Abstract
We have previously reported an enhanced version of sequencing by hybridization (SBH), termed positional SBH (PSBH). PSBH uses partially duplex probes containing single-stranded 3' overhangs, instead of simple single-stranded probes. Stacking interactions between the duplex probe and a single-stranded target allow us to reduce the probe sizes required to 5-base single-stranded overhangs. Here we demonstrate the use of PSBH to capture relatively long single-stranded DNA targets and perform standard solid-state Sanger sequencing on these primer-template complexes without ligation. Our results indicate that only 5 bases of known terminal sequence are required for priming. In addition, the partially duplex probes have the ability to capture their specific target from a mixture of five single-stranded targets with different 3'-terminal sequences. This indicates the potential utility of the PSBH approach to sequence mixtures of DNA targets without prior purification.
Full text
PDF
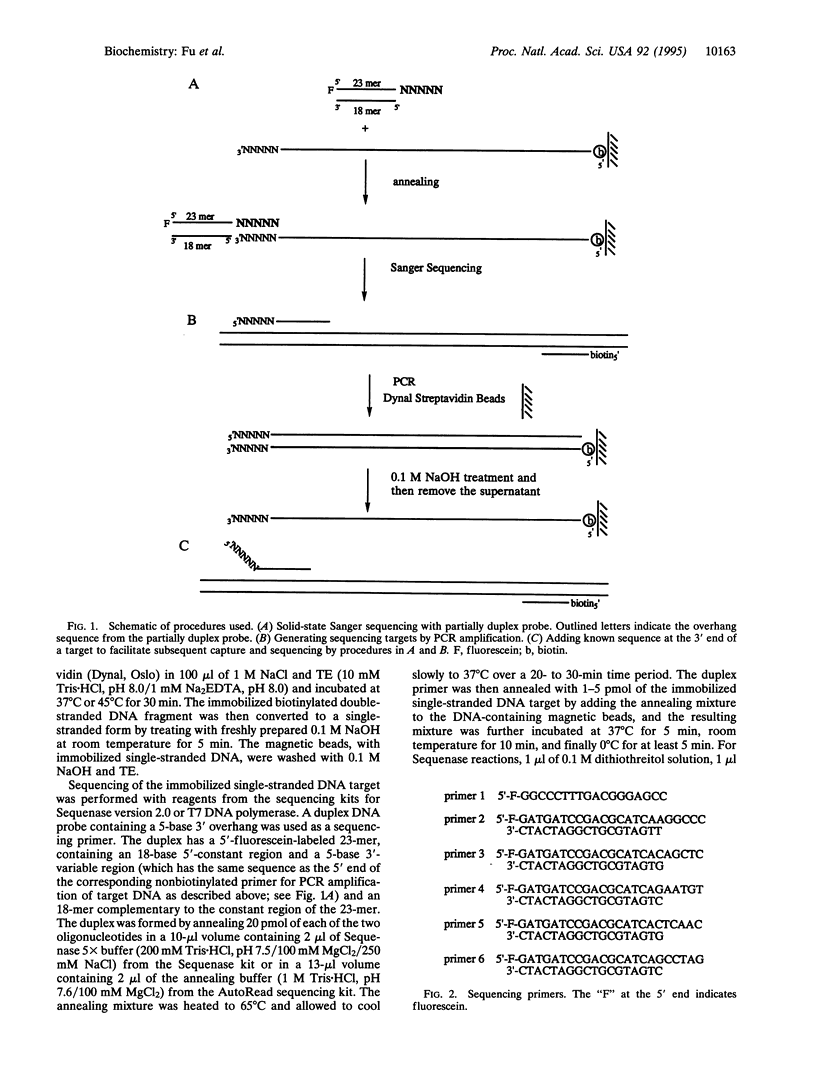




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anderson S. Shotgun DNA sequencing using cloned DNase I-generated fragments. Nucleic Acids Res. 1981 Jul 10;9(13):3015–3027. doi: 10.1093/nar/9.13.3015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bains W. Hybridization methods for DNA sequencing. Genomics. 1991 Oct;11(2):294–301. doi: 10.1016/0888-7543(91)90135-2. [DOI] [PubMed] [Google Scholar]
- Bains W., Smith G. C. A novel method for nucleic acid sequence determination. J Theor Biol. 1988 Dec 7;135(3):303–307. doi: 10.1016/s0022-5193(88)80246-7. [DOI] [PubMed] [Google Scholar]
- Broude N. E., Sano T., Smith C. L., Cantor C. R. Enhanced DNA sequencing by hybridization. Proc Natl Acad Sci U S A. 1994 Apr 12;91(8):3072–3076. doi: 10.1073/pnas.91.8.3072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drmanac R., Drmanac S., Strezoska Z., Paunesku T., Labat I., Zeremski M., Snoddy J., Funkhouser W. K., Koop B., Hood L. DNA sequence determination by hybridization: a strategy for efficient large-scale sequencing. Science. 1993 Jun 11;260(5114):1649–1652. doi: 10.1126/science.8503011. [DOI] [PubMed] [Google Scholar]
- Drmanac R., Labat I., Brukner I., Crkvenjakov R. Sequencing of megabase plus DNA by hybridization: theory of the method. Genomics. 1989 Feb;4(2):114–128. doi: 10.1016/0888-7543(89)90290-5. [DOI] [PubMed] [Google Scholar]
- Hultman T., Ståhl S., Hornes E., Uhlén M. Direct solid phase sequencing of genomic and plasmid DNA using magnetic beads as solid support. Nucleic Acids Res. 1989 Jul 11;17(13):4937–4946. doi: 10.1093/nar/17.13.4937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kieleczawa J., Dunn J. J., Studier F. W. DNA sequencing by primer walking with strings of contiguous hexamers. Science. 1992 Dec 11;258(5089):1787–1791. doi: 10.1126/science.1465615. [DOI] [PubMed] [Google Scholar]
- Kotler L. E., Zevin-Sonkin D., Sobolev I. A., Beskin A. D., Ulanovsky L. E. DNA sequencing: modular primers assembled from a library of hexamers or pentamers. Proc Natl Acad Sci U S A. 1993 May 1;90(9):4241–4245. doi: 10.1073/pnas.90.9.4241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messing J., Crea R., Seeburg P. H. A system for shotgun DNA sequencing. Nucleic Acids Res. 1981 Jan 24;9(2):309–321. doi: 10.1093/nar/9.2.309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pease A. C., Solas D., Sullivan E. J., Cronin M. T., Holmes C. P., Fodor S. P. Light-generated oligonucleotide arrays for rapid DNA sequence analysis. Proc Natl Acad Sci U S A. 1994 May 24;91(11):5022–5026. doi: 10.1073/pnas.91.11.5022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts R. J., Macelis D. REBASE--restriction enzymes and methylases. Nucleic Acids Res. 1994 Sep;22(17):3628–3639. doi: 10.1093/nar/22.17.3628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siemieniak D. R., Slightom J. L. A library of 3342 useful nonamer primers for genome sequencing. Gene. 1990 Nov 30;96(1):121–124. doi: 10.1016/0378-1119(90)90350-z. [DOI] [PubMed] [Google Scholar]
- Strauss E. C., Kobori J. A., Siu G., Hood L. E. Specific-primer-directed DNA sequencing. Anal Biochem. 1986 Apr;154(1):353–360. doi: 10.1016/0003-2697(86)90536-1. [DOI] [PubMed] [Google Scholar]
- Strezoska Z., Paunesku T., Radosavljević D., Labat I., Drmanac R., Crkvenjakov R. DNA sequencing by hybridization: 100 bases read by a non-gel-based method. Proc Natl Acad Sci U S A. 1991 Nov 15;88(22):10089–10093. doi: 10.1073/pnas.88.22.10089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studier F. W. A strategy for high-volume sequencing of cosmid DNAs: random and directed priming with a library of oligonucleotides. Proc Natl Acad Sci U S A. 1989 Sep;86(18):6917–6921. doi: 10.1073/pnas.86.18.6917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szybalski W. Proposal for sequencing DNA using ligation of hexamers to generate sequential elongation primers (SPEL-6). Gene. 1990 May 31;90(1):177–178. doi: 10.1016/0378-1119(90)90458-4. [DOI] [PubMed] [Google Scholar]