Abstract
The Killer-cell Immunoglobulin-like Receptor (KIR) gene complex has considerable biomedical importance. Patterns of polymorphism in the KIR region include variability in the gene content of haplotypes and diverse structural arrangements. Droplet digital PCR (ddPCR) was used to identify different haplotype motifs and to enumerate KIR copy number variants (CNVs). ddPCR detected a variety of KIR haplotype configurations in DNA from well-characterized cell lines. Mendelian segregation of ddPCR-estimated KIR2DL5 CNVs was observed in Gambian families and CNV typing of other KIRs was shown to be accurate when compared to an established quantitative PCR method.
Background
A highly polymorphic gene complex on chromosome 19q13.4 encodes the Killer-cell Immunoglobulin-like Receptors (KIRs) [1]. As natural killer (NK) cell receptors, their roles in modulating outcomes of infectious diseases [2,3], transplantation [4] and pregnancy [5] are subjects of intense research, but their genotyping and subsequent use in association studies and histocompatibility matching are made difficult by the underlying structural and sequence complexity in the KIR region. Both haplotype structural variations [6] and KIR gene copy number variations (CNVs) [7,8] contribute to the diversity of the KIR system. The highly homologous KIR genes are arranged closely head-to-tail, an arrangement that facilitates non-reciprocal recombination (misalignment and unequal crossovers during meiosis), which can delete, duplicate or recombine genes [9] (Figure 1). The implications of this are that KIR genes may be present or absent from a haplotype, that those KIR genes that are present on a haplotype may occur multiple times, and that the structural arrangement of the KIR genes may vary between individuals with otherwise identical gene contents and/or gene copy numbers.
Figure 1.
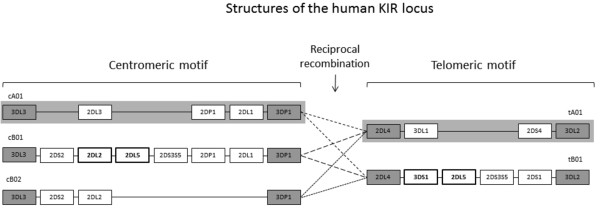
Organization of the KIR locus on the human chromosome region 19q13.4. KIR haplotypes are composed of different centromeric (cA01, cB01 and cB02) and telomeric motifs (tA01, tB01) [10]. Each motif has a different content and arrangement of genes. Framework genes, which are found at the ends and near the middle of the locus on nearly all haplotypes [11] are shaded grey. The KIR2DL5 gene can be located in either the centromeric or telomeric motif, adjacent to KIR2DL2 and KIR3DS1, respectively, or on both motifs of a single haplotype. Different centromeric motifs can be paired with different telomeric motifs through the central reciprocal recombination hotspot between KIR3DP1 and KIR2DL4, as indicated by different dashed lines. KIR haplotypes can be classified into two categories [12]: group A haplotypes (shaded grey), composed of cA01 and tA01 motifs only, have fixed gene-content with one activating gene (KIR2DS4); group B haplotypes (unshaded), comprising at least one motif of type cB01, cB02 or tB01, have variable gene content between framework genes and more than one activating KIR locus.
The KIR complex is divisible into two variable motifs that are defined by their orientation towards the centromeric (Cen) or telomeric (Tel) regions of the chromosome [13]. KIR2DL5 was the first KIR gene that was widely recognized as segregating to more than one locus [14,15] and is common in most human populations. At the KIR2DL5C locus, KIR2DL5 is often (though not exclusively) found to be adjacent to KIR2DL2[13]. At the KIR2DL5T locus, KIR2DL5 is frequently found to be adjacent to KIR3DS1[13]. In some haplotypes, KIR2DL5 is present, whilst both KIR2DL2 and KIR3DS1 are absent [16]. An individual haplotype might have zero (Figure 1, cA01 ~ tA01 conformation), one (Figure 1, cB01 ~ tA01, cA01 ~ tB01 and cB02 ~ tB01 conformations), or two (Figure 1, cB01 ~ tB01 conformation) copies of KIR2DL5 (EMBL accession AY320039; Gassner, C., Williams, L.M., Yamashita, T., Selvakumar, A., Dupont, B. and Geraghty, D.E., unpublished data) and might possess either, both, or neither of the KIR2DL2 ~ KIR2DL5C and KIR3DS1 ~ KIR2DL5T gene arrangements (Figure 1). Consequently, an individual might have up to four copies of KIR2DL5.
Detailed characterization of KIR haplotypes is possible using a combination of pedigree analyses, high-resolution molecular typing and likelihood estimations [17,18]. These approaches are costly, not easily scalable to high throughputs and are frequently not completely accurate. All current methods rely, at least in part, on the specification a priori of known haplotype structures. These haplotype maps are derived from a small number of fully sequenced KIR haplotypes [13,17,19] and also from observed patterns of perfect linkage disequilibrium between key pairs of KIR genes [10,15,18,20,21]. These assumptions appear to hold true in most Caucasian populations [10,17,20-22] but may be invalid in the wider global population, especially in African populations [23]. Empirical approaches to the definition of haplotype structural diversity in global populations are required, but options are currently limited to pedigree analysis [20,21] and sequencing of single chromosomes [11,24,25]. Molecular haplotyping is possible in single-molecule processes, including digital PCR [26-30], and recent reports have highlighted how CNVs can be enumerated using droplet digital PCR (ddPCR) [31,32].
In this application of digital PCR, a limiting number of DNA target molecules are stochastically confined by a microfluidic device [33] into a large number of droplet PCR nano-reactors (volume 10-9 L) that contain either zero or one copy of the PCR target. The ddPCR reaction may be a duplex test that simultaneously detects two targets using fluorescent probes. After PCR is complete, the droplets are passed in single-file through a flow cytometric device, which determines the qualitative end-points of PCR by assaying the presence or absence of hydrolysis probe-derived fluorescence signals. Counts of PCR-positive and PCR-negative droplets are made and these are converted into an accurate measure of the number of target entities (copies/volume) in the total PCR volume without the need to refer to calibration curves or reference samples [34-36].
When there are two ddPCR targets that are not physically linked (either because they originate on different chromosomes or if intra-chromosomal linkage has been cut or sheared during or subsequent to DNA extraction), then two independent stochastic DNA confinement processes occur and these may overlap. In the absence of linkage, the result is that a droplet may contain zero, one, or both targets. In the presence of linkage, the confinement processes are not independent and the frequency of 'double positive' droplets is substantially higher than is observed in the absence of linkage. When one of the targets is an unlinked gene of invariant copy number, then the ratio (corrected for diploidy) between the gene of interest (for example, a KIR gene) and the invariant gene is a direct measure of the gene of interest copy number. In this proof of principle study, we show that ddPCR can be used to perform molecular haplotype analysis and CNV enumeration in the KIR system.
Methods
Droplet digital PCR
ddPCR was carried out using the QX100 Droplet Digital PCR system (Bio-Rad Laboratories, Hemel Hempstead, UK). ddPCR reactions were 22 μl aqueous volumes that contained final concentrations of 1X ddPCR supermix (Bio-Rad), 0.3 μM each primer and probe and 10 to 30 ng of genomic DNA. Droplet generation and droplet reading for ddPCR were carried out according to the manufacturer’s instructions using Bio-Rad reagents. The thermal cycling profile was 95°C:10′00′′ followed by 43 cycles of (95°C:0′15′′|50°C:1′00′′).
KIR2DL5 linkage analysis
Ten specimens of DNA derived from International Histocompatibility Workshop (IHW) cell lines were selected because standard quantitative PCR (qPCR) had indicated that they possessed at least one copy of KIR2DL5 and either or both of KIR2DL2 and KIR3DS1. DNA was extracted using the QIAamp DNA Blood Midi/Maxi kit (QIAGEN, Manchester, UK), which prepares DNA fragments up to 50 kb in length (QIAamp Blood Midi/Maxi kit handbook, 04/2010, QIAGEN). The presence of fragments longer than 23 kb was confirmed by gel electrophoresis and direct comparison to λHindIII/φX174HaeIII molecular weight markers. Two separate ddPCR assays were performed. Each of these was designed to detect one of the haplotypes KIR2DL2 ~ KIR2DL5C (approximately 21.8 kb) or KIR3DS1 ~ KIR2DL5T (approximately 19.9 kb). Sequences of the KIR-specific oligonucleotides (Table S1 in Additional file 1) are identical to those used previously [10]. PCR probes were labeled with either 6-carboxyfluorescein (FAM; excitation 492 nm, emission 517 nm) or 6-carboxy-2,4,4,5,7,7-hexachlorofluorescein succinimidyl ester (HEX; excitation 535 nm, emission 553 nm). The probes hybridized to KIR2DL2 exon 4, KIR3DS1 exon 4 and KIR2DL5 exon 9. Specimens were tested both before and after being subjected to restriction endonuclease digestion with 5U EcoRI (New England Biosciences, Hitchin, UK).
ddPCR data were collected and analyzed with QuantaLife® software V2.0 (BioRad Laboratories). Crosshair gating was used to split the data into four quadrants in a procedure analogous to that applied widely in flow cytometry analysis. Double negative (FAM-HEX-) droplets contained neither KIR gene target. HEX-positive droplets (FAM-HEX+) contained only KIR2DL5. FAM-positive droplets (FAM+HEX-) contained either KIR2DL2 or KIR3DS1 (depending on the assay) and double positive (FAM+HEX+) droplets contained both KIR2DL5 and either of KIR2DL2 or KIR3DS1. The primary measure of linkage between the two genes in the assay is the 'linkage' score, which is automatically calculated by the QuantaLife® software. The linkage score reports the estimated total number of molecules (copies/μl) in the assay that carry both target genes in linkage. Dividing this number by the total concentration of KIR3DS1 or KIR2DL2 determines the percentage of those genes in the test aliquot that are physically linked to KIR2DL5. This normalizes the linkage score for differences in the input amount of DNA between assays and we refer herein to %L as the normalized linkage score. After restriction endonuclease digestion, %L should be significantly decreased.
KIR2DL5 CNV analysis
Seven genomic DNA samples came from the UCLA international KIR exchange programme [37] and these were selected because they were recently extracted, high molecular weight DNA preparations from bulk cultures. Aliquots of the KIR exchange DNAs are also widely available in the KIR research community. Twenty-seven DNA samples were archival specimens that came from five Gambian nuclear families that were collected as part of another study (Medical Research Council Unit, The Gambia, study number SCC1177v2). The Ethics Committee of the Gambian Government/Medical Research Council Unit, and the ethics committee of the London School of Hygiene and Tropical Medicine approved the collection and genotyping of the Gambian samples. Individual written informed consent was obtained from all adult participants. Consent was obtained from a parent/guardian on behalf of those subjects aged <18 years who gave assent. All samples were anonymized. The Gambian specimens were collected in accordance with the tenets of the declaration of Helsinki.
An internal reference gene, the invariant single copy per haplotype ribonuclease P/MRP 30 kDa subunit (RPP30) gene, was assayed on the HEX channel as previously described [38]. Sequences of the KIR-specific oligonucleotides are identical to those used previously [10] (Additional file 1). KIR2DL5 was assayed on the FAM channel. Counts of KIR-positive droplets (FAM+HEX- and FAM+HEX+) and of RPP30-positive droplets (FAM-HEX+ and FAM+HEX+) were used to estimate the number of copies per microliter of each target in the reaction [35]. Estimates of the KIR target were divided by the estimates of the RPP30 target, then multiplied by two to correct for diploidy in KIR CNV estimates. Clustering of raw CNV data into bins representing zero, one, two, three or four copies was performed by application of mclust, which permits a Gaussian finite mixture modeling via the expectation-maximization algorithm [39]. KIR2DS3 and KIR2DS5 genotyping was performed using a validated PCR, using sequence-specific primers (PCR-SSP) approach [40]. CNV variation in eight KIR genes (KIR2DL1, KIR3DL1, KIR2DS1, KIR2DS2, KIR2DS3, KIR2DS5, KIR3DS1 and KIR3DP1; Figures S1 to S8 in Additional file 1) was tested using the same method for 19 samples from a number of sources. There were insufficient data to apply mclust to these data, so clustering into bins was performed by eye with the aid of the figures. qPCR and ddPCR results were compared using Cohen’s Kappa statistic.
Results
Genetic linkage analysis
The experimentally determined evidence for the presence or absence of each haplotype motif is shown in Figure 2. There was evidence for the presence of KIR2DL2 ~ KIR2DL5C arrangement in the undigested DNA preparations of the cell lines HO104, HO301, JESTHOM, LBF (aka LBUF), WJR076 and WT24, whilst the remaining cell lines did not possess any copies of KIR2DL2. The KIR3DS1 ~ KIR2DL5T haplotype was present in BOLETH, COX, HO104, HOM2, JESTHOM, LBF, MCF and WT24. HO301 did not have KIR3DS1. Evidence for linkage was partially or totally abrogated in the EcoRI digested DNA preparations.
Figure 2.
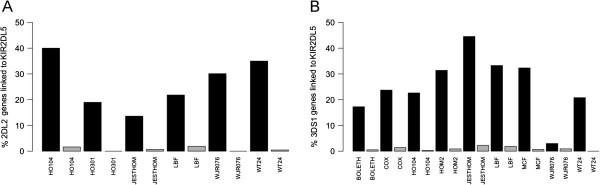
Experimentally determined signatures of linkage between (A) KIR2DL2 and KIR2DL5 and (B) KIR3DS1 and KIR2DL5 for the 10 IHW cell lines. Evidence of linkage (black bars) in DNA sample preparations (fragment size >23 kb), expressed as the percentage of KIR2DL2 or KIR3DS1 genes that were linked to a KIR2DL5 gene. Linkage was abrogated by EcoRI restriction endonuclease digestion (gray bars).
WJR076
In the case of WJR076, there was definitive evidence for the presence of the KIR2DL2 ~ KIR2DL5 haplotype (%L = 30, falling to %L = 0% after EcoRI digestion), which demonstrates that sample quality was sufficient to detect linkage over ~22 kb. In another aliquot from the same DNA source, there was some evidence for the presence of KIR3DS1 ~ KIR2DL5 (%L = 2.9, falling to %L = 0.9% after EcoRI digestion). This may indicate linkage across a much greater distance.
Copy number variation
Based on the ddPCR analysis, KIR2DL5 was present at between zero and three copies in the specimens tested. The raw CNV estimates of the UCLA KIR exchange samples fit closely to integers (Figure 3A), whilst systematic underestimates of copy number were observed in samples from the Gambian families (Figure 3B). The colours of the points in Figure 3B reflect the results of applying mclust to the KIR2DL5 CNV data. KIR2DL5 CNVs segregated through the Gambian families in patterns that were compatible with Mendelian inheritance. KIR2DL5 co-segregated with KIR2DS3 and KIR2DS5 (Figure 4), which are usually found in tight linkage disequilibrium with KIR2DL5[20]. One sample (F-1268) was assigned to the zero copy cluster by the mclust approach, but had a non-zero (0.268, confidence interval (CI) 0.182 to 0.353) CNV estimate, suggesting a true copy number of one. The CNV testing of eight additional KIR genes is shown in Figures S1 to S8 and Table S2 in Additional file 1).
Figure 3.
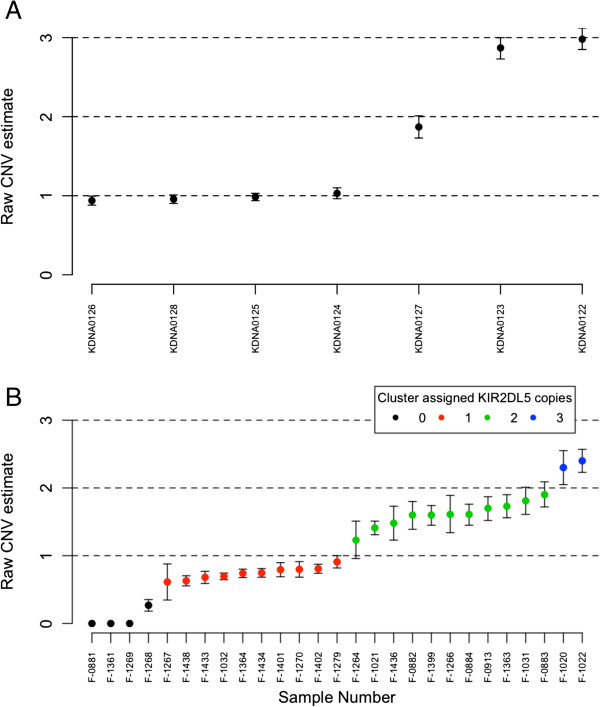
KIR CNV estimates in (A) UCLA International KIR exchange samples and (B) Gambian family samples. Colour-coded points indicate cluster membership (0 copies (n = 3), 1 copy (n = 10), 2 copies (n = 12), 3 copies (n = 2)).
Figure 4.
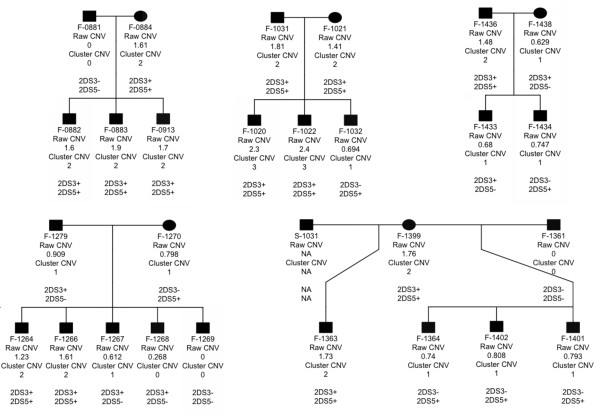
Segregation of KIR2DL5 in five Gambian families. Patterns of segregation are consistent with Mendelian inheritance and with segregation of the linked KIR2DS3 and KIR2DS5 genes. In most world populations, KIR2DL5 is in perfect linkage disequilibrium with KIR2DS3 and KIR2DS5. In Gambia, these genes are in almost perfect linkage disequilibrium (Roberts C.h., Molina, S., Makalo, P., Joof, H., Harding-Esch, E.M., Burr, S.E., Mabey, D.W., Bailey, R.L., Burton, M.J. and Holland, M.J., in press). The absence of both KIR2DS3 and KIR2DS5 implies that KIR2DL5 will be absent. The presence of both genes indicates that at least two KIR2DL5 copies are present. The co-segregation of KIR2DL5 with the KIR2DS3 and KIR2DS5 genes is expected and was observed in these pedigrees. To protect the anonymity of participant families, all F1 subjects have been changed to ‘male’.
Discussion
Recent advances in KIR genotyping [10,17] have confirmed that CNVs are common in KIR genes and may relate directly to NK cell repertoire [41] and surface expression levels of the KIR proteins [42] with potential impact on NK cell function. We developed a next generation digital PCR assay for enumeration of KIR CNVs and resolution of KIR haplotype structures by genetic linkage analysis.
We have shown how ddPCR can be used to detect the presence or absence of the KIR3DS1 ~ KIR2DL5 (spanning approximately 21.8 kb) and KIR2DL2 ~ KIR2DL5 (spanning approximately 19.9 kb) groupings (Figure 2). Evidence for linkage was found in our ability to enumerate the percentage of KIR2DL5 genes that were linked to another assayed gene. The application of EcoRI restriction endonuclease digestion abrogated these signatures of linkage and demonstrated the robustness of the method.
Proximity between the assayed genes, the variable copy number of each target and the average length of the DNA fragment in the tested material are all able to influence the %L score and they are likely to do so in a complex and unpredictable manner. When the distance between two targets is very high, the potential for mechanical shearing is increased and high %L scores are less likely. Low sample quality will reduce the maximum achievable %L score, whilst gene copy numbers will impose absolute limits in all specimens. For instance, a specimen with two copies of KIR2DL5 (one on each chromosome), but a single copy of KIR2DL2, will have a maximum possible %L of 50.
In the case of one sample in our study, WJR076, we detected a weak %L score that was reduced by treatment with EcoRI. We suspected that this may be explained by the presence of an unusual extended haplotype and investigated this further. qPCR copy number data for WJR076 were consistent with the carriage of an extended KIR haplotype carrying a duplication of the KIR2DL4 and KIR3DL1/S1 loci. We subsequently ascertained by PCR-SSP that WJR076 carries the hybrid gene KIR2DL5/3DP1 (known as KIR3DP1*004), which has been previously identified on various extended KIR haplotypes [9,10,43]. Our KIR2DL5 ddPCR assay targets KIR2DL5 exon 9, which is absent from KIR3DP1*004, so would not detect the portion of KIR2DL5 (promoter region to exon 2) that forms part of the hybrid gene. The extended haplotype is, therefore, predicted to take the form KIR2DL2 ~ KIR2DL5 ~ […] ~ KIR3DS1, where the KIR2DL2 ~ KIR2DL5 conformation is in the centromeric motif and the KIR3DS1 gene is in the extended region (Figure 5). This example illustrates how in future this approach will be useful for identifying and characterizing specimens, which may have novel haplotype structures. With improvements in DNA extraction procedures to yield longer unbroken fragments (>100 kb) the ddPCR technique will be able to link genes across the entire length of KIR haplotypes; however, we might expect very low %L scores in these samples, even when great care is taken during DNA preparation. The longest fragments will always be in the minority and there is likely to be a maximum feasible length of DNA fragment that can survive the mechanical forces involved in generating and manipulating the ddPCR droplets.
Figure 5.
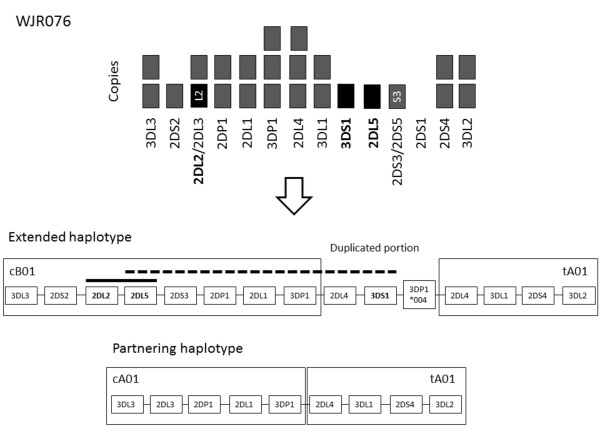
The predicted gene carriage and resolved arrangements for WJR076 haplotypes. Showing the constituent haplotype motifs: cB01:tA01 within the extended KIR B haplotype and cA01:tA01 forming a standard KIR A haplotype. The extended haplotype has the 2DL5/3DP1 hybrid gene (KIR3DP1*004) and duplication of the KIR2DL4 and KIR3DL1/S1 loci. Linkage analysis supported close relationship of KIR2DL2 ~ KIR2DL5 (indicated by a solid black line) and long-distance linkage between KIR2DL5 and KIR3DS1 (dashed line) on the extended haplotype.
Pedigree analysis confirmed that ddPCR was an effective way to carry out CNV enumeration of KIR2DL5. We tested DNA samples of different origins, which had been extracted using different methods and some of which (the Gambian specimens) had been subjected to multiple freeze/thaw cycles and stored for prolonged periods of time. We found that there was variation in how well the estimated CNV values fit to integers (Figure 3) and that this was correlated with the source of the DNA (Figure 3A,B) as well as being influenced by the primer sets (Figures S4 and S7 in Additional file 1) and by sample to sample variation (Figures S1 to S8 in Additional file 1; note that specimen 10865 diverges from the integers in several assays, but 10834 fits closely to integers). We subjected aliquots of the UCLA specimens to 15 freeze-thaw cycles, but were unable to affect changes in the measured CNV estimates (data not shown). There were no systematic differences in the input amounts of DNA, although the measured concentrations of RPP30 were systematically lower in the Gambian samples. This probably reflects that they came from buccal swab specimens and that a significant proportion of the total DNA is likely to be of bacterial origin. The CEPH and local donor DNA specimens (QPQ, NNA, CFF, AHS, JKN, RTC) had no systematic concentration differences from the UCLA samples, but still failed to fit to integers. The common features between the Gambian and CEPH/local donor specimens were their age and that they had been previously archived at -20°C so we might suspect that acid hydrolysis may be a factor for consideration. Further and careful investigation will be needed to identify the reasons for the variation in fit to integers, but a simple clustering approach was still able to discern the groups of samples with zero, one, two and three copies of KIR2DL5.
Assay-specific optimization will be necessary in order to fully establish ddPCR-based KIR CNV tests as routine tools, but the ddPCR CNV data compared well to a conventional qPCR method. Six assays had perfect agreement (Cohen’s Kappa = 1) between the two methods (KIR2DL1, KIR2DS1, KIR2DS2, KIR2DS3, KIR2DS5, KIR3DS1), whilst there were disagreements between the results of two assays, KIR3DL1 (Kappa = 0.9, 95% CI 0.71 to 1.00) and KIR3DP1 (Kappa = 0.74, 95% CI 0.49 to 0.99).
Conclusions
The definition and characterization of structural haplotypes in the KIR region provide a significant challenge and one that has not previously been addressed using a simple, scalable, cost-effective and reproducible method. The next generation PCR approach described here makes it possible for the first time to routinely define KIR structural haplotypes in a way that is not possible using other PCR methods. In addition we have shown that ddPCR has potential for development as a method for KIR CNV enumeration.
We have shown that both the clustering method (Figures 3 and 4) and the use of raw data (Figure 4, kappa analysis) can achieve accurate CNV estimates. Clustering should be generally favored over the more subjective approach, but the example of the F-1268 specimen, which mclust very obviously placed in the wrong cluster, indicates that there will continue to be a role for human inspection in the classification process.
For several of the assays that we describe (Figures S1 to S3 and S6 in Additional file 1), the interpretation of ddPCR KIR CNV data was made easier by the simultaneous testing of a plurality of specimens. This is because sufficient data were generated to identify discrete clusters of data points (either by eye or using statistical clustering methods) even though the raw CNV estimates did not reproducibly fit to integers. In a clinical setting it is unlikely that large numbers of specimens will always need to be tested simultaneously and the variation between DNA from different sources may cause additional problems. A simple solution might be to utilize a supervised learning method wherein a large set of specimens are initially tested in situ at the clinic or centre, in order to train the algorithm for classification in a way that works given the operating procedure and in the context of the specific primer/probe set and how, on average, it performs given the average sample that will be tested at that specific centre. In application, it is also recommended that a set of control samples of known copy number are included in each run to be used as calibrators in the analysis and external quality control.
The gene linkage application will complement KIR genotype data and aid understanding of the population genetic and structural diversity of the KIR gene region. An immediate utility is that it allows KIR 'AB' and 'BB' genotypes to be distinguished, which are currently grouped together as 'Bx' because of uncertainty in KIR copy numbers. The KIR 'Bx' genotype has previously been associated with increased relapse-free survival after unrelated haematopoietic cell transplant [44] and the discrimination of 'AB' from 'BB' genotypes will make it possible to perform a more refined investigation of this association, including the determination of the underlying genetic model. The KIR 'AB' genotype has also been shown to predict higher in vitro cytokine responses to pathogen-associated signals in human KIR+ CD56dim NK cells [45]. The assay described here will make it possible to specifically identify 'AB' individuals and to undertake genetic association studies that model this behaviour. Histocompatibility matching algorithms for haematopoietic stem cell transplants already take account of KIR genotypes [44] and the ability to perform assays such as those we describe on a high throughput platform will facilitate the routine consideration of KIR CNV diversity and haplotype diversity in the selection of appropriate donors. The affordability (less than $5 US per test) and scalability (one operator can run four 96-well plates in a normal working day) of the ddPCR assay makes it an excellent candidate platform for high throughput KIR CNV genotyping/haplotyping for transplantation registries and for use in large-scale genetic association and population genetics studies.
Abbreviations
%L: percent linkage, the normalized linkage score; CI: confidence interval; CNV: copy number variation; ddPCR: droplet digital PCR; FAM: 6-carboxyfluorescein; HEX: 6-carboxy-2,4,4,5,7,7-hexachlorofluorescein succinimidyl ester; IHW: International Histocompatibility Workshop; KIR: Killer-cell Immunoglobulin-like Receptor; NK: natural killer; PCR: polymerase chain reaction; PCR-SSP: PCR, using sequence specific primers; qPCR: quantitative PCR.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
ChR, JAT and WJ conceived the study. ChR, JAT, WJ and JJ performed the experiments. ChR and JAT analyzed the data. JT and MJH contributed reagents and samples. ChR, JAT, JT and MJH wrote the manuscript. All authors read, revised and approved the manuscript.
Supplementary Material
Oligonucleotide primers and probes used in this study. Table S2. Summary of KIR CNV genotypes in 19 samples, tested using qPCR and ddPCR. Table S3. Droplet counts and lambda statistics for all tests carried out in this study. Figure S1. Results of ddPCR KIR CNV assay: KIR2DL1. Figure S2. Results of ddPCR KIR CNV assay: KIR2DS1. Figure S3. Results of ddPCR KIR CNV assay: KIR2DS2. Figure S4. Results of ddPCR KIR CNV assay: KIR2DS3. Figure S5. Results of ddPCR KIR CNV assay: KIR2DS5. Figure S6. Results of ddPCR KIR CNV assay: KIR3DL1. Figure S7. Results of ddPCR KIR CNV assay: KIR3DP1. Figure S8. Results of ddPCR KIR CNV assay: KIR3DS1.
Contributor Information
Chrissy h Roberts, Email: chrissy.roberts@lshtm.ac.uk.
Wei Jiang, Email: wj224@cam.ac.uk.
Jyothi Jayaraman, Email: jj329@cam.ac.uk.
John Trowsdale, Email: jt233@mole.bio.cam.ac.uk.
Martin J Holland, Email: drmjh650@gmail.com.
James A Traherne, Email: jat51@cam.ac.uk.
Acknowledgments
We thank Jim Kaufman and Olympe Chazara for critical reading of the manuscript. The IHW cell line DNA samples were kindly donated by the Tissue Typing Laboratories, Cambridge University Hospitals NHS Foundation Trust.
Funding statement
This work was funded by the Wellcome Trust (GR079246MA) and by the Medical Research Council, UK.
References
- Suto Y, Maenaka K, Yabe T, Hirai M, Tokunaga K, Tadok K, Juji T. Chromosomal localization of the human natural killer cell class I receptor family genes to 19q13.4 by fluorescence in situ hybridization. Genomics. 1996;35:270–272. doi: 10.1006/geno.1996.0355. [DOI] [PubMed] [Google Scholar]
- Khakoo SI, Thio CL, Martin MP, Brooks CR, Gao X, Astemborski J, Cheng J, Goedert JJ, Vlahov D, Hilgartner M, Cox S, Little A-M, Alexander GJ, Cramp ME, O’Brien SJ, Rosenberg WMC, Thomas DL, Carrington M. HLA and NK cell inhibitory receptor genes in resolving hepatitis C virus infection. Science. 2004;305:872–874. doi: 10.1126/science.1097670. [DOI] [PubMed] [Google Scholar]
- Martin MP, Gao X, Lee J-H, Nelson GW, Detels R, Goedert JJ, Buchbinder S, Hoots K, Vlahov D, Trowsdale J, Wilson M, O’Brien SJ, Carrington M. Epistatic interaction between KIR3DS1 and HLA-B delays the progression to AIDS. Nat Genet. 2002;31:429–434. doi: 10.1038/ng934. [DOI] [PubMed] [Google Scholar]
- Cooley S, Weisdorf DJ, Guethlein LA, Klein JP, Wang T, Le CT, Marsh SGE, Geraghty D, Spellman S, Haagenson MD, Ladner M, Trachtenberg E, Parham P, Miller JS. Donor selection for natural killer cell receptor genes leads to superior survival after unrelated transplantation for acute myelogenous leukemia. Blood. 2010;116:2411–2419. doi: 10.1182/blood-2010-05-283051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hiby SE, Walker JJ, O’Shaughnessy KM, Redman CWG, Carrington M, Trowsdale J, Moffett A, Shaughnessy KMO. Combinations of maternal KIR and fetal HLA-C genes influence the risk of preeclampsia and reproductive success. J Exp Med. 2004;200:957–965. doi: 10.1084/jem.20041214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu KC, Liu X-RR, Selvakumar A, Mickelson E, O’Reilly RJ, Dupont B. Killer Ig-like receptor haplotype analysis by gene content: evidence for genomic diversity with a minimum of six basic framework haplotypes, each with multiple subsets. J Immunol. 2002;169:5118–5129. doi: 10.4049/jimmunol.169.9.5118. [DOI] [PubMed] [Google Scholar]
- Traherne JA, Martin M, Ward R, Ohashi M, Pellett F, Gladman D, Middleton D, Carrington M, Trowsdale J. Mechanisms of copy number variation and hybrid gene formation in the KIR immune gene complex. Hum Mol Genet. 2010;19:737–751. doi: 10.1093/hmg/ddp538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams F, Maxwell LD, Halfpenny IA, Meenagh A, Sleator C, Curran MD, Middleton D. Multiple copies of KIR 3DL/S1 and KIR 2DL4 genes identified in a number of individuals. Hum Immunol. 2003;64:729–732. doi: 10.1016/S0198-8859(03)00089-2. [DOI] [PubMed] [Google Scholar]
- Martin MP, Bashirova A, Traherne J, Trowsdale J, Carrington M. Cutting edge: expansion of the KIR locus by unequal crossing over. J Immunol. 2003;171:2192–2195. doi: 10.4049/jimmunol.171.5.2192. [DOI] [PubMed] [Google Scholar]
- Jiang W, Johnson C, Jayaraman J, Simecek N, Noble J, Moffatt MF, Cookson WO, Trowsdale J, Traherne JA. Copy number variation leads to considerable diversity for B but not A haplotypes of the human KIR genes encoding NK cell receptors. Genome Res. 2012;22:1845–1854. doi: 10.1101/gr.137976.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson MJ, Torkar M, Haude A, Milne S, Jones T, Sheer D, Beck S, Trowsdale J. Plasticity in the organization and sequences of human KIR/ILT gene families. Proc Natl Acad Sci U S A. 2000;97:4778–4783. doi: 10.1073/pnas.080588597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uhrberg M, Valiante NM, Shum BP, Shilling HG, Lienert-Weidenbach K, Corliss B, Tyan D, Lanier LL, Parham P. Human diversity in killer cell inhibitory receptor genes. Immunity. 1997;7:753–763. doi: 10.1016/S1074-7613(00)80394-5. [DOI] [PubMed] [Google Scholar]
- Hsu KC, Chida S, Geraghty DE, Dupont B. The killer cell immunoglobulin-like receptor (KIR) genomic region: gene-order, haplotypes and allelic polymorphism. Immunol Rev. 2002;190:40–52. doi: 10.1034/j.1600-065X.2002.19004.x. [DOI] [PubMed] [Google Scholar]
- Vilches C, Gardiner CM, Parham P. Gene structure and promoter variation of expressed and nonexpressed variants of the KIR2DL5 gene. J Immunol. 2000;165:6416–6421. doi: 10.4049/jimmunol.165.11.6416. [DOI] [PubMed] [Google Scholar]
- Gómez-Lozano N, Gardiner CM, Parham P, Vilches C. Some human KIR haploytpes contain two KIR2DL5 genes: KIR2DL5A and KIR2DL5B. Immunogenetics. 2002;54:314–319. doi: 10.1007/s00251-002-0476-2. [DOI] [PubMed] [Google Scholar]
- Norman PJ, Carrington CV, Byng M, Maxwell LD, Curran MD, Stephens HA, Chandanayingyong D, Verity DH, Hameed K, Ramdath DD, Vaughan RW. Natural killer cell immunoglobulin-like receptor (KIR) locus profiles in African and South Asian populations. Genes Immun. 2002;3:86–95. doi: 10.1038/sj.gene.6363836. [DOI] [PubMed] [Google Scholar]
- Pyo C-W, Wang R, Vu Q, Cereb N, Yang SY, Duh F-M, Wolinsky S, Martin MP, Carrington M, Geraghty DE. Recombinant structures expand and contract inter and intragenic diversification at the KIR locus. BMC Genomics. 2013;14:89. doi: 10.1186/1471-2164-14-89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maiers M, Gragert L, Klitz W. High-resolution HLA alleles and haplotypes in the United States population. Hum Immunol. 2007;68:779–788. doi: 10.1016/j.humimm.2007.04.005. [DOI] [PubMed] [Google Scholar]
- The Immuno Polymorphism Database (IPD) [ http://www.ebi.ac.uk/]
- Middleton D, Meenagh A, Gourraud PA. KIR haplotype content at the allele level in 77 Northern Irish families. Immunogenetics. 2007;59:145–158. doi: 10.1007/s00251-006-0181-7. [DOI] [PubMed] [Google Scholar]
- Martin MP, Single RM, Wilson MJ, Trowsdale J, Carrington M. KIR haplotypes defined by segregation analysis in 59 Centre d’Etude Polymorphisme Humain (CEPH) families. Immunogenetics. 2008;60:767–774. doi: 10.1007/s00251-008-0334-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Besson C, Roetynck S, Williams F, Orsi L, Amiel C, Lependeven C, Antoni G, Hermine O, Brice P, Ferme C, Carde P, Canioni D, Brière J, Raphael M, Nicolas J-C, Clavel J, Middleton D, Vivier E, Abel L. Association of killer cell immunoglobulin-like receptor genes with Hodgkin’s lymphoma in a familial study. PLoS One. 2007;2:e406. doi: 10.1371/journal.pone.0000406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norman PJ, Abi-Rached L, Gendzekhadze K, Korbel D, Gleimer M, Rowley D, Bruno D, Carrington CVF, Chandanayingyong D, Chang Y-H, Crespí C, Saruhan-Direskeneli G, Fraser PA, Hameed K, Kamkamidze G, Koram KA, Layrisse Z, Matamoros N, Milà J, Park MH, Pitchappan RM, Ramdath DD, Shiau M-Y, Stephens HAF, Struik S, Verity DH, Vaughan RW, Tyan D, Davis RW, Riley EM. et al. Unusual selection on the KIR3DL1/S1 natural killer cell receptor in Africans. Nat Genet. 2007;39:1092–1099. doi: 10.1038/ng2111. [DOI] [PubMed] [Google Scholar]
- Pyo C-W, Guethlein LA, Vu Q, Wang R, Abi-Rached L, Norman PJ, Marsh SGE, Miller JS, Parham P, Geraghty DE. Different patterns of evolution in the centromeric and telomeric regions of group A and B haplotypes of the human killer cell Ig-like receptor locus. PLoS One. 2010;5:e15115. doi: 10.1371/journal.pone.0015115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hou L, Chen M, Ng J, Hurley CK. Conserved KIR allele-level haplotypes are altered by microvariation in individuals with European ancestry. Genes Immun. 2012;13:47–58. doi: 10.1038/gene.2011.52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dear PH, Cook PR. Happy mapping: a proposal for linkage mapping the human genome. Nucleic Acids Res. 1989;17:6795–6807. doi: 10.1093/nar/17.17.6795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim L-S, Tay Y-L, Alias H, Wan K-L, Dear PH. Insights into the genome structure and copy-number variation of Eimeria tenella. BMC Genomics. 2012;13:389. doi: 10.1186/1471-2164-13-389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keyes D. WO 2012/129436: Identification of Linkage Using Multiplex Digital PCR (Patent application) 2012.
- Boulanger J, Muresan L, Tiemann-Boege I. Massively parallel haplotyping on microscopic beads for the high-throughput phase analysis of single molecules. PLoS One. 2012;7:e36064. doi: 10.1371/journal.pone.0036064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ge Q, Liu Z, Bai Y, Zhang D, Yu P, Lu Z. Emulsion PCR-based method to detect Y chromosome microdeletions. Anal Biochem. 2007;367:173–178. doi: 10.1016/j.ab.2007.05.008. [DOI] [PubMed] [Google Scholar]
- Pinheiro LB, Coleman VA, Hindson CM, Herrmann J, Hindson BJ, Bhat S, Emslie KR. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal Chem. 2012;84:1003–1011. doi: 10.1021/ac202578x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belgrader P, Tanner SC, Regan JF, Koehler R, Hindson BJ, Brown AS. Droplet digital PCR measurement of HER2 copy number alteration in formalin-fixed paraffin-embedded breast carcinoma tissue. Clin Chem. 2013;59:991–994. doi: 10.1373/clinchem.2012.197855. [DOI] [PubMed] [Google Scholar]
- Tan Y-C, Cristini V, Lee AP. Monodispersed microfluidic droplet generation by shear focusing microfluidic device. Sensors Actuators B Chem. 2006;114:350–356. doi: 10.1016/j.snb.2005.06.008. [DOI] [Google Scholar]
- Vogelstein B, Kinzler KW. Digital PCR. Proc Natl Acad Sci U S A. 1999;96:9236–9241. doi: 10.1073/pnas.96.16.9236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dube S, Qin J, Ramakrishnan R. Mathematical analysis of copy number variation in a DNA sample using digital PCR on a nanofluidic device. PLoS One. 2008;3:e2876. doi: 10.1371/journal.pone.0002876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalinina O, Lebedeva I, Brown J, Silver J. Nanoliter scale PCR with TaqMan detection. Nucleic Acids Res. 1997;25:1999–2004. doi: 10.1093/nar/25.10.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- The UCLA Immunogenetics Center. [ http://www.hla.ucla.edu/]
- Luo W, Yang H, Rathbun K, Pau C-P, Ou C-Y. Detection of human immunodeficiency virus type 1 DNA in dried blood spots by a duplex real-time PCR assay. J Clin Microbiol. 2005;43:1851–1857. doi: 10.1128/JCM.43.4.1851-1857.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- mclust.Normal Mixture Modelling for Model-Based Clustering, Classification, and Desity Estimation. [ http://www.stat.washington.edu/mclust/]
- Vilches C, Castaño J, Gómez-Lozano N, Estefanía E. Facilitation of KIR genotyping by a PCR-SSP method that amplifies short DNA fragments. Tissue Antigens. 2007;70:415–422. doi: 10.1111/j.1399-0039.2007.00923.x. [DOI] [PubMed] [Google Scholar]
- Béziat V, Traherne JA, Liu LL, Jayaraman J, Enqvist M, Larsson S, Trowsdale J, Malmberg K-J. Influence of KIR gene copy number on natural killer cell education. Blood. 2013;121:4703–4707. doi: 10.1182/blood-2012-10-461442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norman PJ, Abi-Rached L, Gendzekhadze K, Hammond JA, Moesta AK, Sharma D, Graef T, McQueen KL, Guethlein LA, Carrington CVF, Chandanayingyong D, Chang Y-H, Crespí C, Saruhan-Direskeneli G, Hameed K, Kamkamidze G, Koram KA, Layrisse Z, Matamoros N, Milà J, Park MH, Pitchappan RM, Ramdath DD, Shiau M-Y, Stephens HAF, Struik S, Tyan D, Verity DH, Vaughan RW, Davis RW. et al. Meiotic recombination generates rich diversity in NK cell receptor genes, alleles, and haplotypes. Genome Res. 2009;19:757–769. doi: 10.1101/gr.085738.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gómez-Lozano N, Estefanía E, Williams F, Halfpenny I, Middleton D, Solís R, Vilches C. The silent KIR3DP1 gene (CD158c) is transcribed and might encode a secreted receptor in a minority of humans, in whom the KIR3DP1, KIR2DL4 and KIR3DL1/KIR3DS1 genes are duplicated. Eur J Immunol. 2005;35:16–24. doi: 10.1002/eji.200425493. [DOI] [PubMed] [Google Scholar]
- Cooley S, Trachtenberg E, Bergemann TL, Saeteurn K, Klein J, Le CT, Marsh SG, Guethlein LA, Parham P, Miller JS, Weisdorf DJ. Donors with group B KIR haplotypes improve relapse-free survival after unrelated hematopoietic cell transplantation for acute myelogenous leukemia. Blood. 2009;113:726–732. doi: 10.1182/blood-2008-07-171926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korbel DS, Norman PJ, Newman KC, Horowitz A, Gendzekhadze K, Parham P, Riley EM. Killer Ig-like receptor (KIR) genotype predicts the capacity of human KIR-positive CD56dim NK cells to respond to pathogen-associated signals. J Immunol. 2009;182:6426–6434. doi: 10.4049/jimmunol.0804224. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Oligonucleotide primers and probes used in this study. Table S2. Summary of KIR CNV genotypes in 19 samples, tested using qPCR and ddPCR. Table S3. Droplet counts and lambda statistics for all tests carried out in this study. Figure S1. Results of ddPCR KIR CNV assay: KIR2DL1. Figure S2. Results of ddPCR KIR CNV assay: KIR2DS1. Figure S3. Results of ddPCR KIR CNV assay: KIR2DS2. Figure S4. Results of ddPCR KIR CNV assay: KIR2DS3. Figure S5. Results of ddPCR KIR CNV assay: KIR2DS5. Figure S6. Results of ddPCR KIR CNV assay: KIR3DL1. Figure S7. Results of ddPCR KIR CNV assay: KIR3DP1. Figure S8. Results of ddPCR KIR CNV assay: KIR3DS1.


