS. Journal of the Neurological Sciences. 1994;126(1):77–83. doi: 10.1016/0022-510x(94)90097-3. [DOI] [PubMed] [Google Scholar]
- 22.Nakamura S, Wate R, Kaneko S, et al. An autopsy case of sporadic amyotrophic lateral sclerosis associated with the I113T SOD1 mutation. Neuropathology. 2014;34(1):58–63. doi: 10.1111/neup.12049. [DOI] [PubMed] [Google Scholar]
- 23.Lopate G, Baloh RH, Al-Lozi MT, et al. Familial ALS with extreme phenotypic variability due to the I113T SOD1 mutation. Amyotrophic Lateral Sclerosis. 2010;11(1-2):232–236. doi: 10.3109/17482960902898069. [DOI] [PubMed] [Google Scholar]
- 24.Schmidlin T, Kennedy BK, Daggett V. Structural changes to monomeric CuZn superoxide dismutase caused by the familial amyotrophic lateral sclerosis-associated mutation A4V. Biophysical Journal. 2009;97(6):1709–1718. doi: 10.1016/j.bpj.2009.06.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Bruijn LI, Houseweart MK, Kato S, et al. Aggregation and motor neuron toxicity of an ALS-linked SOD1 mutant independent from wild-type SOD1. Science. 1998;281(5384):1851–1854. doi: 10.1126/science.281.5384.1851. [DOI] [PubMed] [Google Scholar]
- 26.Auclair JR, Boggio KJ, Petsko GA, Ringe D, Agar JN. Strategies for stabilizing superoxide dismutase (SOD1), the protein destabilized in the most common form of familial amyotrophic lateral sclerosis. Proceedings of the National Academy of Sciences of the United States of America. 2010;107(50):21394–21399. doi: 10.1073/pnas.1015463107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Chang S-S, Huang H-J, Chen CY-C. High performance screening, structural and molecular dynamics analysis to identify H1 inhibitors from TCM Database@Taiwan. Molecular BioSystems. 2011;7(12):3366–3374. doi: 10.1039/c1mb05320e. [DOI] [PubMed] [Google Scholar]
- 28.Chen C-Y, Chang Y-H, Bau D-T, et al. Ligand-based dual target drug design for H1N1: swine flu—a preliminary first study. Journal of Biomolecular Structure and Dynamics. 2009;27(2):171–178. doi: 10.1080/07391102.2009.10507307. [DOI] [PubMed] [Google Scholar]
- 29.Chang T-T, Sun M-F, Chen H-Y, et al. Screening from the world’s largest TCM database against H1N1 virus. Journal of Biomolecular Structure and Dynamics. 2011;28(5):773–786. doi: 10.1080/07391102.2011.10508605. [DOI] [PubMed] [Google Scholar]
- 30.Huang H-J, Yu HW, Chen C-Y, et al. Current developments of computer-aided drug design. Journal of the Taiwan Institute of Chemical Engineers. 2010;41(6):623–635. [Google Scholar]
- 31.Chou IC, Lin W-D, Wang C-H, et al. Möbius syndrome in a male with XX/XY mosaicism. BioMedicine. 2013;3(2):102–104. [Google Scholar]
- 32.Chou IC, Lin W-D, Wang C-H, et al. Association analysis between Tourette's syndrome and two dopamine genes (DAT1, DBH) in Taiwanese children. BioMedicine. 2013;3(2):88–91. [Google Scholar]
- 33.Lee C-C, Tsai C-H, Wan L, et al. Increased incidence of Parkinsonism among Chinese with β-glucosidase mutation in central Taiwan. BioMedicine. 2013;3(2):92–94. [Google Scholar]
- 34.Wang C-H, Lin W-D, Bau D-T, et al. Appearance of acanthosis nigricans may precede obesity: an involvement of the insulin/IGF receptor signaling pathway. BioMedicine. 2013;3(2):82–87. [Google Scholar]
- 35.Wang C-H, Lin W-D, Tsai F-J. Craniofacial dysmorphism, what is your diagnosis? BioMedicine. 2012;2(2):49–50. [Google Scholar]
- 36.Lin W-Y, Liu H-P, Chang J-S, et al. Genetic variations within the PSORS1 region affect Kawasaki disease development and coronary artery aneurysm formation. BioMedicine. 2013;3(2):73–81. [Google Scholar]
- 37.Chen CY-C. Weighted equation and rules—a novel concept for evaluating protein-ligand interaction. Journal of Biomolecular Structure and Dynamics. 2009;27(3):271–282. doi: 10.1080/07391102.2009.10507315. [DOI] [PubMed] [Google Scholar]
- 38.Tsai T-Y, Chang K-W, Chen CY-C. IScreen: world’s first cloud-computing web server for virtual screening and de novo drug design based on TCM database@Taiwan. Journal of Computer-Aided Molecular Design. 2011;25(6):525–531. doi: 10.1007/s10822-011-9438-9. [DOI] [PubMed] [Google Scholar]
- 39.Huang H-J, Jian Y-R, Chen C-Y. Traditional Chinese medicine application in HIV: an in silico study. Journal of Biomolecular Structure and Dynamics. 2014;32(1):1–12. doi: 10.1080/07391102.2012.745168. [DOI] [PubMed] [Google Scholar]
- 40.Chang S-S, Huang H-J, Chen CY-C. Two birds with one stone? Possible dual-targeting H1N1 inhibitors from traditional Chinese medicine. PLoS Computational Biology. 2011;7(12) doi: 10.1371/journal.pcbi.1002315.e1002315 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Chen K-C, Sun M-F, Yang S-C, et al. Investigation into potent inflammation inhibitors from traditional Chinese medicine. Chemical Biology and Drug Design. 2011;78(4):679–688. doi: 10.1111/j.1747-0285.2011.01202.x. [DOI] [PubMed] [Google Scholar]
- 42.Yang S-C, Chang S-S, Chen H-Y, Chen CY-C. Identification of potent EGFR inhibitors from TCM Database@Taiwan. PLoS Computational Biology. 2011;7(10) doi: 10.1371/journal.pcbi.1002189.e1002189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Yang S-C, Chang S-S, Chen CY-C. Identifying HER2 inhibitors from natural products database. PLoS ONE. 2011;6(12) doi: 10.1371/journal.pone.0028793.e28793 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Tsou YA, Chen KC, Lin HC, Chang SS, Chen CY. Uroporphyrinogen decarboxylase as a potential target for specific components of traditional Chinese medicine: a virtual screening and molecular dynamics study. PLoS ONE. 2012;7(11) doi: 10.1371/journal.pone.0050087.e50087 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chen C-Y, Chen CY-C. Insights into designing the dual-targeted HER2/HSP90 inhibitors. Journal of Molecular Graphics and Modelling. 2010;29(1):21–31. doi: 10.1016/j.jmgm.2010.04.002. [DOI] [PubMed] [Google Scholar]
- 46.Chen CY-C, Chen Y-F, Wu C-H, Tsai H-Y. What is the effective component in suanzaoren decoction for curing insomnia? Discovery by virtual screening and molecular dynamic simulation. Journal of Biomolecular Structure and Dynamics. 2008;26(1):57–64. doi: 10.1080/07391102.2008.10507223. [DOI] [PubMed] [Google Scholar]
- 47.Chen KC, Chang SS, Huang HJ, et al. Three-in-one agonists for PPAR-alpha, PPAR-gamma, and PPAR-delta from traditional Chinese medicine. Journal of Biomolecular Structure and Dynamics. 2012;30(6):662–683. doi: 10.1080/07391102.2012.689699. [DOI] [PubMed] [Google Scholar]
- 48.Chen CY-C. Virtual screening and drug design for PDE-5 receptor from traditional Chinese medicine database. Journal of Biomolecular Structure and Dynamics. 2010;27(5):627–640. doi: 10.1080/07391102.2010.10508577. [DOI] [PubMed] [Google Scholar]
- 49.Chen C-Y, Chang Y-H, Bau D-T, et al. Discovery of potent inhibitors for phosphodiesterase 5 by virtual screening and pharmacophore analysis. Acta Pharmacologica Sinica. 2009;30(8):1186–1194. doi: 10.1038/aps.2009.100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Chen CY-C. Computational screening and design of Traditional Chinese Medicine (TCM) to block phosphodiesterase-5. Journal of Molecular Graphics and Modelling. 2009;28(3):261–269. doi: 10.1016/j.jmgm.2009.08.004. [DOI] [PubMed] [Google Scholar]
- 51.Chen K-C, Chang K-W, Chen H-Y, Chen CY-C. Traditional Chinese medicine, a solution for reducing dual stroke risk factors at once? Molecular BioSystems. 2011;7(9):2711–2719. doi: 10.1039/c1mb05164d. [DOI] [PubMed] [Google Scholar]
- 52.Chen HY, Chang SS, Chan YC, Chen CY. Discovery of novel insomnia leads from screening traditional Chinese medicine database. Journal of Biomolecular Structure and Dynamics. 2014;32(5):776–791. doi: 10.1080/07391102.2013.790849. [DOI] [PubMed] [Google Scholar]
- 53.Chen CY. Mechanism of BAG1 repair on Parkinson’s disease-linked DJ1 mutation. Journal of Biomolecular Structure and Dynamics. 2012;30(1):1–12. doi: 10.1080/07391102.2012.674182. [DOI] [PubMed] [Google Scholar]
- 54.Chang Y-M, Velmurugan BK, Kuo W-W, et al. Inhibitory effect of alpinate Oxyphyllae fructus extracts on Ang II-induced cardiac pathological remodeling-related pathways in H9c2 cardiomyoblast cells. BioMedicine. 2013;3(4):148–152. [Google Scholar]
- 55.Lin P-C, Liu P-Y, Lin S-Z, Harn H-J. Angelica sinensis: a Chinese herb for brain cancer therapy. BioMedicine. 2012;2(1):30–35. [Google Scholar]
- 56.Ho T-Y, Lo H-Y, Li C-C, Chen J-C, Hsiang C-Y. In vitro and in vivo bioluminescent imaging to evaluate anti-Escherichia coli activity of Galla Chinensis. BioMedicine. 2013;3(4):160–166. [Google Scholar]
- 57.Li C-C, Lo H-Y, Hsiang C-Y, Ho T-Y. DNA microarray analysis as a tool to investigate the therapeutic mechanisms and drug development of Chinese medicinal herbs. BioMedicine. 2012;2(1):10–16. [Google Scholar]
- 58.Hsu S-C, Chung J-G. Anticancer potential of emodin. BioMedicine. 2012;2(3):108–116. doi: 10.1016/j.biomed.2012.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Chen CY-C. TCM Database@Taiwan: the world’s largest traditional Chinese medicine database for drug screening in silico. PLoS ONE. 2011;6(1) doi: 10.1371/journal.pone.0015939.e15939 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Wright GS, Antonyuk SV, Kershaw NM, Strange RW, Hasnain SS. Ligand binding and aggregation of pathogenic SOD1. Nature Communications. 2013;4, article 1758 doi: 10.1038/ncomms2750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Accelerys. Discovery Studio Client V2.5. San Diego, Calif, USA: Accelrys; 2009. [Google Scholar]
- 62.Xue B, Dunbrack RL, Williams RW, Dunker AK, Uversky VN. PONDR-FIT: a meta-predictor of intrinsically disordered amino acids. Biochimica et Biophysica Acta: Proteins and Proteomics. 2010;1804(4):996–1010. doi: 10.1016/j.bbapap.2010.01.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Zhao Y, Kao CP, Chang YS, Ho YL. Quality assessment on Polygoni Multiflori Caulis using HPLC/UV/MS combined with principle component analysis. Chemistry Central Journal. 2013;7(1, article 106) doi: 10.1186/1752-153X-7-106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Allinger NL. Conformational analysis. 130. MM2. A hydrocarbon force field utilizing V1 and V2 torsional terms. Journal of the American Chemical Society. 1977;99(25):8127–8134. [Google Scholar]
- 65.Venkatachalam CM, Jiang X, Oldfield T, Waldman M. LigandFit: a novel method for the shape-directed rapid docking of ligands to protein active sites. Journal of Molecular Graphics and Modelling. 2003;21(4):289–307. doi: 10.1016/s1093-3263(02)00164-x. [DOI] [PubMed] [Google Scholar]
- 66.Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M. CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. Journal of Computational Chemistry. 1983;4(2):187–217. [Google Scholar]
- 67.Fletcher R, Reeves CM. Function minimization by conjugate gradients. The Computer Journal. 1964;7(2):149–154. [Google Scholar]
- 68.Press WH, Flannery BP, Teukolsky SA, Vetterling WT, Gould H. Numerical recipes, the art of scientific computing. American Journal of Physics. 1987;55(1):90–91. [Google Scholar]
- 69.Pronk S, Pall S, Schulz R, et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics. 2013;29(7):845–854. doi: 10.1093/bioinformatics/btt055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Tou WI, Chang SS, Lee CC, Chen CY-C. Drug design for neuropathic pain regulation from traditional Chinese medicine. Scientific Reports. 2013;3, article 844 doi: 10.1038/srep00844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Tou WI, Chen CY. May disordered protein cause serious drug side effect? Drug Discovery Today. 2014;19(4):367–372. doi: 10.1016/j.drudis.2013.10.020. [DOI] [PubMed] [Google Scholar]


