Abstract
Six endophytic bacteria of corn roots were identified as Bacillus sp. and as Enterobacter sp, by sequencing of the 16S rRNA gene. Four of the strains, CNPSo 2476, CNPSo 2477, CNPSo 2478 and CNPSo 2480 were positive for the nitrogen fixation ability evaluated through the acetylene reduction assay and amplification of nifH gene. Two Bacillus strains (CNPSo 2477 and CNPSo 2478) showed outstanding skills for the production of IAA, siderophores and lytic enzymes, but were not good candidates as growth promoters, because they reduced seed germination. However, the same strains were antagonists against the pathogenic fungi Fusarium verticillioides, Colletotrichum graminicola, Bipolaris maydis and Cercospora zea-maydis. As an indication of favorable bacterial action, Enterobacter sp. CNPSo 2480 and Bacillus sp. CNPSo 2481 increased the root volume by 44% and 39%, respectively, and the seed germination by 47% and 56%, respectively. Therefore, these two strains are good candidates for future testing as biological inoculants for corn.
Keywords: Molecular phylogeny, 16S rRNA, nifH, Plant growth promotion, Antagonism
Introduction
Literature shows that some bacteria, which live attached to plants have the ability to promote plant growth (Compant et al. [2010]; George et al. [2012]; Beneduzi et al. [2013]). These microorganisms have attracted attention because of the need to reduce the use of chemicals, especially when considering the context of sustainable agriculture and environmental protection (Vale et al. [2010]). One of the strategies is to exploit the benefits that several microorganisms may give to plants when added as inoculants (Lucy et al. [2004]).
Bacteria promoting plant growth can act directly, through one or more mechanisms, including biological nitrogen fixation (Ashraf et al. [2011]; Verma et al. [2013]), phosphate solubilization (Rodriguez et al. [2004]; Krey et al. [2013]), production of hormones such as auxins, gibberellins and zeatin (Cassan et al. [2009]), or act indirectly by means of biological control of pathogens (Wang et al. [2009]). In addition, some studies point out to the preferred action with endophytic microorganisms or adapted isolates, due to easier colonization and the lower risk to introduce exogenous organisms (Enebak et al. [1998]; Khalid et al. [2004]).
Several studies have shown the positive effects of endophytic bacteria inoculation in plants, e.g. sugarcane (Saccharum spp.), leading to increased contribution of biological nitrogen fixation, to promotion of root development, increased biomass and productivity (Oliveira et al. [2003]); soybean (Glycine max (L.) Merr.), with bacteria capable to inhibit growth and sporulation of pathogenic fungi (Assumpção et al. [2009]); tomato (Lycopersicum esculentum L.), with bacteria increasing plant height, leaf area, leaf number, together with fresh and dry plant weight (Barreti et al. [2008]).
Benefits provided by the inoculation of corn (Zea mays L.) with selected strains have also been described, including the promotion of growth due to the increased availability of nutrients, provided by inoculation with B. subtilis in seeds (Canbolat et al. [2006]). In addition, there are reports of strains of Azospirillum brasilense increasing corn yield by 24% to 30%, compared to the non-inoculated control (Hungria et al. [2010]).
However, for the selection of isolates that have a greater chance of being efficient in the process of colonization and growth promotion, the isolates have first to be identified with preliminary in vitro biochemical analysis (Pal et al. [2001]; Berg et al. [2002]; Bernardes et al. [2010]).Taxonomy of selected microorganisms with biotechnological potential in agriculture is thus a key step and, for this reason, sequencing of the 16S rRNA has been successfully used to identify genera such as Bacillus (Porwal et al. In [2009], Deepa et al. [2010]), Rhizobium, Burkholderia (Ferreira et al. [2011]), Enterobacter, Pantoea Serratia, among others (Tian et al. [2009]).
The aim of this study was to evaluate and characterize the properties of endophytic bacteria isolated from corn roots, with the potential to have biotechnological interest for their use in agriculture.
Material and methods
Biological material
The seven bacteria studied belong to the Laboratory of Microorganisms Genetics Collection (LABGEM) of the Federal University of Parana, and are deposited at the Diazotrophic and Plant Growth Promoting Bacteria Culture Collection of Embrapa Soja (WDCC Collection # 1054), with the record numbers from CNPSo 2475 to CNPSo 248. GeneBank accession numbers for the 16S rRNA go from JQ821359 to JQ821365. These bacteria were isolated from corn roots samples by Ikeda et al. ([2013]). For the isolation, corn roots from field trial conducted at Semilia Genetics and Breeding LTDA, located in the southern Brazilian region of Campo Largo-PR, were surface-sterilized (Petrini, [1986]) and aseptically cut, then transferred to Petri dish plates containing solid culture media. Several controls confirmed that the sterilization procedure was effective.
The fungi Cercospora zea-maydis (MMBF 51/08) and Bipolaris maydis (MMBF 46/08) were kindly provided by the Biological Institute of São Paulo, taken from the Micoteca “Mario Barreto Figueiredo”. The Fusarium verticillioides (CMM-1060) and Colletotrichum graminicola (CMM-1061) were deposited in the Culture Collection of Phytopathogenic Fungi “Prof. Maria Menezes”, with GenBank accession numbers for the 18S rRNA of KC335155 and KC335154.
Bacterium identification by analyzing the 16S rRNA gene
Extraction of bacterial DNA
Genomic DNA was obtained using the Raeder and Broda protocol ([1985]), with the following modifications: bacteria were grown overnight in 4 mL of DYGS medium (Rodrigues Neto et al. [1986]) and then centrifuged for 2 min at 10.000 g. Then, a 2 mL pellet of extraction buffer (125 mMol/L Tris–HCl pH 8.0; 2 Mol/L NaCl; 50 mMol/L EDTA pH 8.0 and 1% SDS pH 7.2) was added, preheated at 60°C. After drying, the DNA was re-suspended in 50 μL of ultrapure water.
The integrity was checked by electrophoresis on agarose gel 0.8% (w/v), stained with GelRedTM (Biotium, USA), diluted in a sample buffer for 100X, observed in a UV transluminator (Ultraviolet Benchtop transilluminators) and photo-documentation application (Program Digidoc it Quantification). Purity was evaluated by spectrophotometer (per wavelength) NanoDrop® 2000 (Thermo Scientific).
Amplification and purification of the coding region of the 16S rRNA gene
The amplification was performed by PCR (Polymerase Chain Reaction) with primers fD1 (5′AGA GTTTGATCCTGGCTCAG 3′) and rD1 (5′AAGGAGGTGATCCAGCC 3′) (Weisburg et al. [1991]). The cycle included an initial denaturation at 95°C for 2 min, 30 cycles of 15 s at 94°C, 45 s at 93°C, 45 s at 55°C, 2 min at 72°C, followed by a final extension of 5 min at 72°C. Purification with 7.5 M ammonium acetate was performed according to Menna et al. ([2006]). Ultrapure water was used to re-suspend the DNA. The amount and purity were assessed as described above on agarose gel 1.5% (w/v), adjusting the final concentration at 16 ng/μL.
Sequencing and analysis of the 16S rRNA gene
For the second PCR reaction, the following primers, designed by Prof. Leonardo M. Cruz (Dept. Biochemistry, Federal University of Paraná, Curitiba, PR, Brazil), were used: 362f (5′ CTCCTACGGGAGGCAGCAGTGGGG 3′), 786f (5′ CGAAAGCGTGGGGAGCAAACAGG 3′), in addition to the fD1 primer (Weisburg et al. [1991]). Eighty ng of the purified product from the first PCR reaction, 0.25 mM of primer and 2 μL ET mixture were used for sequencing. Ultrapure water was added to reach the final volume of 10 μL. Amplification was performed for 35 cycles of 30 s at 94°C, 15 s at 50°C and 90 s at 60°C.
After amplification, purification was performed with Sephadex™ filtration gel, G-50 medium (GE® Healthcare) and then the product was submitted to electrophoresis with an automated DNA sequencing analyser, model MegaBACE1000 (Amersham Biosciences®).
The quality of the bases and the fragment assembly were analyzed with the PhredPhrapConsed software (Ewing et al. [1998]). Alignments were carried out with the PRANK software (Löytynoja and Goldman [2010]). The percentage of similarity was obtained from the RDP (Ribosomal Database Project).
The phylogenetic analysis was made using Bayesian inference with the MrBayes 3.1.2 software (Ronquist and Huelsenbeck [2003]) using parallel computing (Altekar [2004]), with four chains. The evolutionary model was selected through the JModelTest program (Posada and Kandral [1998]). The number of generations and “burn-in” were defined by analysis of multiple sequences, with topological stability assessment by the AWTY package (Nylander et al. [2008]) and analyzed by the parametric stability software, Tracer 1.5. With appropriate values of “burn-in”, the final trees were summarized with the software package SumTrees DendroPy 3.8.1 (Sukumaran and Holder [2010]).
Amplification of the nitrogenase gene nifH
The PCR was performed with primers nifHF (5′ TACGGNAARGGSGGNATCGGCAA 3′) and nifHI (5′ AGCATGTCYTCSAGYTCNTCCA 3′) (Boldface letters denote degenerate positions N = A + C + G + T; R = A + G; S = G + C; Y = C + T; W = A + T), as follows: 1.5 mM MgCl2; 20 μM (each) dATP, dCTP, dTTP,dGTP; 0.1 μM (each) primers; 0.04 U Taq DNA polymerase/μL. The modified steps for amplification were: an initial denaturation at 94°C for 2 min, 30 cycles of 1 min 94°C, 1 min at 55°C, 3 min at 72°C, followed by a final extension of 5 min at 72°C (Laguerre et al. [2001]).
With another pair of primers nifH-F (5′ AAAGGYGGWATCGGYAARTCCACCAC 3′) and nifH-R (5′ TTGTTSGCSGCRTACATSGCCATCAT 3′), the following steps were performed in the PCR: a 4-min hot start at 97°C before Taq DNA polymerase was added; 1 cycle of denaturation at 96°C for 20 s, annealing at 65°C for 30 s, and elongation at 72°C for 30 s; 2 cycles of denaturation at 96°C for 20 s, annealing at 62°C for 30 s, and elongation at 72°C for 35 s; 3 cycles of denaturation at 96°C for 20 s, annealing at 59°C for 30 s, and elongation at 72°C for 40 s; 4 cycles of denaturation at 96°C for 20 s, annealing at 56°C for 30 s, and elongation at 72°C for 45 s; 5 cycles of denaturation at 96°C for 20 s, annealing at 53°C for 30 s, and elongation at 72°C for 50 s; 25 cycles of denaturation at 94°C for 20 s, annealing at 50°C for 45 s, and elongation at 72°C for 60 s; and an extension at 72°C for 10 min (Rösch et al. [2002]). Products of the PCR were confirmed in 1.5% (w/v) agarose gels. A Low DNA Mass Ladder (Invitrogen®) and photo-documentation, as described above.
Enzymatic and physiological characterization of bacteria
Biological nitrogen fixation
Biological nitrogen fixation was verified according to the methodology of Döbereiner et al. ([1995]), by using NFb semi-solid medium (Baldani et al. [1986]). The target was the formation of a sub-surface pellicle.
The nitrogen-fixation ability was confirmed by using the ARA (acetylene reduction assay), as described by Boddey ([1987]). Each bacterial isolate was inoculated into 10 mL vials containing the respective 4 mL semi-solid N-free medium NFb (Baldani et al. [1986]), TBNR (Seldin et al. [1984]) and LGD (Döbereiner [1980]). All isolates were incubated at 30°C in the dark. After 24 h, 10% (v/v) of the air phase was replaced with acetylene (Burris, [1972]) and then the vials were incubated again. After the addition of acetylene, three vials for each isolate were sampled over 1 h to determine the amount of ethylene. Ethylene was measured using a Clarus 600 gas chromatograph (PerkinElmer®) with a Col-Elite-Alumina column and a flame ionization detector connected to a chromatography data computer system. Azospirillum brasilense Ab-V5 was used as positive control.
Production of indoleacetic acid
Quantification of indoleacetic acid (IAA) was performed according to Kuss et al. ([2007]). Single bacterial colonies were inoculated in 2 mL of DYGS medium supplemented with 2 mg/mL of tryptophan and incubated at 30°C for 72 h at 180 rpm. IAA was measured by mixing 1 mL broth with 1 mL of Salkowsky’s reagent (Loper and Schroth, [1986]) and incubated for 30 min at room temperature. Bacterial cells were separated from the supernatant by centrifugation at 10,000 rpm for 5 min and absorbance at 530 nm was determined. A standard IAA curve was plotted with commercial auxin (Sigma Aldrich, USA) and correlated with the absorption of the bacterial strains. Values are expressed in μg/mL.
Siderophore production and solubilization of inorganic calcium phosphate
The confirmation of siderophore production was performed according to Schwyn Neilands ([1987]) universal method, with Chrome Azurol S-(CAS) in Petri dishes, with DYGS solid medium. The spot was inoculated with test organism and incubated at 30°C for 3–5 days. Development of a yellow–orange halo around the colony was considered as a positive result.
Solubilization of calcium phosphate was done according to Sylvester-Bradley et al. ([1982]). Bacteria were grown in GY (glucose yeast medium) (10 g glucose, 2 g yeast extract and 15 g agar per liter). Two other solutions were prepared separately, one containing 5 g K2HPO4 in 50 mL of distilled water, and the other containing 10 g CaCl2 in 100 mL of distilled water. These solutions were added to one liter of GY medium just before pouring onto Petri dishes, and together they formed an insoluble layer of calcium phosphate that made the medium opaque. Clear halos around the colony were recorded as a positive result (after 12 days).
Production of ammonia and hydrocyanic acid
Confirmation of the production of ammonia was done by colorimetric reaction, by adding 1 mL Nessler’s reagent to a 72-h-old culture grown in peptone broth and recording the presence of the yellowish brown color (Dye [1962]).
For hydrocyanic acid (HCN), the Bakker and Schippers ([1987]) method was followed in King B medium (King et al. [1954]), supplemented with 4.4 g/L of glycine. HCN production was inferred with change in the color of the filter paper previously dipped in 2% sodium carbonate prepared in 0.05% picric acid, and it was rated visually depending on the intensity of the color change from yellow to dark brown.
Detection of lytic enzymes
Bacteria were seeded in a mineral medium containing 0.5% carboxymethylcellulose as sole carbon source for detection of cellulose activity. Colloidal chitin at 0.08%, obtained according to Moura et al. ([2005]) for the detection of chitinase and 0.5% of laminarin for the detection of β-1,3-glucanase (Renwick et al. [1991]) were used.
The amylolytic and pectinolytic activities were evaluated after Hankin and Anagnostakis ([1975]), in a minimal M9 medium (Sambrook and Russell [2001]) containing 0.5% of yeast extract and 0.2% soluble starch (v/w) to test for amylase, and pectin 0.5% (v/w) to test for pectinase.
The proteolytic activity of strains was verified through the degradation of casein in half-skimmed milk agar (Berg et al. [2002]). The appearance of a halo around the colonies was recorded as positive (after seven days).
Antifungal activity
The paired cultures test was performed according to Shiomi et al. ([2008]), with potato dextrose agar (PDA) medium, using two equidistant grooves and a 5 mm disc of fungi mycelia placed in the center of the plate, grown for five days. The inhibition activity was evaluated by measuring the radius of fungal growth and comparing it with the control that was not treated with bacteria.
Germination of inoculated corn seeds
A bacterial cell suspension was incubated at 30°C for 24 hours at 150 rpm in DYGS medium. Then the suspension was centrifuged at 5.000 g and re-suspended in saline solution (0.85% NaCl). The concentration was adjusted to 2×109 CFU/mL, in a spectrophotometer, to OD540 nm, from an exponential growth curve previously performed. One mL of this solution was inoculated into 100 commercial hybrid corn seeds SX2530 (Semilia LTD), previously surface disinfested (Shiomi et al. [2008]); tissues were also internally disinfected with warm water to eliminate fungi (Daniels [1983]). To confirm the disinfection, samples of seeds were germinated in Petri dishes with PDA medium. The amount of cells in the seed was measured by doing plate counts (CFU). Bacteria were extracted by using 0.085% saline solution agitated by vortexing (Silva and Reis [2004]). Seeds were germinated in an incubator at 28°C, on filter paper moistened with distilled water. Evaluations of the length (cm) and volume (cm3) of the root and of the hypocotyl were made after 4 and 7 days (Cassán et al. [2009]) using the Win-rhizo v.4.0 software (Regent Systems, Quebec, Canada). Statistical analysis were performed on the Assistat 7.6 Beta software, and the data were set to 5% confidence level (p > 0.05).
Results
Molecular phylogeny of the bacterial isolates
After aligning the sequences of the gene 16S rRNA, a region of 1512 bp was used for phylogenetic analysis, along with type strains taken from the RDP. Strain CNPSo 2480 grouped with the Enterobacter branch with 100% Bayesian posterior probability (BPP). This same strain had a 96% BPP and 98% similarity with type strain E. asburiae JCM6051T. The six other bacterial strains belong to the genus Bacillus. Among them, CNPSo 2481 showed 100% BPP and 98% similarity with B. pumilus DSMZ27T (Figure 1).
Figure 1.
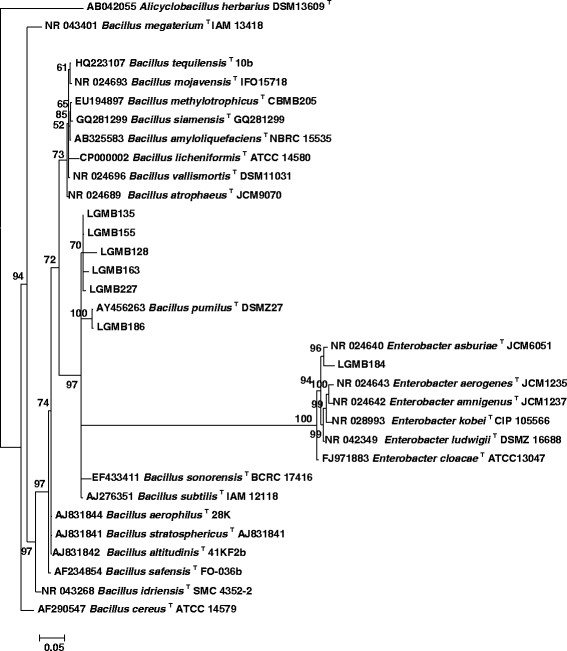
Phylogenetic tree based on the 16S rRNA gene sequence of the isolates in this study and reference strains obtained from the RDP. Used were 1512 bp fragment after alignment. Data generated using Bayesian inference, with outgroup Alicyclobacillus herbarius. Suportes branches less than 50% were not shown.
The sequences of strains CNPSo 2479 (LGMB227), CNPSo 2476 (LGMB163), CNPSo 2477 (LGMB128), CNPSo 2480 (LGMB184), CNPSo 2481 (LGMB186), CNPSo 2475 (LGMB155), CNPSo 2478 (LGMB135) were deposited at the Genbank and received the accession numbers of JQ821359 to JQ821365.
Amplification of nifH gene
Strains Bacillus CNPSo 2477 and CNPSo 2476 were positive for nifH gene amplification (primers nifHF and nifHI), producing an amplified fragment of about 780 bp. Enterobacter sp. CNPSo 2480 was positive for the two pairs of primers tested (nifHF and nifHI; nifH-F and nifH-R). Bacillus sp CNPSo 2478, amplified with the second primer, resulted in a fragment of about 400 pb.
Enzymatic and physiological characterization of bacteria
In relation to the nitrogen fixation capacity evaluated in vitro, all strains were able to grow in NFb semi-solid medium and showed the typical pellicle. But only Enterobacter sp. CNPSo 2480 showed the ability to reduce acetylene to ethylene in NFb N-free semi-solid medium. Bacillus CNPSo 2476, CNPSo 2477and CNPSo 2478 showed the same property in TBNR and LGD.
Only Enterobacter sp. CNPSo 2480 did not produce siderophore and Bacillus sp. CNPSo2481 produced ammonia.
All strains were positive for IAA production with values between 35.1 and 96.3 μg/mL, especially Bacillus CNPSo 2477 and CNPSo 2478, with 105.2 and 105.11 μg/mL respectively.
Lytic enzymes activities were detected in the following strains: cellulase CNPSo 2477, CNPSo 2475 and CNPSo 2476; amylase CNPSo 2477, CNPSo 2478, CNPSo 2475 and CNPSo 2476; pectinase CNPSo 2477, CNPSo 2478, CNPSo 2475, CNPSo 2476 and CNPSo 2479; protease, for degradation of casein CNPSo 2478, CNPSo 2475, CNPSo 2476, CNPSo 2481 and CNPSo 2479. All strains were negative for all phosphate solubilization tests, production of HCN, chitinase and β1,3-glucanase.
Antifungal activity
The seven bacterial strains significantly reduced the growth of fungi, with inhibition above 70% of the following pathogens: CNPSo 2477 (B. maydis, C. graminicola and C. zea-maydis); CNPSo 2478 (B. maydis and C. graminicola); CNPSo 2475 (C. zea-maydis); and CNPSo 2476, CNPSo 2480, CNPSo 2481, CNPSo 2479 (C. graminicola and C. zea-maydis).
Germination of inoculated corn seeds
The inoculation resulted in 107 CFU/seed. The strains Bacillus sp. CNPSo 2477 and CNPSo 2478 reduced the germination percentage by about 27%. In contrast, germination increased to approximately 52% with CNPSo 2475, CNPSo 2480 and CNPSo 2481. CNPSo 2476 and CNPSo 2479 increased germination to about 36.5% and 15.8%, respectively. The bacteria CNPSo 2476, CNPSo 2480 and CNPSo 2481 promoted root volume increases (14.6%, 43.9% and 39.02% respectively). CNPSo 2480 and CNPSo 2481 also promoted an increment in root length. No changes were observed in hypocotyl length and volume.
Discussion
Sequencing of 16S rRNA allows the accurate identification of bacterial genera endophytes from various plant species, including: sugar cane (Ratón et al. [2011]), corn (Pereira et al. [2011]) and rice (Oryza sativa L.) (Beneduzi et al. [2008]). However, for the precise definition of species and even subspecies such as Bacillus subtilis the analysis of other genes, such as gyrA (Chun and Bae [2000]) is needed.
All strains from this study showed an indication of the nitrogen fixation by the formation of the typical pellicule in semi-solid N-free medium. However, in only four strains (CNPSo 2476, CNPSo 2477, CNPSo 2478 and CNPSo 2480) the nifH gene was amplified. The nitrogen fixation genes are found in different phylogenetic groups (Affourtit et al. [2001]). Among them, nifH is one of the oldest and most functional (Rosado et al. [1998]) and its amplification by degenerative primers is a useful tool to confirm fixation potential (Zehr et al. [1996]). However, if amplification does not occur with these primers, it does not mean that the strains are not capable of nitrogen fixation, because the gene may have different nucleotide sequences between species and even within the same species Zehr et al. ([2003]). In our study, the four strains that amplified nifH with different primers were also positive to the ARA test. This result indicated that strains have the possibility to fix nitrogen. Montañez et al. ([2009]) also verified the same situation in bacteria with nitrogen fixation capability by the reduction of acetylene to ethylene and confirmed by PCR the presence of nifH gene. The group of Bacilli was reported to be capable of nitrogen fixation, Ratón et al. ([2011]) and Rana et al. ([2011]) working with ARA-positive strains, indicated suitable environments for searching new inoculants or candidates for biotechnological purposes.
In our study, all strains produced IAA, although in different amounts. Bacteria that produced lower levels (CNPSo 2480 and CNPSo 2481), had bigger influence on root elongation and root volume, characteristics of great interest that provide greater surface area for the absorption of nutrients. The effects of the auxin depend on its concentration, i.e. when it is low it can stimulate growth and when it is high can be inhibitory, same for seed germination (Arshad and Frankenberger [1991]). This may occur because the IAA can modify the endogenous auxin of the plant to an optimal or deleterious level (Patten and Glick [1996]). For example, Sarwar and Kremer ([1995]), by comparing the production of auxin between promoting and inhibiting bacteria growth, noted that the latter produced high levels and inhibited root growth of Convolvulus arvensis; Baranzani and Friedman ([1999]) observed the same effect with lettuce (Lactuca sativa L.). Araujo and Guerreiro ([2010]), working with Bacillus, found that most strains of corn identified as growth promoters did not correspond to those producing higher levels of IAA. The species Pseudomonas fluorescens, whose production of IAA was only 15.63 μg/mL, increased root length in corn plants, according to Hernández-Rodríguez et al. ([2008]).
The ability to produce siderophores has been reported in Bacillus (Pereira and Castro-Silva, [2010]; Kumar and Kumar [2012]) and Enterobacter (Tian et al. [2009]; Arruda et al. [2013]). Out of the strains tested in this study, only Enterobacter sp. (CNPSo 2480) did not produce siderophores. Siderophores produced by bacteria that promote plant growth may act through two mechanisms: directly promoting growth, as iron availability to plants is generally low, therefore organic chelators produced by bacteria will help absorption (Powell et al. [1980]), and indirectly, by inhibiting the availability of iron to pathogens, thereby limiting pathogen growth (Ahmad et al. [2008]).
Lytic enzymes such as β-1,3-glucanases, cellulases, proteases, amylases and chitinases are connected to hyper-parasitic activities (Kim and Chung [2004]) since they have the ability to degrade structural fungal cell walls (Oppenheim and Chet [1992]). Pectinases, as well as cellulases, also assist in penetration of bacteria into the plant host. They can also promote the induction of systemic resistance (Hallmann et al. [1997]). In our study, these enzymes were detected in strains of the genus Bacillus, corroborating Ratón et al. ([2011]). The inhibitory action on fungal pathogens can also be related to the ammonia released by bacteria, as reported by Fravel ([1988]) with Enterobacter cloacae on Verticillium dahliae and Pythium ultimum.
In Brazil, among the field phytopathogens that affect corn, F. verticilliodes is the most common (Peixoto et al. [1998]). The following pathogens are also common: C. graminicola, B. maydis (Pinto [1998]) and C. zea-maydis, and have become more important in recent decades in Brazil (Pereira et al. [2005]). Among the strains tested in our study, Bacillus sp. CNPSo 2477 and CNPSo 2476 were the most efficient in the inhibition against all four fungi tested. They are known to have an antagonist effect against various fungi, for example, F. moniliforme, F. graminearum, M. phaseolina (Pal et al. [2001]), F. oxysporum and Colletotrichum truncatum (Araujo and Guerreiro [2010]). One of the advantages of the biological control strategy using endophytic bacteria is that they can act in the same niche, in direct competition with the pathogens (Bacon and Hinton, [2002]).
Based on the 16 s rRNA sequence, the endophytic bacteria from our study were identified as Enterobacter asburie and Bacillus pumilus, while the other strains were confirmed as belonging to the genus Bacillus. We found that Bacillus CNPSo 2477 and CNPSo 2476 have good antagonist performance against corn phytopathogens. We also showed that Bacillus sp. CNPSo 2481 and Enterobacter sp. CNPSo 2480, have shown the ability to promote plant growth, leading to increases in the percentage of seed germination and improving the early development of seedlings. Therefore, these strains are very promising for use as inoculants in corn. Future studies are needed to test the biotechnological potential of these strains under field conditions, in the hope that they will contribute as an alternative source of biological fertilizer and biological control. It is also interesting to investigate which mechanisms would be related to fungal inhibition activity detected in our study.
Competing interests
All authors declare have no competing interests.
Contributor Information
Vivian Jaskiw Szilagyi-Zecchin, Email: vivian.szilagyi@gmail.com.
Angela Cristina Ikeda, Email: vivian.szilagyi@gmail.com.
Mariangela Hungria, Email: hungria@cnpso.embrapa.br.
Douglas Adamoski, Email: vivian.szilagyi@gmail.com.
Vanessa Kava-Cordeiro, Email: vivian.szilagyi@gmail.com.
Chirlei Glienke, Email: vivian.szilagyi@gmail.com.
Lygia Vitória Galli-Terasawa, Email: lterasawa@ufpr.br.
Acknowledgments
Our thanks go to: Semilia Genetics and Breeding LTDA, specially Dr. Francisco Terasawa Junior, for experiments in obtaining fungal strains. Also, the team of the Biotechnology Laboratory of Embrapa Soybean Soil, namely Jacqueline Delamuta and Renan Ribeiro, for support with the sequencing. Renata Rodrigues Gomes and Maicon Wons Andrew Fernandes, for their help in the antagonism tests. This work had financial support from the National Council for Scientific and Technological Development (CNPq), Project Repensa (562008/2010-2) and Coordination for the Improvement of Higher Education Personnel (CAPES).
References
- Affourtit J, Zehr JP, Paerl HW. Distribution of nitrogen-fixing microrganisms along the Neuser river estuary, North Carolina. Microb Ecol. 2001;4:114–123. doi: 10.1007/s002480000090. [DOI] [PubMed] [Google Scholar]
- Ahmad F, Ahmad I, Khan MS. Screening of free-living rhizospheric bacteria for their multiple plant growth promoting activities. Microbiol Res. 2008;4:73–181. doi: 10.1016/j.micres.2006.04.001. [DOI] [PubMed] [Google Scholar]
- Altekar G, Dwarkadas S, Huelsenbeck JP, Ronquist F. Parallel metropolis coupled Markov chain Monte Carlo for Bayesian phylogenetic inference. Bioinformatics. 2004;4:407–415. doi: 10.1093/bioinformatics/btg427. [DOI] [PubMed] [Google Scholar]
- Araujo FF, Guerreiro RT. Bioprospecção de isolados de Bacillus promotores de crescimento de milho cultivado em solo autoclavado e natural. Ciênc Agrotec. 2010;4(4):837–844. doi: 10.1590/S1413-70542010000400007. [DOI] [Google Scholar]
- Arruda L, Beneduzi A, Martins A, Lisboa B, Lopes C, Bertolo F, Vargas LK. Screening of rhizobacteria isolated from maize (Zea mays L.) in Rio Grande do Sul State (South Brazil) and analysis of their potential to improve plant growth. Appl Soil Ecol. 2013;4:15–22. doi: 10.1016/j.apsoil.2012.09.001. [DOI] [Google Scholar]
- Arshad M, Frankenberger WT. Microbial production of plant hormones. Plant Soil. 1991;4:1–8. doi: 10.1007/BF00011893. [DOI] [Google Scholar]
- Ashraf MA, Rasool M, Mirza MS. Nitrogen fixation and indole acetic acid production potential of bacteria isolated from rhizosphere of sugarcane (Saccharum officinarum L.) Adv Biol Res. 2011;4(6):348–355. [Google Scholar]
- Assumpção LC, Lacava PT, Dias ACF, Azevedo JL, Menten JOM. Diversidade e potencial biotecnológico da comunidade bacteriana endofítica de semente de soja. Pesq Agropec Bras. 2009;4:503–510. doi: 10.1590/S0100-204X2009000500010. [DOI] [Google Scholar]
- Bacon CW, Hinton DM. Endophytic and biological control potential of Bacillus mojavensis and related species. Biological Control. 2002;4(3):274–284. doi: 10.1006/bcon.2001.1016. [DOI] [Google Scholar]
- Bakker AW, Schippers B. Microbial cyanide production in the rhizosphere in relation to potato yield reduction and Pseudomonas spp-mediated plant growth-stimulation. Soil Biol Biochem. 1987;4:451–457. doi: 10.1016/0038-0717(87)90037-X. [DOI] [Google Scholar]
- Baldani JI, Baldani VLD, Seldin L, Döbereiner J. Characterization of Herbaspirillum seropedicae gen nov, sp nov, a root-associated nitrogen-fixing bacterium. Int J Syst Bacteriol. 1986;4:86–93. doi: 10.1099/00207713-36-1-86. [DOI] [Google Scholar]
- Barazani O, Friedman J. Is IAA the major root growth factor secreted from plantgrowth-mediating bacteria. J Chem Ecol. 1999;4(10):2397–2406. doi: 10.1023/A:1020890311499. [DOI] [Google Scholar]
- Barretti PB, Souza RM, Pozza EA. Bactérias endofíticas como agentes promotores do crescimento de plantas de tomateiro e de inibição in vitro de Ralstonia solanacearum. Ciênc Agrotec. 2008;4:731–739. doi: 10.1590/S1413-70542008000300005. [DOI] [Google Scholar]
- Beneduzi A, Peres D, Vargas LK, Bodanese–Zanettini MH, Passaglia LMP. Evaluation of genetic diversity and plant growth promoting activities of nitrogen–fixing Bacilli isolated from rice fields in South Brazil. Appl Soil Ecol. 2008;4:311–320. doi: 10.1016/j.apsoil.2008.01.006. [DOI] [Google Scholar]
- Beneduzi A, Moreira F, Costa PB, Vargas LK, Lisboa BB, Favreto R, Baldani JI, Passaglia LMP. Diversity and plant growth promoting evaluation abilities of bacteria isolated from sugarcane cultivated in the South of Brazil. Appl Soil Ecol. 2013;4:94–104. doi: 10.1016/j.apsoil.2012.08.010. [DOI] [Google Scholar]
- Berg G, Roskot N, Steidle A, Eberl L, Zock A, Smalla K. Plant-dependent genotypic and phenotypic diversity of antagonistic rhizobacteria isolated from different Verticillium host plants. Appl Environ Microbiol. 2002;4(7):3328–3338. doi: 10.1128/AEM.68.7.3328-3338.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernardes FS, Patrício FRA, Santos AS, Freitas SS. Indução de resistência sistêmica por rizobactérias em cultivos hidropônicos. Summa Phytopathol. 2010;4(2):115–121. doi: 10.1590/S0100-54052010000200002. [DOI] [Google Scholar]
- Boddey RM. Methods for quantification of nitrogen fixation associated with gramineae. Crit Rev Plant Sci. 1987;4:209–266. doi: 10.1080/07352688709382251. [DOI] [Google Scholar]
- Burris RH. Nitrogen fixation assaymethods and techniques. Meth Enzymol. 1972;4:415–431. doi: 10.1016/0076-6879(72)24088-5. [DOI] [PubMed] [Google Scholar]
- Canbolat M, Bilen S, Çakmakçi R, Sahin F, Aydi A. Effect of plant growth-promoting bacteria and soil compaction on barley seeding growth, nutrient uptake, soil properties and rhizosphere microflora. Biol Fertil Soils. 2006;4(3):350–357. doi: 10.1007/s00374-005-0034-9. [DOI] [Google Scholar]
- Cassán F, Maiale S, Masciarelli O, Vidal A, Luna V, Ruiz O. Cadaverine production by Azospirillum brasilense and its possible role in plant growth promotion and osmotic stress mitigation. Eur J Soil Biol. 2009;4:12–19. doi: 10.1016/j.ejsobi.2008.08.003. [DOI] [Google Scholar]
- Chun J, Bae KS. Phylogenetic analysis of Bacillus subtilis and related taxa based on partial gyrA gene sequences. Antonie van Leeuwenhoek. 2000;4:123–127. doi: 10.1023/A:1026555830014. [DOI] [PubMed] [Google Scholar]
- Compant S, Clément C, Sessitsch A. Plant growth-promoting bacteria in the rhizo and endosphere of plants: their role, colonization, mechanisms involved and prospects for utilization. Soil Biol Biochem. 2010;4:669–678. doi: 10.1016/j.soilbio.2009.11.024. [DOI] [Google Scholar]
- Daniels BA. Elimination of Fusarium moniliforme from corn seed. Plant Disease. 1983;4(6):609–611. doi: 10.1094/PD-67-609. [DOI] [Google Scholar]
- Deepa CK, Dastager SG, Pandey A. Plant growth-promoting activity in newly isolated Bacillus thioparus (NII-0902) from Western ghat forest, India. World J Microbiol Biotechnol. 2010;4(12):2277–2283. doi: 10.1007/s11274-010-0418-3. [DOI] [Google Scholar]
- Döbereiner J. In: Methods for evaluating biological nitrogen fixation. Bergersen FJ, editor. Wiley, Chichester; 1980. Forage grasses and grain crops; pp. 535–555. [Google Scholar]
- Döbereiner J, Baldani VLD, Baldani JI. Como isolar e identificar bactérias diazotróficas de plantas não-leguminosas. Embrapa, Brasília; 1995. [Google Scholar]
- Dye DW. The inadequacy of the usual determinative tests for identification of Xanthomonas sp. New Zeal J Sci. 1962;4:393–416. [Google Scholar]
- Enebak SA, Wei G, Kloepper JW. Effects of plant growth promoting rhizobacteria on loblolly and slash pine seedlings. Forest Sci. 1998;4:139144. [Google Scholar]
- Ewing B, Hillier L, Wendl MC, Green P. Basecalling of automated sequencer traces using phred I Accuracy assessment. Genome Res. 1998;4(3):175–185. doi: 10.1101/gr.8.3.175. [DOI] [PubMed] [Google Scholar]
- Ferreira PAA, Bomfeti CA, Soares BL, Moreira FMS. Efficient nitrogen-fixing Rhizobium strains isolated from amazonian soils are highly tolerant to acidity and aluminium. World J Microbiol Biotechnol. 2011. [DOI] [PubMed]
- Fravel DR. Role of antibiosis in the biocontrol of plant diseases. Ann Rev Phytopathol. 1988;4:75–91. doi: 10.1146/annurev.py.26.090188.000451. [DOI] [Google Scholar]
- George P, Gupta A, Gopal M, Thomas L, Thomas GV. Multifarious beneficial traits and plant growth promoting potential of Serratia marcescens KiSII and Enterobacter sp. RNF 267 isolated from the rhizosphere of coconut palms (Cocos nucifera L.) World J Microb Biot. 2012;4(1):109–117. doi: 10.1007/s11274-012-1163-6. [DOI] [PubMed] [Google Scholar]
- Hallmann J, Quadt-Hallmann A, Mahaffee WF, Kloepper JW. Bacterial endophytes in agricultural crops. Can J Microbiol. 1997;4:895–914. doi: 10.1139/m97-131. [DOI] [Google Scholar]
- Hankin L, Anagnostakis SL. The use of solid media for detection of enzyme production by fungi. Mycologia. 1975;4:597–607. doi: 10.2307/3758395. [DOI] [Google Scholar]
- Hernandez-Rodriguez A, Heydrich-Perez M, Acebo-Guerrero Y, Velazquez-Del Valle MG, Hernandez-Lauzardo NA. Antagonistic activity of Cuban native rhizobacteria against Fusarium verticillioides (Sacc.) Nirenb. in maize (Zea mays L.) Appl Soil Ecol. 2008;4(2):180–186. doi: 10.1016/j.apsoil.2007.12.008. [DOI] [Google Scholar]
- Hungria M, Campo RJ, Souza EM, Pedrosa FO. Inoculation with selected strains of Azospirillum brasilense and A. lipoferum improves yields of maize and wheat in Brazil. Plant Soil. 2010;4:413–425. doi: 10.1007/s11104-009-0262-0. [DOI] [Google Scholar]
- Ikeda AC, Bassani LL, Adamoski D, Stringari D, Kava-Cordeiro V, Glienke C, Steffens MBR, Hungria M, Galli-Terasawa LV. Morphological and genetic characterization of endophytic bacteria isolated from roots of different maize genotypes. Microb Ecol. 2013;4:154–160. doi: 10.1007/s00248-012-0104-0. [DOI] [PubMed] [Google Scholar]
- Khalid A, Arshad M, Zahir ZA. Screening plant growth promoting rhizobacteria for improving growth and yield of wheat. J Appl Microbiol. 2004;4:473–480. doi: 10.1046/j.1365-2672.2003.02161.x. [DOI] [PubMed] [Google Scholar]
- Kim PI, Chung KC. Production of an antifungal protein for control of Colletotrichum lagenarium by Bacillus amyloliquefaciens MET0908. FEMS Microbiol Lett. 2004;4(1):177–183. doi: 10.1111/j.1574-6968.2004.tb09530.x. [DOI] [PubMed] [Google Scholar]
- King EO, Ward MK, Raney DE. Two simple media for the demonstration of pyocyanin and fluorescin. J Lab Clin Med. 1954;4(2):301–307. [PubMed] [Google Scholar]
- Krey T, Vassilev N, Baum C, Eichler-Löbermann B. Effects of long-term phosphorus application and plant-growth promoting rhizobacteria on maize phosphorus nutrition under field conditions. Eur J Soil Biol. 2013;4:124–130. doi: 10.1016/j.ejsobi.2012.12.007. [DOI] [Google Scholar]
- Kumar MV, Kumar A. Plant growth promoting and phytostimulatory potential of Bacillus subtilis and Bacillus amyloliquefaciens. JABS. 2012;4(7):509–519. [Google Scholar]
- Kuss AV, Kuss VV, Lovato T, Ml F. Fixação de nitrogênio e produção de ácido indol acético in vitro por bactérias diazotróficas endofíticas. PAB. 2007;4:1459–1465. [Google Scholar]
- Laguerre G, Nour SM, Macheret V, Sanjuan J, Drouin P, Amarger N. Classification of rhizobia based on nodC and nifH gene analysis reveals a close phylogenetic relationship among Phaseolus vulgaris symbionts. Microbiol. 2001;4:981–993. doi: 10.1099/00221287-147-4-981. [DOI] [PubMed] [Google Scholar]
- Loper JE, Schroth MN. Influence of bacterial sources of indole-3-acetic acid on root elongation of sugar beet. Phytopathology. 1986;4:386–389. doi: 10.1094/Phyto-76-386. [DOI] [Google Scholar]
- Löytynoja A, Goldman D. webPRANK: a phylogenyaware multiple sequence aligner with interactive alignment browser. BMC Bioinformatics. 2010. [DOI] [PMC free article] [PubMed]
- Lucy M, Reed E, Glick BR. Application of free living plant growth-promoting rhizobacteria. Antonie van Leewenhoek. 2004;4:1–25. doi: 10.1023/B:ANTO.0000024903.10757.6e. [DOI] [PubMed] [Google Scholar]
- Menna P, Hungria M, Barcellos FG, Bangel EV, Hess PN, Martínez-Romero E. Molecular phylogeny based on the 16S rRNA gene of elite rhizobial strains used in brazilian commercial inoculants. Syst Appl Microbiol. 2006;4:315–332. doi: 10.1016/j.syapm.2005.12.002. [DOI] [PubMed] [Google Scholar]
- Montañez A, Abreu C, Gill PR, Hardarson G, Sicardi M. (2009) Biological nitrogen fixation in maize (Zea mays L.) by 15 N isotope-dilution and identification of associated culturable diazotrophs. Biol Fertil Soils. 2009;4:253–263. doi: 10.1007/s00374-008-0322-2. [DOI] [Google Scholar]
- Moura JM, Ferreira AFC, Silva FMM, Rizzi J, Pinto LAA. Obtenção de quitina a partir de carapaças de siri (Maia squinado): uso de um planejamento experimental na etapa de desmineralização. Vetor. 2005;4(1):717. [Google Scholar]
- Nylander JAA, Wilgenbush JC, Warren DL, Swofford DL. AWTY (Are We There Yet?): A system for graphical exploration of MCMC convergence in Bayesian phylogenetics. Bioinformatics. 2008;4:581–583. doi: 10.1093/bioinformatics/btm388. [DOI] [PubMed] [Google Scholar]
- Oliveira ALM, Canuto EL, Reis VM, Baldani JI. Response of micropropagated sugarcane varieties to inoculation with endophytic diazotrophic bacteria. Braz J Microbiol. 2003;4:59–61. doi: 10.1590/S1517-83822003000500020. [DOI] [Google Scholar]
- Oppenheim AB, Chet I. Cloned chitinases in fungal plant-pathogen control strategies. Trends Biotechnol. 1992;4:392–394. doi: 10.1016/0167-7799(92)90281-Y. [DOI] [Google Scholar]
- Pal KK, Tilak KVBR, Saxena AK, Dey R, Singh CS. Suppression of maize root diseases caused by Macrophomina phaseolina, Fusarium moniliforme and Fusarium graminearum by plant growth promoting rhizobacteria. Microbiol Res. 2001;4:209–223. doi: 10.1078/0944-5013-00103. [DOI] [PubMed] [Google Scholar]
- Patten CL, Glick BR. Bacterial biosynthesis of indole-3-acetic acid. Can J Microbiol. 1996;4:207–220. doi: 10.1139/m96-032. [DOI] [PubMed] [Google Scholar]
- Peixoto AR, Torres SB, Karasawa N. Qualidade sanitária de sementes de milho produzidas no submédio São Francisco. Rev Bras Sementes. 1998;4(1):12–15. [Google Scholar]
- Pereira BA, Castro-Silva MA. Rizobactérias formadoras de endósporos associadas a Tibouchina urvilleanade áreas impactadas por rejeitos da mineração do carvão. Rev Bras Ciênc Solo. 2010;4:563–567. doi: 10.1590/S0100-06832010000200030. [DOI] [Google Scholar]
- Pereira AOP, Camargo RV, Camargo LEA. In: Manual de Fitopatologia - Doenças de Plantas cultivadas, 2º vol. Kimati H, Amorim L, Bergamim-Filho A, Camargo LEA, editor. Ceres, Piracicaba; 2005. Doenças do milho (Zea mays) pp. 477–488. [Google Scholar]
- Pereira P, Ibáñez F, Rosenblueth M, Etcheverry M, Martínez-Romero E. Analysis of the bacterial diversity associated with the roots of maize (Zea mays L) through culture-dependent and culture-independent methods. ISRN Ecology. 2011.
- Petrini O. In: Fokkema NJ, Heuvel J. Den V, editor. Cambridge University, Cambridge, Microbiology of the phyllosphere. Cambridge; 1986. Taxonomy of endophytic fungi of aerial plant tissues; pp. 75–87. [Google Scholar]
- Pinto NFJA. Patologia de sementes de milho, Circular Técnica 29. Embrapa, Sete Lagoas; 1998. [Google Scholar]
- Porwal S, Lal S, Cheema S, Kalia VC. Phylogeny in aid of the present and novel microbial lineages: diversity in Bacillus. PLoS ONE. 2009;4(2):e4438. doi: 10.1371/journal.pone.0004438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Posada D, Krandal KA. MODELTEST: testing the model of DNA substitution. Bioinformatics. 1988;4:817–818. doi: 10.1093/bioinformatics/14.9.817. [DOI] [PubMed] [Google Scholar]
- Powell KA, Faull JL, Renwick A. In: Biological control of soil borne plant pathogens. Horny D, editor. CAB International Wallingford, United Kingdom; 1990. The commercial and regulatory challenge; pp. 445–463. [Google Scholar]
- Raeder U, Broda P. Rapid preparation of DNA from filamentous fungi. Lett Appl Microbiol. 1985;4:17–20. doi: 10.1111/j.1472-765X.1985.tb01479.x. [DOI] [Google Scholar]
- Rana A, Saharan B, Joshi M, Prasanna R, Kumar K, Nain L. Identification of multi-trait PGPR isolates and evaluating their potential as inoculants for wheat. Ann Microbiol. 2011;4:893–900. doi: 10.1007/s13213-011-0211-z. doi:1 0.1007/s13213-011-0211-z. [DOI] [Google Scholar]
- Ratón TMO, Yano R, Gámez OR, Floh EIS, Díaz MJS, Barbosa HR. Isolation and characterisation of aerobic endospore forming Bacilli from sugarcane rhizosphere for the selection of strains with agriculture potentialities. World J Microbiol Biotechnol. 2011. [DOI] [PubMed]
- Renwick A, Campbell R, Coe S. Assessment of in vivo screening systems for potencial biocontrol agents o Gaeumannomyces graminis. Plant Pathol. 1991;4:524–532. doi: 10.1111/j.1365-3059.1991.tb02415.x. [DOI] [Google Scholar]
- Rodrigues Neto J, Malavolta Júnior VA, Victor O. Meio simples para isolamento e cultivo de Xanthomonas campestrispv citri tipo B. Summa Phytopatol. 1986;4:16. [Google Scholar]
- Rodriguez H, Gonzalez T, Goire I, Bashan Y. Gluconic acid production and phosphate solubilization by the plant growth-promoting bacterium Azospirillum spp. Naturwissenschaften. 2004;4:552–555. doi: 10.1007/s00114-004-0566-0. [DOI] [PubMed] [Google Scholar]
- Ronquist F, Huelsenbeck JP. MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;4:1572–1574. doi: 10.1093/bioinformatics/btg180. [DOI] [PubMed] [Google Scholar]
- Rosado AS, Duarte GF, Seldin L, VanElsas JD. Genetic diversity of nifH gene sequences in Paenibacillus azotofixans strains and soil samples analyzed by denaturing gradient gel electrophoresis of PCR amplified gene fragments. Appl Environ Microbiol. 1998;4:2770–2779. doi: 10.1128/aem.64.8.2770-2779.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rösch C, Mergel A, Bothe H. Biodiversity of denitrifying and dinitrogen-fixing bacteria in an acid forest soil. Appl Environ Microbiol. 2002;4(8):3818–3829. doi: 10.1128/AEM.68.8.3818-3829.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sambrook J, Russell D. Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York; 2001. [Google Scholar]
- Sarwar M, Kremer RJ. Enhanced suppression of plant growth through production of L-tryptophan-derived compounds by deleterious rhizobacteria. Plant Soil. 1995;4:261–269. doi: 10.1007/BF00011328. [DOI] [Google Scholar]
- Schwyn B, Neilands JB. Universal chemical assay for the detection and determination of siderophores. Anal Biochem. 1987;4:4756. doi: 10.1016/0003-2697(87)90612-9. [DOI] [PubMed] [Google Scholar]
- Seldin L, van Elsas JD, Penido EGC. Bacillus azotofixans sp. nov., a nitrogen-fixing species from Brazilian soils and grass roots. Int J Syst Bacteriol. 1984;4:451–456. doi: 10.1099/00207713-34-4-451. [DOI] [Google Scholar]
- Shiomi HF, Melo IS, Minhoni MTA. Seleção de bactérias endofíticas com ação antagônica a fitopatógenos. Scientia Agraria. 2008;4:535–538. [Google Scholar]
- Silva MF, Reis VM. Técnicas usadas para quantificar a população de bactérias diazotróficas inoculadas em sementes de milho (Zea mays) Embrapa, Seropédica; 2004. [Google Scholar]
- Sukumaran J, Holder MT. DendroPy: a Python library for phylogenetic computing. Bioinformatics. 2010;4:1569–1571. doi: 10.1093/bioinformatics/btq228. [DOI] [PubMed] [Google Scholar]
- Sylvester-Bradley R, Asakawa N, La Torraca S, Magalhães FMM, Oliveira L, Pereira RM. Levantamento quantitativo de microrganismos solubilizadores de fosfatos na rizosfera de gramíneas e leguminosas forrageiras na Amazônia. Acta Amaz. 1982;4:15–22. [Google Scholar]
- Tian F, Ding Y, Zhu H, Yao L, Du B. Genetic diversity of siderophore-producing bacteria of tobacco rhizosphere. Braz J Microbiol. 2009;4(2):276–284. doi: 10.1590/S1517-83822009000200013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vale M, Seldin L, Araújo FF, Lima R. Maheshwari DK (ed) Plant growth and health promoting bacteria Springer, Berlin; 2010. Plant growth promoting rhizobacteria: fundamentals and applications; pp. 21–43. [Google Scholar]
- Verma JP, Yadav J, Tiwari KN, Kumar A. Effect of indigenous Mesorhizobium spp. and plant growth promoting rhizobacteria on yields and nutrients uptake of chickpea (Cicer aritenium L.) under sustainable agriculture. Ecol Eng. 2013;4:282–286. doi: 10.1016/j.ecoleng.2012.12.022. [DOI] [Google Scholar]
- Wang S, Huijun W, Junqing Q, Lingli M, Jun L, Yanfei X, Xuewen G. Molecular mechanism of plant growth promotion and induced systemic resistance to tobacco mosaic virus by Bacillus spp. J Microbiol Biotechnol. 2009;4(10):1250–1258. doi: 10.4014/jmb.0901.008. [DOI] [PubMed] [Google Scholar]
- Weisburg WG, Barns SM, Pelletier DA, Lane DJ. 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol. 1991;4:697–703. doi: 10.1128/jb.173.2.697-703.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zehr JP, Capone DG. Problems and promises of assaying the genetic potential for nitrogen fixation in the marine environment. Microb Ecol. 1996;4:263–281. doi: 10.1007/BF00183062. [DOI] [PubMed] [Google Scholar]
- Zehr JP, Jenkins BD, Short SM, Steward GF. Nitrogenase gene diversity and microbial community structure: A cross-system comparison. Environ Microbiol. 2003;4:539–554. doi: 10.1046/j.1462-2920.2003.00451.x. [DOI] [PubMed] [Google Scholar]


