Abstract
Background
Aberrant hypermethylation of gene promoter regions is a primary mechanism by which tumor suppressor genes become inactivated in breast cancer. Epigenetic inactivation of the protein tyrosine phosphatase receptor-type O gene (PTPRO) has been described in several types of cancer.
Results
We screened primary breast cancer tissues for PTPRO promoter hypermethylation and assessed potential associations with pathological features and patient outcome. We also evaluated its potential as a breast cancer biomarker. PTPRO methylation was observed in 53 of 98 (54%) breast cancer tissues but not in adjacent normal tissue. Among matched peripheral blood samples from breast cancer patients, 33 of 98 (34%) exhibited methylated PTPRO in plasma. In contrast, no methylated PTPRO was observed in normal peripheral blood from 30 healthy individuals. PTPRO methylation was positively associated with lymph node involvement (P = 0.014), poorly differentiated histology (P = 0.037), depth of invasion (P = 0.004), and HER2 amplification (P = 0.001). Multivariate analysis indicated that aberrant PTPRO methylation could serve as an independent predictor for overall survival hazard ratio (HR): 2.7; 95% CI: 1.1-6.2; P = 0.023), especially for patients with HER2-positive (hazard ratio (HR): 7.5; 95% CI: 1.8-31.3; P = 0.006), but not in ER + and PR + subpopulation. In addition, demethylation induced by 5-azacytidine led to gene reactivation in PTPRO-methylated and -silenced breast cancer cell lines.
Conclusions
Here, we report that tumor PTPRO methylation is a strong prognostic factor in breast cancer. Methylation of PTPRO silences its expression and plays an important role in breast carcinogenesis. The data we present here may provide insight into the development of novel therapies for breast cancer treatment. Additionally, detection of PTPRO methylation in peripheral blood of breast cancer patients may provide a noninvasive means to diagnose and monitor the disease.
Keywords: Protein tyrosine phosphatase receptor-type O (PTPRO), Methylation, Breast cancer, Clinical outcome, Biomarker
Background
Breast cancer is one of the most common cancers among women worldwide, and its incidence, unfortunately, continues to rise. Breast tumor is a heterogeneous disease derived from different molecular subtypes and displaying varied clinical behavior [1]. Considerable efforts have been made to improve survival via early diagnosis and treatment with targeted therapies [2]. However, the limited success of current therapeutic modalities has led to calls for new prognostic tools and for the development of additional targeted therapies [3].
Promoter hypermethylation is a type of epigenetic alteration associated with gene silencing. In cancer, many tumor suppressor genes are inactivated in this way. Hypermethylation of key tumor suppressors is a key contributor to breast tumorigenesis and acts in concert with genetic alterations to drive disease progression [4]. Epigenetic modifications of tumor DNA may have prognostic significance for breast cancer patients and provide targets for treatment because they are potentially reversible. Epigenetic changes may also serve as markers for early detection of the disease. As an example, screening for RASSF1A hypermethylation in serum has been proposed as a form of surveillance to detect early stage breast cancer [5].
In recent years, there has been considerable interest in better understanding the role of tyrosine phosphorylation in cancer [6-11], especially since this post-translational modification helps regulate diverse cellular processes, including proliferation, differentiation, metabolism, cell-to-cell communication, transcription, and survival [12]. Phosphorylation is a dynamic process that is positively regulated by protein tyrosine kinases (PTKs) and negatively regulated by protein tyrosine phosphatases (PTPs). More than 80% of oncogenes encode PTKs [13]; in contrast, many PTPs have been described to function as tumor suppressors [14]. For example, the tyrosine phosphatase PTPN2 activates TP53 and induces apoptosis in human tumor cells [15]. Another phosphatase, PTP1B, negatively regulates insulin signaling via dephosphorylation of insulin receptor kinase [16]. Computational analysis of the human genome identified 38 classical PTP genes, 19 of which mapped to regions frequently deleted in human cancers. Thirty of these protein phosphatases have been implicated in tumorigenesis [17], further demonstrating their potential roles as tumor suppressors.
Protein tyrosine phosphatase receptor-type O (PTPRO) is classified as a receptor-type PTP of the R3 subtype [18] and exhibits characteristics of a tumor suppressor in multiple cancers [19]. Several PTPRO variants have been described due to use of distinct transcriptional start sites and to alternative splicing; while many lymphoid-derived cells express a truncated PTPRO isoform, most epithelial tissues, including the breast, express the full-length form [19]. Previous studies have reported methylation-mediated down-regulation of PTPRO expression in breast cancer and other tumor types, such as rat hepatocellular carcinoma, human chronic lymphocytic leukemia, human lung cancer, esophageal carcinoma [6-11,20]. Hypermethylation of PTPRO occurs frequently in esophageal carcinoma and may be a potential biomarker of the disease [20]. A recent study also revealed that acute lymphoblastic leukemia patients with PTPRO methylation showed increased rates of relapse and chemoresistance [9].
More recently, a tumor suppressive role for PTPRO in breast cancer has emerged. Tumor-specific PTPRO promoter methylation was documented in primary human breast cancer cases [10]. The authors of this study also found that PTPRO expression was reduced upon treatment with estrogen but increased by treatment with the anti-estrogen Tamoxifen. Furthermore, ectopic expression of PTPRO in non-expressing MCF-7 cells sensitized them to the growth suppressive effects of Tamoxifen. PTPRO methylation has been further confirmed to be clinically relevant in breast cancer, particularly in HER2-amplified patients. Huang et al. showed that overall survival is significantly worse in HER2-positive patients with methylated PTPRO compared to tumors lacking methylation of this promoter region [21]. Another study found that low expression of PTPRO correlated with reduced survival for HER2-positive breast cancer patients [11]. It is possible that the pronounced impact of PTPRO specifically in HER2-positive disease could be due to the fact that HER2 itself is a direct substrate of PTPRO phosphatase activity [11]; specifically, loss of PTPRO was shown to increase HER2 phosphorylation and HER2-induced proliferation and transformation of breast cancer cell lines. Taken together, these data support a role for PTPRO as a tumor suppressor in breast cancer and suggest that its methylation and expression may have prognostic significance in the disease.
In the current study we investigated the methylation status of PTPRO in primary human breast cancer from fresh frozen specimens with the aim of defining the frequency of this epigenetic aberration in the disease. We examined the methylation status of PTPRO in primary breast tumors and matched peripheral blood samples and determined if promoter methylation was associated with decreased gene expression in breast cancer cell lines. We also examined associations between PTPRO methylation and several clinicopathological parameters, including patient outcome.
Methods
Tumor samples
Between 2006 and 2009, we obtained 98 tumor samples and matched pre-operative peripheral blood samples from women undergoing surgery for primary invasive breast carcinoma at ShenZhen Maternal and Child Health Hospital, an affiliate of Southern Medical University in China. None of the patients had received any pre-operative treatment, including chemotherapy or radiotherapy. This is a well-characterized series of patients under the age of 74 years (median, 46 years). The median follow-up time of patients in the study was 60 months (range 43–70 months). All patients were treated uniformly at a single institution.
Pathologic characteristics, including histological grade, histological tumor type, tumor size, and lymph node involvement were routinely assessed; several patient characteristics, including age and family history of cancer and menopause, were also recorded. Survival data were maintained prospectively. At the end of the study period, 39 (40%) patients had died because of disease recurrence. In total, 98% of node-positive and 82% of node-negative patients received adjuvant systemic therapy consisting of either hormone therapy alone or hormone therapy plus chemotherapy.
Tumor samples were immediately frozen in liquid nitrogen and stored at −80°C until use. All tumors were confirmed histopathologically and their clinical features were classified based on the TNM system of the International Union Against Cancer [22]. Corresponding adjacent non-cancerous tissues were also obtained from surgical resections. Peripheral venous blood samples from breast tumor patients were collected in EDTA-containing tubes and immediately centrifuged at 2500 g for 15 min to prepare plasma. The plasma samples were stored at −80°C until further processing. Peripheral blood samples from an additional 30 healthy volunteers were used as normal controls. Estrogen receptor (ER), progesterone receptor (PgR), and human epidermal growth factor receptor 2 (HER2) immunohistochemistry was performed on TMA sections as previously described [23].
Approval for the use of human tissues and clinical information was obtained from the Committee for Ethical Review of Research involving Human Subjects at Southern Medical University. All patients provided written informed consent for sample collection prior to surgery.
Cell culture and treatment
Human breast cancer cell lines MCF-7, MDA-MB-231, and Hs578t (provided by Dr. Qi T. Yan, Southern Medical University, Guangzhou, China), were maintained in DMEM supplemented with 5% fetal bovine serum and 1 mM nonessential amino acids in a 5% CO2 incubator. Normal human mammary epithelial cells (HMEC 48R; provided by Dr. Qi T. Yan, Southern Medical University, Guangzhou, China) were maintained in MEGM (Cambrex Corp., USA) as previously described [10]. To confirm that methylation of the PTPRO promoter in breast cancer cell lines was responsible for its suppression, MCF-7 and MDA-MB-231 cells were treated with 5-azacytidine (5-AzaC, Sigma Chemical Co., HK), a DNA-hypomethylating agent, according to the following conditions: 1 μmol/L for 72 h for MCF-7 cells, and 2.5 μmol/L for 96 h for MDA-MB-231 cells. The response of different cell lines to demethylating agents probably varies due to different drug sensitivities as well as different kinetics of association/dissociation of chromatin remodelers with specific genes. All cells used in this study were between passages 8 and 11.
DNA extraction and bisulfite modification
Genomic DNA from primary tumors and plasma was extracted using a QIAamp DNA Mini Kit (Qiagen, Germany) and QIAamp DNA blood Mini Kit (Qiagen, Germany). Gene methylation status was evaluated using sodium bisulfite modification of DNA and subsequent methylation-specific PCR (MSP), essentially as previously described [24-26]. DNA (1–2 μg) from each sample was subjected to bisulfite modification using EpiTect 96 Bisulfite Kits according to the manufacturer’s instructions (Qiagen, Germany). Bisulfite-modified DNA was typically immediately used for PCR.
Methylation-specific PCR analysis
Primer sequences for PCR amplification of methylated and unmethylated alleles of PTPRO were previously published [10] and are listed in Table 1. Primers were synthesized by Shenggong (Shenggong Biotech, Shanghai, China). Primers were designed to amplify 170 bp (methylated) or 201 bp (unmethylated) regions of the CpG island within the PTPRO promoter [8]. For each reaction, 3 μl of sodium bisulfite- converted DNA was added to a total volume of 50 μl of PCR mix (EpiTect MSP Kits, Qiagen, Germany) according to the manufacturer’s instructions. Briefly, samples were initially incubated at 95°C for 10 min. This was followed by 35 cycles of denaturation at 95°C for 15 s, annealing at 55°C for 30 s, and extension at 72°C for 30 s; finally, there was one round of extension at 72°C for 10 min. An additional 15 cycles of denaturing (30 s at 94°C), annealing (15 s at 50.4°C), and extension (30 s at 72°C) were required for blood samples. PCR products were analyzed by electrophoresis on 2% agarose gels. Primers for unmethylated PTPRO (Table 1) were used to confirm the presence of DNA in each sample following bisulfite modification. This control was run for each sample on the same day that MSP analysis was carried out for the PTPRO gene. Breast tumor samples previously identified as DNA hypermethylated were used as positive controls. For each PCR assay, experimental reactions were accompanied by a black reaction (no DNA), a negative control reaction (blood DNA), and a positive control reaction (breast cancer DNA).
Table 1.
PCR primer sequences for methylation analysis of PTPRO
| Primers | Sequences | Product size |
|---|---|---|
| PTPRO-forward |
5′-CTCCACCCAAATCACTCTTCGCAG-3′ |
268 bp |
| PTPRO-reverse |
5′-ACCATTGTTGAGACGGCTATGAACG-3′ |
|
| 18 s rRNA-forward |
5′-TCAAGAACGAAAGTCGGAGG-3′ |
110 bp |
| 18 s rRNA- reverse |
5′-GGACATCTAAGGGCATCACA-3′ |
|
| MSP-methylated-forward |
5′-CGTTTTTGGAGGATTTCGGGC-3′ |
170 bp |
| MSP-methylated- reverse |
5′-AAAACACGACTACGCTAACG-3′ |
|
| MSP-unmethylated-forward |
5′-ATGTTTTTGGAGGATTTTGGGT-3′ |
201 bp |
| MSP-unmethylated- reverse | 5′-ATACCCCATCACTACACAAACA-3′ |
Bisulfite genomic sequencing
Bisulfite-converted DNA was used to PCR amplify the PTPRO CpG island from -208 bp to +236 bp with respect to the transcription start site as described earlier; ref. [7,8,19]. The PCR product was purified using a gel extraction kit (Qiagen, Germany). The purified PCR product was used for bisulfite sequencing and was cloned into the pDrive vector according to the instructions of the PCR cloning kit (Qiagen, Germany). Ten randomly selected clones were subjected to automated sequencing. Direct sequencing was performed using the Thermo Sequenase Radiolabeled terminator cycle sequencing kit (Qiagen, Germany) with the primer hGlepp1-BS-F3 (5′-TAGGGGGATTGGAAAGGTAG-3′) following the manufacturer’s protocol.
RNA isolation and reverse transcription PCR analysis
Total RNA was isolated using the RNeasy Mini kit (Qiagen, Germany). Reverse transcription of deoxyribonuclease-treated RNA (1 μg) was carried out according to instructions provided with the QuantiTect Reverse Transcription kit (Qiagen, Germany). Semi-quantitative PCR for PTPRO expression was performed. 0.2 mM of each primer was added to a 25 μl PCR reaction mixture. Cycling conditions were as follows: denaturation at 94°C, annealing at 54.5°C (for PTPRO) or 65°C (for 18S rRNA), and extension at 72°C. For PTPRO, a total of 32 cycles were run, and for18S rRNA, 25 cycles were used. The PCR products were separated on 2% agarose, stained with ethidium bromide, and imaged under UV light using Bio-rad Quantity One software. 18S rRNA transcripts in each sample were also amplified as internal controls for normalization. Gene-specific primers used for amplification of PTPRO and 18S rRNA are listed in Table 1.
Statistical analysis
The χ2 test was used to determine associations between methylation of PTPRO and various phenotypic or molecular features of breast cancer. Fisher’s exact test was used when individual cell numbers were less than 5. All P values were derived from two-tailed statistical tests and significance was assumed at P < 0.05. Kaplan–Meier analysis was used to assess cumulative survival probabilities, and differences were evaluated using the log-rank test. Multinominal logistic regression analyses were used to assess the hazard model for the survival of breast cancer patients. All analyses were performed using the SPSS 19.0 (Chicago, IL, US) statistical software package.
Results
Frequent methylation of PTPRO in primary tumors and peripheral blood samples
Methylation of the PTPRO promoter has been demonstrated in different tumor types, including breast cancer [7,8,10]. These observations, along with the growth-suppressive properties not only of PTPRO [7,8] but of PTPs in general [27], prompted us to further investigate PTPRO methylation status in a large series of human breast tumors. Genomic DNA isolated from tumor tissue, surrounding normal tissue, and matched peripheral blood samples (n = 98) was subjected to MSP analysis. Among the 98 primary breast tumor specimens investigated, 53 (54%) showed hypermethylation of PTPRO. Methylation of this gene was not observed in any adjacent normal tissues. 33 of 98 (34%) patients exhibited detectable levels of methylated PTPRO in matched plasma. No methylated PTPRO was observed in normal peripheral blood samples from 30 healthy individuals. The χ2 test revealed that PTPRO methylation in plasma was significantly correlated to that in tumor tissue (r = 0.435; P < 0.0001, Table 2). Representative MSP results from primary tumors are shown in Figure 1a, b.
Table 2.
Association of methylation of PTPRO gene between tumor tissues and plasma
|
Tumor tissues |
Plasma |
||
|---|---|---|---|
| M | U | Total | |
| M |
29 |
24 |
53 |
| U |
4 |
41 |
45 |
| Total | 33 | 65 | 98 |
M: Methylated; U: Unmethylated.
Figure 1.
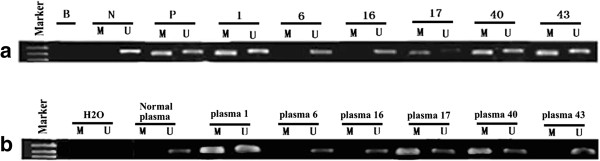
Representative MSP results for methylation of the PTPRO gene. (a) primary breast tumors; (b) matched peripheral blood samples. Numbers indicate the sample number. B, blank (no DNA); N, negative control; P, positive control; M, methylated; U, unmethylated.
Clinicopathological significance of PTPRO methylation in breast tumors and peripheral blood samples
Associations between PTPRO methylation and clinicopathological and molecular features of breast tumors in this study are shown in Table 3. The strongest correlation was between PTPRO methylation and HER2 amplification (P = 0.001). PTPRO methylation was also significantly more frequent in node positive (P = 0.014), poorly differentiated (P = 0.037), and stage III (P = 0.004) tumors. The highest frequency of PTPRO methylation (84%) was seen in late stage tumors. Trends were also observed for more frequent PTPRO methylation in ER- and PR-negative tumors but no associations were apparent with patient age, tumor size, histological tumor type, or TP53 mutation. Methylation of PTPRO in plasma was more frequent in those with HER2 amplification (P = 0.018). For all other clinical and pathological parameters, there was no statistically significant correlation associated with methylation of PTPRO in plasma (Table 3).
Table 3.
Associations between PTPRO methylation and clinicopathological features of breast cancer
|
Characteristics |
No. |
PTPRO
methylation |
|||||
|---|---|---|---|---|---|---|---|
| Tumor tissue (%) | χ 2 | P value | plasma (%) | χ 2 | P value | ||
| Total |
98 |
53 (54) |
|
|
33 (34) |
|
|
| Age |
|
|
|
|
|
|
|
| < 45 years |
45 |
22 (49) |
|
|
16 (36) |
|
|
| ≥ 45 years |
53 |
31 (59) |
0.903 |
0.417 |
17 (32) |
0.132 |
0.716 |
| Nodal involvement |
|
|
|
|
|
|
|
| Negative |
61 |
27 (44) |
|
|
18 (30) |
|
|
| Positive |
37 |
26 (70) |
6.273 |
0.014 |
15 (40) |
0.935 |
0.334 |
| Stage |
|
|
|
|
|
|
|
| I/II |
79 |
37 (47) |
|
|
24 (30) |
|
|
| III |
19 |
16 (84) |
8.616 |
0.004 |
9 (47) |
1.979 |
0.159 |
| Histological type |
|
|
|
|
|
|
|
| Non-ductal |
23 |
12 (52) |
|
|
7 (30) |
|
|
| Ductal |
75 |
41 (55) |
1.000 |
0.510 |
26 (35) |
0.141 |
0.707 |
| Tumour size |
|
|
|
|
|
|
|
| ≤20 mm |
47 |
26 (55) |
|
|
12 (26) |
|
|
| >20 mm |
51 |
27 (53) |
0.056 |
0.842 |
21 (40) |
2.234 |
0.135 |
| Histological grade |
|
|
|
|
|
|
|
| Well/mod diff. |
60 |
27 (45) |
|
|
18 (30) |
|
|
| Poorly diff. |
38 |
26 (68) |
5.139 |
0.037 |
15 (40) |
0.935 |
0.334 |
| ER status |
|
|
|
|
|
|
|
| Negative |
24 |
16 (67) |
|
|
10 (42) |
|
|
| Positive |
74 |
37 (50) |
2.027 |
0.167 |
23 (31) |
0.909 |
0.340 |
| PR status |
|
|
|
|
|
|
|
| Negative |
29 |
20 (69) |
|
|
10 (35) |
|
|
| Positive |
69 |
33 (48) |
3.674 |
0.076 |
23 (33) |
0.012 |
0.912 |
|
HER2 status |
|
|
|
|
|
|
|
| Normal |
51 |
19 (37) |
|
|
12 (24) |
|
|
| Amplified |
47 |
34 (72) |
12.124 |
0.001 |
21 (46) |
5.570 |
0.018 |
|
TP53 status |
|
|
|
|
|
|
|
| Normal |
59 |
30 (51) |
|
|
17 (29) |
|
|
| Mutant | 39 | 23 (59) | 0.624 | 0.535 | 16 (41) | 1.568 | 0.211 |
Prognostic significance of tumor tissue and plasma PTPRO methylation
Univariate analysis examined clinicopathologic parameters including PTPRO methylation and their association with overall survival end points. The results showed that survival was significantly worse in patients with lymph node involvement (P = 0.0001), late stage tumors (P = 0.0001), poorly differentiated tumors (P = 0.033), larger tumors (P = 0.019), positive HER2 amplification (P = 0.022), TP53 mutation (P = 0.012) and PTPRO methylation (hazard ratio (HR): 3.8; 95% CI: 1.9-7.5; P = 0.0001; Table 4). We then stratified all patients into subpopulations according to ER, PR and HER2 status. In ER- positive, PR- positive and HER2-positive patients, the methylated PTPRO group show significantly worse overall survival compared to those of unmethylated PTPRO (P = 0.001, P = 0.012 and P = 0.010, respectively, Table 4). Kaplan-Meier curves for overall tumor group and the above subgroups according to PTPRO methylation are shown in Figure 2. As shown, tumor tissue PTPRO methylation was associated with significantly worse cancer-specific survival in the overall tumor group (log-rank test P = 0.0001; Figure 2a). Subgroup analysis revealed that PTPRO methylation also showed significant prognostic value within the ER + (P = 0.0001), PR + (P = 0.007), and HER2-amplified (P = 0.003) patient groups (Figure 2b, c, and d, respectively).
Table 4.
Univariate and multivariate cox proportional hazard model for the survival of breast cancer patients
|
Variable |
Univariate analysis |
Multivariate analysis |
||||
|---|---|---|---|---|---|---|
| Hazard ratio | 95% CI | P | Hazard ratio | 95% CI | P | |
| Tumor size (large vs small) |
2.4 |
1.2-4.9 |
0.019 |
|
|
|
| Lymph node status (pos. vs neg.) |
4.9 |
2.4-9.8 |
0.0001 |
4.0 |
1.6 - 9.9 |
0.003 |
| Histological grade (poor vs well) |
2.1 |
1.1-4.0 |
0.033 |
|
|
|
| Stage (III vs I/II) |
3.2 |
1.7-5.9 |
0.0001 |
|
|
|
|
HER2 status (amp. vs wildtype) |
2.2 |
1.1-4.2 |
0.022 |
|
|
|
|
TP53 (mutant vs wildtype) |
2.3 |
1.2-4.3 |
0.012 |
|
|
|
| Tumor tissue PTPRO methylation (yes vs no) |
3.8 |
1.9-7.5 |
0.0001 |
2.7 |
1.1- 6.2 |
0.023 |
| ER + group tumor tissue PTPRO methylation (yes vs no) |
3.9 |
1.7-8.7 |
0.001 |
2.8 |
1.0-8.4 |
0.060 |
| PR + group tumor tissue PTPRO methylation (yes vs no) |
3.1 |
1.2-7.4 |
0.012 |
3.2 |
0.8-11.9 |
0.091 |
| HER2+ group tumor tissue PTPRO methylation (yes vs no) | 5.0 | 1.8-16.8 | 0.010 | 7.5 | 1.8-31.3 | 0.006 |
Figure 2.
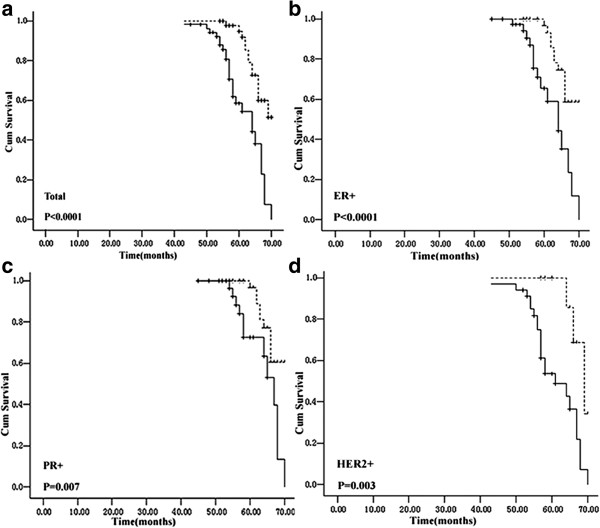
Kaplan–Meier survival analysis for breast cancer patients with (solid line) PTPRO tumor methylation or without (dotted line). (a) overall group; (b) ER+; (c) PR+; (d) HER2-amplified subgroup.
To confirm the significance of this finding, we performed multivariate analysis, treating methylated-PTPRO as a factor with tumor size, lymph node metastasis, histological grade, stage, HER2 status and TP53 status for their impact on overall survival. After adjustment for these convariates, methylated-PTPRO was identified as an independent predictor for overall survival in all tumor group (hazard ratio (HR): 2.7; 95% CI: 1.1-6.2; P = 0.023) and HER2+ subpopulation (hazard ratio (HR): 7.5; 95% CI: 1.8-31.3; P = 0.006), but not in ER + and PR + subpopulation. Similarly, lymph node metastasis also had an independent association with overall survival in this patient series. We also analyzed the potential prognostic value of plasma PTPRO methylation but no significant data were obtained.
PTPRO expression is inversely correlated with methylation status
We next sought to determine the relationship between PTPRO methylation and gene expression in a panel of breast cancer cell lines (MCF-7, MDA-MB-231, and Hs578t) and in normal human mammary epithelial cells (HMEC, 48R). In normal mammary epithelial cells, PTPRO is expressed at appreciable levels and its promoter region is not methylated; in contrast, PTPRO expression was relatively low in two (MCF-7, MDA-MB-231) of the three breast cancer cell lines examined and its promoter was methylated (Figure 3a, b).
Figure 3.
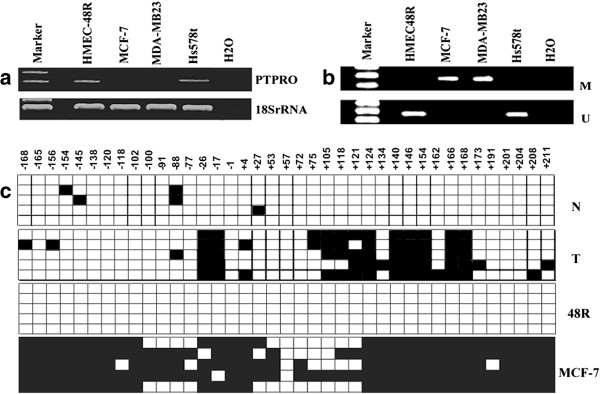
PTPRO is methylated in breast cancer cell lines but not in normal breast epithelial cells. (a) Expression of PTPRO in normal human mammary epithelial cells (48R) and human breast cancer cell lines Hs578t, MCF-7, and MDA-MB-231. Total RNA isolated from cell lines was subjected to RT-PCR analysis using PTPRO-specific primers. 18S rRNA was used as an internal loading control. (b) MSP analysis of PTPRO methylation status in breast cancer cell lines. HMESC48R was used as a normal control. M, methylated; U, unmethylated. (c) PTPRO CpG island from randomly selected breast tumor tissue and its matched normal tissue; also shown are HMEC 48R and MCF-7 cells, all of which were subjected to BS genomic sequencing. Each solid square represents a methylated cytosine and an open square represents unmethylated cytosine in a CpG dinucleotide. Each row corresponds to a single clone. N, normal corresponding adjacent non-cancerous tissue; T, tumor tissue.
We performed bisulfite genomic sequencing from ten pairs of breast tumor tissue and matched normal tissue, one representative HMEC (48R), and one breast cancer cell line (MCF-7) to determine if the CpG Island located in the promoter of PTPRO was differentially methylated. Complete bisulfite conversion was confirmed by the presence of substituted thymine for all cytosine residues at non-CpG sites. We detected hypermethylation of CpGs in both PTPRO-silenced tumors and in MCF-7 cells. In contrast, the PTPRO-expressing cell line HMEC 48R and matched normal breast tissue exhibited low levels or no methylation of the PTPRO promoter—strongly supporting the MSP results (Figure 3c).
To confirm that hypermethylation of the PTPRO promoter was responsible for its suppression, both MCF-7 and MDA-MB-231 cells (hypermethylated PTPRO promoter; silenced mRNA expression) were treated with 5-AzaC at a final concentration of 1 μM for MCF-7 cells and 2.5 μM for MDA-MB-231 cells. Re-expression of PTPRO in both cell lines was observed after exposure to this demethylating agent for 72 h and 96 h, respectively (Figure 4a). Moreover, the MSP result showed that unmethylated PTPRO alleles increased after 5-AzaC treatment (Figure 4b). These data further support the notion that methylation of the PTPRO CpG island plays an important role in gene silencing.
Figure 4.
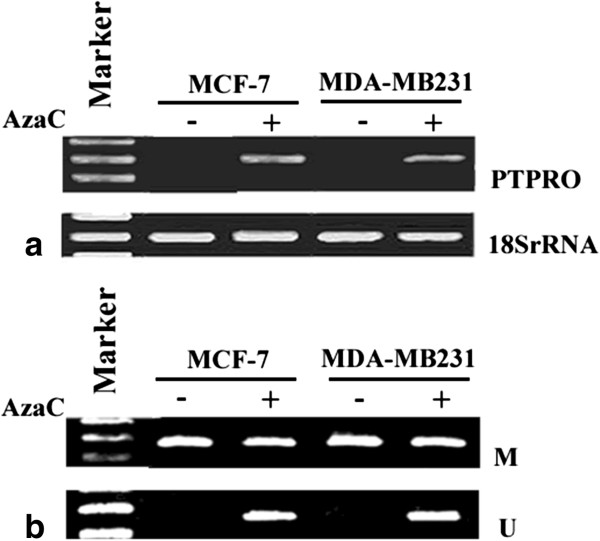
Re-expression of PTPRO following treatment with 5-AzaC. (a) Breast cancer cell lines MCF-7 and MDA-MB-231 were treated with 1 μM 5-AzaC for 72 h and 2.5 μM 5-AzaC for 96 h, respectively. Total RNA from cells was subjected to RT-PCR to amplify PTPRO mRNA. 18S rRNA was used for normalization; (b) MSP analysis of PTPRO methylation status in breast cancer cell lines with or without 5-AzaC treatment. M, methylated; U, unmethylated.
Discussion
Although protein tyrosine kinases have long been recognized as key players in oncogenesis, the role of protein tyrosine phosphatases in the initiation and progression of cancer is only now gaining increased attention [27-29]. In this study, the breast cancer series investigated here for DNA methylation is well characterized and conventional pathological indicators, including nodal involvement, histological grade, tumor size, and stage, all show the expected prognostic significance. PTPRO methylation was detected in two of three breast cancer cell lines and in 53 of 98 (54%) primary human breast cancer specimens; however, no PTPRO methylation was observed in adjacent normal tissue. This result is within the range (52% to 81%) reported in previous studies of human cancers [7-10,20,30]. The rather high frequency of methylation suggests that PTPRO is a common target for epigenetic silencing in breast tumors and that it may contribute to the development of this tumor type. As reported previously, demethylation of the PTPRO promoter resulted in gene re-expression [31]. These observations demonstrate growth-suppressor characteristics of PTPRO that are typical of a classical tumor suppressor gene.
Aberrant hypermethylation of tumor suppressor genes is an important epigenetic event in the development and progression of many human cancers and may serve as a biomarker for disease detection at early stages [32-34]. In this study, we detected PTPRO methylation in the plasma of 34% (33/98) of patients; this value was significantly correlated with PTPRO methylation detected in tumor tissue. Such a high correlation confirmed that peripheral blood samples could potentially be used to assist the detection and diagnosis of breast cancer. Moreover, this assay appears to be robust and highly specific; no methylated PTPRO was detected in plasma from breast cancer patients without primary tumor methylation or from normal healthy control peripheral blood samples. These findings are consistent with results published by Huang et al. [21] who also examined PTPRO methylation in peripheral blood samples from breast cancer cases. Among 24 matched plasma samples, PTPRO was aberrantly methylated in 11 (45.8%) cases. Importantly and consistent with our findings, no methylation was observed in normal control plasma samples from 10 healthy individuals. These data help confirm that PTPRO methylation in plasma samples may provide a robust, specific, non-invasive means for early detection of breast cancer. However, the frequency of PTPRO methylation detected in plasma was lower than in cancer tissues and less association of methylation were found in plasma with clnicopathological data. This might due to fewer tumors DNA releasing in the circulation, or poor quality of DNA when extracted from peripheral blood, whose impact factors include acquisition condition, storage time, human factor, etc. Our method of detecting PTPRO methylation from plasma may not be extremely robust. The more standard conditions and a larger series of breast cancer patients should be involved for more understanding the molecular mechanism and clinical behavior of these tumors, as well as provide targets for better diagnosis and therapy. For sure, a more robust method must be used if this is translated to clinical application.
In agreement with You et al. [20], we found a strong correlation between PTPRO methylation and tumor stage (Table 2), with 84% of stage III tumors found to be methylated. Similar to Huang et al. [21], PTPRO methylation correlated with higher histological grade. The current study is the first to report an association between PTPRO methylation and positive lymph node status and HER2 amplification in breast cancer. We also observed more frequent PTPRO methylation in ER-negative and PR-negative patient groups, possibly due to the association between these features and poor prognosis. Interestingly, Ramaswamy et al. [10] found that positive PTPRO expression was associated with improved response to tamoxifen; these results are consistent with previous reports of protein tyrosine phosphatase gene (PTPG) [35,36]. Therefore, estrogen-mediated suppression of PTPRO and the methylation of this gene may play important roles in estrogen-induced tumorigenesis. While interesting, each of the above associations with PTPRO methylation requires confirmation in larger studies. Moreover, it remains to be established whether the characteristic aggressive phenotype is linked to methylation via silencing of gene expression or through other mechanisms.
The significant associations between PTPRO methylation and nodal involvement, poorly differentiated histology, stage III tumors, and HER2 amplification suggest that PTPRO expression may be involved in breast tumor invasion. Given these aforementioned correlations, it is not surprising that PTPRO methylation served as a prognostic indicator of worse outcome. Although PTPRO methylation was weakly associated with ER- and PR- status, these factors had no prognostic value in the current tumor series (data not shown). Similar to Huang et al. [21], unmethylated PTPRO was significantly associated with favorable outcome in ER + and PR + subgroups, as well as in patients with HER2 amplification. As reported, activation of ER results in multiple downstream effects [37]. Recent studies indicate that ERβ expression is decreased in human neoplastic breast tissue, suggesting that ERβ may be an inhibitor of tumorigenesis [38-40]. For clinically apparent tumors, the proposed tumor-associated factors may help protect against tumor progression. Thus, according to prior studies, inactive PTPRO might be a stimulating factor during tumorigenesis, explain the ineffection of endocrine therapy and more precise subpopulations could be stratified to decide whether the patients with ER-positive need a regimen containing tamoxifen.
In a univariate model including strong prognostic factors such as nodal status, histological grade, tumor size, stage, HER2 amplification, and TP53 mutation status, PTPRO methylation of overall tumors, ER+, PR + and HER2+ group was found to be predictive of poorer outcome for breast cancer. Multivariate analysis identified methylated-PTPRO as an independent predictor for overall survival (P = 0.023), expecially in HER2+ subpopulation (P = 0.006). In contrast to our findings, Huang et al. reported that PTPRO methylation only correlated with higher histological grade but not with any other clinical parameters assessed [21]. This could be due to differences in sample size or to differences in sample processing. For example, while we used fresh tumor tissue, Huang et al. made use of formalin-fixed paraffin-embedded samples. Despite this, the trend is still in the same direction. That is, PTPRO methylation and low expression are associated with worse prognostic features, especially for HER2-positive patients. Further supporting our claim is work from another group showing that the receptor tyrosine kinase ErbB2/HER2 is a direct substrate of PTPRO, and low levels of PTPRO expression correlated with reduced survival of HER2-positive breast cancer patients [11]. This may also help explain why plasma PTPRO methylation was only significantly associated with HER2 amplification. The data we present here, in conjunction with earlier work, establish PTPRO as a likely tumor suppressor in breast cancer. Moreover, PTPRO methylation status might predict response to anti-HER-targeted therapies in HER2-positive patients, even provide extensive survival benefits or improve the efficiency of targeted drugs due to active PTPRO. To further study, patients who receive targeted therapy are required.
Conclusion
In summary, our results confirm that PTPRO methylation is detected at a high frequency in breast cancer, occurring at a higher rate than either TP53 mutation or HER2 amplification. Positive associations with nodal involvement, poorly differentiated histology, and HER2 amplification indicate that PTPRO methylation may contribute to an aggressive breast tumor phenotype. This was particularly evident for ER+, PR+, and HER2-amplified breast cancer subgroups, which all showed that PTPRO methylation in tumor tissues was a strong prognostic factor. Methylated-PTPRO could serve as an independent predictor for overall survival, expecially in HER2-positive breast cancer patients. Changes in protein tyrosine phosphatase activity likely play an important role in breast carcinogenesis and may provide a useful target for the development of novel therapies.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
SL and RL developed the study and drafted the manuscript. YC and LR participated in sample collection and data analysis. LX and HW carried out the molecular genetic studies and participated in sequence alignment. *RL participated in the design of the study and its coordination and helped draft the manuscript. All authors read and approved the manuscript.
Contributor Information
Shao-ying Li, Email: charlenesyli@126.com.
Rong Li, Email: ynlirong@vip.sina.com.
Yu-li Chen, Email: cheng.yuli@wo.com.cn.
Li-kuang Xiong, Email: xionglk@sina.cn.
Hui-lin Wang, Email: 155320028@qq.com.
Lei Rong, Email: 652121691@qq.com.
Rong-cheng Luo, Email: 273334556@qq.com.
Acknowledgements
We thank Professor Barry Iacopetta from the School of Surgery, University of Western Australia for his critical reading of the manuscript. We also thank Dr. Qi T. Yan for his gracious gift of cell lines. The authors are grateful to Professor Tasneem Motiwala for information on primer sequences. This work was supported by the Science and Technology Planning Project of Shenzhen China Grant 201103049 (SY Li).
References
- Simpson PT, Reis-Filho JS, Gale T, Lakhani SR. Molecular evolution of breast cancer. J Pathol. 2005;205:248–254. doi: 10.1002/path.1691. [DOI] [PubMed] [Google Scholar]
- Curigliano G, Spitaleri G, Dettori M, Locatelli M, Scarano E, Goldhirsch A. Vaccine immunotherapy in breast cancer treatment:Promising, but still early. Expert Rev Anticancer Ther. 2007;7:1225–1241. doi: 10.1586/14737140.7.9.1225. [DOI] [PubMed] [Google Scholar]
- Emens LA, Reilly RT, Jaffee EM. Augmenting the potency of breast cancer vaccines: Combined modality immunotherapy. Breast Dis. 2004;20:13–24. doi: 10.3233/bd-2004-20103. [DOI] [PubMed] [Google Scholar]
- Dworkin AM, Huang TH, Toland AE. Epigenetic alterations in the breast: Implications for breast cancer detection, prognosis and treatment. Semin Cancer Biol. 2009;19:165–171. doi: 10.1016/j.semcancer.2009.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hesson LB, Cooper WN, Latif F. The role of RASSF1A methylation in cancer. Dis Markers. 2007;23:73–87. doi: 10.1155/2007/291538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motiwala T, Ghoshal K, Das A, Majumder S, Weichenhan D, Wu YZ, Holman K, James SJ, Jacob ST, Plass C. Suppression of the protein tyrosine phosphatase receptor type O gene (PTPRO) by methylation in hepatocellular carcinomas. Oncogene. 2003;22:6319–6331. doi: 10.1038/sj.onc.1206750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motiwala T, Majumder S, Kutay H, Smith DS, Neuberg DS, Lucas DM, Byrd JC, Grever M, Jacob ST. Methylation and silencing of protein tyrosine phosphatase receptor type O in chronic lymphocytic leukemia. Clin Cancer Res. 2007;13:3174–3181. doi: 10.1158/1078-0432.CCR-06-1720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motiwala T, Kutay H, Ghoshal K, Bai S, Seimiya H, Tsuruo T, Suster S, Morrison C, Jacob ST. Protein tyrosine phosphatase receptor-type O (PTPRO) exhibits characteristics of a candidate tumor suppressor in human lung cancer. Proc Natl Acad Sci U S A. 2004;101:13844–13849. doi: 10.1073/pnas.0405451101. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- Hogan LE, Meyer JA, Yang J, Wang J, Wong N, Yang W, Condos G, Hunger SP, Raetz E, Saffery R, Relling MV, Bhojwani D, Morrison DJ, Carroll WL. Integrated genomic analysis of relapsed childhood acute lymphoblastic leukemia reveals therapeutic strategies. Blood. 2011;118:5218–5226. doi: 10.1182/blood-2011-04-345595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramaswamy B, Majumder S, Roy S, Wang J, Wong N, Yang W, Condos G, Hunger SP, Raetz E, Saffery R, Relling MV, Bhojwani D, Morrison DJ, Carroll WL. Estrogen-mediated suppression of the gene encoding protein tyrosine phosphatase PTPRO in human breast cancer: mechanism and role in tamoxifen sensitivity. Mol Endocrinol. 2009;23:176–187. doi: 10.1210/me.2008-0211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu M, Lin G, Arshadi N, Kalatskaya I, Xue B, Haider S, Nguyen F, Boutros PC, Elson A, Muthuswamy LB, Tonks NK, Muthuswamy SK. Expression profiling during mammary epithelial cell three-dimensional morphogenesis identifies PTPRO as a novel regulator of morphogenesis and ErbB2-mediated transformation. Mol Cell Biol. 2012;32:3913–3924. doi: 10.1128/MCB.00068-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunter T. Signaling–2000 and beyond. Cell. 2000;100:113–127. doi: 10.1016/S0092-8674(00)81688-8. [DOI] [PubMed] [Google Scholar]
- Fischer EH. Cell signaling by protein tyrosine phosphorylation. Adv Enzyme Regul. 1999;39:359–369. doi: 10.1016/S0065-2571(98)00014-4. [DOI] [PubMed] [Google Scholar]
- Laczmanska I, Sasiadek MM. Tyrosine phosphatases as a superfamily of tumor suppressors in colorectal cancer. Acta Biochim Pol. 2011;V58N4:467–470. [PubMed] [Google Scholar]
- Gupta S, Radha V, Sudhakar C, Swarup G. A nuclear protein tyrosine phosphatase activates p53 and induces caspase-1-dependent apoptosis. FEBS Lett. 2002;532:61–66. doi: 10.1016/S0014-5793(02)03628-1. [DOI] [PubMed] [Google Scholar]
- Salmeen A, Andersen JN, Myers MP, Tonks NK, Barford D. Molecular basis for the dephosphorylation of the activation segment of the insulin receptor by protein tyrosine phosphatase 1B. Mol Cell. 2000;6:1401–1412. doi: 10.1016/S1097-2765(00)00137-4. [DOI] [PubMed] [Google Scholar]
- Alonso A, Sasin J, Bottini N, Friedberg I, Friedberg I, Osterman A, Godzik A, Hunter T, Dixon J, Mustelin T. Protein tyrosine phosphatases in the human genome. Cell. 2004;117:699–711. doi: 10.1016/j.cell.2004.05.018. [DOI] [PubMed] [Google Scholar]
- Andersen JN, Mortensen OH, Peters GH, Drake PG, Iversen LF, Olsen OH, Jansen PG, Andersen HS, Tonks NK, Møller NP. Structural and evolutionary relationships among protein tyrosine phosphatase domains. Mol Cell Biol. 2001;21:7117–7136. doi: 10.1128/MCB.21.21.7117-7136.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacob ST, Motiwala T. Epigenetic regulation of protein tyrosine phosphatases: potential molecular targets for cancer therapy. Cancer Gene Ther. 2005;12:665–672. doi: 10.1038/sj.cgt.7700828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- You YJ, Chen YP, Zheng XX, Meltzer SJ, Zhang H. Aberrant methylation of the PTPRO gene in peripheral blood as a potential biomarker in esophageal squamous cell carcinoma patients. Cancer Lett. 2012;315:138–144. doi: 10.1016/j.canlet.2011.08.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang YT LIFF, Ke C, Li Z, Li ZT, Zou XF, Zheng XX, Chen YP, Zhang H. PTPRO promoter methylation is predictive of poorer outcome for HER2-positive breast cancer: indication for personalized therapy. J Transl Med. 2013;11:245. doi: 10.1186/1479-5876-11-245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sobin LH, Wittekind C. International Union Against Cancer (UICC), 5th ed., TNM classification of malignant tumors. Baltimore (MD): Wiley-Liss; 1997. pp. 54–58. [Google Scholar]
- Abd El-Rehim DM, Bal G, Pinder SE, Rakha E, Paish C, Robertson JF, Macmillan D, Blamey RW, Ellis IO. High-throughput protein expression analysis using tissue microarray technology of a large well-characterised series identifies biologically distinct classes of breast cancer confirming recent cDNAexpression analyses. Int J Cancer. 2005;116:340–350. doi: 10.1002/ijc.21004. [DOI] [PubMed] [Google Scholar]
- Li SY, Rong MN, Iacopetta B. DNA hypermethylation in breast cancer and its association with clinicopathological features. Cancer Lett. 2006;237:272–280. doi: 10.1016/j.canlet.2005.06.011. [DOI] [PubMed] [Google Scholar]
- Herman JG, Graff JR, Myohanen S, Nelkin BD, Baylin SB. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci U S A. 1996;93:9821–9826. doi: 10.1073/pnas.93.18.9821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paulin R, Grigg G, Davey MW, Piper AA. Urea improves efficiency of bisulphate-mediated sequencing of 50-methylcytosine in genomic DNA. Nucleic Acids Res. 1998;26:5009–5010. doi: 10.1093/nar/26.21.5009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motiwala T, Jacob ST. Role of protein tyrosine phosphatases in cancer. Prog Nucleic Acid Res Mol Biol. 2006;81:297–329. doi: 10.1016/S0079-6603(06)81008-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tonks NK. Protein tyrosine phosphatases: from genes, to function, to disease. Nat Rev Mol Cell Biol. 2006;7:833–846. doi: 10.1038/nrm2039. [DOI] [PubMed] [Google Scholar]
- Ostman A, Hellberg Cand Bohmer FD. Protein-tyrosine phosphatases and cancer. Nat Rev Cancer. 2006;6:307–320. doi: 10.1038/nrc1837. [DOI] [PubMed] [Google Scholar]
- Hsu SH, Motiwala T, Roy S, Claus R, Mustafa M, Plass C, Freitas MA, Ghoshal K, Jacob ST. Methylation of the PTPRO gene in human hepatocellular carcinoma and identification of VCP as its substrate. J Cell Biochem. 2013;114:1810–1818. doi: 10.1002/jcb.24525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motiwala T, Majumder S, Ghoshal K, Kutay H, Datta J, Roy S, Lucas DM, Jacob ST. PTPROt inactivates the oncogenic fusion protein BCR/ABL and suppresses transformation of K562 cells. J Biol Chem. 2009;284:455–464. doi: 10.1074/jbc.M802840200. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- Jin Z, Cheng Y, Olaru A, Kan T, Yang J, Paun B, Ito T, Hamilton JP, David S, Agarwal R, Selaru FM, Sato F, Abraham JM, Beer DG, Mori Y, Shimada Y, Meltzer SJ. Promoter hypermethylation of CDH13 is a common, early event in human esophageal adenocarcinogenesis and correlates with clinical risk factors. Int J Cancer. 2008;123:2331–2336. doi: 10.1002/ijc.23804. [DOI] [PubMed] [Google Scholar]
- Jin Z, Hamilton JP, Yang J, Mori Y, Olaru A, Sato F, Ito T, Kan T, Cheng Y, Paun B, David S, Beer DG, Agarwal R, Abraham JM, Meltzer SJ. Hypermethylation of the AKAP12 promoter is a biomarker of Barrett’sassociated esophageal neoplastic progression. Cancer Epidemiol Biomarkers Prev. 2008;17:111–117. doi: 10.1158/1055-9965.EPI-07-0407. [DOI] [PubMed] [Google Scholar]
- Jin Z, Olaru A, Yang J, Sato F, Cheng Y, Kan T, Mori Y, Mantzur C, Paun B, Hamilton JP, Ito T, Wang S, David S, Agarwal R, Beer DG, Abraham JM, Meltzer SJ. Hypermethylation of tachykinin-1 is a potential biomarker in human esophageal cancer. Clin Cancer Res. 2007;13:6293–6300. doi: 10.1158/1078-0432.CCR-07-0818. [DOI] [PubMed] [Google Scholar]
- Liu S, Sugimoto Y, Sorio C, Tecchio C, Lin YC. Function analysis of estrogenically regulated protein tyrosine phosphatase (PTPs) in human breast cancer cell line MCF-7. Oncogene. 2004;23:1256–1262. doi: 10.1038/sj.onc.1207235. [DOI] [PubMed] [Google Scholar]
- Zheng J, Kulp SK, Zhang Y, Sugimoto Y, Dayton MA, Govindan MV, Brueggemeier RW, Lin YC. 17-Estradiol-regulated expression of protein tyrosine phosphatase gene in cultured human normal breast and breast cancer cells. Anticancer Res. 2000;20:11–19. [PubMed] [Google Scholar]
- Henderson BE, Feigelson HS. Hormonal carcinogenesis. Carcinogenesis. 2000;21:427–433. doi: 10.1093/carcin/21.3.427. [DOI] [PubMed] [Google Scholar]
- Dotzlaw H, Leygue E, Watson PH, Murphy LC. Estrogen receptor messenger RNA expression in human breast tumor biopsies: relationship to steroid receptor status and regulation by progestins. Cancer Res. 1999;59:529–532. [PubMed] [Google Scholar]
- Iwao K, Miyoshi Y, Egawa C, Ikeda N, Noguchi S. Quantitative analysis of estrogen receptor-β mRNA and its variants in human breast cancers. Int J Cancer. 2000;88:733–736. doi: 10.1002/1097-0215(20001201)88:5<733::AID-IJC8>3.0.CO;2-M. [DOI] [PubMed] [Google Scholar]
- Roger P, Sahla ME, Makela S, Gustafsson JA, Baldet P, Rochefort H. Decreased expression of estrogen receptor-β protein in proliferative preinvasive mammary tumors. Cancer Res. 2001;61:2537–2541. [PubMed] [Google Scholar]


