Abstract
Intestinal stem cells (ISCs) in the adult Drosophila midgut can respond to tissue damage and support repair. We used genetic manipulation to increase the number of ISC-like cells in the adult midgut and performed gene expression profiling to identify potential ISC regulators. A detailed analysis of one of these potential regulators, the zinc-finger protein Charlatan, was carried out. MARCM clonal analysis and RNAi in precursor cells showed that loss of Chn function caused severe ISC division defects, including loss of EdU incorporation, phosphorylated histone 3 staining and expression of the mitotic protein Cdc2. Loss of Charlatan also led to a much reduced histone acetylation staining in precursor cells. Both the histone acetylation and ISC division defects could be rescued by the simultaneous decrease of the Histone Deacetylase 2. The overexpression of Charlatan blocked differentiation reversibly, but loss of Charlatan did not lead to automatic differentiation. The results together suggest that Charlatan does not simply act as an anti-differentiation factor but instead functions to maintain a chromatin structure that is compatible with stem cell properties, including proliferation.
Keywords: Drosophila, Intestine, Stem cells, Chromatin, Charlatan, Zinc-finger protein
INTRODUCTION
Tissue homeostasis requires a balance between removing dead cells and producing new cells. An imbalance between the two processes can lead to tissue degeneration, inflammation and cancer. Replenishment of lost cells in an adult tissue is usually mediated by resident stem cells. Elucidating the mechanisms that regulate stem cell-mediate homeostasis and regeneration may provide insights into the development of therapeutics for various diseases (Nakada et al., 2011).
The mammalian intestine is essentially a monolayer of epithelial cells with fast cell turnover rate and the tissue integrity is maintained by active stem cells located near the crypt base (Rizk and Barker, 2012). Two overlapping or inter-converting stem cell populations give rise to progenitor cells in the transit-amplifying zone that can replenish cells of various lineages along the crypt-villus axis (Buczacki et al., 2013; Takeda et al., 2011; Tian et al., 2011; Yan et al., 2012).
The adult Drosophila midgut is equivalent to the mammalian stomach and small intestine. The midgut epithelium is largely a monolayer of enterocytes (ECs) and does not have crypt-villus structure. Approximately 1000 intestinal stem cells (ISCs) are distributed evenly along the basal side of the epithelium (Micchelli and Perrimon, 2006; Ohlstein and Spradling, 2006). An ISC divides to generate a renewed ISC and an enteroblast (EB), which ceases division and starts to differentiate. The ISC-EB asymmetry is governed by the Delta-Notch signaling, with high levels of Delta in the renewed ISC activating Notch signaling in the neighboring EB (Bardin et al., 2010; Ohlstein and Spradling, 2007) (see Fig. 1I). Depending on the strength of stimulation on the Notch pathway, the EB may differentiate to become an EC (in wild-type gut 90% of the time) or enteroendocrine cell (EE) (10% of the time) (Micchelli and Perrimon, 2006; Ohlstein and Spradling, 2006, 2007).
Fig. 1.
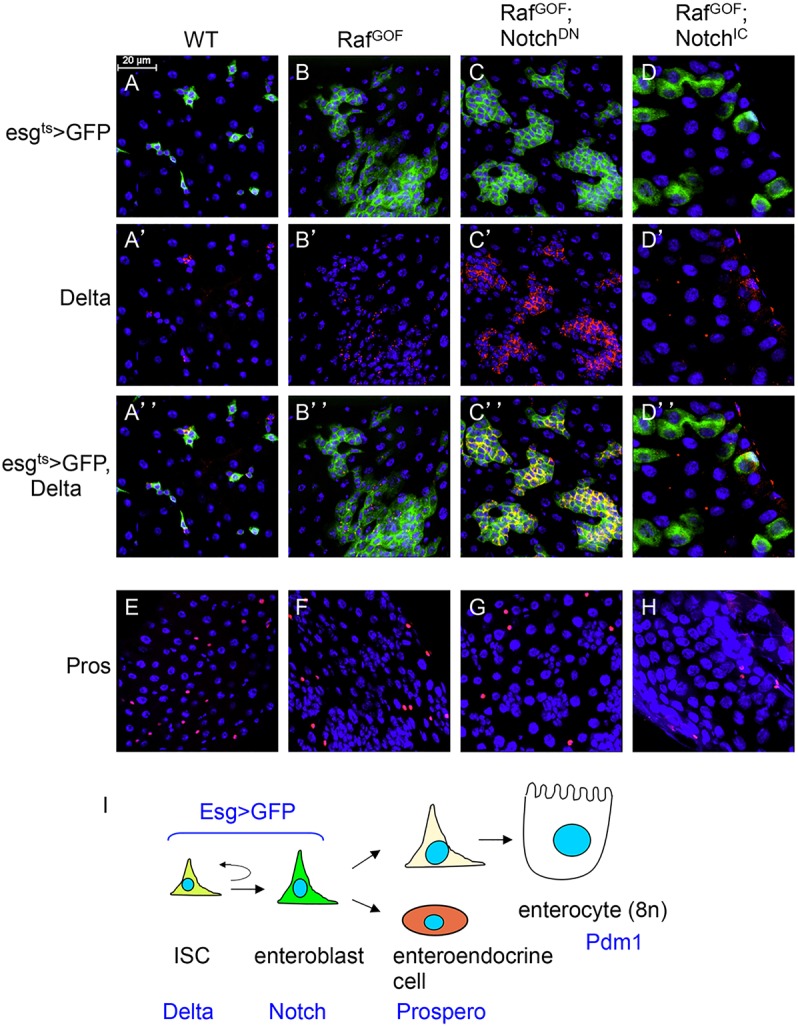
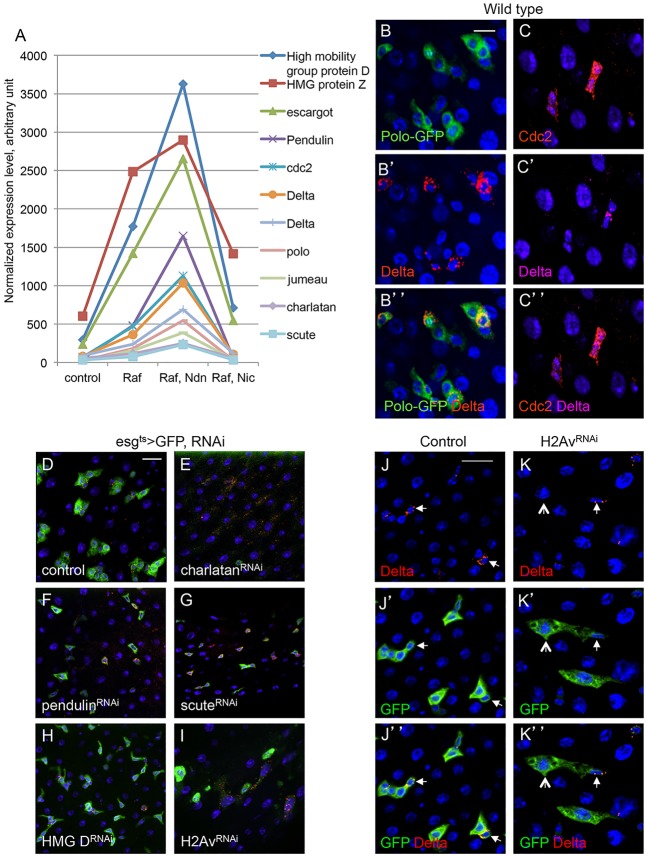
Genetic conversion of precursors into stem-like cells. (A-H) The driver line genotype is esg-Gal4, UAS-GFP; tubulinGal80ts (esgts>GFP). This driver was crossed with w– as wild-type (WT) control. The other genotypes include UAS-RafGOF located on the X chromosome, together with UAS-NotchDN or UAS-NotchIC. The progenies of the crosses were kept at 18°C. After hatching for 5 days, the flies of the correct genotype combinations were transferred to 29°C for 36 h, which would inactivate the Gal80ts repressor and allow Gal4 to function and initiate UAS-driven expression. The guts were then dissected and subjected to immunofluorescent staining with antibodies for Delta or Prospero. The red is Delta staining, which is mostly punctate and cytoplasmic (A′-D′). The red nuclear staining is Prospero (E-H). Blue staining is DAPI for nuclear DNA; green is GFP. Scale bar in A for A-H: 20 μm. (I) The cell types and markers in the adult midgut. ISC, intestinal stem cell; EB, enteroblast; EE, enteroendocrine cell; EC, enterocyte. Delta, Su(H)LacZ (Notch pathway target gene), Prospero and Pdm1 are markers for the respective cell types. The esg>GFP is expressed in both ISC and EB.
The cells surrounding ISC constitute the niche and secrete growth factors to regulate ISC maintenance and activity. The visceral muscle and mature ECs are sources of Wingless, insulin-like peptides, epidermal growth factor receptor (EGFR) ligands and JAK-STAT pathway ligands called Unpaired (Upd) (Biteau and Jasper, 2011; Buchon et al., 2010; Jiang et al., 2010; Lin et al., 2008; O'Brien et al., 2011; Ragab et al., 2011; Xu et al., 2011). Moreover, the differentiating EBs contribute Upd, Wingless and EGFR ligands to regulate intestinal homeostasis (Cordero et al., 2012; Jiang et al., 2010; Zhou et al., 2013). Recent reports also reveal the secretion of Decapentaplegic/BMP from trachea and ECs to regulate ISC activity (Guo et al., 2013; Li et al., 2013a,b; Tian and Jiang, 2014). Hedgehog signal coming from multiple cell types serves a negative regulatory function in ISC division (Li et al., 2014). Other conserved signaling pathways, including JNK, p38, PVF2 and Hippo, are also required for the regulation of ISCs during homeostasis, tissue damage and aging (Biteau et al., 2008; Bond and Foley, 2012; Jiang et al., 2009; Karpowicz et al., 2010; Park et al., 2009; Ren et al., 2010; Shaw et al., 2010; Staley and Irvine, 2010).
Many ISC-intrinsic factors have been shown to regulate asymmetry and renewal. Osa (as part of the SWI/SNF complex) and the Brahma chromatin remodeling complex regulate Delta expression and ISC proliferation (Jin et al., 2013; Zeng et al., 2013). The histone deubiquitinase Scrawny and growth regulators [including Myc, Target of Rapamycin (TOR) and tuberous sclerosis complex (TSC)] are also required for ISC growth and division (Amcheslavsky et al., 2011; Buszczak et al., 2009; Ren et al., 2013). To search for other intrinsic ISC regulators, we used genetic manipulation to increase the number of ISC-like cells in the adult midgut and performed gene expression profiling. The results revealed many possible proteins that can modulate the function of ISCs or precursor cells (defined as ISCs+EBs). We carried out an in depth analysis of the zinc-finger protein Charlatan (Chn) because its loss of function shows a severe defect in ISC division. Our data further demonstrate that Chn regulation of chromatin is essential for ISC division.
RESULTS
Genetic conversion of precursors into stem-like cells in the adult midgut
Although sorting individual cell types from adult midgut for genomic analysis is possible (Dutta et al., 2014), the number of ISCs present in the midgut is low and the available ISC-specific Delta-Gal4 driver exhibits weak expression (Zeng et al., 2010). Moreover, stem cells may change their properties if dissociated from the niche. Therefore, we artificially increased the pool of ISC-like cells in the midgut by expressing via Gal4/UAS a gain-of-function Raf (RafGOF) and a dominant-negative Notch (NotchDN) (Bardin et al., 2010; Markstein et al., 2014; Micchelli and Perrimon, 2006; Ohlstein and Spradling, 2006, 2007). RafGOF increases the proliferation of ISC, whereas NotchDN skews the asymmetry to produce precursor cells that are mostly Delta positive (Delta+). The escargot promoter-Gal4 together with the tubulin-Gal80ts (esgts>) provide temporal control, and were used to drive the expression in precursor cells; the UAS-mCD8GFP was included to mark the cells expressing the constructs. In control guts, after a temperature shift to 29°C for 36 h, the GFP+ cells appeared frequently as small clusters with usually one Delta+ cell per cluster, representing the ISC and EB cell nests (Fig. 1A-A″). In guts that overexpressed RafGOF, there was a substantial increase in GFP+ precursor cells, indicating high proliferation (Fig. 1B-B″). Overexpression of NotchDN together with RafGOF resulted in not only high numbers of GFP+ cells but also Delta+ staining of these ISC-like cells (Fig. 1C-C‴). By contrast, overexpression of the constitutively active Notch intracellular domain (NotchIC) together with RafGOF caused the formation of larger GFP+ cells that had no Delta staining, indicating a loss of stem cell fate and an increase in differentiation (Fig. 1D-D‴). The number of EEs, as revealed by the Prospero staining, was not affected within the time course of these experiments (Fig. 1E-H). These results demonstrate that we can create fly intestines with highly increased number of Delta+ ISC-like cells or with mostly differentiated cells that have no Delta staining.
Identification of transcripts differentially expressed in the midguts with varying number of ISC-like cells
We used dissected guts, from the cardia to the midgut/hindgut junction excluding malpighian tubules and crops, from the above-described genetic combinations for gene expression profiling. Total RNA was isolated from control, RafGOF, RafGOF+NotchDN and RafGOF+NotchIC fly guts. Then cRNA probes were prepared and hybridized to high-density GeneChip Drosophila Genome 2.0 Arrays (Affymetrix). This type of microarray contains 18,880 probe sets and is able to measure the expression of 18,500 transcripts and variants. Four independent RNA isolation and microarray hybridization were performed.
The expression profile of each genetic background was compared with that of the esgts>GFP control. Based on four stringent criteria (see Materials and Methods), a total of 1127 transcripts with significantly increased or decreased expression were identified. To investigate the relationships among the transcripts that showed significant changes, we first carried out a two-dimensional hierarchical clustering analysis (Fig. 2A). This type of clustering algorithm arranges genes or samples according to their similarity in expression profiles across all of the genes or all of the experiments, such that samples or genes with similar expression patterns are clustered together (Eisen et al., 1998; He et al., 2003). The analysis reveals extensive similarity in all three experimental samples, suggesting that many may be potential Raf target genes because RafGOF was used in all three experimental strains. The genes that showed opposite expression in the NotchDN versus NotchIC samples may represent Notch pathway target genes.
Fig. 2.
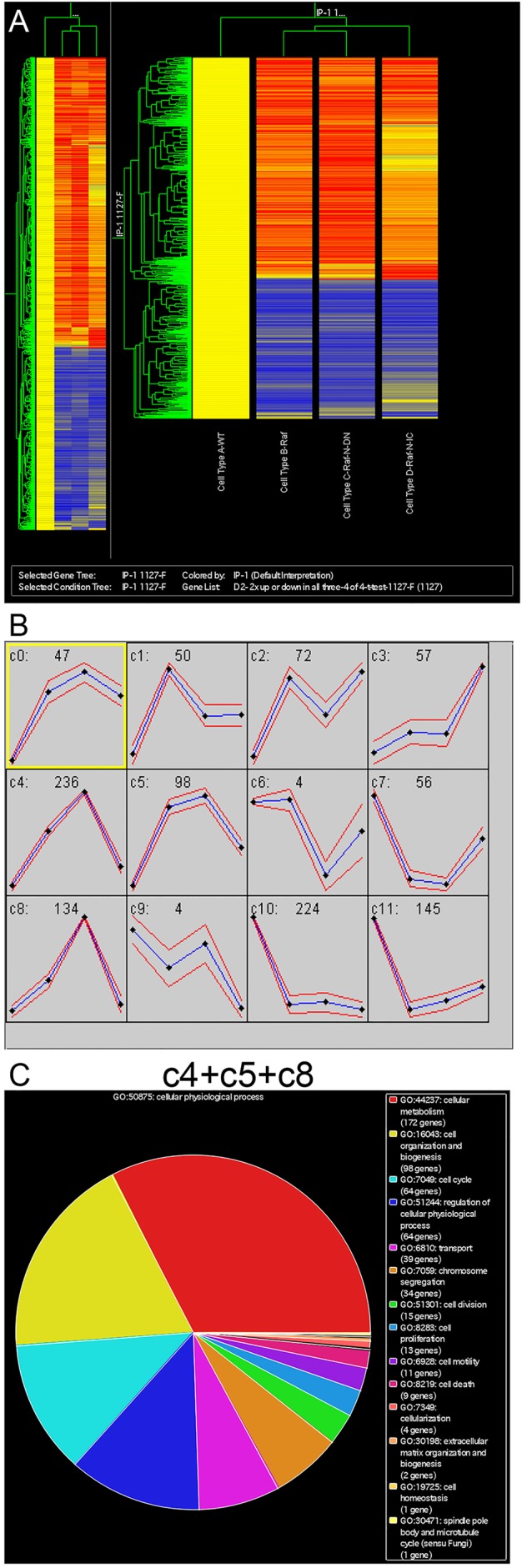
Identification of transcripts that are differentially expressed in the midguts with varying number of ISC-like cells. (A) RNA isolated from adult midguts of the genotypes described in Fig. 1 were used for microarray analysis. 1127 genes showed significant increases or decreases in expression when compared with the wild type. These 1127 genes were analyzed by using a hierarchical clustering algorithm (GeneSpring) in two dimensions, according to gene and experiment vectors. In this analysis, the intensity signal of each transcript in the Raf, Raf+NotchDN and Raf+NotchIC backgrounds was normalized to that in the wild-type background. The resulting expression ratios from four independent replicates were averaged, transformed to base two logarithms and clustered using the standard correlation coefficient as a distance metric. Genes were plotted along the vertical axis, with the gene cluster on the left. Experimental samples are plotted along the horizontal axis, with the experiment cluster tree at the top of the plot. Color coding is used to represent the fold-change in expression. Red indicates transcripts with increased expression levels in the experimental condition relative to the wild-type sample; blue indicates transcripts with reduced expression levels. (B) Self-organizing map (SOM) analysis of the 1127 genes. The intensity signal of each transcript in the experimental sample was normalized to that in the wild-type sample. The resulting expression ratios from four independent replicates were averaged, normalized by standardizing each row (gene) to mean=0 and variance=1. Transcripts were then grouped into 12 clusters (a 3×4 SOM). For each cluster, the y-axis represents the relative expression levels and the horizontal axis represents different experimental conditions in the order (from left to right): 1, wild type; 2, Raf; 3, Raf+NotchDN; 4, Raf+NotchIC. In each cluster, the mean expression level is represented as a blue line and standard deviation is the red line. (C) Gene Ontology analysis of the transcripts from clusters c4, c5 and c8 of SOM in B. The transcripts are classified into different categories based on their involvement in different physiological processes.
We then derived self-organizing maps (SOM) for our data sets (Fig. 2B). The SOM algorithm clusters genes or transcripts into distinct groups or classes based on the similarity of their relative expression profiles in the different genetic backgrounds (Tamayo et al., 1999). Such analysis yielded 12 expression clusters (c0 to c11). The first noteworthy pattern is low expression in wild-type guts, higher in RafGOF alone, highest in RafGOF+NotchDN and lower in RafGOF+NotchIC. Clusters c4, c5 and c8 match this description the best, and they may represent ISC- or precursor-enriched genes. Based on gene ontology analysis, we can further classify most of these genes as involving in cellular metabolism, cell organization, cell cycle, cellular transport and chromosome segregation (Fig. 2C).
A total of 468 genes are included in clusters c4, c5 and c8. Based on our own scientific experience and the preference of higher fold change, we have selected a set of 63 genes that may represent novel or serve as examples of ISC regulators (Table 1). The expression profile of all 1127 genes grouped as clusters is available at http://dx.doi.org/10.6084/m9.figshare.855619.
Table 1.
Representative genes from a total of 468 genes in clusters c4, c5 and c8
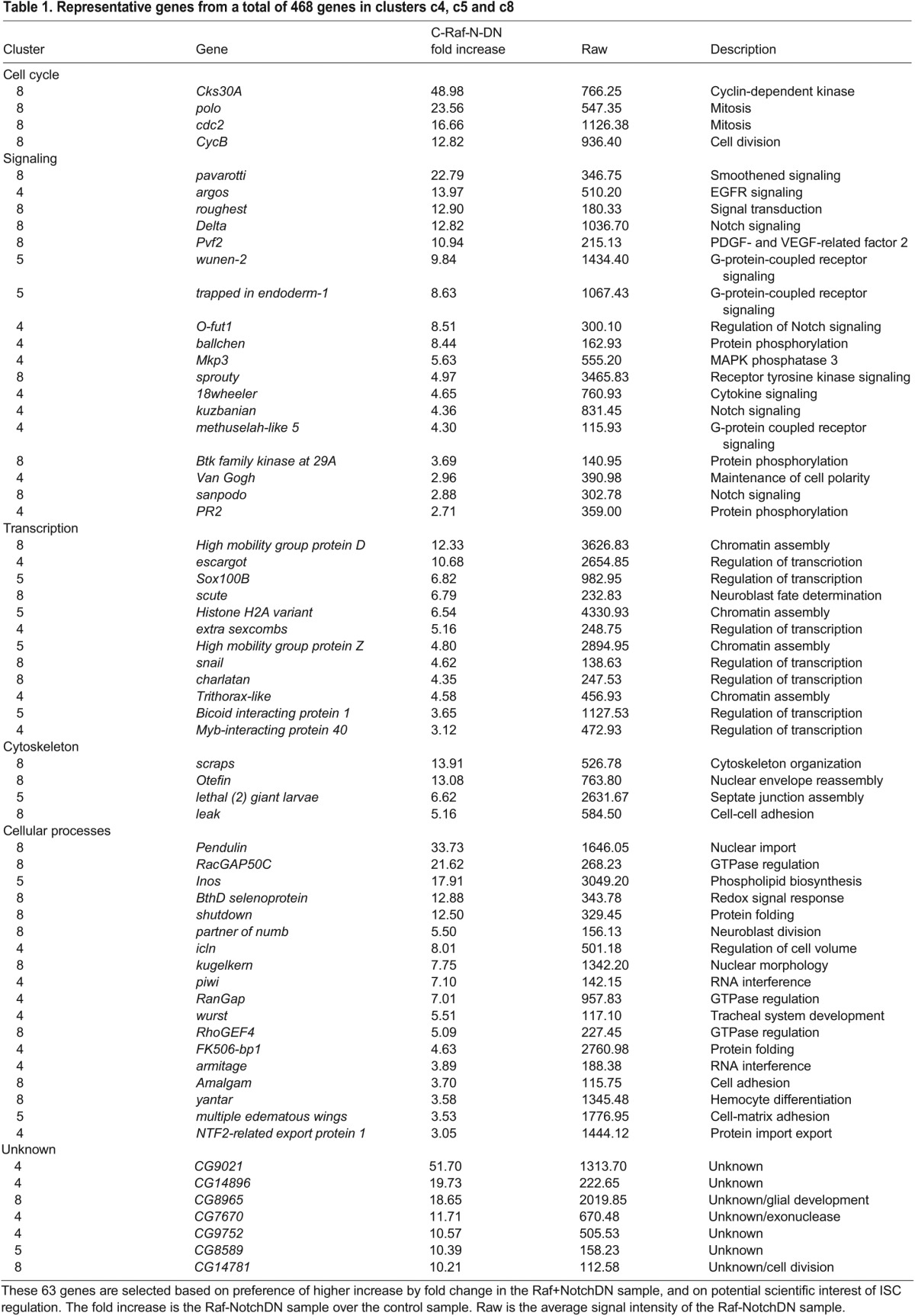
Expression and function of representative genes associated with ISCs or precursors
Delta protein is a well-characterized ISC-specific marker in the Drosophila midgut, whereas esg is well known for its expression in the precursors. From our analysis, both esg and Delta showed their expression profiles belonging to the clusters c4, c5 and c8. Furthermore, ISCs are the only dividing cells in the adult Drosophila midgut and many of the genes in the c4, c5 and c8 clusters are cell cycle-associated genes, including Polo, Cdc2, Pendulin, Cyclin B and string (http://dx.doi.org/10.6084/m9.figshare.855619 and Fig. 3A). By using a protein trap line (FlyTrap CC01326), we confirmed that the Polo-GFP fusion protein was detectable by GFP antibody staining in Delta+ ISCs, as well as in the neighboring EBs (Fig. 3B). Cdc2 is another essential component for mitotic entry, and the Cdc2 protein was detectable using antibody staining in Delta+ ISCs and in adjacent EBs (Fig. 3C). These results further suggest that the transcripts in the c4, c5 and c8 clusters are associated with precursor cells in the midgut.
Fig. 3.
Expression and function of representative genes associated with ISCs or precursors. (A) The plot shows normalized expression of selected genes from SOM clusters c4, c5 and c8, in which the well-known ISC/precursor cell markers Delta and escargot are included. Average values from the four microarray experiments are shown. Genes in this set have diverse molecular functions. (B-C″) Expression analysis of the cell-cycle regulators Polo and Cdc2 in wild-type midguts. The Polo-GFP is a FlyTrap line in which the GFP-coding sequence is fused with the endogenous Polo-coding sequence. The expression was detected by anti-GFP immunofluorescence staining. The Polo-GFP was detected in Delta+ cells and neighboring EBs. Cdc2 antibody staining for the endogenous protein was similarly present in both Delta+ ISCs and adjacent EBs. (D-I) Functional analysis by RNAi of potential ISC regulators. Transgenic RNAi flies for chn, pendulin, scute, HMG D and H2Av were crossed with esgts>GFP. Control was a cross with w– flies. Adult flies ∼5 days old were placed at 29°C for 3 days more and the midguts were dissected for imaging. (J-K″) H2Av RNAi midguts stained for Delta expression showing that ISCs still had Delta expression (arrows), but GFP+ cell nests had lower cell numbers and the neighboring EB appeared to be larger in size (arrowheads in K).
We used the esg-Gal4 driver to perform RNAi knockdown of 10 of the transcripts identified from clusters c4, c5 and c8. Four lines that targeted RhoGEF4, kugelkern, ballchen and roughest did not show detectable phenotype, whereas six RNAi lines showed mild to severe phenotype (Fig. 3D-I). The knockdown of the transcript for the zinc-finger protein Charlatan (Chn) resulted in the most severe loss of the esg>GFP signal that marked the precursors (Fig. 3E). Pendulin as an Armadillo repeat-containing protein has similarity with Importin and may play a role in nuclear import during proliferation (Küssel and Frasch, 1995); the knockdown of pendulin led to an observable reduction of the esg>GFP signal (Fig. 3F). Scute has been shown to be an upstream activator of Chn in the developing peripheral neurons (Escudero et al., 2005; Reeves and Posakony, 2005), and knockdown of scute caused a mild defect in the midgut precursors (Fig. 3G). The high-mobility group proteins (HMG D and Z) function to regulate the compaction of chromatin (Ragab et al., 2006); the knockdown of HMG D alone caused only a mild disorganization, although we did not attempt the double RNAi for D and Z (Fig. 3H). Another chromatin protein Histone 2A variant (H2Av) serves as a signal for the recruitment of repair proteins at the site of DNA damage (Madigan et al., 2002; Park et al., 2012). The knockdown of H2Av showed a severe disruption of midgut organization (Fig. 3I). Further analysis of H2Av RNAi-induced phenotype showed that the Delta+ ISCs (arrows in Fig. 3J,K) did not divide as efficiently so the cell nests were smaller, whereas the EBs had bigger cell size, indicating abnormal differentiation (open arrows in Fig. 3K). Therefore, a sample of the genes identified from the profiling have biological functions in ISCs or precursors.
Chn is required cell-autonomously for ISC division
We performed an in-depth analysis of chn because the RNAi of this transcript exhibited a severe defect in ISCs/EBs. Chn regulates peripheral neuron development by forming a positive regulatory loop with the achaete-scute complex genes (Escudero et al., 2005; Kania et al., 1995; Reeves and Posakony, 2005). The predicted Chn protein has 1108 amino acids and contains five C2H2-type zinc fingers. Chn has no extensive homology with any mammalian proteins, but the sequence of the Chn zinc fingers has 35% identity and 55% similarity to those of the human zinc-finger protein 462. Meanwhile, the functional property of Chn as a transcriptional repressor in photoreceptor neurons resembles that of another human zinc-finger protein, NRSF/REST (Escudero et al., 2005; Tsuda et al., 2006).
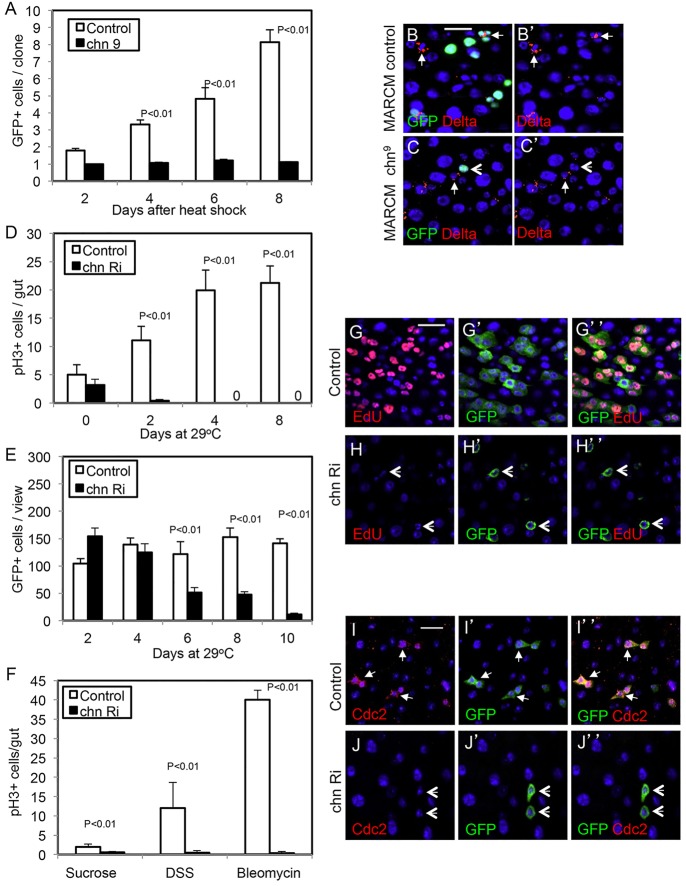
A strong loss-of-function allele chn9 (Tsuda et al., 2006) was used to perform a clonal analysis by the mosaic analysis of repressible cell marker (MARCM) technique. chn9 mutant ISCs marked by GFP expression remained as single cells, whereas wild-type clones had increased cell numbers over the same period of 8 days (Fig. 4A). Moreover, chn9 mutant cells remained small and the Delta staining was not detectable, within 2 days of clone induction (Fig. 4B,C). We also analyzed in further detail the adult midgut precursors after chn RNAi using the esgts> driver. The number of mitotic cells in the midgut, as revealed by staining for phosphorylated histone 3 (pH3) was reduced after 2 days. A concomitant gradual decrease of GFP+ cells that represent precursors was observed (Fig. 4D,E). This mitotic defect was still dominant when two different damaging agents were used to stimulate ISC division (Fig. 4F). These defects were corroborated with the lack of EdU incorporation, which reveals both ISC replication and EB endoreplication, and the loss of staining of the mitotic marker Cdc2 (Fig. 4G-J). The inclusion of a transgenic chn cDNA rescued the RNAi-induced Delta staining and cell proliferation phenotype (supplementary material Fig. S1A-D). Together, these results demonstrate that Chn has an intrinsic function in ISCs to maintain their ability to divide.
Fig. 4.
Chn is required cell-autonomously for ISC division. (A-C′) Wild-type and chn9 mutant MARCM clones were generated after 37°C heat shock of the flies to induce FLP expression and mitotic recombination. The flies were returned to 29°C for the number of days indicated and midguts were dissected for analysis. (A) The average number of GFP+ cells per clone in chn9 mutant clones (black bars) is significantly lower than in wild-type clones (white bars) at various time points. Data are mean±s.e.m. (B,B′) A 4-day-old wild-type clone; Delta staining indicates ISCs (arrows). (C,C′) A 4-day-old chn9 mutant clone, which shows no detectable Delta staining (arrowheads). (D) The control flies were esgts>GFP crossed with w– flies, and the chn Ri were esgts>GFP crossed with UAS-chnRNAi flies. The strong reduction in mitotic activity in chn RNAi fly midguts becomes apparent as early as 2 days after shifting to 29°C. Data are mean±s.e.m. (E) Similar experiments were carried out and GFP+ cells were counted from microscopic images taken from posterior midguts with a 40× objective. (F) The control and chn RNAi flies were fed in 5% sucrose, or supplemented with 3% DSS or 25 μg/ml bleomycin. After 3 days of feeding, the guts were dissected and stained for pH3. (G-H′) The control and chn RNAi flies were incubated in 29°C for a total of 8 days. EdU was added to a 5% sucrose solution for feeding the flies for the last 24 h. Guts were dissected and used for EdU detection using the Click-iT detection kit. Approximately 90% of GFP+ cells in control guts also contained EdU signal. No EdU signal was detected in GFP+ cells of chn RNAi midguts (arrowheads in H-H″). (I-J″) Similar experiments were performed and the guts were stained for the mitotic regulator Cdc2. The staining was absent from the GFP+ cells after chn RNAi (arrowheads in J-J″).
High levels of Chn antagonize differentiation reversibly
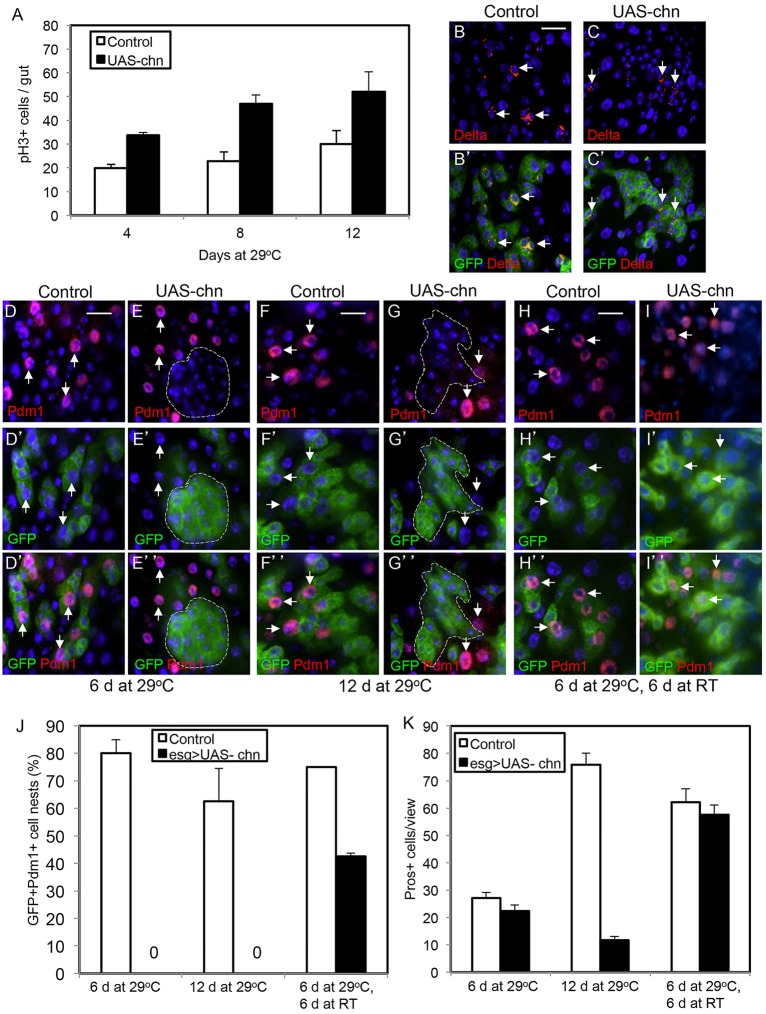
We then tested whether increasing the expression level of Chn might increase ISC division. A UAS-chn transgene (Tsuda et al., 2006) was crossed with the esgts>GFP and adult flies were shifted to 29°C to allow Gal4-driven expression of Chn. There was an approximately 1.5-fold increase of pH3+ cells in the midguts at every time point examined (Fig. 5A). The number of GFP+ cells also increased, whereas the number of Delta+ cells remained similar (Fig. 5B,C, arrows). These results suggest that increased expression of Chn promotes stem cell division but does not change the stem cell number by altering the asymmetric division.
Fig. 5.
High level of Chn antagonizes differentiation reversibly. (A-C) The control flies were esgts>GFP crossed with w–flies, and the UAS-chn were esgts>GFP crossed with a fly strain containing a UAS-chn transgene. Midguts were dissected from the flies after incubation at 29°C for the indicated time. Staining for pH3 was performed and the number of positive cells is shown in A. Data are mean±s.e.m. The Delta staining in both control and UAS-chn guts after the flies were incubated at 29°C for 4 days are shown in B and C (some examples of Delta+ cells are indicated by arrows). (D-I″) The control and UAS-chn flies were incubated at 29°C for 6 (D-E″) or 12 (F-G″) days. The downshift experiments were carried out by incubating at 29°C for 6 days and at room temperature (∼23°C) for 6 days (H-I″). The guts were dissected and stained for the EC differentiation marker Pdm1. The arrows indicate examples of Pdm1+ nuclear staining. The encircled clusters of GFP+ cells in the UAS-chn samples contain no detectable Pdm1 staining. In the downshift experiments, many of the GFP+ cells became Pdm1+. (J) Quantification of GFP+ cells clusters that also contained at least 1 Pdm1+ cell, also expressed as a percentage of all GFP+ cells clusters. UAS-chn expression after incubation at 29°C suppressed all the expression of Pdm1 in GFP+ cells, and the differentiation marker could be detected again when the Chn expression was stopped at room temperature for 6 days. Data are mean±s.e.m. (K) The same experiments were performed and stained for the EE marker Prospero. The Prospero+ cells do not overlap with the esg>GFP+ cells. Graph shows that the number of Prospero+ cells after 29°C incubation is similar in control and UAS-chn guts because they were already present before temperature shift. Incubation for 12 days allowed more EEs to accumulate in control guts, but the accumulation was inhibited when Chn was overexpressed. The downshift experiment showed that the differentiation of EE resumed when Chn overexpression was stopped after 6 days. Data are mean±s.e.m.
Tissue homeostasis requires the coordination of stem cell division and daughter cell differentiation. Although overexpression of Chn increased the number of GFP+ daughter cells, these cells had relatively small nuclei and were tightly associated with each other (Fig. 5C,C′). This suggests that they behave as precursor-like cells and may be arrested at an early stage of differentiation. The marker Pdm1 is expressed in late EBs and mature ECs (Fig. 5D,F, arrows) (Zhou et al., 2013). The GFP+ cells in Chn overexpression guts did not express Pdm1 (Fig. 5E, encircled). This arrest of differentiation could continue for 12 days (Fig. 5G, encircled). If we downshifted to the permissive 23°C from day 6 to 12 to allow the Gal80ts to function again to inhibit Gal4-driven Chn expression, then a substantial number of GFP+ cells became Pdm1+ (Fig. 5H-J for quantification). By using the EE marker Prospero, we also found that the number of Prospero+ cells was lower after Chn overexpression but increased again after temperature downshift (Fig. 5K). These results demonstrate that high levels of Chn can block differentiation, but the process can resume when Chn overexpression is halted.
Loss of Chn does not lead to precocious differentiation
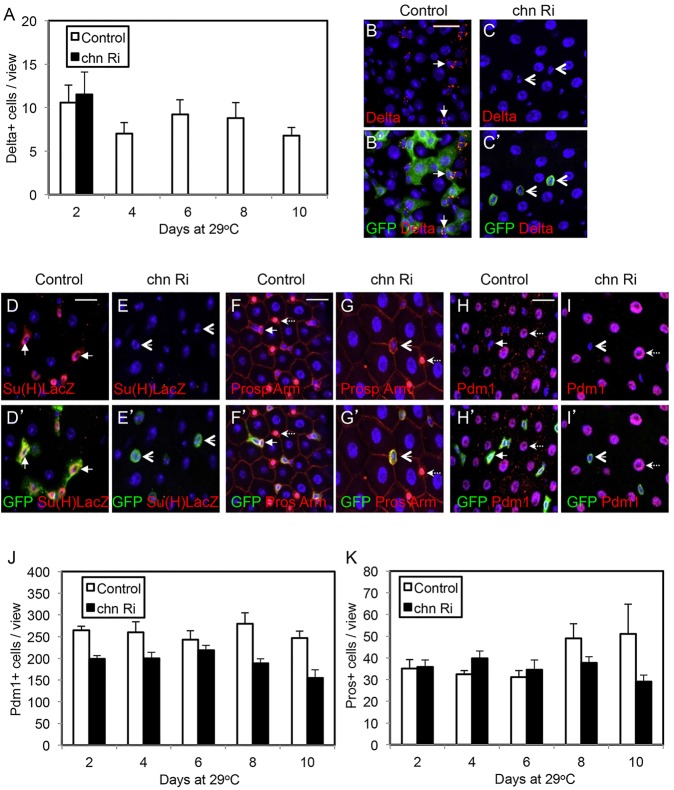
Some stem cell maintenance factors function by antagonizing differentiation (Fuchs and Chen, 2013; Stergiopoulos and Politis, 2013). Even though we show in Fig. 5 that overexpressed Chn can block differentiation, if the endogenous Chn acts by such a mechanism, the loss of Chn should lead to differentiation at the expense of stem cell number. We carried out staining with antibodies for various cell fate markers. Similar to that in the chn9 mutant clones, the expression of Delta rapidly disappeared after esgts>chn RNAi (Fig. 6A-C). The GFP+ cells after chn RNAi remained small and did not show the Su(H)LacZ staining, suggesting that they are not differentiating EBs (Fig. 6D,E). Moreover, the overall number of Pdm1+ cells or Prospero+ cells in chn RNAi guts did not increase at any time point examined and decreased slightly over time (Fig. 6F-K). These results suggest that, in the absence of Chn, the ISCs merely lose their stem cell properties but do not automatically enter differentiation. The data shown in Figs 4-6 together suggest a model in which the normal function of Chn is to promote ISC and precursor cell properties, while differentiation is governed by a separate set of factors that are not directly regulated but can be antagonized by Chn.
Fig. 6.
Loss of Chn does not cause precocious differentiation. The control flies were esgts>GFP crossed with w– flies, and the chn Ri were esgts>GFP crossed with UAS-chnRNAi flies. (A) The hatched flies were transferred to 29°C for the days as indicated and guts were dissected and stained for Delta. The number of Delta+ cells is significantly reduced after induction of chn RNAi (black bars) compared with control midguts (white bars). No Delta+ cells in chn RNAi samples were detected after day 4. Data are mean±s.e.m. The samples in B-C′ are from flies incubated at 29°C for 6 days. The arrows indicate examples of Delta+ staining and arrowheads indicate GFP+ cells with no Delta staining after chn RNAi. (D-E′) Similar experiments after an 8-day incubation at 29°C. The size of the GFP+ cells in RNAi midguts remained small but no expression of the EB reporter Su(H)LacZ was detected (arrowheads). (F-G′) The esg>GFP+ cells (arrows) and the nuclear Prospero staining (dashed arrows) normally do not overlap in control guts. After chn RNAi, the GFP+ cells were still separate from Prospero+ cells (arrows versus dashed arrows). (H-I′) Similarly, the esg>GFP+ cell nests (arrows) did not express Pdm1, which was detectable in differentiated ECs (dashed arrows). After chn RNAi, the GFP+ cells still did not express Pdm1 (arrowheads). (J,K) Quantification of differentiated cells in control and chn RNAi guts. Images under a 40× objective were taken from the posterior midguts of control and RNAi flies. Graph shows the average numbers of Pdm1+ ECs and Prospero+ EEs at different time points after incubation of the flies at 29°C. Data are mean±s.e.m.
Chn regulates the chromatin of precursor cells
Previous reports show that Chn represses Delta gene expression in the eye imaginal discs and binds to promoter sequences of Delta and other genes in vitro (Tsuda et al., 2006; Yamasaki et al., 2011). However, the direct repression of Delta by Chn does not correlate with our midgut phenotypic analysis, because Chn is required positively within the ISCs for Delta expression. Moreover, Chn is essential for ISC division, whereas Delta is required for establishing ISC/EB asymmetry but not for ISC division (Ohlstein and Spradling, 2007). Therefore, we surmised that Chn might regulate other ISC properties that indirectly influence Delta expression.
To search for other possible mechanisms, we stained the midgut with various markers. The JAK-STAT is a major growth regulatory pathway in the midgut and the 10XSTAT-GFP reporter in individual precursor cells did not show an obvious decrease after chn RNAi (supplementary material Fig. S1E-J). Meanwhile, the acetylated histone 3 (AcH3) staining that was normally higher in precursor cells became undetectable after chn RNAi (Fig. 7A,B). HP-1 is associated not only with heterochromatin and gene silencing but also euchromatin to positively regulate transcription (Agarwal et al., 2007; Kwon and Workman, 2011). In the midgut, HP-1 was detected in all cell types, mainly as one strong focus per cell (Fig. 7C), as previously described (Lee et al., 2011). In esgts>chn RNAi midguts, the HP-1 staining in the precursor cells became weaker and was present as more foci (Fig. 7D). Higher magnification confocal images revealed that this abnormal HP-1 staining correlated closely with the DNA staining (supplementary material Fig. S2A,B). In comparison, the HP-1 staining remained similar to that in control samples when the Myo1A-Gal4 driver was used to induce chn RNAi in ECs (Fig. 7E,F). Moreover, we stained for H3K4me3, another chromatin marker associated with transcriptional initiation. This marker showed only a slight decrease after esgts>chn RNAi (supplementary material Fig. S2C-D). These results suggest that Chn is required specifically in precursor cells, but not in mature ECs, to maintain a normal chromatin structure, especially that marked by histone acetylation.
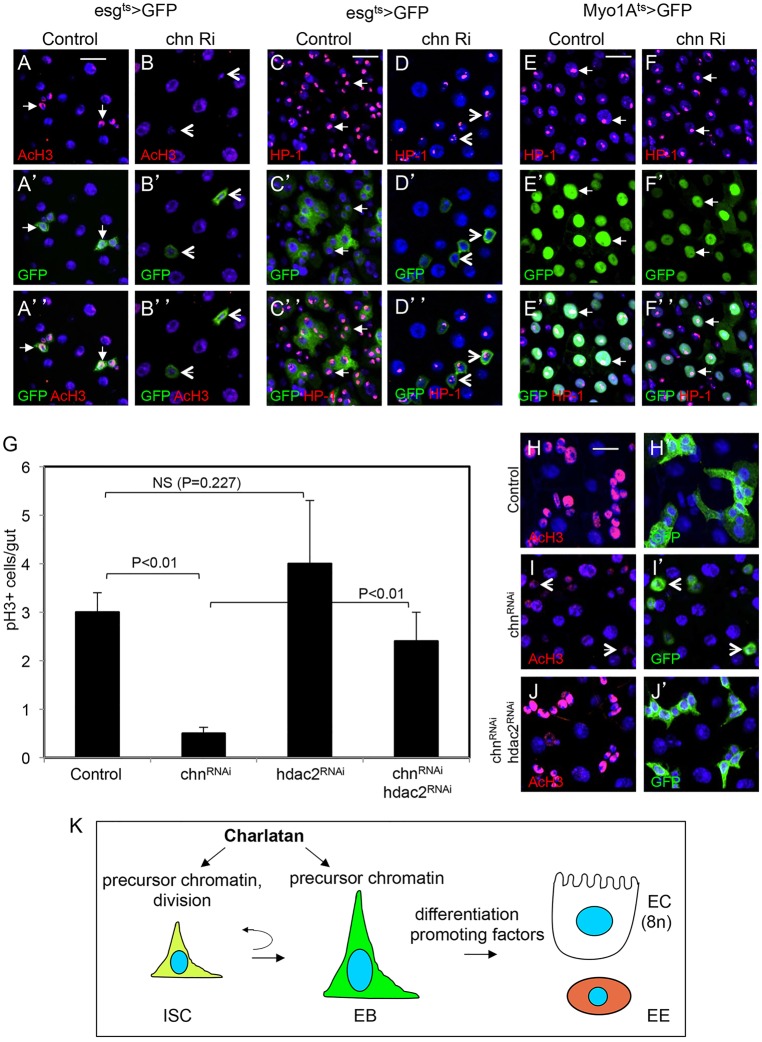
Fig. 7.
Chn regulates the chromatin of precursor cells. (A-D) The control flies were esgts>GFP crossed with w– flies, and the chn Ri were esgts>GFP crossed with UAS-chnRNAi flies. Five-day-old flies were transferred to 29°C for 4 days and the midguts analyzed by staining for acetylated histone 3 (AcH3) (A-B″) and Heterochromatin Protein 1 (HP1) (C-D″). Arrows indicate normal cells with positive staining and arrowheads indicate chn RNAi cells with no AcH3 staining or with abnormal HP-1 staining. (E-F″) Myo1Ats>GFP crossed with w– was the control, in which the HP-1 staining was detected in all cells, including enterocytes (arrows). When this enterocyte driver was crossed with chn RNAi, the HP-1 staining appeared to be similar to control samples (arrows). Flies were incubated for 8 days at 29°C before dissection and staining. (G) Quantification of the number of mitotic cells in the midgut with the indicated RNAi constructs driven by the esgts>GFP. The progenies of the crossed flies were allowed to hatch and age for 5 days and were then shifted to 29°C for 4 days before dissection and staining for pH3. NS is not significant, with P>0.05. Data are mean±s.e.m. (H-J′) Representative confocal images of midguts from the indicated RNAi experiments after staining for AcH3. Arrowheads in I,I′ indicate examples of GFP+ RNAi cells with no AcH3 staining. The staining reappeared in J when hdac2 was also knocked down. (K) A model for stem cell maintenance by Chn. Chn is required to maintain a chromatin structure compatible with being precursor cells. Loss of Chn, however, does not cause automatic differentiation, indicating that loss of stemness and initiation of differentiation can be a multi-step process.
We assessed the functional importance of the reduced histone acetylation observed in the Chn-depleted precursor cells. Various transgenic RNAi constructs targeting the transcripts of different histone deacetylases (HDACs) were crossed together with the chn RNAi construct (supplementary material Fig. S2E). There was a significant rescue of the mitotic cell count, particularly when HDAC2 was also knocked down (supplementary material Fig. S2E and Fig. 7G). The antibody staining for overall histone acetylation was visibly improved after the double knockdown of HDAC2 and chn (Fig. 7H-J). The HP-1 and DNA staining was also visibly improved after the double knockdown (supplementary material Fig. S2F-H). Because reducing the expression of an HDAC is sufficient to rescue most of the chn RNAi-induced midgut phenotypes, it suggests that Chn-mediated regulation of histone acetylation is an event upstream of other chromatin modification and is essential for ISC division.
DISCUSSION
We have used microarray analysis of genetically transformed adult Drosophila midguts to identify and characterize potential ISC/precursor cell regulators. A total of 1127 genes showed significant changes in at least one of the modified genetic backgrounds when compared with control midguts (http://dx.doi.org/10.6084/m9.figshare.855619). Analysis of the data in the various backgrounds has identified a group of genes (in the clusters c4, c5 and c8 in Fig. 2) that should have enriched expression in ISCs or precursors. Examples of these genes are Delta and escargot, two genes already well known to exhibit this expression pattern. Because ISCs are the only mitotic cells in the midgut, it is logical to expect that mitotic regulators should have enriched expression in ISCs. A substantial number of genes in clusters c4, c5 and c8 are indeed involved in cell cycle or mitotic regulation. Moreover, the RafGOF used probably pushed the cell cycle to become much faster and further increased the presence of these mitosis-associated transcripts in the ISC-like cells. Overall, the analyses presented in this report suggest that our approach has identified potentially important ISC or precursor regulators.
In depth analysis of one of the genes on the list, Chn, demonstrates its function to modulate the chromatin that is essential for ISC division and for EB precursor state (Fig. 7K). Previous reports suggest that Chn as a zinc-finger protein binds to the enhancers of achaete-scute complex genes, as well as other genes, including Delta (Escudero et al., 2005; Kania et al., 1995; Reeves and Posakony, 2005; Tsuda et al., 2006; Yamasaki et al., 2011). Further results suggest that Chn can function during development as either an activator or repressor of transcription in a context-dependent manner (Yamasaki et al., 2011). However, the binding sequences of Chn are highly divergent. Therefore, Chn may also have another means by which to regulate gene expression. We show that the prominent histone H3 acetylation in midgut precursor cells is dependent on Chn. Furthermore, the rescue of this acetylated H3 pattern by knocking down HDAC2 correlates with the rescue of ISC division.
A recent study reported that the epigenetic stability of Drosophila follicle stem cells increases during differentiation (Skora and Spradling, 2010). In the same study, the overall epigenetic status of differentiated cells was found to be more consistent than that of stem cells. Therefore, the differential effect of chromatin modulation in precursor cells but not in ECs after chn knockdown may be due to differences in the epigenetic flexibility of ISCs/precursor cells versus differentiated cells. It is possible that a chromatin structure in precursor cells allows Chn to bind to and maintain the chromatin in a precursor cell-specific state. When Chn is absent, the chromatin structure is altered to an extent that some important stem cell properties such as stem cell division are compromised and Delta expression is lost.
A proposed scenario for stem cell maintenance is that stem cells are kept in a state that prohibits them from differentiation (Fuchs and Chen, 2013; Stergiopoulos and Politis, 2013). Therefore, the loss of ‘stemness’ would directly lead to differentiation and therefore loss of stem cells. However, our analysis of Chn argues that stemness and initiation of differentiation can be a multi-step process. Although loss of Chn causes loss of stem cell properties required for ISC division, these chn− ISCs do not enter differentiation automatically. Meanwhile, overexpression of Chn does not convert precursor cells into stem cells but can halt differentiation, and temperature-shift experiments demonstrate that this is a reversible process. This suggests that there is a separate set of differentiation-promoting factors that are antagonized by the overexpressed Chn. The expression of these differentiation promoting factors are under separate controls, perhaps established during the asymmetric division to form the EBs. Overall, our analyses demonstrate that Chn in the adult midgut is a novel regulator required for maintaining a precursor cell chromatin structure that is compatible with ISC division.
MATERIALS AND METHODS
Drosophila stocks and feeding experiments
RNAi lines of chn, scute, pendulin, HMG D, H2AvD, HMG Z, RhoGEF4, ballchen, roughest and kugelkern were from Bloomington Stock Center (TRiP #26779, 26206, 27692, 31344, 28966, 26219, 31178, 31350, 28672 and 28750, respectively). UAS-chn and chn9 mutant lines were as described previously (Tsuda et al., 2006). Other stocks have been previously described (Amcheslavsky et al., 2011, 2009). Viability tests and feeding experiments were as previously described (Amcheslavsky et al., 2011, 2009). For lineage analysis, GFP-marked intestinal stem cell clones from MARCM were generated as previously described (Amcheslavsky et al., 2009; Lee and Luo, 2001). Fly stocks were crossed to generate the following genotype: hsFLP,UASGFP,tubGal4;FRT42Dchn9/FRT42D,tubGal80;+/+. The final cross and offsprings were maintained at 18°C. To induce MARCM clones, flies were heat shocked in a 37°C water bath for 45 min. The flies were then kept at 29°C for an additional 1-8 days, as indicated in the figure legends, before dissection.
Gut dissection
For gut dissection, female flies were used routinely because of their bigger size, but male flies were also used to check the phenotypes. The entire gastrointestinal tract was pulled from the posterior end directly into fixation medium containing 1×PBS and 4% formaldehyde (Mallinckrodt Chemicals). Guts were fixed in this medium for 3 h, except for Delta staining where the fixation was for 0.5 h. Subsequent rinses, washes and incubations with primary and secondary antibodies were carried out in a solution containing 1×PBS, 0.5% BSA and 0.1% Triton X-100. For RNA isolation, the malpighian tubules, crops, esophagus and rectum were removed, leaving only the midgut. Approximately 15 midguts were put into a 1.5 ml centrifuge tube on dry ice and frozen immediately for subsequent RNA isolation.
Immunostaining and fluorescent microscopy
For EdU labeling experiments, wild-type and chn RNAi flies were fed on 100 μM EdU (Invitrogen) in a 5% sucrose solution for 24 h. Guts were dissected and EdU incorporation was detected using Alexa Fluor 555 heat-shock assay (Click iT, Invitrogen) following the manufacturer's instruction. The immunostaining procedure has been described previously (Amcheslavsky et al., 2011, 2009). The following antisera were used: anti-Delta (monoclonal, 1:100 dilution, DSHB, Iowa), anti-Prospero (monoclonal, 1:100, DSHB), anti-Armadillo (monoclonal, 1:100, DSHB), anti-HP1 (monoclonal, 1:50, DSHB), anti β-galactosidase (rabbit polyclonal, 1:50,000, 55976; Cappel, MP Biomedicals), anti-Cdc2 (rabbit polyclonal, 1:500, sc-53; Santa Cruz Biotechnology), anti-acetylated-Histone 3 (rabbit polyclonal, 1:500, 06-599; Millipore), anti H3K4me3 (rabbit monoclonal, 1:2000, C42D8; Cell Signaling), anti-GFP (rabbit polyclonal, 1:1000, A11122; Molecular Probes) and anti-Pdm1 (1:500, from Dr Xiaohang Yang, Zhejiang University, Hang Zhou, China). Secondary antibodies were used in 1:2000 dilutions as follows: goat anti-mouse IgG conjugated to Alexa 488, Alexa 568 or Alexa 555 (A11017, A11019, A21422; Molecular Probes), and goat anti-rabbit IgG conjugated to Alexa 488 or Alexa 555 (A11070, A21428; Molecular Probes). DAPI in Vectashield (Vector Labs) was used at 1:1 dilution in PBS. Images were taken by a Nikon Spinning Disk Confocal microscope (UMass Medical School Imaging Core Facility).
Microarray methodology and data analysis
High-density oligonucleotide GeneChip Drosophila Genome 2.0 Arrays (Affymetrix 900531) containing 18,880 probe sets/chip were used to analyze expression profiles of wild-type and experimental genetic backgrounds (RafGOF, RafGOF+NotchDN and RafGOF+NotchIC). In these microarrays, each probe set contains 14 pairs of oligonucleotide to measure the level of transcription of each annotated sequence. In total, the array uses more than 500,000 data points to measure the expression of 18,500 transcripts and variants. Four hybridization controls (bioB, bioC and bioD from Escherichia coli, and cre from P1 bacteriophage), five poly-A controls (dap, lys, phe, thr and trp from Bacillus subtilis) and three housekeeping gene controls (actin 42A, glyceraldehyde-3-phosphate dehydrogenase 2, eukaryotic initiation factor 4a) were included for quality control purposes. cRNA sample preparation, array hybridization and scanning followed protocols as recommended by the manufacturer (http://www.affymetrix.com/index.affx). The hybridization intensity signal of each transcript was determined using the Affymetrix Microarray Suite 5.0 software package. Intensity values were scaled such that the overall fluorescence intensity of each array was equivalent. The GeneSpring (version 7.30; http://www.agilent.com) and GENECLUSTER (version 2.0; http://www-genome.wi.mit.edu/cancer/software/genecluster2/gc2.html) software packages were used to identify transcripts that were differentially expressed, and to conduct hierarchical two-dimensional clustering and self-organizing map analyses. The criteria for significant changes are: first, the signal values (after the initial flooring to an arbitrary value of 11 units) of a specific transcript in the wild-type and experimental conditions had to have a relative change of at least twofold; second, the absolute signal change must have at least 50 units; third, these changes had to be reproducible in all four independent replicate experiments; fourth, these changes had to demonstrate statistically significant P values ≤0.05. As a result, RNAs represented by 1127 probe sets showed significantly changed expression in at least one of the genetically modified backgrounds. Microarray data are available in the NCBI Gene Expression Omnibus (GSE58351).
Supplementary Material
Acknowledgements
We acknowledge the Bloomington Drosophila Stock Center and FlyTrap for providing fly stocks. Y.T.I. is a member of the UMass DERC (DK32520), a member of the UMass Center for Clinical and Translational Science (UL1TR000161) and a member of the Guangdong Innovative Research Team Program (No. 201001Y0104789252). L.T. thanks Dr Lim Young-Mi for preparing DNA and antibody. We thank Juerg Straubhaar for help with bioinfomatics.
Footnotes
Competing interests
The authors declare no competing financial interests.
Author contributions
A.A., Y.N., Q.L. and Y.T.I. designed and carried out the experiments. F.H. analyzed the microarray data, performed the two-dimensional hierarchical clustering analysis and derived the self-organizing map. L.T. provided the Charlatan mutant strain, overexpression strain and antibody, and designed the clonal analysis and rescue experiment. M.M. provided the Raf, Notch transgenic strains and analyzed these constructs in the midgut. A.A. and Y.T.I. wrote the manuscript and all authors amended the manuscript.
Funding
The work in Y.T.I.’s laboratory is supported by a National Institutes of Health grant [DK83450]. Deposited in PMC for release after 12 months.
Supplementary material
Supplementary material available online at http://dev.biologists.org/lookup/suppl/doi:10.1242/dev.106237/-/DC1
References
- Agarwal N., Hardt T., Brero A., Nowak D., Rothbauer U., Becker A., Leonhardt H., Cardoso M. C. (2007). MeCP2 interacts with HP1 and modulates its heterochromatin association during myogenic differentiation. Nucleic Acids Res. 35, 5402-5408 10.1093/nar/gkm599 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amcheslavsky A., Jiang J., Ip Y. T. (2009). Tissue damage-induced intestinal stem cell division in Drosophila. Cell Stem Cell 4, 49-61 10.1016/j.stem.2008.10.016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amcheslavsky A., Ito N., Jiang J., Ip Y. T. (2011). Tuberous sclerosis complex and Myc coordinate the growth and division of Drosophila intestinal stem cells. J. Cell Biol. 193, 695-710 10.1083/jcb.201103018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bardin A. J., Perdigoto C. N., Southall T. D., Brand A. H., Schweisguth F. (2010). Transcriptional control of stem cell maintenance in the Drosophila intestine. Development 137, 705-714 10.1242/dev.039404 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biteau B., Jasper H. (2011). EGF signaling regulates the proliferation of intestinal stem cells in Drosophila. Development 138, 1045-1055 10.1242/dev.056671 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biteau B., Hochmuth C. E., Jasper H. (2008). JNK activity in somatic stem cells causes loss of tissue homeostasis in the aging Drosophila gut. Cell Stem Cell 3, 442-455 10.1016/j.stem.2008.07.024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bond D., Foley E. (2012). Autocrine platelet-derived growth factor-vascular endothelial growth factor receptor-related (Pvr) pathway activity controls intestinal stem cell proliferation in the adult Drosophila midgut. J. Biol. Chem. 287, 27359-27370 10.1074/jbc.M112.378018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buchon N., Broderick N. A., Kuraishi T., Lemaitre B. (2010). Drosophila EGFR pathway coordinates stem cell proliferation and gut remodeling following infection. BMC Biol. 8, 152 10.1186/1741-7007-8-152 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buczacki S. J. A., Zecchini H. I., Nicholson A. M., Russell R., Vermeulen L., Kemp R., Winton D. J. (2013). Intestinal label-retaining cells are secretory precursors expressing Lgr5. Nature 495, 65-69 10.1038/nature11965 [DOI] [PubMed] [Google Scholar]
- Buszczak M., Paterno S., Spradling A. C. (2009). Drosophila stem cells share a common requirement for the histone H2B ubiquitin protease scrawny. Science 323, 248-251 10.1126/science.1165678 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cordero J. B., Stefanatos R. K., Scopelliti A., Vidal M., Sansom O. J. (2012). Inducible progenitor-derived Wingless regulates adult midgut regeneration in Drosophila. EMBO J. 31, 3901-3917 10.1038/emboj.2012.248 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dutta D., Xiang J., Edgar B. A. (2014). RNA expression profiling from FACS-isolated cells of the Drosophila intestine. Curr. Protoc. Stem Cell Biol. 27, Unit 2F 2 [DOI] [PubMed] [Google Scholar]
- Eisen M. B., Spellman P. T., Brown P. O., Botstein D. (1998). Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. U.S.A. 95, 14863-14868 10.1073/pnas.95.25.14863 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Escudero L. M., Caminero E., Schulze K. L., Bellen H. J., Modolell J. (2005). Charlatan, a Zn-finger transcription factor, establishes a novel level of regulation of the proneural achaete/scute genes of Drosophila. Development 132, 1211-1222 10.1242/dev.01691 [DOI] [PubMed] [Google Scholar]
- Fuchs E., Chen T. (2013). A matter of life and death: self-renewal in stem cells. EMBO Rep. 14, 39-48 10.1038/embor.2012.197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo Z., Driver I., Ohlstein B. (2013). Injury-induced BMP signaling negatively regulates Drosophila midgut homeostasis. J. Cell Biol. 201, 945-961 10.1083/jcb.201302049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- He F., Li X., Spatrick P., Casillo R., Dong S., Jacobson A. (2003). Genome-wide analysis of mRNAs regulated by the nonsense-mediated and 5′ to 3′ mRNA decay pathways in yeast. Mol. Cell 12, 1439-1452 10.1016/S1097-2765(03)00446-5 [DOI] [PubMed] [Google Scholar]
- Jiang H., Patel P. H., Kohlmaier A., Grenley M. O., McEwen D. G., Edgar B. A. (2009). Cytokine/Jak/Stat signaling mediates regeneration and homeostasis in the Drosophila midgut. Cell 137, 1343-1355 10.1016/j.cell.2009.05.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang H., Grenley M. O., Bravo M. J., Blumhagen R. Z., Edgar B. A. (2010). EGFR/Ras/MAPK signaling mediates adult midgut epithelial homeostasis and regeneration in Drosophila. Cell Stem Cell 8, 84-95 10.1016/j.stem.2010.11.026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin Y., Xu J., Yin M.-X., Lu Y., Hu L., Li P., Zhang P., Yuan Z., Ho M. S., Ji H., et al. (2013). Brahma is essential for Drosophila intestinal stem cell proliferation and regulated by Hippo signaling. Elife 2, e00999 10.7554/eLife.00999 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kania A., Salzberg A., Bhat M., D'Evelyn D., He Y., Kiss I., Bellen H. J. (1995). P-element mutations affecting embryonic peripheral nervous system development in Drosophila melanogaster. Genetics 139, 1663-1678 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karpowicz P., Perez J., Perrimon N. (2010). The Hippo tumor suppressor pathway regulates intestinal stem cell regeneration. Development 137, 4135-4145 10.1242/dev.060483 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Küssel P., Frasch M. (1995). Pendulin, a Drosophila protein with cell cycle-dependent nuclear localization, is required for normal cell proliferation. J. Cell Biol. 129, 1491-1507 10.1083/jcb.129.6.1491 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kwon S. H., Workman J. L. (2011). The changing faces of HP1: from heterochromatin formation and gene silencing to euchromatic gene expression. Bioessays 33, 280-289 10.1002/bies.201000138 [DOI] [PubMed] [Google Scholar]
- Lee T., Luo L. (2001). Mosaic analysis with a repressible cell marker (MARCM) for Drosophila neural development. Trends Neurosci. 24, 251-254 10.1016/S0166-2236(00)01791-4 [DOI] [PubMed] [Google Scholar]
- Lee S.-H., Kim I.-J., Kim J.-G., Park J.-S., Kim Y.-S., Yamaguchi M., Kim C.-M., Yoo M.-A. (2011). Regulation of intestinal stem cell proliferation by human methyl-CpG-binding protein-2 in Drosophila. Cell Struct. Funct. 36, 197-208 10.1247/csf.11027 [DOI] [PubMed] [Google Scholar]
- Li H., Qi Y., Jasper H. (2013a). Dpp signaling determines regional stem cell identity in the regenerating adult drosophila gastrointestinal tract. Cell Rep. 4, 10-18 10.1016/j.celrep.2013.05.040 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Z., Zhang Y., Han L., Shi L., Lin X. (2013b). Trachea-derived dpp controls adult midgut homeostasis in Drosophila. Dev. Cell 24, 133-143 10.1016/j.devcel.2012.12.010 [DOI] [PubMed] [Google Scholar]
- Li Z., Guo Y., Han L., Zhang Y., Shi L., Huang X., Lin X. (2014). Debra-mediated ci degradation controls tissue homeostasis in Drosophila adult midgut. Stem Cell Rep. 2, 135-144 10.1016/j.stemcr.2013.12.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin G., Xu N., Xi R. (2008). Paracrine Wingless signalling controls self-renewal of Drosophila intestinal stem cells. Nature 455, 1119-1123 10.1038/nature07329 [DOI] [PubMed] [Google Scholar]
- Madigan J. P., Chotkowski H. L., Glaser R. L. (2002). DNA double-strand break-induced phosphorylation of Drosophila histone variant H2Av helps prevent radiation-induced apoptosis. Nucleic Acids Res. 30, 3698-3705 10.1093/nar/gkf496 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Markstein M., Dettorre S., Cho J., Neumuller R. A., Craig-Muller S., Perrimon N. (2014). Systematic screen of chemotherapeutics in Drosophila stem cell tumors. Proc. Natl. Acad. Sci. U.S.A. 111, 4530-4535 10.1073/pnas.1401160111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Micchelli C. A., Perrimon N. (2006). Evidence that stem cells reside in the adult Drosophila midgut epithelium. Nature 439, 475-479 10.1038/nature04371 [DOI] [PubMed] [Google Scholar]
- Nakada D., Levi B. P., Morrison S. J. (2011). Integrating physiological regulation with stem cell and tissue homeostasis. Neuron 70, 703-718 10.1016/j.neuron.2011.05.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohlstein B., Spradling A. (2006). The adult Drosophila posterior midgut is maintained by pluripotent stem cells. Nature 439, 470-474 10.1038/nature04333 [DOI] [PubMed] [Google Scholar]
- Ohlstein B., Spradling A. (2007). Multipotent Drosophila intestinal stem cells specify daughter cell fates by differential notch signaling. Science 315, 988-992 10.1126/science.1136606 [DOI] [PubMed] [Google Scholar]
- O'Brien L. E., Soliman S. S., Li X., Bilder D. (2011). Altered modes of stem cell division drive adaptive intestinal growth. Cell 147, 603-614 10.1016/j.cell.2011.08.048 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park J. S., Kim Y. S., Yoo M. A. (2009). The role of p38b MAPK in age-related modulation of intestinal stem cell proliferation and differentiation in Drosophila. Aging 1, 637-651 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park J.-S., Lee S.-H., Na H.-J., Pyo J.-H., Kim Y.-S., Yoo M.-A. (2012). Age- and oxidative stress-induced DNA damage in Drosophila intestinal stem cells as marked by Gamma-H2AX. Exp. Gerontol. 47, 401-405 10.1016/j.exger.2012.02.007 [DOI] [PubMed] [Google Scholar]
- Ragab A., Thompson E. C., Travers A. A. (2006). High mobility group proteins HMGD and HMGZ interact genetically with the Brahma chromatin remodeling complex in Drosophila. Genetics 172, 1069-1078 10.1534/genetics.105.049957 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ragab A., Buechling T., Gesellchen V., Spirohn K., Boettcher A.-L., Boutros M. (2011). Drosophila Ras/MAPK signalling regulates innate immune responses in immune and intestinal stem cells. EMBO J. 30, 1123-1136 10.1038/emboj.2011.4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reeves N., Posakony J. W. (2005). Genetic programs activated by proneural proteins in the developing Drosophila PNS. Dev. Cell 8, 413-425 10.1016/j.devcel.2005.01.020 [DOI] [PubMed] [Google Scholar]
- Ren F., Wang B., Yue T., Yun E.-Y., Ip Y. T., Jiang J. (2010). Hippo signaling regulates Drosophila intestine stem cell proliferation through multiple pathways. Proc. Natl. Acad. Sci. U.S.A. 107, 21064-21069 10.1073/pnas.1012759107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ren F., Shi Q., Chen Y., Jiang A., Ip Y. T., Jiang H., Jiang J. (2013). Drosophila Myc integrates multiple signaling pathways to regulate intestinal stem cell proliferation during midgut regeneration. Cell Res. 23, 1133-1146 10.1038/cr.2013.101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rizk P., Barker N. (2012). Gut stem cells in tissue renewal and disease: methods, markers, and myths. Wiley Interdiscip. Rev. Syst. Biol. Med. 4, 475-496 10.1002/wsbm.1176 [DOI] [PubMed] [Google Scholar]
- Shaw R. L., Kohlmaier A., Polesello C., Veelken C., Edgar B. A., Tapon N. (2010). The Hippo pathway regulates intestinal stem cell proliferation during Drosophila adult midgut regeneration. Development 137, 4147-4158 10.1242/dev.052506 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skora A. D., Spradling A. C. (2010). Epigenetic stability increases extensively during Drosophila follicle stem cell differentiation. Proc. Natl. Acad. Sci. U.S.A. 107, 7389-7394 10.1073/pnas.1003180107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staley B. K., Irvine K. D. (2010). Warts and Yorkie mediate intestinal regeneration by influencing stem cell proliferation. Curr. Biol. 20, 1580-1587 10.1016/j.cub.2010.07.041 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stergiopoulos A., Politis P. K. (2013). The role of nuclear receptors in controlling the fine balance between proliferation and differentiation of neural stem cells. Arch. Biochem. Biophys. 534, 27-37 10.1016/j.abb.2012.09.009 [DOI] [PubMed] [Google Scholar]
- Takeda N., Jain R., LeBoeuf M. R., Wang Q., Lu M. M., Epstein J. A. (2011). Interconversion between intestinal stem cell populations in distinct niches. Science 334, 1420-1424 10.1126/science.1213214 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamayo P., Slonim D., Mesirov J., Zhu Q., Kitareewan S., Dmitrovsky E., Lander E. S., Golub T. R. (1999). Interpreting patterns of gene expression with self-organizing maps: methods and application to hematopoietic differentiation. Proc. Natl. Acad. Sci. U.S.A. 96, 2907-2912 10.1073/pnas.96.6.2907 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian A., Jiang J. (2014). Intestinal epithelium-derived BMP controls stem cell self-renewal in Drosophila adult midgut. Elife 3, e01857 10.7554/eLife.01857 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian H., Biehs B., Warming S., Leong K. G., Rangell L., Klein O. D., de Sauvage F. J. (2011). A reserve stem cell population in small intestine renders Lgr5-positive cells dispensable. Nature 478, 255-259 10.1038/nature10408 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuda L., Kaido M., Lim Y.-M., Kato K., Aigaki T., Hayashi S. (2006). An NRSF/REST-like repressor downstream of Ebi/SMRTER/Su(H) regulates eye development in Drosophila. EMBO J. 25, 3191-3202 10.1038/sj.emboj.7601179 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu N., Wang S. Q., Tan D., Gao Y., Lin G., Xi R. (2011). EGFR, Wingless and JAK/STAT signaling cooperatively maintain Drosophila intestinal stem cells. Dev. Biol. 354, 31-43 10.1016/j.ydbio.2011.03.018 [DOI] [PubMed] [Google Scholar]
- Yamasaki Y., Lim Y.-M., Niwa N., Hayashi S., Tsuda L. (2011). Robust specification of sensory neurons by dual functions of charlatan, a Drosophila NRSF/REST-like repressor of extramacrochaetae and hairy. Genes Cells 16, 896-909 10.1111/j.1365-2443.2011.01537.x [DOI] [PubMed] [Google Scholar]
- Yan K. S., Chia L. A., Li X., Ootani A., Su J., Lee J. Y., Su N., Luo Y., Heilshorn S. C., Amieva M. R., et al. (2012). The intestinal stem cell markers Bmi1 and Lgr5 identify two functionally distinct populations. Proc. Natl. Acad. Sci. U.S.A. 109, 466-471 10.1073/pnas.1118857109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeng X., Chauhan C., Hou S. X. (2010). Characterization of midgut stem cell- and enteroblast-specific Gal4 lines in drosophila. Genesis 48, 607-611 10.1002/dvg.20661 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeng X., Lin X., Hou S. X. (2013). The Osa-containing SWI/SNF chromatin-remodeling complex regulates stem cell commitment in the adult Drosophila intestine. Development 140, 3532-3540 10.1242/dev.096891 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou F., Rasmussen A., Lee S., Agaisse H. (2013). The UPD3 cytokine couples environmental challenge and intestinal stem cell division through modulation of JAK/STAT signaling in the stem cell microenvironment. Dev. Biol. 373, 383-393 10.1016/j.ydbio.2012.10.023 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.