Abstract
Nanotechnology represents an area of particular promise and significant opportunity across multiple scientific disciplines. Ongoing nanotechnology research ranges from the characterization of nanoparticles and nanomaterials to the analysis and processing of experimental data seeking correlations between nanoparticles and their functionalities and side effects. Due to their special properties, nanoparticles are suitable for cellular-level diagnostics and therapy, offering numerous applications in medicine, e.g. development of biomedical devices, tissue repair, drug delivery systems and biosensors. In nanomedicine, recent studies are producing large amounts of structural and property data, highlighting the role for computational approaches in information management. While in vitro and in vivo assays are expensive, the cost of computing is falling. Furthermore, improvements in the accuracy of computational methods (e.g. data mining, knowledge discovery, modeling and simulation) have enabled effective tools to automate the extraction, management and storage of these vast data volumes. Since this information is widely distributed, one major issue is how to locate and access data where it resides (which also poses data-sharing limitations). The novel discipline of nanoinformatics addresses the information challenges related to nanotechnology research. In this paper, we summarize the needs and challenges in the field and present an overview of extant initiatives and efforts.
1. Introduction
Over the last 20 years, nanotechnology has pushed back the frontiers of science and has especially revolutionized the field of medicine (Duncan and Gaspar 2011). This novel scientific field deals with materials at the nano-dimension (macromolecules that range in size from 1 to 100 nanometers), offering breakthroughs in a wide range of scientific areas, such as physics, chemistry, microbiology and materials science, but also posing many challenges (Lemley 2005).
At the nanoscale, physical, chemical and functional properties differ from those at the cell, tissue or organ level (McNeil 2011). Unique characteristics emerge from nanoparticles, based on their size—they are small enough to interact with receptors with high specificity and effectiveness and large enough to carry drugs at the molecular level—and also on their quantum properties: density, stability or unexpected optical properties such as high absorption of solar radiation (Scholes and Rumbles 2006) or superparamagnetism (Sato et al 2011), for instance. All this represents a dramatic turnaround from a few years ago for the scientific community. In medicine, worldwide efforts to foster these novel technologies promises to bring us new insights into the development of biomarkers and personalized treatments, but also raises concerns about inefficacy in the management and analysis of all of this new information to address potential hazards and risks. In this sense, society’s perception of nanotechnology requires strong consideration since it could influence industry oversight of future advancements in the area.
One current issue regarding nanoparticles is potential toxicity, a field that is still obscure to researchers. It is unclear how and what toxicity some nanoparticles have and there is a lack of extensive regulation for new products derived from them. What may be harmless at the macro or chemical level may have different effects in the body than the nanoparticle form. Possibly the largest time- and money-related costs involved in developing nanomaterials may not be in making them work, but in determining their side effects and biological impacts.
Computational methods and tools offer novel paradigms to materials discovery and design, improving properties prediction or material selection capabilities that could increase product performance and reduce time from drug discovery to marketing. The novel discipline of nanoinformatics deals with all of these information challenges related to nanotechnology research and, specifically, with nanomedicine (Maojo et al 2012a). Several nanoinformatics initiatives are currently identifying data gaps and research priorities in the area and developing collaborative nanoinformatics environments where research institutions can publish descriptions of their data sources and make them publicly available over the Internet.
In this paper, we present an overview of the needs and challenges in nanoinformatics, as well as the extant initiatives and international efforts in the field. All these initiatives are fostering scientific advancement and knowledge dissemination in the nanomedical area and closing the gaps between basic research and clinical applications (de la Iglesia et al 2011).
1.1. Applications of nanoparticles in medicine
Groundbreaking advances in nanoscience continue to provide incremental insights into medicine. Among numerous potential applications, such as biosensors, biological carriers for drug delivery or structures for tissue repair, we illustrate in this section some examples of the more innovative research in the field of nanomedicine. We can differentiate two contexts in nanotechnology applications: diagnostics (based on molecular biosensing) and therapeutics, which follow tailored strategies.
As biomarkers and imaging contrast agents, nanoparticles can be decorated with specific diagnostic agents targeted to specific receptors (Mailänder and Landfester 2009), which allows monitoring of their distribution (Haun et al 2011). This property has many diagnostic biosensing applications (Yeri and Gao 2011). The large surface area of nanomaterials, chemically functionalized with targeting ligands, provides excellent levels of sensitivity for the detection of molecular targets, such as DNA, proteins, pathogens, tumor cells or enzymes (Kumbar et al 2008). For instance, scientists at the Northwestern University Feinberg School of Medicine have developed a bio-barcode assay (Thaxton et al 2009) for detecting prostate-specific antigen (PSA) based on the use of gold nanoparticles and have recently defined nanocombinatorics (Giam et al 2012), a new lithography technique that generates nanoscale biological patterns. In the NanoSystems Biology Cancer Center at the California Institute of Technology, researchers have developed a nano-based, integrated platform with two different applications: cell sorting and multiplex detection of DNA and proteins (Bailey et al 2007). Photoacoustic molecular imaging is another potential application of nanoparticles as molecular markers. It consists of marking biological targets with acoustic detectors, such as carbon nanotubes and gold nanoparticles. This promising technique has been successfully applied in mice by the Molecular Imaging Program at Stanford for multimodal brain tumor imaging (Kircher et al 2012) and for the diagnosis of retinal diseases (de la Zerda et al 2012).
As delivery systems, nanoparticles are suitable for systematically releasing drugs to a specific target (cell or tissue), increasing their efficiency and improving the characterization of the treatment response. Nanocarriers can be seen as sophisticated mechanisms by which nano-sized objects can be transported through the body to target specific cells and penetrate cellular and nuclear membranes. Advanced design of nanomaterials has demonstrated that engineered nanoparticles can evade immune detection, facilitating the targeted delivery of a specific drug. Nanoparticles hide their payloads, protecting drugs from several defense mechanisms and biological barriers, such as particle disposal by phagocytic cells or renal plasma clearance (Zuckerman et al 2012). Nanoparticles can also be used for gene delivery (Jin et al 2009), leading to deactivation of genes related to a specific illness (e.g. cancer). The nanoparticle preserves its genetic payload from degrading enzymes and the immune system and carries it into the target cells, as well. In addition, advances in nanotechnology are being applied to in vitro regenerative medicine: tissue regeneration (Veetil and Ye 2009), cardiovascular implants (Ghanbari et al 2011, de Mel et al 2012), cell therapy (Perán et al 2012), wound healing and delivery of exogenous growth factors (Koria 2012) are examples.
With regards to cancer research—one of the main areas where nanotechnology discoveries are being applied—biological mechanisms are complex and drug interactions offer very different results between in vitro and in vivo assays. However, with the use of nanoparticles, diagnostics and therapeutics could be greatly improved for certain types of tumors, such as those tumors currently treated with toxic drugs—i.e. lung (Upadhyay et al 2012) or breast cancer (Xin et al 2012)—or gene therapy—ovarian (Fathabadi et al 2012) and brain cancer (Liu et al 2012). For instance, researchers at the University of Melbourne (Australia) are using certain engineered nanostructures (polymers, micelles and polymersomes) as carriers to deliver drugs that target cancer cells causing minimal biological side effects (Kamphuis et al 2010, Shimoni et al 2012). With the same objective, the continuous efforts of the National Cancer Institute (NCI) to apply nanotechnology-based advances to early diagnosis and effective therapy against cancer (Service 2005, 2010, Shukla et al 2011, Ptak et al 2010) is also worthy of mention. There are other remarkable examples of innovative nanomedical research. At the University of North Carolina, researchers are conducting experiments that use the novel PRINT technique (Perry et al 2011) for nanoparticle replication and mimicry (Parrott et al 2012, Wang et al 2012), obtaining new particles that can be tuned with controlled properties, e.g. elasticity, size or shape. Such advances could be applied to the development of nanoparticles for use in the emerging area of theranostics (Perry et al 2011, Kelkar and Reineke 2011, Kiessling et al 2012), a new paradigm that combines diagnostics and therapy for individual patients in one step.
However, there is an urgent need to shorten the timeline between the acquisition of fundamental knowledge about nanomaterials, the publication of related studies and assays in scientific media and their clinical application. Social concerns about nanoparticles toxicity or nanotoxicity (the potential negative health impact of exposure to nanomaterials) could be a barrier for the agile and efficient transfer of nanotechnology from the bench to the bedside (Bellare 2011). This could be addressed by supporting responsible nanotechnology development, providing high-quality information through a better exchange system and sharing available toxicity and physico-chemical data to efficiently determine the association between exposure to nanomaterials and side effects. In both the diagnostic and treatment contexts, pharmacokinetic behaviour and potential risks pose by nanoparticle-based drugs should be studied in-depth, considering both genotypic and phenotypic studies. Currently, in most cases, early clinical trials on nanoparticles are not revealing new toxic effects from the nanoparticle itself or its components. Therefore, it would be necessary to differentiate between the toxicity of the container and that of its payload (i.e. surface versus core), which are cornerstones in determining and monitoring the efficacy and adverse health effects of new nanomaterials. Indeed, in the case of nanomedicines, the development and approval of new drugs does not need to take so long and clinical trials are not necessarily vital in some cases, particularly when considering that nanomaterials are generally derived from drugs previously authorized by the U.S. Food and Drug Administration (FDA) (Etheridge et al 2013) or the European Medicines Agency (EMA) (Karalis and Macheras 2012). There are many drugs already approved or under commercial use, such as those for therapy against cancer—Daunoxome® (Petre and Dittmer 2007), Doxil® (Pearce et al 2012) and Caelyx® (Li et al 2012), among others—, bone substitutes such as CarriGen®4, dental composites such as FilTek® (Moosavi et al 2012), or platforms as the Verigene® System (Buxton 2009), which is extensively used in hospitals for diagnostic purposes. Considering the myriad of new and increasingly rapid results generated by nanomedical research nowadays, it is obvious that the development of new tools for managing all this information has become crucial for advancing the field.
2. Information in nanomedicine
Information delivery channels are rapidly evolving, generating an increasing need to manage available data and resources. Specifically, discovery and design of new materials (Bikiaris 2007, Majzoub and Rönnebro 2012, Huh and Kwon 2011, Mentovich et al 2012) and recent advances in personalized medicine (Jain 2011) raise new challenges in the analysis and integration of the massive heterogeneous data generated by nanomedical research.
Unfortunately, researchers are facing many difficulties in their daily tasks related to information retrieval and management in nanomedicine. The number of sources is huge, and is increasing every day, but the available information is not always what is required while, in other cases, the data are not comprehensive or the quality is not sufficiently high. In addition, professionals currently working on nanotechnology have different roles and professional profiles, such as researchers, program managers, policy makers, infrastructure providers, clinicians, students and workers. Thus, the type of information to be managed is very complete and of broad scope, but also heterogeneous with regards to format and content. Information about researchers, research programs, tools, nanomaterial data and current projects is distributed among numerous sources, leading to a considerable disconnect between resources and users currently performing clinical, environmental, medical and basic research. Thus, how to manage nanoinformation is still the open question for computer scientists. Figure 1 shows the top terms appearing in nanomedical research publications, demonstrating the variety of concepts (from tissues to cells to particles) embraced by nanomedicine.
Figure 1.
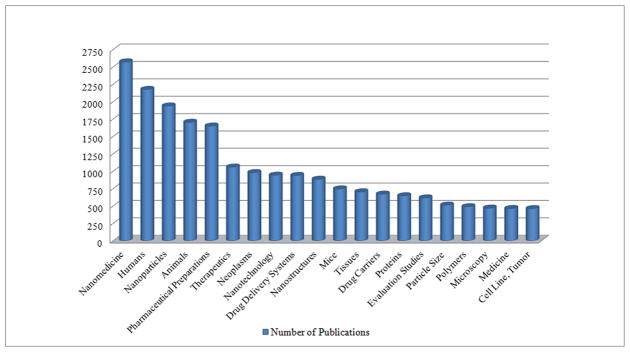
Top terms that appear in research publications related to nanomedicine
Over the last year, there has been a proliferation of computational resources related to the area of nanotechnology and, specifically, nanomedicine. As proposed elsewhere (Baker et al 2010, Maojo et al 2010, 2012b), nanoinformatics deals with how to extract useful information from all available data relevant to nanotechnology. Because this novel discipline is in its early stages, research on nanoinformatics currently faces many challenges: i) the definition of semantics in the field; ii) the standardization and integration of data; iii) the interoperability of resources; iv) the lack of consistent annotations; and iv) the necessary integration of data analysis tools with databases. Building cooperation among these areas may facilitate the development of useful and effective nanoinformatics methods and tools.
2.1. From data to knowledge
The inference of knowledge from raw data sets involves several steps: i) data collection; ii) data analysis and annotation; iii) information extraction; and, finally, iv) data validation by experts in the field. With regards to data collection, we currently face serious restrictions for accessing nanomaterial data, such as the unwillingness to share experiments and results or the existence of proprietary data that is not publicly available. In addition, some of the experimental results reported in the scientific literature contain numerical errors or are not properly described—in terms of protocols, experiment parameters or techniques—, so it is not possible to reproduce the experiment. Therefore, in order to extract high-quality information from raw data sets, a huge effort is needed to validate and curate existing repositories, deleting invalid and redundant data.
Knowledge inference also involves classifying and ranking information into categories and requires the definition of a specific semantic domain, based on standard terminologies and vocabularies. Specifically, knowledge discovery in nanomedicine should be driven by two means: i) biological and clinical problems, and ii) research data and models. Biological and clinical problems raise the target questions and challenges to be addressed: what tasks need to be supported and which resources—data and services—are needed. With regards to the second point, we find data modeling to be an issue: semantics requires a solid context to be defined in order to achieve data consistency between research results and predictive models.
Informatics tools aim to define this context by providing different techniques and methodologies—models, taxonomies and ontologies—to ensure data quality and harmonization and to interpret the meaning of data, turning it into usable information and knowledge. Additionally, computational methods to store, organize and manage the huge volumes of digital data generated during research on nanomedicine could be used to: i) integrate structural and functional characterization research data with multiple types of “nano” information (with clinical trial data, for instance); and ii) find, through the development of data analysis methods, potential associations between the size, structure, composition and physical and chemical properties of nanoparticles, as well as the extant correlations between in vitro and in vivo assays.
2.2. Information standards
Information standards are the basis of any scientific discipline. Using a common formalism to represent the elements within a domain guarantees the creation of dynamic, interoperable information systems that share data models linked to standard terminologies. Due to the wide range and heterogeneity of fields contributing to nanotechnology, global consensus in semantics—nomenclatures, code sets, descriptions, classifications and specifications of nanomaterials and related procedures, for example—would facilitate the interoperability of data, ensuring data quality, consistency and normalization. Examples of major formal standards in nanotechnology, developed and approved by the main international standards bodies and organizations (Murashov and Howard 2008, 2011, Hatto 2009, Weidl et al 2010) are listed below, in table 1.
Table 1.
Summary of the main current nanotechnology standards.
| American Society for Testing and Materials (ASTM) – Nanotechnology Standardsa
| ||||
|---|---|---|---|---|
| Terminology/Nomenclature | Measurement/Characterization | Materials Specifications | EHS | |
| E2490 - 09 | X | |||
| E2524 - 08 | X | |||
| E2525 - 08 | X | |||
| E2526 - 08 | X | |||
| E2578 - 07(2012) | X | |||
| E2834 - 12 | X | |||
| E2859 - 11 | X | |||
| E2535 - 07 | X | |||
| E2456 - 06 | X | |||
| British Standards Institution (BSI) – Nanotechnology British Standardsb
| ||||
|---|---|---|---|---|
| Terminology/Nomenclature | Measurement/Characterization | Materials Specifications | EHS | |
| PAS 71 | X | |||
| PAS 130 | X | |||
| PAS 131 | X | |||
| PAS 132 | X | |||
| PAS 133 | X | |||
| PAS 134 | X | |||
| PAS 135 | X | |||
| PAS 136 | X | |||
| PD 6699-1 | X | |||
| PD 6699-2 | X | |||
| PD 6699-3 | X | |||
| European Committee for Standardization (CEN) – Nanotechnologies CEN/TC 352c
| ||||
|---|---|---|---|---|
| Terminology/Nomenclature | Measurement/Characterization | Materials Specifications | EHS | |
| CEN ISO/TS 27687:2009 | X | |||
| EN ISO 10801:2010 | X | |||
| EN ISO 10808:2010 | X | |||
| EN ISO 29701:2010 | X | |||
| Federal Agency for Technical Regulations and Metrology (Russia)d
| ||||
|---|---|---|---|---|
| Terminology/Nomenclature | Measurement/Characterization | Materials Specifications | EHS | |
| GOST R 8.628-2007 | X | |||
| GOST R 8.629-2007 | X | |||
| GOST R 8.630-2007 | X | |||
| GOST R 8.631-2007 | X | |||
| GOST R 8.635-2007 | X | |||
| GOST R 8.636-2007 | X | |||
| GOST R 8.644-2008 | X | |||
| Institute of Electrical and Electronics Engineers – Nanotechnology Standardse
| ||||
|---|---|---|---|---|
| Terminology/Nomenclature | Measurement/Characterization | Materials Specifications | EHS | |
| P1650™-2005 | X | |||
| P1690™ | X | |||
| International Electrotechnical Commission – Nanotechnology standardization for electrical and electronic products and systemsf
| ||||
|---|---|---|---|---|
| Terminology/Nomenclature | Measurement/Characterization | Materials Specifications | EHS | |
| IEC/TC 113 | X | X | X | |
| ISO - International Organization for Standardizationg
| ||||
|---|---|---|---|---|
| Terminology/Nomenclature | Measurement/Characterization | Materials Specifications | EHS | |
| ISO/TC 229 | X | X | X | X |
| ISO/TC 24/SC4 | X | |||
| ISO/TC 201 | X | |||
| ISO/TC 202 | X | |||
| TS Z 0027:2010 | X | |||
| Occupational Safety & Health Administrationh
| ||||
|---|---|---|---|---|
| Terminology/Nomenclature | Measurement/Characterization | Materials Specifications | EHS | |
| 1904 | X | |||
| 1910.132 | X | |||
| 1910.133 | X | |||
| 1910.134 | X | |||
| 1910.138 | X | |||
| 1910.141 | X | |||
| 1910.1200 | X | |||
| 1910.1450 | X | |||
| United Working Group for Nanomaterials Standardization (China)i
| ||||
|---|---|---|---|---|
| Terminology/Nomenclature | Measurement/Characterization | Materials Specifications | EHS | |
| GB/T 19619-2004 | X | |||
| GB/T 13221-2004 | X | |||
| GB/T 19587-2004 | X | |||
| GB/T 19588-2004 | X | |||
| GB/T 19589-2004 | X | |||
| GB/T 19590-2004 | X | |||
| GB/T 19591-2004 | X | |||
There are many other worldwide initiatives actively involved in the study and development of nanotechnology standards, such as the International Alliance for NanoEHS Harmonization5, the Nanosciences African Network6, the Asia Nano Forum7, the Bureau International des Poids et Mesures8, the International Union of Pure and Applied Chemistry9, the Organisation for Economic Co-operation and Development10 and the Versailles Project on Advanced Materials and Standards11, among many others.
2.3. Information sources
The variety of nanomedical data sources constitutes another main issue for information management in nanomedicine. Usually, these sources are disparate and heterogeneous, providing different types of data in different formats. These sources are also physically separate, forcing researchers to spend research time locating them. In addition, information sources are of different natures: structured (databases and knowledgebases, for example) and unstructured (wikis, technical reports and scientific publications are examples). Structured sources, where data are well-defined and semantics are implicit, could be easily mined with automatic methods that are already available for the scientific community: standard queries to databases, for instance. On the other hand, unstructured sources provide data in free text format, making it difficult to automate the process of information extraction. Results from nanomedical research are usually reported in scientific publications and technical reports, resulting in large volumes of unstructured text. As can be seen in Figure 2, the number of publications in the field has grown considerably in the last two decades. This led to the urgent need to develop advanced natural language processing (NLP) techniques and text mining methods to convert these unstructured sources to machine-interpretable format. This issue has been partially addressed in the biomedical domain through biomedical informatics approaches, and should be addressed in the same way in the nanomedical context, by developing and applying specific nanoinformatics methods and techniques (de la Iglesia et al 2009).
Figure 2.
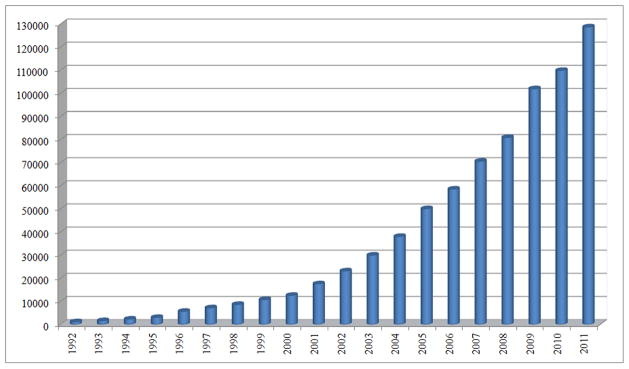
Number of publications with nano-prefixed terms in the title published over the last 20 years (1992–2011), following previous studies (Braun et al 1997, Huang et al 2003, Hullman and Meyer 2003, Lee et al 2006). Source: Web of Knowledge (Thomson Reuters)12.
Although there are several nanomedical data sources, it is necessary to analyze the availability and quality of such data and properly handle the existing inconsistencies and uncertainties. For instance, protocol details are inadequately described in research publications and original datasets are rarely provided in manuscripts or deposited online. Standard methods for data validation should be developed, covering the different aspects of research conducted in nanomedicine, such as material conditions or protocol parameters. In the last five years, the number of web applications offering access to nanomaterial data sources has grown rapidly, so it has become necessary to evaluate and compare the specific features of such sources: i) the scope of the provided data, ii) the level of annotation of nanomaterials with relevant information, iii) the automatic generation of catalogues from research results, and iv) the use of taxonomies and ontologies to classify the available data on nanomaterials. Another important feature of nanomaterial inventories to be evaluated, considering its use by the scientific community, is the search capabilities that it can offer. Searching for nanomaterials by name in a list sorted alphabetically is an arduous task that does not always generate the desired results. Additionally, researchers may not always know the name of the nanomaterial they are looking for (only some properties or the name of manufacturer) and, even when they do know it, it is a good idea to show them other nanomaterials that have similar characteristics. Using ontologies can assist users who are browsing the entire collection by organizing the repositories with a hierarchical category classification.
Another relevant characteristic of nanomedical data is its interdisciplinary nature. Data from chemistry, physics, medicine and other involved fields could be very different in terms of format, measurement units and vocabulary. Additionally, it generally needs to be integrated with multilevel biological data, ranging from the population level to the atomic level. The development of comprehensive data repositories linking and integrating data from diverse domains and different sources—experimental results, scientific literature, clinical trials, Electronic Health Records (EHRs), genomics, proteomics, nanomaterials characterization, imaging and disease information—would lead to the design of more accurate predictive models and simulations of nanomaterials and their biological effects.
3. Available resources for nanomedical researchers
Several initiatives are currently supporting nanoinformatics research worldwide. For instance, since 2001, under the National Nanotechnology Initiative (NNI) (Rocco 2003), 25 governmental agencies are sustaining nanotechnology advancements in the United States. Previous initiatives also connect to activities under the NNI, such as the Global Change Research Program13 and the Networking and Information Technology Research and Development14, as well as other new initiatives, including the Materials Genome Initiative (National Science and Technology Council 2011), a material innovation infrastructure launched in 2011, and the Advanced Manufacturing Partnerships. In Europe, diverse projects aim at raising the levels of nanotechnology knowledge across the European Union Member States by supporting the analysis and dissemination of results from nanotechnology research, as we will see later in this manuscript.
The set of available nanoinformatics resources comprises databases, tools and web services useful for storing, organizing, managing and analyzing the plethora of data generated during nanotechnology research. In the next sections, we will analyze these resources, providing a comprehensive picture of the ongoing research efforts and advances in the field of nanoinformatics.
3.1. Data sources
Nowadays, there are several resources providing free access to nanomedical data. These repositories contain large amounts of heterogeneous information related to nanomedicine, including data about samples and experiments, specific studies and related protocols, nanoparticle interactions, exposure pathways, delivery targets, manufacturers, etc. Additionally, many research institutes and groups have developed elaborate nanoparticle classifications based on different parameters: nanomaterial application domain, size, function, target and toxicity, among others. The variety of ways used to categorize nanomaterials is the main barrier to standardizing common classifications. Therefore, the creation of a standard taxonomy for the classification of nanomaterials is a key element to be considered. The use of standardized data across systems and services would facilitate the updating and integration of data from many different sources. Below, we provide some examples of the interesting initiatives that are providing access to sources of information on nanomedicine and nanomaterials, as well as a comparison of their specific features (table 2).
Table 2.
Comparison of some data resources on nanotechnology and type of data provided.
| Resource | Database | Nanomaterials Characterization | Protocols | Assays | Reports Technical | Scientific Publications | Toxicity | Tools Modeling | Interactions |
|---|---|---|---|---|---|---|---|---|---|
| Nanomaterials Registry | X | X | X | X | X | X | |||
| NIL | X | X | |||||||
| ICON Nano-EHS database | X | X | X | X | |||||
| NBI Knowledgebase | X | X | |||||||
| CSN | X | X | X | ||||||
| caNanoLab | X | X | X | X | |||||
| TOXNET | X | ||||||||
| NanoExPERT | X | X | X | X | |||||
| MICAD | X | X | |||||||
| NCL | X | X | X | X |
The Nanomaterial Registry (Ostraat et al 2012) gives access to curated information on the interactions of nanomaterials with biological and environmental systems, including links to related publications, modeling tools, computational results and manufacturing guidance. This initiative, created and maintained by Research Triangle Institute International, is sponsored by the National Institute of Biomedical Imaging and Bioengineering, the National Institute of Environmental Health Sciences and the NCI. Users can search this database for information about a specific nanomaterial or compare the properties of multiple nanomaterials. Users can also browse by nanomaterial type (metals, fullerenes or dendrimers, for example), size, shape (one, two or three dimensions) and surface area. Each record contains information about the physicochemical and structural characteristics of the nanomaterial, including: size (particle size and distribution), aggregation or agglomeration state, surface (area, charge, chemistry and reactivity), shape, composition, purity, solubility and stability. Information about the environmental and biological interactions of the nanomaterial is also provided. The Nanomaterial Registry is an ongoing project; its first beta version was released to the public at the end of June 201215 and is updated as new information about nanomaterials is published. This resource will also offer, in future releases, tools for predictive modeling of nanomaterials. Another relevant data source is the Nanoparticle Information Library (NIL)16, an ongoing project for building a web-based nanoparticle repository, developed by the National Institute for Occupational Safety and Health (NIOSH) and the Oregon Nanoscience and Microtechnologies Institute through the Safer Nanomaterials and Nanomanufacturing Initiative17. A user can browse nanotechnology reports from a set of 88 nanoparticle records (at the time of writing). Each record contains the nanoparticle’s material data, such as core element, structure or production method, the nanoparticle’s image, an abstract describing the chemical and physical properties of the nanoparticle and information about pertinent scientific publications and other related data, such as its commercial availability or the sponsoring company or institution. These records also provide links to websites with extended information about the described nanoparticles. With the same goal, the International Council on Nanotechnology is developing the ICON Environmental, Health and Safety (EHS) database18. The ICON Nano-EHS database is a repository of papers related to nanoparticles. At the time of this writing, it contains over 7,000 records. In this repository, the user can perform searches based on a particular characteristic of the nanoparticle—particle type, exposure pathway, method of study, exposure or hazard target, risk exposure group and production method—or based on related publications, such as paper type, target audience and content emphasis. This initiative also provides an analysis tool offering two types of analyses: i) a simple distribution analysis (pie chart), which compares categories within a specified time range and, ii) a time progressive distribution analysis (histogram), which compares categories over a specified time range and data grouping period. Another valuable resource is the Nanomaterial-Biological Interactions (NBI) Knowledgebase19, an initiative funded by Oregon Nanoscience and Microtechnologies Institute (ONAMI) and several U.S. agencies: the National Science Foundation (NSF), the National Institutes of Health (NIH), the United States Environmental Protection Agency and the Air Force Research Laboratory. The NBI Knowledgebase is a web-accessible electronic library that allows users access to precise and standardized data element definitions. It serves as a repository for annotated data on nanomaterial characterization (e.g. purity, electronic and photonic properties, size, shape, charge, composition, chemical functionalization and agglomeration state), synthesis methods and nanomaterial-biological interactions (beneficial, benign or deleterious) defined at multiple levels of biological organization (e.g. molecular, cellular or organismal), thus providing the framework upon which to conduct species, route, dose and scenario extrapolations and to identify key data required to predict the biological interactions of nanomaterials. Computational and data mining tools are currently being developed and incorporated into the NBI Knowledgebase to facilitate this work.
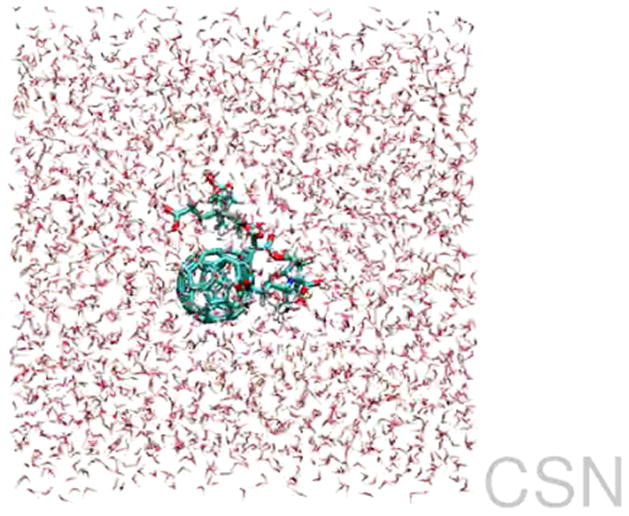
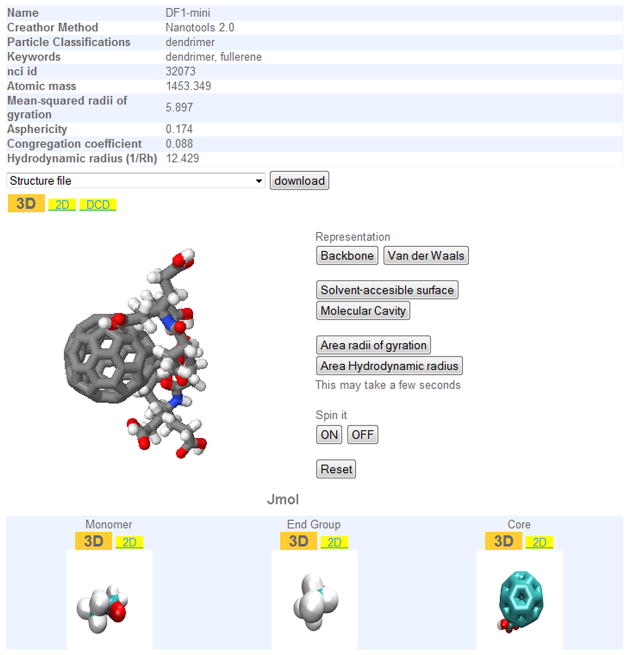
The comparative analysis of structural complexities in nanoparticles and nanocompounds is a fundamental factor in understanding the particular properties and behaviours of nanomaterials. Regulatory approval of new engineered materials should be obtained only after a rigorous review of data from structural and functional analyses. Thus, the development of accurate models and simulations would be an essential step for the translation of results from the bench to the bedside, reducing the number of necessary phases required in clinical trials and, subsequently, the time-to-market of these manufactured nanomaterials. Following this structural approach, the Collaboratory for Structural Nanobiology (CSN)20 initiative, coordinated by one of the co-authors, provides a database of chemical and structural models of nanoparticles such as dendrimers, buckyballs, nanotubes and metallic particles. Each nanoparticle is associated with its correspondent structure file, similar to those offered by the Protein Data Bank (PDB), and related research data, such as molecular weight, simplified molecular input line entry specification (SMILES) or studies on van der Waals forces. The platform also provides several tools for the two-dimensional (2D) and three-dimensional (3D) visualization of nanoparticles (figure 3), as well as representative videos of dynamic simulations of such nanoparticles (figure 4). This kind of approach could lead to future platforms upon which to share structural and predictive models.
Figure 3.
Example of nanoparticle record in the CSN: a 3D representation of a fullerene (DF1-mini)
Figure 4.
Video simulating the water molecular dynamics of DF1-mini in the CSN
Another interesting data source for nanomedicine is caNanoLab21, a portal for sharing information about nanomaterial characterizations resulting from cancer-related nanotechnology research, funded by the NCI. It supports the annotation of nanomaterials with data from in vitro assays and allows linking to remote characterization protocols. At the time of writing, the caNanoLab Portal (v1.5.3) provides access to 41 protocols and 886 samples from different sources, supplying information about the sample, such as composition, function and characterizations, and providing access to more than 1,000 publications related to the stored samples. In addition, the portal supplies the data submission date in the search results. It is an open-source initiative, so users are able to download the source code22 and try it in their own systems.
Also accessible through the web, the Toxicology Data Network (TOXNET)23 provides access to toxicology data for about 5,000 chemicals, including nanomaterials. TOXNET comprises several databases from the U.S. National Library of Medicine and some of them—the Hazardous Substances Data Bank (HSDB) and TOXLINE—allow general and specific searches about nanoparticles and nanostructures. Records can be queried by substance name or Chemical Abstract Service (CAS) number. The HSDB provides chemical records about nanotoxicity by means of descriptions of physicochemical properties, toxicity studies, pharmacology, manufacturing, environmental and health safety, pharmacodynamics, etc. On the other hand, TOXLINE provides bibliographic records of scientific papers related to nanomaterials’ toxicity. All of this information is presented to the user in free text, but can also be integrated with other applications via its web services Application Program Interface (API). NanoExPERT24, the Nanomaterials Experiment-based Predictor of Environmental Risk and Toxicity, is another tool that provides fate and toxicity predictions for nanoparticles, as well as a database containing experimental results from peer-reviewed journals. With respect to images databases, the Molecular Imaging and Contrast Agent Database (MICAD)25, developed by the National Center for Biotechnology Information (NCBI) at the NIH, stores imaging data about a range of nanomaterials, such as nanobubbles, quantum dots and gold nanoparticles.
Finally, it is worth mentioning the Nanotechnology Characterization Laboratory (NCL)26 for conducting preclinical studies to characterize nanomaterials that are used for in vivo diagnostics and therapy against cancer. This outstanding nanotechnology institution, developed through a collaboration between the NCI, the FDA and the National Institute of Standards and Technology, provides access to a set of high-quality technical reports regarding liposomes, fullerenes and dendrimers. There are also other available databases dedicated to collecting transversal data about nanotechnology, such as the database of nanoscience journal publications developed by Braun (Braun et al 1997) and the database of nanotechnology patents, developed by Meyer (Meyer 2000, 2001).
3.2. Standards, terminologies, taxonomies and ontologies
In order to address the organization and homogenization of the huge volume of information that nanotechnology research is producing nowadays, scientists need support from lexical and semantic tools, such as terminologies, vocabularies, nomenclatures, code and synonyms sets, lexicons, thesauri, ontologies, taxonomies and classifications. Specifically, there is an urgent demand for guidance in labeling and annotation of nanomaterials, as well as for the production of electronic tools for standards development and revision. Besides, since ontologies are meeting urgent needs in other areas, such as biomedicine (Maojo et al 2011), it is reasonable to think about their applications in the nanomedical field. Ontology is a model describing the concepts of a specific domain, as well as their categories and relations. In nanoinformatics, ontologies can be considered a semantic framework representing the nanotechnology domain. This model may contain a myriad of chemical, physical and nanotechnology concepts that can be used as controlled terminology for data management and analysis, allowing further nanoinformatics tasks, such as the annotation of experimental data or the integration of heterogeneous data sources, to be undertaken. Considering the extensive vocabulary and the wide range of names and terms for the same concepts in this area, ontologies would provide a mechanism to standardize the representation of the nanomedical domain that could be very useful for the dissemination of information and knowledge that exists in this field in a machine-readable format and to express the metadata related to nanomaterials in a semantically interoperable way.
In 1999, Freitas proposed a primer vocabulary for nanomedicine in the Nanomedicine Book Glossary (Freitas 1999), included in his nanomedicine book series27, followed by a partial taxonomy of nanomedical technologies, published in 2005 (Freitas 2005). That same year, Tanaka developed a preliminary ontology for nanoscience, a meta-level ontology for nanoscale research (Tanaka 2005), based on an analysis of the subject headings and controlled index terms of papers published in journals related to materials science, physics, chemistry, engineering, medicine and biology.
Some popular indexes, such as the Library of Congress Classification28, the Dewey Decimal Classification29 and the Bliss Bibliographic Classification (Thomas 1997) are already indexing nanoscience. There also are biomedical and material science indexes that include nanotechnology, such as Medical Subject Headings (MeSH), a controlled vocabulary for indexing publications (Lowe and Barnett 1994), and the NCI Thesaurus (Sioutus et al 2007)30, developed by the NCI Center for Bioinformatics as part of the Enterprise Vocabulary Services Project, among others. Additionally, standardized terminologies for nanotechnology and nanosciences can be accessed through the websites of the main international organizations for standards, as noted in previous sections of this review, e.g. the Standard Terminology Relating to Nanotechnology included in ASTM E2456–0631 (published by ASTM International, formerly known as the American Society for Testing and Materials). Other non-standardized glossaries and vocabularies, developed by research institutions and companies, can be found on the web, such as the Glossary for NanoBiotechnology32, developed by JPK Instruments, and the Glossary of Nanotechnology Terms33, created by the UK Institute of Nanotechnology. The Dictionary of Nanotechnology34 expands such glossaries with single-level classification, offering a nanotechnology vocabulary that contains terms collected from different scientific sites and references and classified into different categories: general, nanomaterial and nanoelectronic, nanomedical, nanobio and bionano, carbon nanostructure, nanocharacterization, nanofabrication and molecular nanotech.
Several general taxonomies for nanotechnology can also be found on the Internet. For instance, the Nanotechnology Taxonomy developed by Codesta35, which divides nanotechnology products based on their manufacturing process by using three criteria: manufactured state, chemistry of fabrication and means of arrangement, and the Nanomedicine Taxonomy developed by the Canadian NanoBusiness Alliance36, which classifies terms by their area of application (e.g. biopharmaceuticals, implantable devices, surgical aids, and so on). Another example is the taxonomy developed by Wong et al (2007) for classifying patents based on their area of application: i) instrumentation, ii) chemical processes and materials, iii) medical and biotechnology and iv) nanoelectronics. In a similar direction, the United States Patent and Trademark Office included in its classification a hierarchical terminology named Class 977 “Nanotechnology”37 to categorize existing patents in the field. Nanosafe Inc.38 also provides a logical classification of nanoparticles based on their elemental composition, properties and toxicity in support of defense-related nanotechnology applications.
The InterNano initiative, a project of the National Nanomanufacturing Network funded by the NSF, is developing a general taxonomy39 that defines the nanotechnology domain. The InterNano taxonomy brings together content about nanomanufacturing from a variety of sources and is divided in eight main categories, titled: i) Areas of Application, ii) Environment, Health and Safety, iii) Informatics and Standards, iv) Nanomanufacturing Characterization Techniques, v) Nanomanufacturing Processes, vi) Nanoscale Objects and Nanostructured Materials, vii) Social and Economic Impact and viii) Tool Development. Many of these categories, in turn, are divided into several subcategories down to three levels of granularity. The taxonomy can be navigated in three different ways: as a hierarchical tree, as a tag cloud or as graph with nodes, presenting the relationships between the extant terms of the taxonomy. The database contains over 1,000 elements and each element of the taxonomy is tagged and linked to the portal’s contents and repositories, which include descriptions, scientific publications, technical reports, techniques, methodologies and patent information related to manufacturing processes for nanostructured materials and devices. It also includes lists of organizations and contributors to these processes, as well as usage statistics. Users can submit their own processes by uploading the related information in standard forms, facilitating the dissemination of this knowledge to organizations and laboratories.
The NanoParticle Ontology (NPO)40, primarily developed by Thomas et al (2011) and currently supported by the NCI’s cancer Biomedical Informatics Grid (caBIG®) Nanotechnology Working Group (Nano WG), provides the frame of reference for representing the cancer nanotechnology domain. A user can organize and annotate knowledge underlying the preparation, chemical composition and characterization of nanomaterials involved in cancer research by browsing and navigating the ontology’s hierarchies of classes. Currently, NPO researchers are focused on support for ISA-TAB-Nano41, an extension of the standard Investigation/Study/Assay (ISA) format42 developed by the European Bioinformatics Institute, and other annotation projects. The NPO is also being updated to include a broader range of nanotechnology concepts. The continuous advancements in the development of this ontology translate to great steps forward for nanotechnology standardization and data interoperability, paving the way towards a global ontology for nanomedicine.
Despite of the myriad varieties of glossaries, vocabularies and taxonomies focused on nanotechnology, there is an urgent need for a unique controlled vocabulary for the nano domain and, specifically, a standard specification, to facilitate the import/export of data on nanomaterials and their characterizations to/from nanotechnology resources. A current major effort to reach consensus in nomenclature and categorization is ISA-TAB-Nano, a standard, approved by ASTM International, which aims to represent and share information, including characterization data, about nanomaterials. Developed by the NCI’s caBIG® Nano WG, this standard provides a framework for describing investigations, studies, materials and assays through plain text files. In addition, ISA-TAB-Nano supports nanomaterials and concepts from the NPO, facilitating annotation and integration with this ontology.
3.3. Modeling and simulation tools
Creating in silico models and simulation environments at the nanoscale could assist in the prediction and assessment of risks of engineered nanomaterials; for instance, modeling differences in toxicity response due to changes in particle properties. In this respect, it would be necessary to determine which characteristics are most important for building a high-quality model, and to include these characteristics in accurate modeling and simulation tools. It would also be fundamental to abandon the current theoretical approach and consider factors such as the dependencies between the nanoparticle’s properties and the system where they are embedded or the behaviour and time-scale of nanoparticles in the human body (degradation, toxicity, side effects, etc). For this purpose, extended knowledge about the biological identity of engineered nanomaterials (function, related biological effects, domain of application, relationships between particle shapes and toxicity, etc) would be a requisite (Lynch et al 2009).
Also to be developed are modeling and simulation tools that would allow researchers to predict new properties and potential structures to acquire knowledge about the nature and behaviour of nanoparticles. These data could include: i) nanomanufacturing models for the virtual design of engineered nanomaterials considering various requirements simultaneously: nanomaterial physicochemical characterization, functionalization of the nanomaterial surface and design and control of synthesis processes, among others; ii) models of pharmacokinetics and pharmacodynamics; iii) models of nanoparticles’ life cycle in the body: biodistribution, metabolism disposal, immunogenicity, clearance, phagocytosis, etc; and iv) quantitative structure-activity relationships (QSARs) for modeling nanoparticle structure, nanotoxicity and nanoparticle interactions in biological environments at the tissue, cellular, molecular and atomic level. A good example is the work carried out at the UCLA Center for the Environmental Implications of Nanomaterials (CEIN). CEIN researchers have developed nano-QSARs (Puzyn et al 2011) for predicting nanoparticle toxicity (Liu et al 2013) as well as a set of web-based High-Throughput Data Analysis Tools for rapid analysis of nanotoxicity data43.
3.4. Educational tools
Due to the novel and heterogeneous character of nanoinformatics, the development of interdisciplinary and cross-disciplinary education programs integrating nanomedicine, biology, materials science, physics, chemistry and informatics has become a priority (Greenberg 2009). Over the last few years, several educational resources and repositories have emerged to give training support to researchers working in the field.
For instance, nanoHUB44 (Qiao et al 2006) is a web portal for educational purposes developed by the Network for Computational Nanotechnology and sponsored by NSF under the NNI. This modeling and simulation framework offers real-time demonstrations of a myriad of extant computational resources in nanotechnology, materials science, physics and chemistry. NanoHUB does not allow the direct upload of experimental data, but users can run specific simulations and models by selecting the values for the different parameters requested by each resource. Users can also act as contributors, uploading their own resources and making them accessible to the vast scientific community supported by nanoHUB. Additionally, the portal provides access to courses and learning material—such as extensive lecture notes, quizzes, exams, homework and exercises—video lectures, animations, presentations about the available resources, online seminars, related scientific publications and information about events and job opportunities. There are also question-and-answer forums for interacting with professors and other students. Similar educational frameworks for e-learning on nanotechnology (e.g. Nano4Me45) are gaining force in the Web.
The NCL is also conducting educational activities by publishing and publicly disseminating the recent advances in preclinical studies on nanomaterials by means of scientific bibliographies and conferences, news, open forums and press notes in scientific media. There are also plenty of web pages and other media oriented to the general public that serve to distribute this information. For instance, Nanowerk46 and NanoTechWeb47 provide access to nanotechnology articles and news oriented to scientists and to the general public.
In spite of the didactic initiatives mentioned above, it is still necessary to develop and sustain additional educational resources and an international network of skilled professionals, as well as the supporting infrastructure and tools to advance training in nanotechnology. This involves the organization of workshops at international scientific events that engage students, technicians and trainees in nanotechnology environmental, health and safety considerations, and the establishment of a network of websites for student recruitment and dissemination of information, e.g. webinars and courses.
3.5. Working groups on nanoinformatics
In order to integrate and disseminate information on global nanotechnology research efforts, it is necessary to create and sustain international groups of interest by identifying experts in the field among networks, research organizations, public agencies and the nanotechnology industry. Working groups provide an excellent opportunity to bring together individuals representing a broad range of perspectives to accomplish a common goal. While scientific communities in long-established areas, such as bioinformatics, have demonstrated the high value of collaborative approaches to advance science, such efforts by the newer nanoinformatics community are less developed. However, this is changing quickly through the organization of joint workshops and publications, as well as the creation of working groups within in various sub-areas under development. These working groups are composed of scientists from different laboratories, research institutions and universities, committed to working together and sharing complementary capabilities to enhance nanotechnology research.
For instance, the caBIG® Nano WG48 is developing concerted activities and several pilot projects49. Decisions and processes within the group are consensus-driven and representatives from the participant groups meet regularly, generally once a week. Within this set of pilots, it is worthwhile to highlight the Meta-ontology Pilot50, related to the NPO, and the ISA-TAB-Nano pilot51, related to the standard previously described in this paper. In Europe, there are multiple collaborative initiatives, such as QNano52, a research infrastructure for nanomaterial safety and quality testing; NanoImpact.net53, a network concentrating on the health and environmental impact of nanomaterials; the EU NanoSafety cluster54, a group of European Commission-funded projects working on nanotoxicology and nanosafety issues and involved in the preparation of new research topics for the next European-funded Framework Programme (Horizon 202055); the European Technology Platform on Nanomedicine56, a group of industrial and academic stakeholders working for the establishment of a shared vision an research lines in the area; the European Technology Platform for Micro- and NanoManufacturing57, a collaborative platform for discussing topics of interest to industrial micro- and nanomanufacturing; MODENA58, an action modeling nanomaterial toxicity; and NANOfutures59, a shared environment for collaboration between industry, academy and society on nanotechnology issues which is currently creating a roadmap in the area, among many others. These European groups are working cooperatively to achieve their individual scientific goals, as well as to meet the common objective to advance nanotechnology research at the European level. In the international context, it is particularly worthwhile to mention the ongoing work of the US-EU dialogue, “bridging nanoEHS research”60, which is supporting nanotechnology through the establishment of worldwide Communities of Research.
4. Cyberinfrastructures for nanomedical data management: current limitations and future prospects
The existing nanotechnology platforms, data resources and tools are powerful, but some aspects are already being improved. For instance, usually each service provider’s interface is different—both in the subset of data presented and in structure—and each institution promotes its own web sites, services and data formats. This presents a challenge to nanomedical researchers, who typically need to link data from multiple online sources to generate a data set for further analysis. The development of new standards and specifications, like the ISA-TAB-Nano and the NPO, is directed at harmonizing access to nanotechnology data.
Another issue is that the development of inventories of nanomaterials has been done without automating the process, thus consuming a big part of researchers’ time. Inventories are manually maintained by trained personnel who conduct periodic updates based on the new published material on nanoparticles. In fact, some of these data sources are not pure databases, but web pages containing lists of nanomaterial descriptions. In some cases, users can contribute by uploading new data. Nevertheless, now that relevant resources are widely distributed over the Web and data are made available in different formats, manual integration has become almost infeasible and could lead to the publication of non-comprehensive indexes that include only a portion of the available data about nanomaterials. To replace the manual development of nanomaterials repositories, there are some quite innovative approaches that could generate indexes of nanomaterials in an automatic manner. These approaches would facilitate the initial implementation of the index and, above all, allow its continuous update and curation, adding new information and deleting the obsolete data. Due to the rapid growth of information about new experiments and results in nanomedicine, the automation of the creation and update process is an important element to be considered.
Joining data sources together is quite problematic, as they frequently have insufficient or nonexistent metadata describing the nanomaterials. An important consequence of the lack of metadata is that some existent nanomaterials cannot be found in all the registries. Besides, sometimes the information provided by the different sources is not directly compatible with other sources and tools. These issues are being addressed by applying information technology methods and techniques and, therefore, a comprehensive infrastructure is needed to support an interoperable network of data and analytical services. Transforming the current efforts to develop databases and tools for nanomedical research into a dedicated nanoinformatics research infrastructure requires an understanding of the needs of the scientific community, the need for efficient and high-quality management of information at the nano level and an understanding of the strengths and weakness of experimental results and clinical data. This multidisciplinary, open and free infrastructure should provide a new way for scientists to immediately access and use data from nanomedical research, saving them from investing time in fruitless web searches. With such an infrastructure, researchers can focus on moving forward with their research, not on acquiring the necessary resources for data analysis and management.
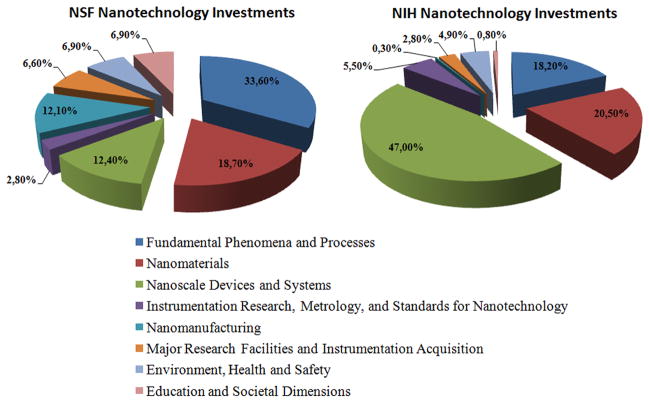
For instance, in the United States, the NNI61 has led to public funding for nanotechnology research (see figure 5). According to a recent NNI statement62, published on May 14, 2012, the establishment of a Nanotechnology Knowledge Infrastructure is a priority research line for the forthcoming years. The Subcommittee on Nanoscale Science, Engineering and Technology, within the National Science and Technology Council’s Committee on Technology, announced, through the publication of a White Paper63, the fourth Nanotechnology Signature Initiative to stimulate the development of and improvements in the tools and methods required to access, organize and gather breakthroughs from huge amounts of data generated by nanotechnology, beginning to weigh in on just what data really need to be intelligently managed and stored.
Figure 5.
Comparison of the NNI investment distribution (2011–2013) into the different nano areas between two US governmental agencies: the NSF (covering general research on nanotechnology) and the NIH (focused on nanomedicine). Source: NNI Dashboard64
4.1. Previous successes in the biomedical domain
Computational resources and infrastructures for biomedicine have successfully given support to address a major part of the challenges existing in the field of biomedicine. For instance, the development of databases, tools and services has accelerated advancements and findings in the omics fields. Researchers have benefited from these resources, adapting their professional skills to the new technological requirements for managing biomedical information. This has been the case for numerous existing omics databases that have proliferated in recent years, providing a common framework for research on genomics [e.g. Online Mendelian Inheritance in Man (Hamosh et al 2005), Ensembl GenBank (Benson et al 2012) and Artemis (Rutherford et al 2000)]; proteomics [e.g. PRoteomics IDEntifications Database (Martens et al 2005), Database of Interacting Proteins (Xenarios et al 2002), Swiss-Prot (Bairoch and Apweiler 2000) and Universal Protein Resource (Bairoch et al 2005)]; epigenomics [e.g. NCBI Epigenomics]; metabolomics [e.g. Human Metabolome Database (Wishart et al 2007); and others.
An exemplary model to emulate would be the development strategy for the PDB (Berman et al 2000), the outstanding, worldwide protein structure database. The PDB has grown over the decades, since its creation in 1971, as a result of international collaboration between researchers who have deposited their protein structures in the database. This open strategy—where researchers can submit their own work, following a set of standards and guidelines, and research institutions act as storage, data processing and distribution centers for PDB data (Berman et al 2003)—led to the cooperative development, population and curation of this database.
In biomedicine, several application development projects have emerged for constructing and managing computational pipelines from existing bioinformatics databases and tools. A good example is iTools65 (Dinov et al 2008), a system for classification, categorization and integration of different computational biology resources. This system has several functionalities, such as storage, searching, traversal and retrieval of resources and their corresponding metadata description. iTools infrastructure includes an ontology of computational biology resources, a database of descriptions of computational biology resources and an infrastructure for integration and interoperability of meta-algorithms for biomedical computing. Each resource is annotated with metadata that includes a short description, its authors, version, date and stage, the ontology classification of the resource, a set of keywords and the implementation language. In the ontologies’ area, it is worthwhile to mention the BioPortal (Noy et al 2009), a compendium of ontologies for life sciences that is constantly updated with new ontologies and new versions of the existent ones. Regarding the indexing of bioinformatics databases and tools, there are some remarkable ontologies attempting to generate a standardized classification of bioinformatics resources, like the Biomedical Resource Ontology66 (Musen et al 2008), a controlled terminology defining bioinformatics resources types under development by NIH researchers who have an interest in the classification of such resources. Medical imaging is another field that has grown considerably due to successful advances in the techniques for image acquisition, analysis, storage and annotation. A renowned resource in this field that should be mentioned is Digital Imaging and Communications In Medicine (DICOM)67, the standard for exchanging medical images.
All of these resources are highly accessed by biomedical researchers on a daily basis, which indicates the relevant role for computational tools and infrastructures in conducting and reporting experimental research. While it is certainly correct that investigators typically do not control the systems in the sense of design, administration, etc, they do maintain the data. Thus, these sorts of approaches should be followed in the nanomedical area to develop and publish nanoinformatics resources.
4.2. Requirements for developing a nanoinformatics infrastructure
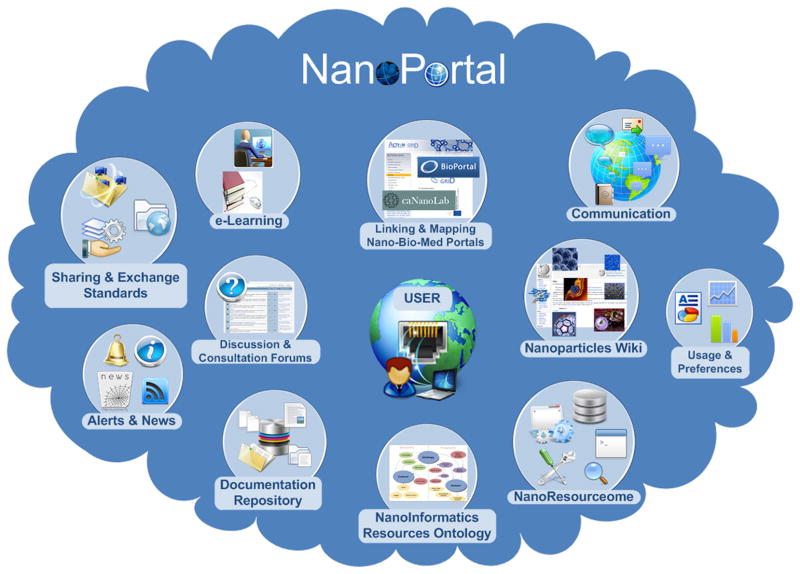
This section describes the desired elements of a hypothetical, multidisciplinary, open infrastructure, a research framework including a set of applications enabling easy access to a myriad of nanomaterials, enhancing cooperation among different institutions and researchers (see figure 6). This infrastructure should provide users with uniform access to a wide set of sources of nanomaterial data, not only collecting and integrating information from different databases but also offering information (metadata) about the source of the data. The nanoportal should offer services for sharing data and resources between researchers—e.g. scientific articles, demos, access to nanomedicine databases, ontologies, terminologies. It would also serve as dissemination channel for characterization data and aim at looking across characterization profiles and techniques. In order to accelerate the discussion and validation of nanomedicine research data, the infrastructure should support services for data annotation, data curation and data quality rating, which could improve the quality of published experimental results. Curation will rely on the literature descriptions as well as in registered experiments and results. The infrastructure should also help researchers to keep track of their experiments (techniques, parameters, protocols details) and then apply computational techniques and methods to analyze and inference knowledge from such data. We believe that such an open-environment approach could foster the development of novel integrative tools, accessible over the Internet, to share nanotechnology knowledge and resources through a geographically distributed virtual community.
Figure 6.
The NanoPortal: A hypothetical, multidisciplinary, open infrastructure enabling uniform access to a myriad of resources on nanotechnology
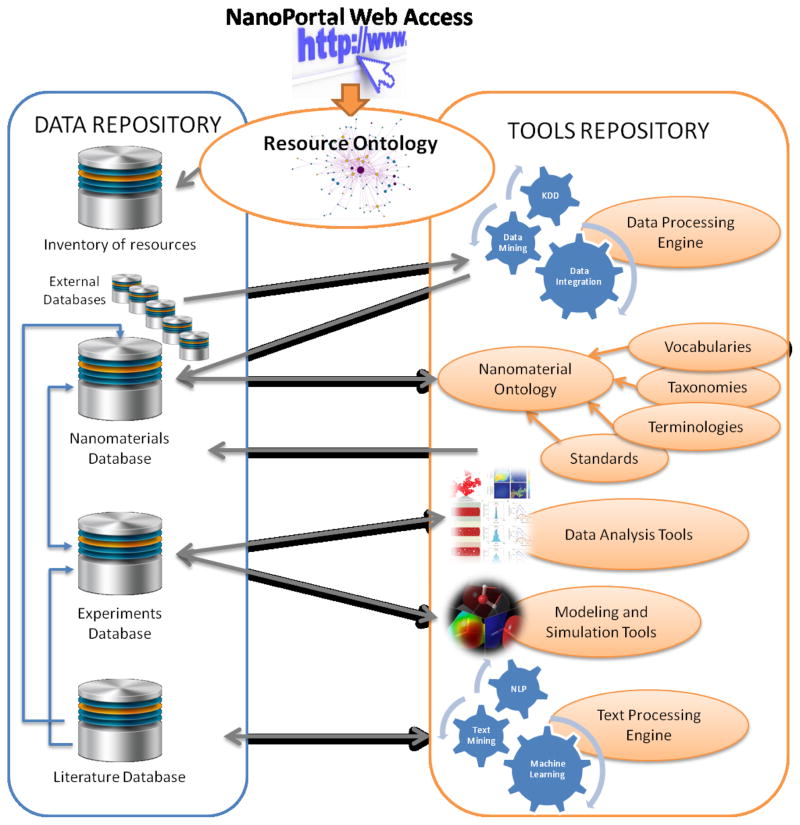
This infrastructure could be divided into two different parts (Figure 7). The first one would consist of a Data Repository to support the exchange of nano research data, publish data generated by the research community and make these data freely available. All these data would be stored in a database to be queried over the Internet. Each nanomaterial would appear annotated with its corresponding information and the whole data set would be wrapped by an upper-level ontology. The second part would be the Tool Repository containing tools and services which would be classified into several categories: acquisition, calibration and quality assurance; data upload, storage and management; analysis and processing; data sharing and collaboration; visualization and “atlasing” tools; and standards, terminologies and ontologies. For this purpose, the automatic assignment of categories to the nanomaterials could be carried out with the use of text mining and NLP techniques to extract relevant information from papers describing them. Clustering engines could also help in this categorization by the application of standard mechanisms and algorithms, such as normalization, feature selection, consensus clustering and multidimensional scaling, among others (García-Remesal et al 2010). Nanomaterials would be clustered into sets so as to discover useful patterns (García-Remesal et al 2013). Data and tools gathered in the repositories should be committed to an open-source policy and the software should be freely available, allowing users to share their tools.
Figure 7.
Schematic representation of the main components of the infrastructure and their interconnections
The creation of such a nanoinformatics platform would also involve the mandatory development of the following elements:
Terminologies, thesauri and vocabularies defining the nanotechnology domain and, specifically, the nanomedical semantic context: The proposed data model would use nanomedical domain information captured in terminologies, thesauri and taxonomies, which provide information related to nanomaterials, such as chemical descriptors, toxicity, size and structure. The organized description of nanomaterials through these properties will enable users to make advanced searches, such as all nanomaterials that perform a specific function or have a concrete composition.
Standards that will allow data and systems interoperability: Having a controlled terminology is a key factor for indexing and integrating nano sources; therefore, the platform should be supported by a set of well-established international standards.
Taxonomies and ontologies modeling the scientific knowledge existent in the nanomedical domain, including the design and implementation of services for searching and managing such taxonomies and ontologies: For organizational purposes, the ontology could be separated into two different components: the domain ontology, an annotation vocabulary that would include descriptions of nanomaterial data types and their relationships, and the service ontology, which would describe the operational features of services within the domain ontology. The final goal of this ontology would be to capture the semantics of the relationship between the different nanomaterials, providing a contextual framework to share data and develop interoperable resources using this information.
Databases of related resources, bibliography, data sources, etc., including services for generating use statistics and monitoring user preferences: As the nanotechnology field grows, it has become harder to find and access all the knowledge that is relevant to the discipline. In addition, non-specific search engines –such as Google— produce too general and massive results, so, in most cases, researchers are unable to find the best resource to obtain the target information necessary for their research. Therefore, it is necessary to solve the gap between the rising information needs of nanomedical researchers and the growing number of information sources available through the web, and create inventories indexing, describing and linking the existing data. As we have seen before, over the last years, several research groups and laboratories have developed directories of resources which work as “yellow pages” for the biomedical scientific community. The same approach should be followed by the nanomedical community, promoting the dissemination and sharing of the currently available resources in the field. Then, it would be needed to describe potential applications of these indexes, including the automatic orchestration of nanoinformatics workflows, or novel methods to integrate public-online nanomedical databases and tools.
Tools for the analysis and visualization of experimental results, accompanied by reference data sets for testing new algorithms.
Tools for nanotechnology-oriented simulation and modeling.
Communication services between users, including the development of tools for knowledge and resources exchange among researchers working in the area: This brokerage system would allow users to register their own nanomaterials and tools and provide an inventory for use by others in acquiring the desired data by means of a syntactically and semantically matching process, by entering words or phrases. This is a flexible novel paradigm already applied in biomedicine, which successfully integrates knowledge obtained from databases using Semantic Web technologies, sharing a common ontology and that can be accessed through normalized URLs.
Documentation repositories providing researchers with intelligent search tools, as well as services for version control.
Resources and tools for management of recent contents, alerts and news published in the area.
Learning tools and consultation forums.
Finally, this hypothetical cyber-infrastructure should also support different views for different type of users, for instance: i) small data users versus massive data users; ii) novice users versus expert users; iii) researcher’s view versus patient’s view versus administration’s view versus clinician’s view and iv) basic research versus translational research.
5. Conclusions
As discussed above, nanomedical research is producing huge volumes of information due to the numerous advances in the nanotechnology field. Due to the fact that the nature of this information is heterogeneous and has different levels of granularity, researches are facing many challenges in analyzing all these data. Consequently, the use of computational methods for the design and development of innovative analysis tools has progressively gained higher relevance, requiring new publicly accessible resources that facilitate the cooperative work between research groups working in the field.
In this paper, we have analyzed the urgent need to use automatic approaches to efficiently manage the new data generated during nanomedical research. This knowledge must be organized and structured, by means of the creation of standard domain models, and annotated by labeling data using existing terminologies, vocabularies and ontologies. The constant publication of new results in the area also demands a continuous update of data resources, which cannot be done manually, thus resources providing new methods for automatically updating these resources are crucial.
One of the main challenges in nanoinformatics is the variety and heterogeneity of the nanomedical domain, where users have very different needs, depending on their training and professional role. In order to understand the current problems and challenges and to develop the required applications to address them, nanoinformatics researchers should have a fluid communication with the scientific community conducting research in the area. This research involves professionals with varied profiles working in very different areas: chemistry, physics, medicine, the pharmaceutical industry, among others (including the materials science and engineering community), working closely with technology developers and manufacturers. It is necessary to consider social factors, as well, since they affect the acceptance of nanomedicine by consumers and its impact upon them in a global way. With this in mind, a great effort should go into undertaking and analyzing nanotoxicity studies, taking into account the results of previous studies, and the results, both positive and negative, managed as a complete information set and made easily available. Computational methods for modeling and simulation in nanomedicine are vital for understanding the behaviour and interactions of engineered nanoparticles, as well as their biological effects within the human body and the environment.
Finally, due to the large number of new nanomaterials that are being produced by manufacturing processes, an open approach is necessary with regards to the publication of new research data. A major portion of the information about nanoparticles studies, assays and clinical trials can be found in the scientific literature, but there is a lack of publicly available databases and repositories of structured information. In order to accelerate advancements and improvements in nanomedicine and nanoinformatics, easy availability of experimental and clinical data is essential, allowing for inference of information from data and the subsequent integration of research results. Thereby, current informatics methods and techniques for text mining and natural language processing can help to characterize and classify nanomaterials, as well as identify data correlations from current research results.
By adopting such a big-picture approach, we can foster a community of integrated and coordinated research groups, facilitating the creation and support of interdisciplinary teams. Scientific groups collaborating through a nano-platform or cyber-infrastructure should include basic and translational scientists, clinical researchers and computer scientists. This kind of platform would also promote the establishment of public–private partnerships across the different areas involved, leading to long-term sustainability of funded research programs. The definitive establishment of nanotechnology in medicine and industry depends on all of these factors where, as we have seen, nanoinformatics is critical to improving the attainment of research results.
Acknowledgments
The work of the authors in this area of research has been partially funded by the European Commission (the ACTION-Grid Support Action, FP7-224176), the Spanish Ministry of Economy and Competitiveness (FIS/AES PS09/00069, RETICS COMBIOMED RD07/0067/0006, Ibero-NBIC CYTED 209RT0366) and the Consejo Social of the Universidad Politécnica de Madrid.
This work has also been funded in part with funds from the NCI-NIH (Contract No. HHSN261200800001E).
Footnotes
CarriGen®, porous bone substitute material composed of a proprietary nanocrystalline calcium phosphate formulation that mimics the crystalline mineral structure of human bone. ETEX products. https://secure.etexcorp.com/products_overview.html.
The contents of this publication do not necessarily reflect the views or policies of the DHHS, nor does mention of trade names, commercial products, or organizations imply endorsement by the U.S. Government.
References
- Bailey RC, Kwong GA, Radu CG, Witte ON, Heath JR. DNA-encoded antibody libraries: a unified platform for multiplexed cell sorting and detection of genes and proteins. J Am Chem Soc. 2007;129:1959–67. doi: 10.1021/ja065930i. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bairoch A, Apweiler R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 2000;28:45–8. doi: 10.1093/nar/28.1.45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bairoch A, Apweiler R, Wu CH, Barker WC, Boeckmann B, Ferro S, Gasteiger E, Huang H, Lopez R, Magrane M, Martin MJ, Natale DA, O’Donovan C, Redaschi N, Yeh LS. The Universal Protein Resource (UniProt) Nucleic Acids Res. 2005;33:D154–9. doi: 10.1093/nar/gki070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker NA, Fritts M, Guccione S, Paik DS, Pappu RV, Patri A, Rubin D, Shaw SY, Thomas DG. Nanotechnology Informatics White Paper. National Cancer Institute; 2009. caBIG. February 2009. Non-peer Reviewed Manuscript. [Google Scholar]
- Bellare JR. Nanotechnology and nanomedicine for healthcare: challenges in translating innovations from bench to bedside. J Biomed Nanotechnol. 2011;7:36–7. doi: 10.1166/jbn.2011.1189. [DOI] [PubMed] [Google Scholar]
- Benson DA, Karsch-Mizrachi I, Clark K, Lipman DJ, Ostell J, Sayers EW. GenBank. Nucleic Acids Res. 2012;40:D48–53. doi: 10.1093/nar/gkr1202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berman HM, Henrick K, Nakamura H. Announcing the worldwide Protein Data Bank. Nature Structural Biology. 2003;10:980. doi: 10.1038/nsb1203-980. [DOI] [PubMed] [Google Scholar]
- Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE. The Protein Data Bank. Nucleic Acids Res. 2000;28:235–42. doi: 10.1093/nar/28.1.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bikiaris D, Koutris E, Karavas E. New aspects in sustained drug release formulations. Recent Pat Drug Deliv Formul. 2007;1(3):201–13. doi: 10.2174/187221107782331629. [DOI] [PubMed] [Google Scholar]
- Braun T, Schubert A, Zsindely S. Nanoscience and nanotechnology on the balance. Scientometrics. 1997;38:321–325. [Google Scholar]
- Buxton DB. Current status of nanotechnology approaches for cardiovascular disease: a personal perspective. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2009;1:149–55. doi: 10.1002/wnan.8. [DOI] [PubMed] [Google Scholar]
- de la Zerda A, Bodapati S, Teed R, May SY, Tabakman SM, Liu Z, Khuri-Yakub BT, Chen X, Dai H, Gambhir SS. Family of Enhanced Photoacoustic Imaging Agents for High-Sensitivity and Multiplexing Studies in Living Mice. ACS Nano. 2012;6:4694–701. doi: 10.1021/nn204352r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Mel A, Chaloupka K, Malam Y, Darbyshire A, Cousins B, Seifalian AM. A silver nanocomposite biomaterial for blood-contacting implants. J Biomed Mater Res A. 2012;100:2348–57. doi: 10.1002/jbm.a.34177. [DOI] [PubMed] [Google Scholar]
- de la Iglesia D, Chiesa S, Kern J, Maojo V, Martin-Sanchez F, Potamias G, Moustakis V, Mitchell JA. Nanoinformatics: new challenges for biomedical informatics at the nano level. Stud Health Technol Inform. 2009;150:987–91. [PubMed] [Google Scholar]
- de la Iglesia D, Maojo V, Chiesa S, Martin-Sanchez F, Kern J, Potamias G, Crespo J, Garcia-Remesal M, Keuchkerian S, Kulikowski C, Mitchell JA. International efforts in nanoinformatics research applied to nanomedicine. Methods Inf Med. 2011;50:84–95. doi: 10.3414/ME10-02-0012. [DOI] [PubMed] [Google Scholar]
- Dinov ID, et al. iTools: a framework for classification, categorization and integration of computational biology resources. PLoS One. 2008;3:e2265. doi: 10.1371/journal.pone.0002265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duncan R, Gaspar R. Nanomedicine(s) under the microscope. Mol Pharm. 2011;8:2101–41. doi: 10.1021/mp200394t. [DOI] [PubMed] [Google Scholar]
- Etheridge ML, Campbell SA, Erdman AG, Haynes CL, Wolf SM, McCullough J. The big picture on nanomedicine: the state of investigational and approved nanomedicine products. Nanomedicine. 2013;9:1–14. doi: 10.1016/j.nano.2012.05.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fathabadi EG, Shelling AN, Al-Kassas R. Nanocarrier systems for delivery of siRNA to ovarian cancer tissues. Expert Opin Drug Deliv. 2012;9:743–54. doi: 10.1517/17425247.2012.683173. [DOI] [PubMed] [Google Scholar]
- Freitas RA., Jr . Nanomedicine, Volume I: Basic Capabilities. Georgetown, TX: Landes Bioscience; 1999. Available at: http://www.nanomedicine.com/NMI/Glossary.htm. [Google Scholar]
- Freitas RA., Jr What is nanomedicine? Nanomedicine. 2005;1:2–9. doi: 10.1016/j.nano.2004.11.003. [DOI] [PubMed] [Google Scholar]
- García-Remesal M, Cuevas A, Pérez-Rey D, Martín L, Anguita A, de la Iglesia D, de la Calle G, Crespo J, Maojo V. PubDNA Finder: a web database linking full-text articles to sequences of nucleic acids. Bioinformatics. 2010;26:2801–2. doi: 10.1093/bioinformatics/btq520. [DOI] [PubMed] [Google Scholar]
- García-Remesal M, García-Ruiz A, Pérez-Rey D, de la Iglesia D, Maojo V. Using nanoinformatics methods for automatically identifying relevant Nanotoxicology entities from the literature. Biomed Res Int. 2013;2013:410294. doi: 10.1155/2013/410294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghanbari H, Cousins BG, Seifalian AM. A nanocage for nanomedicine: polyhedral oligomeric silsesquioxane (POSS) Macromol Rapid Commun. 2011;32:1032–46. doi: 10.1002/marc.201100126. [DOI] [PubMed] [Google Scholar]
- Giam LR, Massich MD, Hao L, Shin Wong L, Mader CC, Mirkin CA. Scanning probe-enabled nanocombinatorics define the relationship between fibronectin feature size and stem cell fate. Proc Natl Acad Sci U S A. 2012;109:4377–82. doi: 10.1073/pnas.1201086109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greenberg A. Integrating nanoscience into the classroom: perspectives on nanoscience education projects. ACS Nano. 2009;3:762–9. doi: 10.1021/nn900335r. [DOI] [PubMed] [Google Scholar]
- Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA. Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 2005;33:D514–7. doi: 10.1093/nar/gki033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hatto P. International standards for risk management in nanotechnology. Nat Nanotechnol. 2009;4:205. doi: 10.1038/nnano.2009.25. [DOI] [PubMed] [Google Scholar]
- Haun JB, Yoon TJ, Lee H, Weissleder R. Molecular detection of biomarkers and cells using magnetic nanoparticles and diagnostic magnetic resonance. Methods Mol Biol. 2011;726:33–49. doi: 10.1007/978-1-61779-052-2_3. [DOI] [PubMed] [Google Scholar]
- Huang Z, Chen H, Yip A, Ng G, Guo F, Chen ZK, Roco MC. Longitudinal patent analysis for nanoscale science and engineering: Country, institution and technology field. Journal of Nanoparticle Research. 2003;5:333–363. [Google Scholar]
- Huh AJ, Kwon YJ. “Nanoantibiotics”: a new paradigm for treating infectious diseases using nanomaterials in the antibiotics resistant era. J Control Release. 2011;156:128–45. doi: 10.1016/j.jconrel.2011.07.002. [DOI] [PubMed] [Google Scholar]
- Hullman A, Meyer M. Publications and patents in nanotechnology: An overview of previous studies and the state of the art. Scientometrics. 2003;58:507–527. [Google Scholar]
- Jain KK. Nanobiotechnology and personalized medicine. Prog Mol Biol Transl Sci. 2011;104:325–54. doi: 10.1016/B978-0-12-416020-0.00008-5. [DOI] [PubMed] [Google Scholar]
- Jin S, Leach JC, Ye K. Nanoparticle-mediated gene delivery. Methods Mol Biol. 2009;544:547–57. doi: 10.1007/978-1-59745-483-4_34. [DOI] [PubMed] [Google Scholar]
- Kamphuis MM, Johnston AP, Such GK, Dam HH, Evans RA, Scott AM, Nice EC, Heath JK, Caruso F. Targeting of cancer cells using click-functionalized polymer capsules. J Am Chem Soc. 2010;132:15881–3. doi: 10.1021/ja106405c. [DOI] [PubMed] [Google Scholar]
- Karalis V, Macheras P. Current regulatory approaches of bioequivalence testing. Expert Opin Drug Metab Toxicol. 2012;8:929–42. doi: 10.1517/17425255.2012.690394. [DOI] [PubMed] [Google Scholar]
- Kelkar SS, Reineke TM. Theranostics: combining imaging and therapy. Bioconjug Chem. 2011;22(10):1879–903. doi: 10.1021/bc200151q. [DOI] [PubMed] [Google Scholar]
- Kiessling F, Fokong S, Koczera P, Lederle W, Lammers T. Ultrasound microbubbles for molecular diagnosis, therapy, and theranostics. J Nucl Med. 2012;53:345–8. doi: 10.2967/jnumed.111.099754. [DOI] [PubMed] [Google Scholar]
- Kircher MF, de la Zerda A, Jokerst JV, Zavaleta CL, Kempen PJ, Mittra E, Pitter K, Huang R, Campos C, Habte F, Sinclair R, Brennan CW, Mellinghoff IK, Holland EC, Gambhir SS. A brain tumor molecular imaging strategy using a new triple-modality MRI-photoacoustic-Raman nanoparticle. Nat Med. 2012;18:829–34. doi: 10.1038/nm.2721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koria P. Delivery of growth factors for tissue regeneration and wound healing. BioDrugs. 2012;26:163–75. doi: 10.2165/11631850-000000000-00000. [DOI] [PubMed] [Google Scholar]
- Lee LL, Chan CK, Ngaim M, Ramakrishna S. Nanotechnology patent landscape 2006. Nano. 2006;1:101–113. [Google Scholar]
- Lemley MA. Patenting nanotechnology. Stanford Law Review. 2005;61:601–630. [PubMed] [Google Scholar]
- Li Y, Zheng X, Cao Z, Xu W, Zhang J, Gong M. Self-assembled peptide (CADY-1) improved the clinical application of doxorubicin. Int J Pharm. 2012;434:209–14. doi: 10.1016/j.ijpharm.2012.06.003. [DOI] [PubMed] [Google Scholar]
- Liu S, Guo Y, Huang R, Li J, Huang S, Kuang Y, Han L, Jiang C. Gene and doxorubicin co-delivery system for targeting therapy of glioma. Biomaterials. 2012;33:4907–16. doi: 10.1016/j.biomaterials.2012.03.031. [DOI] [PubMed] [Google Scholar]
- Liu R, Zhang HY, Ji ZX, Rallo R, Xia T, Chang CH, Nel A, Cohen Y. Development of Structure-Activity Relationship for Metal Oxide Nanoparticles. Nanoscale. 2013 doi: 10.1039/C3NR01533E. Accepted Manuscript 3 May 2013. [DOI] [PubMed] [Google Scholar]
- Lowe HJ, Barnett GO. Understanding and using the medical subject headings (MeSH) vocabulary to perform literature searches. JAMA. 1994;271:1103–8. [PubMed] [Google Scholar]
- Lynch I, Salvati A, Dawson KA. Protein-nanoparticle interactions: What does the cell see? Nat Nanotechnol. 2009;4:546–7. doi: 10.1038/nnano.2009.248. [DOI] [PubMed] [Google Scholar]
- Mailänder V, Landfester K. Interaction of nanoparticles with cells. Biomacromolecules. 2009;10:2379–400. doi: 10.1021/bm900266r. [DOI] [PubMed] [Google Scholar]
- Majzoub EH, Rönnebro ECE. Methodology of materials discovery in complex metal hydrides using experimental and computational tools. Materials Science and Engineering Reports. 2012;73:15–26. [Google Scholar]
- Maojo V, Crespo J, García-Remesal M, de la Iglesia D, Perez-Rey D, Kulikowski C. Biomedical ontologies: toward scientific debate. Methods Inf Med. 2011;50:203–16. doi: 10.3414/ME10-05-0004. [DOI] [PubMed] [Google Scholar]
- Maojo V, et al. The ACTION-Grid White Paper: Linking Biomedical Informatics, Grid Computing and Nanomedicine. 2010 Available at: http://www.action-grid.eu/index.php?url=whitepaper. Non-peer Reviewed Manuscript.
- Maojo V, Fritts M, de la Iglesia D, Cachau RE, Garcia-Remesal M, Mitchell JA, Kulikowski C. Nanoinformatics: a new area of research in nanomedicine. Int J Nanomedicine. 2012a;7:3867–90. doi: 10.2147/IJN.S24582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maojo V, et al. Nanoinformatics: developing new computing applications for nanomedicine. Comput Sci Eng. 2012b;94:521–539. doi: 10.1007/s00607-012-0191-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martens L, Hermjakob H, Jones P, Adamski M, Taylor C, States D, Gevaert K, Vandekerckhove J, Apweiler R. PRIDE: the proteomics identifications database. Proteomics. 2005;5:3537–45. doi: 10.1002/pmic.200401303. [DOI] [PubMed] [Google Scholar]
- McNeil SE. Characterization of nanoparticles intended for drug delivery. In: McNeil SE, editor. Methods in Molecular Biology. Vol. 697. New York: Humana Press, Springer Science+Business Media; 2011. pp. 3–8. [Google Scholar]
- Mentovich ED, Livanov K, Prusty D, Sowwan M, Richter S. DNA-nanoparticle assemblies go organic: Macroscopic polymeric materials with nanosized features. J Nanobiotechnology. 2012;10:21. doi: 10.1186/1477-3155-10-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer M. Patent citations in a novel field of technology: what can they tell about Interactions of emerging communities of science and technology? Scientometrics. 2000;48:151–178. [Google Scholar]
- Meyer M. Patent citation analysis in a novel field of technology: an exploration of nano-science and nano-technology. Scientometrics. 2001;51:163–183. [Google Scholar]
- Moosavi H, Zeynali M, Pour ZH. Fracture resistance of premolars restored by various types and placement techniques of resin composites. Int J Dent. 2012;2012:973641. doi: 10.1155/2012/973641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murashov V, Howard J. The US must help set international standards for nanotechnology. Nat Nanotechnol. 2008;3:635–6. doi: 10.1038/nnano.2008.323. [DOI] [PubMed] [Google Scholar]
- Murashov V, Howard J. Nanotechnology Standards. In: Murashov V, Howard J, editors. Nanostructure science and technology. New York: Springer Science+Business Media; 2011. pp. 1–20. [Google Scholar]
- Musen MA, Shah NH, Noy NF, Dai BY, Dorf M, Griffith N, Buntrok J, Jonquet C, Montegut MJ, Rubin DL. BioPortal: Ontologies and Data Resources with the Click of a Mouse. AMIA Annu Symp Proc. 2008;6:1223–1224. [PubMed] [Google Scholar]
- National Science and Technology Council. Materials Genome Initiative for Global Competitiveness. White Paper June 2011. 2011 Available at: http://www.whitehouse.gov/sites/default/files/microsites/ostp/materials_genome_initiative-final.pdf.
- National Science and Technology Council. A National Strategic Plan for Advanced Manufacturing, February 2012. 2012 Available at: http://www.whitehouse.gov/sites/default/files/microsites/ostp/iam_advancedmanufacturing_strategicplan_2012.pdf.
- Noy NF, Shah NH, Whetzel PL, Dai B, Dorf M, Griffith N, Jonquet C, Rubin DL, Storey MA, Chute CG, Musen MA. BioPortal: ontologies and integrated data resources at the click of a mouse. Nucleic Acids Res. 2009;37:W170–3. doi: 10.1093/nar/gkp440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ostraat ML, Mills K, Murry D, Guzan K. Nanomaterial Registry: An authoritative resource for assessing environmental and biological interactions of nanomaterials. Tech Connect WORLD: Summit and Innovation Showcase 2012; Santa Clara, CA. 18–21 June 2012.2012. [Google Scholar]
- Parrott MC, Finniss M, Luft JC, Pandya A, Gullapalli A, Napier ME, Desimone JM. Incorporation and Controlled Release of Silyl Ether Prodrugs from PRINT Nanoparticles. J Am Chem Soc. 2012;134:7978–82. doi: 10.1021/ja301710z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pearce TR, Shroff K, Kokkoli E. Peptide targeted lipid nanoparticles for anticancer drug delivery. Adv Mater. 2012;24:3803–22. 3710. doi: 10.1002/adma.201200832. [DOI] [PubMed] [Google Scholar]
- Perán M, García MA, López-Ruiz E, Bustamante M, Jiménez G, Madeddu R, Marchal JA. Functionalized nanostructures with application in regenerative medicine. Int J Mol Sci. 2012;13:3847–86. doi: 10.3390/ijms13033847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perry JL, Herlihy KP, Napier ME, Desimone JM. PRINT: a novel platform toward shape and size specific nanoparticle theranostics. Acc Chem Res. 2011;44:990–8. doi: 10.1021/ar2000315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petre CE, Dittmer DP. Liposomal daunorubicin as treatment for Kaposi’s sarcoma. Int J Nanomedicine. 2007;2:277–88. [PMC free article] [PubMed] [Google Scholar]
- Ptak K, Farrell D, Panaro NJ, Grodzinski P, Barker AD. The NCI Alliance for Nanotechnology in Cancer: achievement and path forward. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2010;2:450–60. doi: 10.1002/wnan.98. [DOI] [PubMed] [Google Scholar]
- Puzyn T, Rasulev B, Gajewicz A, Hu X, Dasari TP, Michalkova A, Hwang HM, Toropov A, Leszczynska D, Leszczynski J. Using nano-QSAR to predict the cytotoxicity of metal oxide nanoparticles. Nat Nanotechnol. 2011;6:175–8. doi: 10.1038/nnano.2011.10. [DOI] [PubMed] [Google Scholar]
- Qiao W, McLennan M, Kennell R, Ebert DS, Klimeck G. Hub-based simulation and graphics hardware accelerated visualization for nanotechnology applications. IEEE Trans Vis Comput Graph. 2006;12:1061–8. doi: 10.1109/TVCG.2006.150. [DOI] [PubMed] [Google Scholar]
- Roco MC. Nanotechnology: convergence with modern biology and medicine. Curr Opin Biotechnol. 2003;14:337–46. doi: 10.1016/s0958-1669(03)00068-5. [DOI] [PubMed] [Google Scholar]
- Rutherford K, Parkhill J, Crook J, Horsnell T, Rice P, Rajandream MA, Barrell B. Artemis: sequence visualization and annotation. Bioinformatics. 2000;16:944–5. doi: 10.1093/bioinformatics/16.10.944. [DOI] [PubMed] [Google Scholar]
- Sato K, Yokosuka S, Takigami Y, Hirakuri K, Fujioka K, Manome Y, Sukegawa H, Iwai H, Fukata N. Size-tunable silicon/iron oxide hybrid nanoparticles with fluorescence, superparamagnetism, and biocompatibility. J Am Chem Soc. 2011;133:18626–33. doi: 10.1021/ja202466m. [DOI] [PubMed] [Google Scholar]
- Scholes GD, Rumbles G. Excitons in nanoscale systems. Nat Mater. 2006;5:683–96. doi: 10.1038/nmat1710. [DOI] [PubMed] [Google Scholar]
- Service RF. Materials and biology. Nanotechnology takes aim at cancer. Science. 2005;310:1132–4. doi: 10.1126/science.310.5751.1132. [DOI] [PubMed] [Google Scholar]
- Service RF. Nanotechnology. Nanoparticle Trojan horses gallop from the lab into the clinic. Science. 2010;330:314–5. doi: 10.1126/science.330.6002.314. [DOI] [PubMed] [Google Scholar]
- Shimoni O, Postma A, Yan Y, Scott AM, Heath JK, Nice EC, Zelikin AN, Caruso F. Macromolecule functionalization of disulfide-bonded polymer hydrogel capsules and cancer cell targeting. ACS Nano. 2012;6:1463–72. doi: 10.1021/nn204319b. [DOI] [PubMed] [Google Scholar]
- Shukla GC, Haque F, Tor Y, Wilhelmsson LM, Toulmé JJ, Isambert H, Guo P, Rossi JJ, Tenenbaum SA, Shapiro BA. A boost for the emerging field of RNA nanotechnology. ACS Nano. 2011;5:3405–18. doi: 10.1021/nn200989r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sioutos N, de Coronado S, Haber MW, Hartel FW, Shaiu WL, Wright LW. NCI Thesaurus: a semantic model integrating cancer-related clinical and molecular information. J Biomed Inform. 2007;40:30–43. doi: 10.1016/j.jbi.2006.02.013. [DOI] [PubMed] [Google Scholar]
- Tanaka M. CAIS/ACSI 2005: Data, Information, and Knowledge in a Networked World. The University of Western Ontario; London, Ontario: 2005. Toward a Proposed Ontology for Nanoscience. 2005. [Google Scholar]
- Thaxton CS, Elghanian R, Thomas AD, Stoeva SI, Lee JS, Smith ND, Schaeffer AJ, Klocker H, Horninger W, Bartsch G, Mirkin CA. Nanoparticle-based bio-barcode assay redefines “undetectable” PSA and biochemical recurrence after radical prostatectomy. Proc Natl Acad Sci U S A. 2009;106:18437–42. doi: 10.1073/pnas.0904719106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas AR. Bibliographical classification: the ideas and achievements of Henry E. Bliss. Cataloging & Classification Quarterly. 1997;25:51–104. [Google Scholar]
- Thomas DG, Pappu RV, Baker NA. NanoParticle Ontology for cancer nanotechnology research. J Biomed Inform. 2011;44:59–74. doi: 10.1016/j.jbi.2010.03.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Upadhyay D, Scalia S, Vogel R, Wheate N, Salama RO, Young PM, Traini D, Chrzanowski W. Magnetised thermo responsive lipid vehicles for targeted and controlled lung drug delivery. Pharm Res. 2012;29:2456–67. doi: 10.1007/s11095-012-0774-9. [DOI] [PubMed] [Google Scholar]
- Veetil JV, Ye K. Tailored carbon nanotubes for tissue engineering applications. Biotechnol Prog. 2009;25:709–21. doi: 10.1002/bp.165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Byrne JD, Napier ME, DeSimone JM. Engineering nanomedicines using stimuli-responsive biomaterials. Adv Drug Deliv Rev. 2012;64:1021–30. doi: 10.1016/j.addr.2012.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weidl T, Klein G, Zöllner R. In: International Handbook on Regulating Nanotechnologies. Hodge GA, Bowman DM, Maynar AD, editors. Cheltenham, UK: Edward Elgar Publishing, Inc; 2010. pp. 462–83. [Google Scholar]
- Wishart DS, et al. HMDB: the Human Metabolome Database. Nucleic Acids Res. 2007;35:D521–6. doi: 10.1093/nar/gkl923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong PK, Ho YP, Chan CK. Internationalization and evolution of application areas of an emerging technology: The case of nanotechnology. Scientometrics. 2007;70:715–737. [Google Scholar]
- Xenarios I, Salwínski L, Duan XJ, Higney P, Kim SM, Eisenberg D. DIP, the Database of Interacting Proteins: a research tool for studying cellular networks of protein interactions. Nucleic Acids Res. 2002;30:303–5. doi: 10.1093/nar/30.1.303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xin G, Zhao X, Duan X, Ning Q, Meng M, Meng D, Liu L. Antitumor effect of a generation 4 polyamidoamine dendrimer/cyclooxygenase-2 antisense oligodeoxynucleotide complex on breast cancer in vitro and in vivo. Cancer Biother Radiopharm. 2012;27:77–87. doi: 10.1089/cbr.2011.1028. [DOI] [PubMed] [Google Scholar]
- Yeri A, Gao D. Biosensing Using Nanoelectromechanical Systems. In: Hurst SJ, editor. Biomedical Nanotechnology Methods and Protocols, Methods in Molecular Biology. Vol. 726. Hatfield, Hertfordshire, UK: Humana Press, Springer Science+Business Media; 2011. pp. 119–140. [Google Scholar]
- Zuckerman JE, Choi CH, Han H, Davis ME. Polycation-siRNA nanoparticles can disassemble at the kidney glomerular basement membrane. Proc Natl Acad Sci USA. 2012;109:3137–42. doi: 10.1073/pnas.1200718109. [DOI] [PMC free article] [PubMed] [Google Scholar]