Analysis of anthocyanin methyltransferase genes reveals genetic mechanisms underlying the evolution of anthocyanins with diverse structures.
Abstract
Anthocyanins are a chemically diverse class of secondary metabolites that color most flowers and fruits. They consist of three aromatic rings that can be substituted with hydroxyl, sugar, acyl, and methyl groups in a variety of patterns depending on the plant species. To understand how such chemical diversity evolved, we isolated and characterized METHYLATION AT THREE2 (MT2) and the two METHYLATION AT FIVE (MF) loci from Petunia spp., which direct anthocyanin methylation in petals. The proteins encoded by MT2 and the duplicated MF1 and MF2 genes and a putative grape (Vitis vinifera) homolog Anthocyanin O-Methyltransferase1 (VvAOMT1) are highly similar to and apparently evolved from caffeoyl-Coenzyme A O-methyltransferases by relatively small alterations in the active site. Transgenic experiments showed that the Petunia spp. and grape enzymes have remarkably different substrate specificities, which explains part of the structural anthocyanin diversity in both species. Most strikingly, VvAOMT1 expression resulted in the accumulation of novel anthocyanins that are normally not found in Petunia spp., revealing how alterations in the last reaction can reshuffle the pathway and affect (normally) preceding decoration steps in an unanticipated way. Our data show how variations in gene expression patterns, loss-of-function mutations, and alterations in substrate specificities all contributed to the anthocyanins’ structural diversity.
Distinct plant species produce presumably hundreds of thousands of secondary metabolites, which are low-Mr compounds that occur in some specific groups only (Pichersky and Gang, 2000). Flavonoids, for example, are a major family of secondary metabolites found in virtually all vascular plants that are involved in protection against UV-light and pathogens, signaling to nodulating bacteria, hormone transport, pollen germination, and the coloration of flowers and fruits. This functional diversity is paralleled by a large chemical diversity, as today over 9,000 flavonoids with distinct structures have been identified in variety of species (Buer et al., 2010). How the ability to synthesize such a vast number of different compounds has evolved is largely unknown but presumably involved gene duplications and divergence of encoded proteins and gene expression patterns (Pichersky and Gang, 2000).
Flavonoid biosynthesis is arguably one of the best studied pathways in plants, in particular the early steps that generate the flavonoid scaffolds of the major subclasses found in angiosperms, including chalcones, flavones, flavonols, flavanols (i.e. flavan-3-ols), proanthocyanidins, and anthocyanins (Winkel-Shirley, 2001). However, much less is known about the subsequent modifications (decorations), such as glycosylation, acylation, and methylation, which cause most of the structural variation (Grotewold, 2006). Grape (Vitis vinifera), for example, synthesizes very simple anthocyanins bearing one or two methyl groups and a single (acetylated) sugar, whereas Arabidopsis (Arabidopsis thaliana) accumulates more complex anthocyanins with seven sugar and acyl moieties but no methyl groups (Brouillard et al., 2003; Saito et al., 2013), and Petunia spp. pigments its petals with methylated anthocyanins that bear an acylated rutinoside moiety (Ando et al., 1999; Slimestad et al., 1999).
Petunia hybrida was generated 200 years ago by interspecific crosses between various accessions/subspecies of Petunia inflata and Petunia axillaris. P. inflata subspecies have bee-pollinated flowers with violet petals containing 3′5′-methylated anthocyanins (malvidins), while P. axillaris accessions have moth-pollinated flowers with a white corolla limb that lacks anthocyanins and a variegated corolla tube containing anthocyanins that lack one or both methyl groups (Quattrocchio et al., 1999; Hoballah et al., 2007). Classical breeding generated P. hybrida cultivars, each containing a different mix of the parental species genomes, with a wide variety of flower colors resulting from the introgression of mutant alleles from the parental species (Quattrocchio et al., 1999) and new mutations that arose in P. hybrida (Kroon et al., 1994; van Houwelingen et al., 1998).
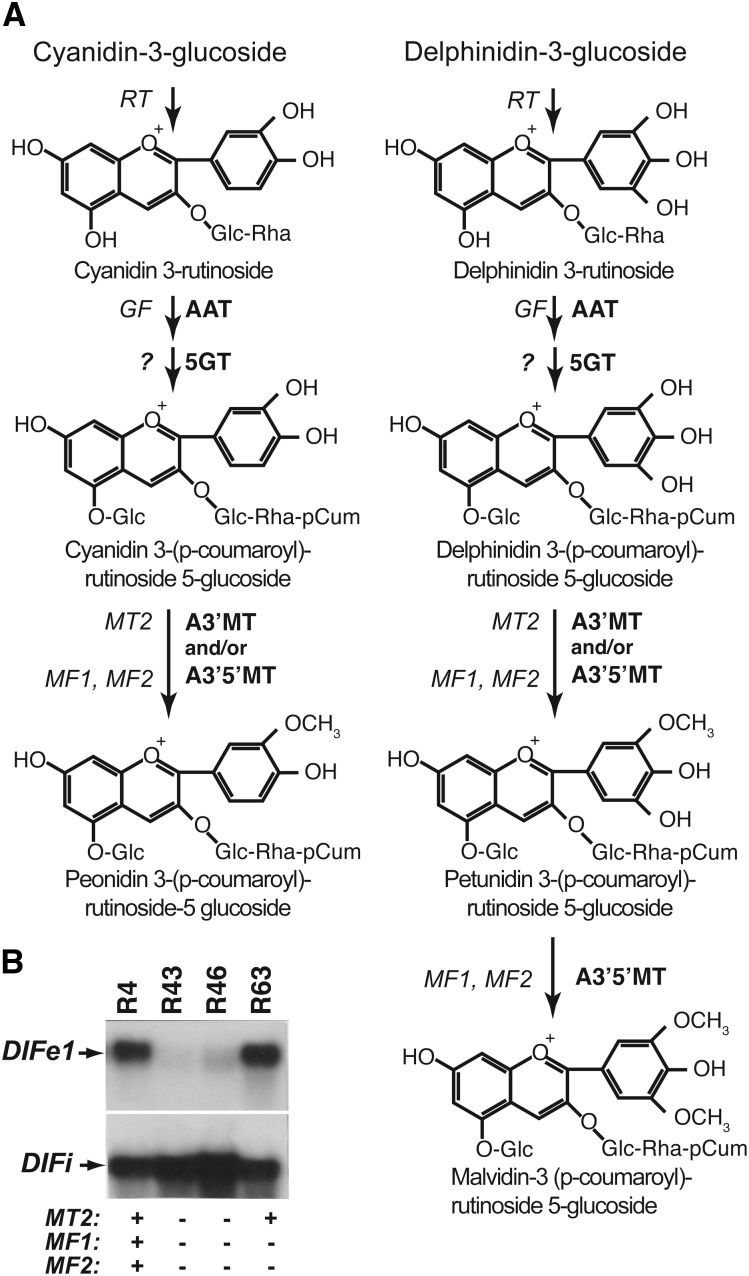
Analysis of segregating populations of unrelated P. hybrida cultivars identified several loci controlling the decoration of anthocyanins and suggested a model for their sequential action (Wiering, 1974; Wiering and De Vlaming, 1977; de Vlaming et al., 1984). This model (Fig. 1A) was substantiated by enzymological analyses in the 1980s (Jonsson et al., 1982, 1983, 1984a, 1984b) and, more recently, the isolation of some of the genes (Brugliera et al., 1994; Kroon et al., 1994; Yamazaki et al., 2002). The current view is that anthocyanin-3 glucosides are in the petal limb sequentially (1) rhamnosylated by a sugar-transferase encoded by the RHAMNOSYLATION AT THREE (RT) locus (Wiering, 1974; de Vlaming et al., 1984; Brugliera et al., 1994; Kroon et al., 1994), (2) acylated by an anthocyanin acyltransferase (AAT) encoded by GLUCOSYLATION AT FIVE (GF; Wiering and De Vlaming, 1973; Provenzano, 2011), (3) 5-glucosylated by a 5-glucosyltransferase (5GT; Jonsson et al., 1984c; Yamazaki et al., 2002), and (4) methylated in the 5′ and/or 3′ position by anthocyanin methyltransferases (AMTs; Fig. 1A).
Figure 1.
Role of loci and enzymes required for methylation of anthocyanins in Petunia spp. A, Diagram depicting the modification of simple anthocyanin 3-glucosides by subsequent rhamnosylation, 5-glucosylation, acylation, and methylation. Enzymes involved in each reaction are indicated on the right of the arrows and genetic loci (italics) controlling the reaction on the left. B, RNA gel-blot analysis of DIFe1 expression in petals of four inbred Petunia spp. lines that are homozygous for functional (+) or mutant alleles (–) of MT2, MF1, and MF2. As a control for RNA loading and integrity, the filter was stripped and rehybridized with DIFi.
Genetic analyses suggested that three or four partially redundant loci control the methylation of anthocyanins in Petunia spp. petals (Wiering, 1974; Wiering and De Vlaming, 1977) and the activity/expression of multiple AMT isoforms with different isoelectric points (Jonsson et al., 1983, 1984b). METHYLATION AT THREE (MT) directs methylation of the 3′ hydroxyl group and METHYLATION AT FIVE (MF) directs methylation of both the 3′ and 5′ hydroxyl group. Segregation ratios of 3′5′ methylation in certain crosses and linkage to markers on chromosomes III and V suggested that two MF loci (MF1 and MF2) might exist (Wiering and De Vlaming, 1977) but could also be explained by the translocation of a single MF gene in some lines. Because it was originally thought that 3′ methylation by an MT gene is essential for subsequent 5′ methylation by MF1/MF2 (Wiering, 1974), two MT loci were postulated: one, MT1, tightly linked with MF1 on chromosome III, and another, MT2, linked with MF2 on chromosome V (Wiering, 1974; Wiering and De Vlaming, 1977). However, analyses of partially purified AMTs from petals (Jonsson et al., 1984b) and experiments with an MF2 transgene (this paper) demonstrated that a single enzyme (anthocyanin 3′5′-methyltransferase [A3′5′MT]) can methylate both the 3′ and 5′ position, proving the premise for postulating MT1 incorrect. Moreover, molecular data, presented here, also indicate that MT and MF genes constitute a family of only three genes. Hence, we consider the MT1 locus for this paper as nonexistent.
ANTHOCYANIN1 (AN1), AN2, and AN11 are Helix-Loop-Helix (HLH), MYB, and WD40 proteins that activate transcription of genes involved in anthocyanin synthesis and vacuolar acidification (Koes et al., 2005). A differential hybridization screen identified complementary DNAs (cDNAs) from seven so-called DIFFERENTIAL (DIF) genes that were down-regulated in an1 petals (Kroon et al., 1994). DIFa, DIFf, DIFg, and DIFi encode, respectively, anthocyanidin synthase, a cytochrome b5 required for hydroxylation by flavonoid 3′,5′ hydroxylase (de Vetten et al., 1999), a sugar-transferase encoded by RT (Kroon et al., 1994), and a glutathione-transferase encoded by AN9 (Alfenito et al., 1998). To identify additional enzymes involved in the decoration of anthocyanins, we analyzed the remaining DIF genes. Recent data show that DIFi originates from the GF locus and encodes an acyl-transferase (Provenzano, 2011; S. Provenzano, R. Koes, and F. Quattrocchio, unpublished data), while DIFe1 encodes an AMT (this article).
Here, we show that DIFe1 and two paralogous genes, DIFe2a and DIFe2b, encode O-methyltransferases that are expressed in petals under the control of AN1, AN2, and AN11 and are located at the MT2, MF1, and MF2 locus, respectively. Complementation experiments with mt2 mf1 mf2 mutants in distinct genetic backgrounds revealed remarkable differences in the substrate specificities of these Petunia spp. AMTs and a putative anthocyanin O-methyltransferase from grape (VvAOMT1; Hugueney et al., 2009). Our data provide insight into the molecular genetic mechanisms underlying the evolution of distinct anthocyanin structures.
RESULTS
Identification of a Putative AMT Gene from P. hybrida
RNA gel-blot analysis showed that expression of mRNAs hybridizing to DIFe1 is down-regulated in lines with mutations in all MT and MF genes (Fig. 1B). Hence, we examined whether a His-tagged DIFe1 protein expressed in Escherichia coli can methylate delphinidin 3-glucoside (D3G) or delphinidin 3-rutinoside (D3R) in vitro, using, as a methyl donor, 14C-S-adenosyl-l-Met (SAM). Homogenates from DIFe1-expressing cells converted D3R and D3G into petunidin and malvidin derivatives, as judged by thin-layer chromatography. No activity was seen with homogenates from untransformed cells or when SAM was omitted (Supplemental Table S1). In further experiments, we analyzed reaction products by HPLC (Fig. 2A). In these assays, DIFe1 converted D3R, D3G, and delphinidin 3,5-diglucoside with similar efficiency. In all cases, the major products were petunidin derivatives, while malvidin derivatives constituted a smaller but substantial fraction. The D3R and D3G substrates contained low amounts of cyanidin derivatives as impurities, which were converted by DIFe1 into peonidin. These data suggest that DIFe1 has strong anthocyanin 3′-methyltransferase (A3′MT) activity and a low A3′5′MT activity.
Figure 2.
Methylation of anthocyanins by DIFe1 and DIFe2 in vitro and in vivo. A, Anthocyanidins (percentage of total) after incubation of SAM and the 3-glucoside (3G), 3-rutinoside (3R), or 3,5 diglucoside (3,5G) of delphinidin with crude extracts of E. coli expressing DIFe1, the empty vector, or no extract at all (none). Mal, Malvidin; Pet, petunidin; Del, delphinidin; Peo, peonidin; Cya, cyanidin. B, Phenotype of a flower from a control plant and MAC:asDIFe1 transformant A. C, Composition of anthocyanidins in flowers of independent MAC:asDIFe1 and MAC:asDIFe2 lines and a control plant without transgene.
To examine DIFe1 function in vivo, we expressed antisense DIFe1 RNA from a transgene (MAC:asDIFe1) driven by the MAC promoter (Comai et al., 1990) in P. hybrida F1 hybrid V23 × R51 (VR). VR is dominant for all known anthocyanin and flavonol genes and has violet petals. One of five MAC:asDIFe1 transformants (A) produced petals with a dull reddish color (Fig. 2B). In VR flowers, 80% of the anthocyanins are malvidin derivatives, whereas the main anthocyanins in the dull reddish MAC:asDIFe1 petals were delphinidin derivatives, showing that DIFe1 is involved in anthocyanin methylation in vivo (Fig. 2C).
DIFe1 Is Part of a Small Gene Family
Screening a petal cDNA library from P. hybrida ‘Old Glory Blue’ (OGB) with DIFe1 yielded two clones. One (E20) was identical to the original DIFe1 cDNA from V26, except that its 5′ untranslated region (UTR) was 25 nucleotides longer and its 3′ UTR region was 96 nucleotides shorter. The latter might result from promiscuous priming of oligo(dT) on an A-rich region in the 3′ UTR during cDNA synthesis or polyadenylation of the mRNA at distinct sites. The coding sequence of the second cDNA (E33) shared 82% nucleotide identity with DIFe1 but was disrupted by a 2-bp deletion, indicating that it originated from a paralogous gene, named DIFe2, and that OGB contains a mutant allele (dife2OGB).
To study the role of DIFe2 in vivo, we generated transgenic VR plants containing a MAC:asDIFe2 transgene that expresses the antisense RNA. One of five MAC:asDIFe2 transformants displayed a flower color phenotype similar to that of MAC:asDIFe1 that was also associated with a strong reduction of anthocyanin methylation (Fig. 2C). The high similarity of the DIFe1 and DIFe2 sequences and their RNA interference phenotypes, together with the analysis of mutant alleles of these genes (see below), suggests that RNA interference silenced both genes (and even a third gene, see below) simultaneously.
We assessed the number of DIFe paralogs in the lines V30 (MT2 mf1 mf2), R78 (mt2 mf1 mf2), and M1 (MT2 MF1 MF2) by DNA gel-blot analysis (Fig. 3). The DIFe2 cDNA probe detected three EcoRI fragments and at least two BamHI fragments in each line using low-stringency washing conditions. One of these EcoRI fragments was presumed to contain DIFe1, because it was eliminated after high-stringency washing and matched in size exactly to a fragment hybridizing to DIFe1 under stringent conditions. The DIFe2 alleles of lines V30 and M1 lack EcoRI sites (see below), indicating that the two other EcoRI fragments detected by DIFe2 probe under high-stringency conditions contain two distinct genes that are both highly similar to DIFe2.
Figure 3.
DNA gel-blot analysis of DIFe genes/paralogs in P. hybrida. DNA from P. hybrida lines M1, V30, and R78 was digested with EcoRI (E) or BamHI (B), size separated, hybridized to DIFe2OGB cDNA, washed at low stringency (2× SSC, 25°C; left) and subsequently at high stringency (0.1× SSC 68°C; middle), or hybridized with the DIFe1 cDNA and washed using high-stringency conditions (right). Fragments hybridizing to DIFe1 and DIFe2 at high stringency are marked with white and black arrowheads, respectively.
When we amplified DNA from the P. hybrida lines V30 and M1 or the parental species P. inflata S6, we found that most primers complementary to the DIFe2OGB cDNA amplified two distinct genes with exon sequences that are nearly identical, except for a few single nucleotide polymorphisms (SNPs). Because this low number of SNPs is similar to that observed between alleles from different P. hybrida accessions, we could not distinguish which of these two genes is allelic to the DIFe2OGB cDNA and therefore designated them DIFe2a and DIFe2b. DIFe2a and DIFe2b are not alleles of one locus but paralogs, because we could amplify both genes from a variety of lines that were inbred for more than 30 generations.
DIFe Genes Encode Proteins with Similarity to O-Methyltransferases
Plant O-methyltransferases constitute a large family made up of two major clades, A and B, consisting of two and three subgroups respectively (Lam et al., 2007). The DIFe proteins belong to the A1 subfamily (Supplemental Fig. S1; Supplemental Table S2), which contains caffeoyl-Coenzyme A O-methyltransferases (CCoA-MTs) and an AMT from grape known as VvAOMT1 (Hugueney et al., 2009), also known as flavonol and anthocyanin O-methyltransferase (Lücker et al., 2010). Other (putative) flavonoid methyltransferases, such as the Arabidopsis flavanol 3′methylase AtOMT1 (Saito et al., 2013), and enzymes with similar activities from Catharanthus roseus and Mentha × piperita, belong to the B1 and B2 groups (Lam et al., 2007; Supplemental Fig. S1) and are only distantly related to DIFe proteins.
A more detailed analysis, focused on proteins of the A1 subfamily (Fig. 4; Supplemental Table S3), shows that DIFe proteins are very similar and probably orthologous to three grape paralogs, including VvAOMT1, and closely related (predicted) proteins from other Asterids (e.g. Solanum lycopersicum and Solanum tuberosum) and Rosids (e.g. Fragaria vesca and Prunus persica), the major clades in eudicots. The relationship between these proteins resembles that of the species, consistent with the idea that they are orthologous. Interestingly, the two most similar proteins from the Rosid Arabidopsis (At1g67980 and At1g67990) fall just outside the DIFe/VvAOMT1 clade, while the next most similar protein (At4g26220) groups with methyltransferases from Mesembryanthemum crystallinum (Mg2+-dependent phenylpropanoid and flavonoid O-methyltransferase [PFOMT]) and Stellaria longipes (both Caryophyllales; Ibdah et al., 2003) and the Asterid cyclamen (Akita et al., 2011) in a distant clade. The Arabidopsis and M. crystalinum and S. longipes enzymes can in vitro methylate a broad but distinct range of substrates, which includes caffeoyl esters, flavones, and flavonols, also at hydroxyl groups other than those in the 3′5′ position (Ibdah et al., 2003; Wils et al., 2013). This suggests that Arabidopsis, which does not methylate anthocyanins (Saito et al., 2013), lacks a true DIFe/VvAOMT1 homolog. We found no clear DIFe/VvAOMT1 homologs in monocot species either; the most similar proteins from maize (Zea mays) and rice (Oryza sativa) fall in the CCoA-MT clade. This suggests that these species, and possibly all monocots, lack true DIFe/VvAOMT1 homolog(s).
Figure 4.
Phylogram of A1-type plant O-methyltransferases. The tree (maximum likelihood) is based on a cured sequenced alignment (Supplemental Table S3). Numbers on branches indicate percentage bootstrap support (500 replicates). For each protein, the name, accession number, and substrates accepted in vitro is given. CafCoA, Caffeoyl-CoA; CafAcid, caffeic acids; CafEster, caffeic acid esters; At, Arabidopsis; Ckm, Cyclamen persicum × purpurascens; Fv, F. vesca; Mc, Mesembryanthemum crystallinum; Nt, Nicotiana tabacum; Os, rice; Pc, Petroselinum crispum; Ph, P. hybrida; Pp, P. persica; Pt, Populus trichocarpa; Sl, S. lycopersicum; St, S. tuberosum; Stl, Stellaria longipes; Vv, V. vinifera; Zm, maize; Zv, Zinnia violacea.
Expression Patterns of DIFe Genes
RNA analyses revealed that DIFe2a and DIFe2b are primarily expressed in petals and ovaries but not, or much less, in anthers, sepals, leaves, stems, and roots (Fig. 5A). The expression of both mRNAs is severalfold higher in the petal limb than in the tube and peaks in developing flower buds (stages 1–4) and rapidly declines after opening of the flower (Fig. 5A), similar to other anthocyanin genes such as DIHYDROFLAVONOL REDUCTASE (DFR) and CHALCONE SYNTHASE (Quattrocchio et al., 1993). Although ovaries of Petunia spp. do not accumulate anthocyanins, they do express structural and regulatory anthocyanin genes for an unknown purpose (Huits et al., 1994; Spelt et al., 2000). DIFe1 mRNAs are also abundant in petals. However, DIFe1 is expressed in the petal limb and tube at a similar level at an almost constant level from developing buds (stage 1/2) to open flowers (stage 6). Furthermore, DIFe1 displays, in contrast to DIFe2a and DIFe2, a low background expression in a wide range of other tissues.
Figure 5.
Expression pattern and regulation of DIFe genes. A, Quantitative RT-PCR analysis of MT2, MF1, and MF2 mRNAs in different tissues of the P. hybrida F1 hybrid M1 × V30. Floral tissues were from buds of three different stages. Stage 1/2, buds up to 20 mm; stage 3/4, 30- 40-mm buds; and stage 5/6, open flowers. GLYCERALDEHYDE 3-PHOSPHATE DEHYDROGENASE (GAPDH) and ACTIN were used as internal controls. B, Real-time PCR analysis of mRNAs expressed in stage 3/4 petals of regulatory mutants. Lines R27 (AN1 AN2 AN11), W225 (an1), and W134 (an11) are isogenic. Line W242 is in a distinct genetic background and harbors a mutable an2 allele (an2mut). A germinal AN2REV revertant in which the transposon had excised from AN2 was used as an isogenic wild type. mRNA levels were normalized using ACTIN, and expression in R27 was set to 1. Primers used for real-time PCR could not distinguish between DIFe2a/MF1 and DIFe2b/MF2.
The WD40, HLH, and MYB factors encoded by AN11, AN1, and AN2, respectively, activate a range of structural anthocyanin genes (Koes et al., 2005) and also DIFe genes (Fig. 5B). The DIFe1 genes are expressed in petals of the line R27, while in isogenic an1 and an11 petals, their expression is strongly down-regulated, similar to the expression of DFR. Because no an2 mutants are available in the R27 background, we used a mutant line (W242) harboring an unstable an2 allele with a transposon insertion and a derived germinal revertant (AN2REV) and found that, in this isogenic background, the an2 mutation also reduced DIFe expression to a similar extent as DFR.
In summary, these data show that both DIFe genes are coexpressed in time and place and coregulated by the AN1-AN2-AN11 complex with other anthocyanin genes such as DFR.
Analysis of Mutant DIFe Alleles
To determine whether DIFe1, DIFe2a, and DIFe2b are encoded or regulated by the MT or MF loci, we analyzed their expression in stage 3/4 petal limbs of different P. hybrida genotypes (Fig. 6A). In general, expression of DIFe genes seemed reduced in mt, mf1, or mf2 mutants, but it was difficult to link the observed variation in DIFe gene expression to mt2, mf1, and/or mf2 mutations with certainty, in part, because of the divergent genetic backgrounds of the lines used. DIFe1 mRNA expression is strongly reduced in all accessions containing a mutant mt2 allele but is hardly or not at all affected by mf1 and/or mf2 mutations. In V30 (mf1 mf2), expression of DIFe2a was reduced, suggesting that DIFe2a may be encoded/regulated by MF1 or MF2. In V23 petals, all three DIFe genes seem normally expressed, and expression of DIFe2b seems even somewhat enhanced, indicating that the mf2V32 mutation does not eliminate/reduce the mRNA levels of any of the DIFe genes.
Figure 6.
Structure of wild-type and mutant alleles of MT2 (DIFe1), MF1 (DIFe2b), and MF2 (DIFe2a). A, Quantitative RT-PCR analysis of DIFe mRNAs in genotypes with mt, mf1, and/or mf2 alleles. B, Structure of DIFe1/MT2 locus in P. hybrida (Ph), P. inflata (Pi), and P. axillaris (Pa) lines with different MT2 MF1 MF1 genotypes (+, homozygous for dominant allele; –, homozygous for recessive nonfunctional allele; and ×, homozygous dominant for MF1 and/or MF2). C, Structure of DIFe2a/MF1 and DIFe2b/MF2 locus in P. hybrida (Ph), P. inflata (Pi), and P. axillaris (Pa) lines with different genotypes. Coding and noncoding sequences (introns are not drawn to scale) are indicated by rectangles and lines, respectively, and start and stop codons by white and black circles. Numbers denote the number of nucleotides in (functional) exons and introns. Lesions found in mutant alleles are indicated above the gene maps.
To determine whether the DIFe genes are located at MT2 or the MF loci, we analyzed DIFe1, DIFe2a, and DIFe2b alleles in mutant P. hybrida lines and inbred accessions of the two parental species P. inflata and P. axillaris. The DIFe1 alleles of the P. hybrida mt2 mf1 mf2 lines R78, V32, and M29 all contain an approximately 2.4-kb insertion in intron 5, compared with DIFe1 from M1 (MT2, MF1, and MF2), V30 (MT2, mf1, and mf2) or P. inflata line S6 (Fig. 6B; Supplemental Fig. S2). This insertion can account for the reduced DIFe1 mRNA levels in R78 petals (Fig. 6A) and consists of a tandem repeat of a novel approximately 1,210-bp transposon-like element, designated dTPH11, which has 125-bp imperfect terminal inverted repeats having nine mismatches and is flanked by a 9-bp target site duplication. The P. axillaris lines S1 and S26, which accumulate delphinidins in their petal tubes and lack active MT and MF alleles, also contain this double dTPH11 insertion. The P. axillaris line S2, however, accumulates (3′-methylated) petunidins in the petal tube and contains an active MT2 allele (Jonsson et al., 1984a). The DIFe1S2 allele is identical to that of S6 and M1, except for a few SNPs, and lacks the 2.4-kb insertion or any other obvious lesions, such as nonsense, frameshift, or splice site mutations, indicating that DIFe1S2 is a functional allele (Supplemental Fig. S2). Taken together, this shows that DIFe1 resides at the MT2 locus and that the mutant mt2/dife1 allele arose under natural conditions in P. axillaris and was subsequently introgressed in P. hybrida.
The DIFe2a and DIF2b genes of P. hybrida M1 (MT2 MF1 MF2) and P. inflata S6 have the same intron-exon architecture as DIFe1, though the introns differ in size and sequence (Fig. 6C; Supplemental Figs. S3 and S4). P. hybrida R78 (mt2 mf1 mf2) has a 76-bp deletion at the exon3-intron3 junction of DIFe2a, which can explain the reduced DIFe2a mRNA expression, and a 13-bp deletion in exon 5 of DIFe2b (Fig. 6C; Supplemental Fig. S3). Line V30 (MT2 mf1 mf2) contains the same 13-bp deletion in DIFe2b and multiple insertions and deletions in the coding sequence DIFe2a. This indicates that the two DIFe2 genes do not reside at the MT2 locus but at MF1 and MF2. In P. hybrida R100 (MT2 mf1 MF2), DIFe2a is disrupted by the same 76-bp deletion as in line R78, whereas its DIFe2b allele seemed functional, as it lacks clear lesions such as frameshift, nonsense or, splice site mutations. In P. hybrida V23 (MT MF1 mf2), DIFe2b is disrupted by a 2-bp deletion in exon 4, the same as found in the DIF2eOGB cDNA, while the DIFe2aV23 allele seemed functional. This suggests DIFe2a and DIFe2b are located at the MF1 and MF2 locus, respectively.
The mf1 alleles of P. axillaris S1, S2, and S26 all share a 22-bp deletion in exon 4, the same as in dife2aV30, and an approximately 7-kb insertion in exon 2 with features of a retrotransposon, such as approximately 130-bp terminal direct repeats, and fragments of a putative GROUP SPECIFIC ANTIGEN-POLYMERASE (GAG-POL) gene. Presumably, one or both of these mutations inactivated MF1 in P. axillaris early on, whereas the many other deletions and small insertions that are specific for each P. axillaris line presumably occurred later, after the gene was inactivated. The mf2 allele of P. axillaris S1, S2, and S26 contained two deletions, of 2 and 13 bp, which are identical to the 2- and 13-bp deletions found in the distinct P. hybrida mf2 alleles (Fig. 6C; Supplemental Fig. S4). This suggests that mf2 alleles arose under natural conditions in P. axillaris and were introgressed in P. hybrida, although it remains unclear how two distinct deletions in a single P. axillaris allele ended up in two separate P. hybrida alleles.
Complementation of P. hybrida mt and mf Mutants
Because the wild-type progenitors of mt and mf mutants are not available, their genetic and biochemical characterization relied on comparison with MT or MF lines in unrelated genetic P. hybrida backgrounds (Wiering and De Vlaming, 1977; Jonsson et al., 1983, 1984b), which can be as different as that of distinct Petunia species. To ascertain that the mt2 mf1 mf2 phenotype is due to the mutations in DIFe genes and to compare mutants with the isogenic wild types, we expressed MT2, MF2, and VvAOMT1 from the constitutive 35S promoter in two distinct mt2 mf1 mf2 backgrounds.
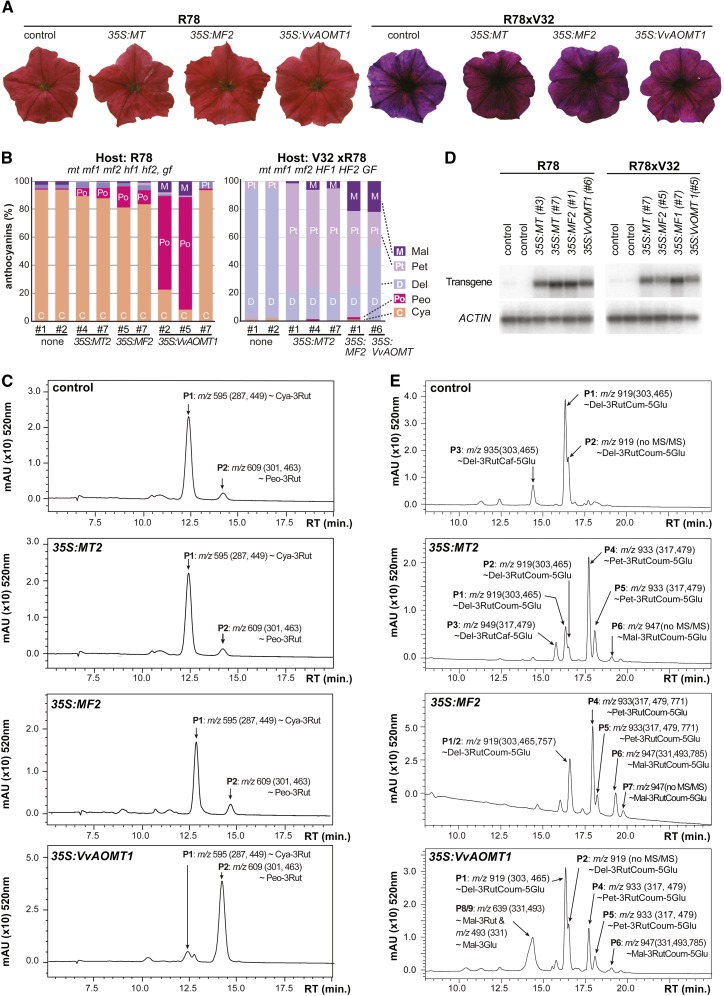
The inbred line R78 (mt2 mf1 mf2) has additional mutations that block 5′ hydroxylation (hf1 hf2) and 5-glycosylation and acylation (gf), resulting in a red flower color (Fig. 7A; Supplemental Table S4). HPLC of the anthocyanidins showed that most anthocyanins in R78 petals are cyanidin derivatives (approximately 94%), while peonidin, delphinidin, petunidin, and malvidin derivatives were found in small amounts only (<2.5% each; Fig. 7B; Supplemental Fig. S5). Both cyanidin and peonidin accumulated as a single anthocyanin species that produced ions with a mass-to-charge ratio (m/z) of 595.2 and 609.2, respectively, in positive mode mass spectroscopy [MS(+)] analysis and ion fragments with m/z values corresponding to loss of a rhamnose or a rutinoside moiety, indicating that they are cyanidin and peonidin 3-rutinosides, respectively (Fig. 7C; Supplemental Fig. S6).
Figure 7.
Complementation of mf and mt mutants by 35S:MT2, 35S:MF2, and 35S:VvAOMT1. A, Flowers of untransformed R78 and R78 × V32 plants and transgenic siblings that express 35S:MT2, 35S:MF2, or 35S:VvAOMT1. B, Composition of anthocyanidins obtained by acid hydrolysis of anthocyanins in petals of transgenic and control plants, as determined by HPLC. Number signs indicate specific transformants. C, LC-MS/MS analysis of anthocyanins in R78 petals (control) and transgenic siblings expressing 35S:MT2, 35S:MF2, or 35S:VvAOMT1. D, Quantitative RT-PCR analysis of transgene expression in distinct transformants. Number signs denote individual transformants. E, LC-MS/MS analysis of anthocyanins in R78 × V32 petals (control) and transgenic siblings expressing 35S:MT2, 35S:MF2, or 35S:VvAOMT1. Diagrams in C and E show LC profiles and, for the major peaks, m/z values of the corresponding ion and, in brackets, subfragments observed in MS/MS(+) spectra (see Supplemental Figs. S4–S9).
Five of eight 35S:MT2 and 10 of 12 35S:MF2 transformants expressed the transgene and produced similar amounts of mRNA (Fig. 7D). Expression of MT2 or MF2 increased the level of peonidins only modestly from approximately 1% to 6.4% ± 1% and 12.5% ± 3.5% of the total amount of anthocyanidins (Fig. 7B). Liquid chromatography (LC) tandem mass spectrometry (MS/MS) showed that the peonidins in these 35S:MT2 and 35S:MF2 petals were 3-rutinosides (Fig. 7C; Supplemental Figs. S5 and S6). AMT enzymes in crude petal extracts from different P. hybrida lines could methylate anthocyanins with acylated 3-rutinoside and 5-Glc moieties at 60 µm concentration (Km in 2–20 µm range, depending on the genotype used), but no conversion of 3-glucosides or 3-rutinosides was seen at all at a 60 µm concentration and only relatively little at 1 mm concentration, suggesting a Km for these substrates in the micromolar range (Jonsson et al., 1982, 1984b). This suggested that MT2 and MF1 methylated only a small fraction of the cyanidin 3-rutinosides in the R78 background because they have a too low affinity/efficiency for these substrates. To test this hypothesis, we expressed in R78 the homologous grape protein VvAOMT1, which can efficiently methylate anthocyanin 3-glucosides and 3-rutinosides in vitro with a Km in the approximately 7 to 40 µm range, depending on the conditions used (Hugueney et al., 2009; Lücker et al., 2010). Four of the seven 35S:VvAOMT1 R78 transformants expressed the transgene at roughly similar levels as 35S:MT2 and 35S:MF2 (Fig. 7D), which caused a very modest change in petal color that was only evident in side-by-side comparisons with control flowers and difficult to capture by photography (Fig. 7A), and increased the amount of peonidins from 1% to about 70% of the total anthocyanins (Fig. 7B). These peonidins accumulated as 3-rutinosides (Fig. 7D; Supplemental Fig. S6). These data show that VvAOMT1 can efficiently methylate cyanidin 3-rutinoside and/or the cyanidin 3-glucoside precursor in vivo, whereas MT2 and MF2 do so with low efficiency/kinetics only.
Next, we introduced the same transgenes into an F1 hybrid (R78 × V32) that is mt2 mf1 mf2 but harbors dominant alleles of HYDROXYLATION AT FIVE1 (HF1), HF2, and GF, resulting in a purple flower color (Fig. 7A; Supplemental Table S4). The major anthocyanins in R78 × V32 petals are delphinidins, occurring in at least three different molecular species (Fig. 7B; Supplemental Fig. S7). The most abundant species (peak 1) produced an MS(+) ion with m/z 919.2 and fragments with m/z 465.1 (delphinidin 5-glucoside) and 303.0 (delphinidin), suggesting it to be a delphinidin 3-(p-coumaroyl)-rutinoside 5-glucoside (Fig. 7B; Supplemental Fig. S5). Peak 2 has the same m/z ratio but a slightly longer retention time and is most likely a stereoisomer with a sugar and/or p-coumaroyl group in a different position. The positive mode tandem mass spectroscopy [MS/MS(+)] spectrum of peak 3, a less abundant anthocyanin, was consistent with delphinidin 3-(caffeoyl)-rutinoside 5-glucoside (Supplemental Fig. S7F).
Thirteen of 20 35S:MT2, five of six 35S:MF2, and three of six 35S:VvAOMT1 transformants in the R78 × V32 background expressed the transgene to similar levels as the R78 transformants (Fig. 7D), which resulted in a slightly more reddish petal color (Fig. 7A). HPLC showed that, in R78 × V32 petals expressing 35S:MT2, most delphinidins are converted into (3′-methylated) petunidins and a small amount of (3′5′-methylated) malvidins, indicating that MT2 has strong A3′MT activity and a slight A3′5′MT activity (Fig. 7B). 35S:MF2 also converted delphinidins into petunidins and malvidins, but the malvidin/petunidin ratio was much higher than in 35S:MT2, suggesting that MF2 has a stronger A3′5′MT activity than MT2. Although VvAOMT1 expression resulted in methylation of only some 50% of the delphinidins, the ratio of malvidin/petunidin products was even higher than in 35S:MF2 petals, indicating that VvOMT1 has, in vivo, both A3′MT and A3′5′MT activity.
LC-MS/MS showed that expression of MT2 or MF2 in R78 × V32 petals yielded several new anthocyanins (peaks 4–7; Fig. 7E). Peak 4 produced a positive molecular ion with m/z 933.3 and fragments with m/z 771.2, 479.1, and 317.1, consistent with petunidin (p-coumaroyl) rutinoside-5-glucoside (Fig. 7E; Supplemental Figs. S8 and S9). This structure is supported by an additional fragment with m/z 623.1 in negative mode mass spectrometry [MS(−)] spectra, originating from the loss of a Glc and p-coumaroyl group (Supplemental Fig. S10). Peak 5 also has m/z 933.3 in MS(+), suggesting that it is a positional isomer. Peaks 6 and 7 were identified in a similar way as positional isomers of malvidin 3-(p-coumaroyl)rutinoside 5-glucoside (Fig. 7E; Supplemental Figs. S8 and S9).
Expression of VvAOMT1 in R78 × V32 petals yielded two isoforms of petunidin 3-(p-coumaroyl)rutinoside 5-glucoside (peaks 4 and 5), similar to MT2 and MF2 expression. Surprisingly, the produced malvidins were not malvidin 3-(p-coumaroyl)rutinoside 5-glucoside (peak 6), as in 35S:MF2, but malvidin 3-glucoside and malvidin 3-rutinoside (peaks 8 and 9; Fig. 7E; Supplemental Fig. S11), which are not normally found in Petunia spp. or mutants. This shows again that VvAOMT1 can, unlike MT2 and MF2, methylate 3-rutinosides and/or 3-glucosides and reveals, unexpectedly, that precocious 3′5′ methylation of rutinosides blocks the acylation and 5-glucosylation by the endogenous Petunia spp. enzymes.
Functional Divergence of Methyltransferases
Analysis of plant methyltransferases belonging to other clades than CCoA-MTs and AMTs suggested that their substrate specificities are determined by few amino acids (Wang and Pichersky, 1999; Zubieta et al., 2001, 2003). The Petunia spp. and grape AMTs and CCoA-MTs share high similarity throughout the protein, suggesting that their divergent substrate specificities also result from relatively small structural differences (Fig. 8A). Most residues of Medicago sativa (Ms) CCoA-MT that are involved in binding the SAM and caffeoyl-CoA substrates or dimerization (Ferrer et al., 2005) are conserved in the AMTs (Fig. 8A). This suggests that AMTs form, like Ms CCoA-MT, dimers with a catalytic site in each monomer, which is consistent with the size of AMT activities observed in crude petal extracts (approximately 50 kD; Jonsson et al., 1984b). Residues 200 to 215 in Ms CCoA-MT form one side of the pocket that holds the caffeoyl group and are conserved among CCoA-MTs (Ferrer et al., 2005) but not in MT2, MF1, and MF2 and VvAOMT1 (Fig. 8A). Residues 50 to 64 constitute a second region where CCoA-MTs diverged from the AMTs.
Figure 8.
Two possible binding modes of delphinidin 3-(p-coumaroyl) rutinoside 5-glucoside. A, Alignment of CCoA-MTs and AMTs. Amino acids and motifs of Ms CCoA-MT involved in dimerization, binding of SAM/S-adenosyl-L-homo-Cys (SAH), the caffeoyl or CoA moiety, and divalent cation are indicated. B, Substrate binding with the 3-(p-coumaroyl)rutinoside chain pointing into the active site and clashing with bulky residues (Tyr-208 and Tyr-212), which are, in the Petunia spp. AMTs, replaced by smaller ones (Gly and Leu). For clarity, the 5-Glc group of the anthocyanin is not shown. C, Side view of B. D, Substrate binding with the 3-(p-coumaroyl)rutinoside chain oriented toward the active site and clashing with bulky residues (Trp-58, Asn-116, and Tyr 208), which are, in the Petunia app. AMTs, replaced by Ala/Gly, Ala, and Gly, respectively. For clarity, the 5-Glc group of the anthocyanin is not shown. E, Side view of D.
To study effects of these alterations on substrate binding, we used the crystal structure of Ms CCoA-MT (Protein Data Bank structure 1SUI; Ferrer et al., 2005) and Molecular Operator Environment software (version 2011, Chemical Computing Group) to superpose the trihydroxylated C-ring of anthocyanin onto the methylated ring of the sinapoyl-CoA product bound to Ms CCoA-MT (Supplemental Fig. S12, A and B). This position of the C-ring forces the 3-Glc or 3-rutinoside chains to orientate toward Tyr-208; other binding conformations lead to steric clashes with (one of the) bulky residues at positions 190, 193, and 238 (Supplemental Fig. S12, B and C), which are conserved in the AMTs (Fig. 8A). In case of a delphinidin 3-(p-coumaroyl)rutinoside substrate without a 5-Glc group, the 3-(p-coumaroyl)rutinoside moiety can adopt an orientation as depicted in Supplemental Figure S12, D and E. However, our computational analysis suggests that the 5-Glc moiety of the MT and MF substrate is orientated as depicted in Supplemental Figure S12F to avoid steric hindrance with N-terminal residues of the enzyme (e.g. Lys-21). This would, in turn, result in intrusion of the 5-Glc moiety into the 3-(p-coumaroyl) rutinoside chain (Supplemental Fig. S12F), which offers a possible explanation for the observed replacement of large bulky active-site residues such as Tyr-208 and Tyr-212 by smaller Gly or Leu residues, to enable occupancy by the 3-(p-coumaroyl)rutinoside group of the corresponding pocket within the active site (Fig. 8, B and C). Upon the Tyr208Gly conversion in the Petunia spp. AMTs, it may even be possible for the substrate to adopt a conformation as shown in Figure 8, D and E. This would require replacement of His-56, Trp-58, and Asn-118 by smaller residues, which is observed in MT2, MF1, and MF2 (Fig. 8A).
DISCUSSION
The pathway, enzymes, and genes involved in the synthesis of simple anthocyanin 3-glucosides is widely conserved and has been studied in great detail in a variety of species. The subsequent decorations with additional sugar, acyl, and methyl groups, which account for most of the structural variation of anthocyanin from different species, has remained largely obscure. The identification of AMT genes and mutants described here shows that alterations in the expression or the substrate specificities of decorating enzymes lead to chemical anthocyanin diversity.
Genetic Control of AMT Activities in Petunia spp. Flowers
Using mutants, we show that the three DIFe genes identified here are essential for methylation of anthocyanins in Petunia spp. and are located in the MT2, MF1, and MF2 loci. The finding that mf1 and mf2 mutants have lesions in two distinct genes (DIFe2a and DIFe2b) proves unequivocally that MF1 and MF2 are closely related paralogs that presumably arose through a recent gene duplication (shortly) before the separation of the P. axillaris and P. inflata lineages. DIFe1 is located at MT2, as all P. hybrida accessions lacking active MT gene(s) contain a nonfunctional dife1 allele and 3′ methylation can be restored by 35S:DIFe1. The existence of a second MT gene (MT1) that should be tightly linked with MF1 was postulated because 3′ methylation was thought to be an essential prerequisite for subsequent 5′ methylation by an MF gene. Given that MF1 and MF2 can produce both 3′- and 3′5′-methylated anthocyanins and that we found no evidence for a fourth DIFe gene, there is no indication for the existence of MT1.
Plant O-methyltransferases constitute a large family with a wide spectrum of substrates (Lam et al., 2007), which were, in most cases, inferred by testing various compounds in in vitro assays or from sequence similarity to enzymes for which in vitro data are available. Our in vivo data show that such results should be interpreted with some caution. For example, the MT2 and MF enzymes exhibit high similarity to proteins annotated as CCoA-MTs but apparently cannot methylate these substrates. Petunia spp. flowers only express activities that can methylate anthocyanins, but not cinnamic acids, and do not contain ferulic or sinapic acids (Jonsson et al., 1982). In vitro assays of methyltransferases often employ substrate concentrations in the 0.1 to 1 mm range (Gang et al., 2002; Ibdah et al., 2003; Kim et al., 2006; Akita et al., 2011). Using similar anthocyanin substrate concentrations (0.4–1 mm), the AMTs in crude petal extracts, which originate from the three genes identified here, and recombinant MT2/DIFe1 protein could methylate anthocyanin 3-glucosides and 3-rutinosides in vitro (Jonsson et al., 1982; Fig. 2). In vivo, however, only little methylation of 3-rutinosides or their 3-glucoside precursors was seen, whereas efficient methylation occurred in petals synthesizing anthocyanin (3-acyl)rutinoside 5-glucosides, which matches well with the Km values determined for AMTs in crude petal extracts (Jonsson et al., 1982). However, even when Km, the catalytic constant (Kcat), and Kcat/Km ratios for potential substrates are known, it remains difficult to ascertain the function in the absence of in vivo data, as (1) the in vivo concentrations of potential substrates (in the cytoplasm) are difficult to determine, (2) the primary in vivo substrate(s) may have escaped attention in the in vitro assays, and (3) enzymes competing for the same substrate are absent.
Gain and Loss of Functional AMT Genes during Evolution
It is thought that the enzymes involved in anthocyanin synthesis evolved from enzymes in primary metabolism (Winkel-Shirley, 2001). The DIFe genes apparently originate from genes encoding methyltransferases involved in phenylpropanoid metabolism, which required relatively small changes in the coding sequence to alter the substrate specificity of the encoded enzymes and changes in cis-regulatory elements that brought them under the control of the MYB-HLH-WD40 complex that also regulates other anthocyanin genes. Given that monocots such as maize and rice do synthesize 3′- and/or 3′5′-methylated anthocyanins (Styles and Ceska, 1972; Yoshimura et al., 2012), but lack clear DIFe1/VvAOMT1 homologs, AMT genes may have evolved multiple times from different ancestral genes.
Subsequent modifications of AMT expression patterns as well as complete inactivation contributed to the diversification of anthocyanins. Petunia species and varieties with functional methylation genes accumulate (3′5′-methylated) malvidins in the petal limb and (3′-methylated) petunidins in the tube. This seems largely due to the divergent expression patterns of MT2, which is about equally expressed in the petal limb and tube, and the MF genes, which are highly expressed in the limb and much less in the tube, indicating that tissue specificity of decoration patterns can arise by alterations in cis-regulatory elements of some AMT genes.
P. axillaris accessions with limited (line S2) or no anthocyanin methylation (S1 and S26) contain AMT genes with disrupting mutations. This indicates that (1) the last common ancestor of P. axillaris and P. inflata possessed functional MF1, MF2, and MT2 alleles and synthesized malvidins and (2) that P. axillaris subsequently lost this capacity through loss-of-function mutations. Because the moth-pollinated P. axillaris flowers lack anthocyanins in the petal limb, where MF genes are mostly expressed, the inactivation of MF1 and MF2 might be due to the decreased selective pressure after limb pigmentation was lost by inactivation of the MYB transcription factor AN2 (Quattrocchio et al., 1999; Hoballah et al., 2007). Because Arabidopsis lacks a clear MT/MF/VvAOMT1 homolog, the absence of methylated anthocyanins in this species (Saito et al., 2013) probably resulted from loss-of-function mutations as well.
Consequences of Altered AMT Substrate Specificities: The Last Will Be First and the First Will Be Last
Other more subtle alterations in the coding sequences, which modified the substrate specificities of the AMTs, contributed to the diversification of the anthocyanins in at least two different ways. Distinct decorations of the anthocyanin could, in theory, take place in any order. Biochemical and genetic data indicate, however, that anthocyanin decoration is, in Petunia spp. (an Asterid), a linear pathway rather than a grid and that different decorations are added in the strict order depicted in Figure 1A. The pathway that modifies anthocyanins in Arabidopsis (a Rosid) is, although entirely different from the Petunia spp. pathway, also more linear than grid shaped (Saito et al., 2013). Interestingly, the anthocyanins of Silene spp., which represent a third clade of eudicots (Caryophyllales) that is distinct from Rosids and Asterids, are glycosylated, rhamnosylated, and acylated in the same way (i.e. in the same position and in the same order) as in Petunia spp., though methylation does not occur (Kamsteeg et al., 1979).
The strict order of modifications is established in two ways. Some enzymes add substituents that are subsequently modified by another enzyme. For example, the rhamnosyltransferase encoded by RT has to add the rhamnose group before it can be acylated. Such directly interdependent reactions can only run in a fixed order that cannot be tinkered with during evolution. For other reactions, their relative order is determined by the substrate specificities of the enzymes, and this is, in principle, more flexible. For example, in Petunia spp., acylation of the rhamnose group is required prior to 5-glucosylation and subsequent methylation because the 5GT does not accept anthocyanins without the acyl group as a substrate (Jonsson et al 1984c; Yamazaki et al., 2002) and MT2, MF1, and MF2 do not accept substrates without the acyl and/or 5-Glc moieties. This suggests that the substrate specificities of late enzymes, such as MT2 and MF1/MF2, were adapted to that of enzymes catalyzing preceding steps, such as AAT and 5GT. It also envisages that alteration of the substrate specificity of AMTs to circumvent (or establish) the necessity for prior acylation or 5-glucosylation may produce a new repertoire of anthocyanins. The finding that VvAOMT1 can, unlike MT2 and MFs, methylate anthocyanin-rutinosides (or their 3-glucoside precursors), demonstrates that substrate specificities of homologous AMTs diverged substantially and explains why Vitis species and varieties synthesize simple 3-glucosides or acetylated 3-glucosides of peonidin, petunidin, and malvidin (Acevedo De la Cruz et al., 2012), while Petunia species or mutants cannot (Wiering, 1974; Ando et al., 1999).
Surprisingly, the malvidin 3-rutinosides and 3-glucosides found in Petunia spp. petals expressing VvAOMT1 are not acylated or 5-glycosylated. This suggests that the (precociously methylated) malvidins derivatives are not accepted by the Petunia spp. AAT or 5GT enzyme as substrates. This implies that alterations in the specificity of one decorating enzyme, even the last enzyme of the pathway, may reshuffle the order in which decorations are added. Such a reshuffling of an entire part of a pathway provides new possibilities, but also constraints, by which evolutionary tinkering with substrate specificities of individual enzymes can create chemical diversity. Moreover, it suggests that the substrate specificities of distinct decorating enzymes are adjusted to one another in unforeseen ways, as late enzymes (in Petunia spp., AMTs) are adapted to earlier enzymes (in Petunia spp., AAT and 5GT) but also vice versa. Analysis of additional decorating enzymes, such as the AAT encoded by GF and homologs in other species may shed further light on this issue.
MATERIALS AND METHODS
Plant Material
Inbred Petunia hybrida lines were derived by inbreeding of cultivars from various breeders in the 1960s and 1970s and maintained by inbreeding (>20 generations) ever since and are, consequently, homozygous for all known genes. Genotypes, as determined by H. Wiering and P. de Vlaming (unpublished data) in the 1970s and 1980s using testcrosses and chromatographic analysis of anthocyanins, are given in Supplemental Table S4. Independently inbred lines are in distinct genetic backgrounds and essentially comprise different mixtures of the parental Petunia inflata and Petunia axillaris genomes. Lines W225 (an1 AN11) and W134 (AN1 an11) derived by transposon insertions and excisions from the isogenic progenitor line R27 (AN1 AN11; de Vetten et al., 1997; Spelt et al., 2000). W240 contains an unstable an2 allele with a transposon insertion, and we used a spontaneous revertant, originating from a germinal transposon excision, as an isogenic AN2+ genotype. (Verweij et al., 2008). OGB is a commercial variety from Ball Seeds (http://www.ballhort.com) and may be heterozygous for certain genes, including anthocyanin genes. Plants were grown and maintained throughout the year in a greenhouse with supplemental artificial lighting (cycles of 16 h light and 8 h darkness) at a (minimum) temperature of 22°C or higher. Tissues used for analysis of gene expression and anthocyanin metabolites were harvested in the summer period (May–September), taking care that, for direct comparisons, tissues were harvested simultaneously.
Isolation and Analysis of DIFe Genes
A cDNA library (1 × 106 plaques) was constructed with a ZAP-cDNA Gigapack III Gold Cloning kit (Stratagene; http://www.genomics-agilent.com) using poly(A+) RNA isolated from petal tissue of P. hybrida OGB and screened by hybridization with a DIFe1 cDNA from V26. Genomic DNA sequences of P. inflata and P. hybrida were obtained using primers complementary to 5′ UTR and 3′ UTR sequences of the DIFe1 and DIFe2 cDNAs from OGB and V26. P. axillaris genomic sequences were obtained using primers (Supplemental Table S5) designed on (preliminary) genome sequence data provided by the Petunia spp. platform.
Enzymatic Activity of DIFe1/MT2
For expression in Escherichia coli, the DIFe1 (MT2) cDNA from OGB was amplified with the primers 1907BamHI-FW (GCATGGATCCACAGGCAAAACCGCCCACCCTG) and 1907PstI-RV (GCATCTGCAGCTAGGAGAG ACGCCTGCAAAG), digested with BamHI and PstI and ligated into the E. coli expression vector pQE30 (Qiagen) to yield pCGP3086. Cell-free extracts were assayed for enzyme activity as described (Jonsson et al., 1983) using 0.5 mm D3G, D3R, or delphinidin 3,5-diglucoside (obtained from Industrial Research Limited, Gracefield Research Centre) and SAM as a methyl donor. Hydrolyzed reaction products (anthocyanidins) were analyzed by HPLC, and those obtained using 14C-SAM (Amersham Pharmacia) were analyzed by thin-layer chromatography and autoradiography, alongside standard samples of petunidin, malvidin, and delphinidin (obtained from Roth).
Phylogenetic Analysis
Phylogenetic trees were calculated using a software suite at http://www.phylogeny.fr (Dereeper et al., 2008). Therefore, sequences were aligned with MUSCLE and used to generate a neighbor-joining tree using BioNJ or, after curation of the alignment (Gblocks), a maximum likelihood tree using PhyML, which were visualized with TreeDyn.
DNA and RNA Gel-Blot Hybridization
DNA isolated from leaves (6 µg) was digested with EcoRI or BamHI, size separated on a 0.9% (w/v) agarose gel, blotted to nylon membranes by capillary transfer, hybridized at 42°C overnight in DIG Easy-Hyb buffer (Roche; http://www.roche-applied-science.com) using a digoxigenin-labeled MF1 or MT2 cDNA probe, washed under low-stringency conditions (2× SSC, 25°C), treated with high-affinity antidigoxigenin antibodies conjugated to alkaline phosphatase, incubated with the chemiluminescent substrate CSPD (disodium 3-(4-methoxyspiro {1,2-dioxetane-3,2′-(5′-chloro)tricyclo [3.3.1.13,7]decan}-4-yl)phenyl phosphate), the membrane, and exposed to x-ray film. Subsequently, the blot was rewashed at high stringency (0.1× SSC, 68°C) and again exposed (1× SSC is 15 mm Na3-citrate and 150 mm NaCl). The same membrane was hybridized twice, checking that any residual signal was completely eliminated from the membrane before the following hybridization. RNA gel-blot analysis was done as described previously (van Tunen et al., 1988).
Quantitative and Real-Time PCR
Real-time PCR was performed using Power SYBER Green (Applied Biosystems; http://appliedbiosystems.com) using primers shown in Supplemental Table S6. Normalization was done based on the expression of ACTIN. Because MF1 (DIFe2a) and MF2 (DIFe2b) mRNAs were difficult to distinguish with primers suitable for real-time PCR, we used a previously described quantitative reverse transcription (RT)-PCR procedure (Souer et al., 2008) instead. This involved the amplification of specific mRNA products with gene-specific primers (Supplemental Table S7) using a reduced number of amplification cycles (20–25, depending on the abundance of the mRNA), to avoid reaching saturation and ensure a linear response, and detection of PCR products by DNA gel-blot hybridization with 32P-labeled cDNA probes and phosphorimaging, which ensures a quantitative detection of PCR products.
Transgenic Plants
The coding sequence of VvAOMT1 was amplified from berry cDNA using primers VvOMT1-F+attB1 and VvOMT1-R+attB2 and MT2 from P. hybrida M1 petal cDNA with primers PhMT-F+attB1 and PhMT-R+attB2, recombined into the Gateway entry vector pDonor221 (Invitrogen), and subsequently recombined into destination vector pK2GW7.0 (Karimi et al., 2002). The PhMF2 coding sequence was amplified from P. hybrida M1 petal cDNA with PhMF-F Topo primer and PhMF-R primer, cloned in pENTR/D Topo (Invitrogen; http://www.lifetechnologies.com), and subsequently recombined into pK2GW7.0. Antisense constructs were made by ligating DIFe1OGB and DIFeOGB2 cDNA in antisense orientation between the MAC promoter and Mannopine synthase 3′ end (Comai et al., 1990). Constructs were introduced into Agrobacterium tumefaciens strain AGL0 or LBA4404 used to infect Petunia spp. leaf discs. Primer sequences are shown in Supplemental Table S8.
Analysis of Anthocyanins by LC Ion Trap Time-of-Flight MS with Photo-Diode Array
About 0.5 g of fresh weight of petals were lyophilized, and their anthocyanins were extracted in 10 volumes of 50% (v/v) acetonitrile containing 0.1% (v/v) trifluoroacetic acid. The extract was filtrated with polyvinylidene difluoride membranes (0.45 μm; Merck Millipore; http://www.millipore.com), and 5 μL of the filtrate was injected to an Inertsil ODS-4 column (4.6 mm × 250 mm; pore size, 5 μm; GL-Sciences Inc., http://www.glsciences.com) for anthocyanin analysis using LCMS-IT-TOF (Shimadzu; http://www.shimadzu.com) attached with a photo diode array (Shimadzu). The solvent system used included a linear gradient elution for 30 min from 90% solvent A (0.1% [v/v] formic acid in water) and 10% solvent B (0.1% [v/v] formic acid and 90% [v/v] acetonitrile in water) to 50% solvent A and then, for 15 min, 100% solvent B. Different gradients were used occasionally. The flow rate was 0.6 mL min–1, and the column temperature was 40°C. The absorption spectra were monitored from 250 to 600 nm. MS and MS/MS were analyzed in m/z ranges of from 250 to 1,000 and from 100 to 1,000, respectively, in a positive mode after electrospray ionization.
Sequence data from this article can be found in the GenBank/EMBL databases under the following accession numbers: KJ639934 (Pi MT2), KJ639935 (Pi MF1), KJ639936 (Pi MF2), KJ676512 (Ph MF1M1), KJ676513 (Ph MTOGB), KJ676514 (Ph MF2M1), and KJ676515 (Ph MTV30).
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure S1. Cladogram of plant O-methyltransferases.
Supplemental Figure S2. Aligned sequences of wild-type and mutant MT2/DIFe1 alleles.
Supplemental Figure S3. Aligned sequences of wild-type and mutant MF1/DIFe2a alleles.
Supplemental Figure S4. Aligned sequences of wild-type and mutant MF2/DIFe2a alleles.
Supplemental Figure S5. HPLC-Photo-Diode Array analysis of anthocyanidins and anthocyanins in petals expressing 35S:MT, 35S:MF2, or 35S:VvAOMT1.
Supplemental Figure S6. LC-MS/MS(+) analysis of anthocyanins in R78 petals and siblings expressing 35S:MT2, 35S:MF2, or 35S:VvAOMT1.
Supplemental Figure S7. LC-MS/MS(+) analysis of anthocyanins in R78 × V32 petals.
Supplemental Figure S8. LC-MS/MS(+) analysis of anthocyanins in R78 × V32 petals expressing 35S:MT2.
Supplemental Figure S9. LC-MS/MS(+) analysis of anthocyanins in R78 × V32 petals expressing 35S:MF2.
Supplemental Figure S10. LC-MS/MS(–) analysis of peak 4 from R78 × V32 petals expressing 35S:MF2.
Supplemental Figure S11. LC-MS/MS(+) analysis of anthocyanins in R78 × V32 petals expressing 35S:VvAOMT1.
Supplemental Figure S12. Structural models of Ms CCoA-MT in complex with anthocyanin substrates.
Supplemental Table S1. Methyltransferase activity in E. coli cells expressing DIFe1.
Supplemental Table S2. Genotypes of Petunia spp. lines and derived F1 hybrids.
Supplemental Table S3. Primers used for amplification of MT2, MF1, and MF2 alleles.
Supplemental Table S4. Primer combinations used for quantitative RT-PCR.
Supplemental Table S5. Primers used for real-time PCR.
Supplemental Table S6. Primers used for construction of fusion genes.
Supplemental Table S7. Sequence alignment used for the phylogenetic tree in Supplemental Figure S1 (Fasta format).
Supplemental Table S8. Sequence alignment used for the phylogenetic tree in Figure 4 (Fasta format).
Supplementary Material
Acknowledgments
We thank Pieter Hoogeveen, Martina Meesters, Daisy Kloos, and Gianna Kalc for plant care and Lisa Keam, Noelene Treloar, and Belinda Sheehan for technical assistance.
Glossary
- AAT
anthocyanin acyltransferase
- 5GT
5-glucosyltransferase
- AMT
anthocyanin methyltransferase
- A3′MT
anthocyanin 3′-methyltransferase
- A3′5′MT
anthocyanin 3′5′-methyltransferase
- cDNA
complementary DNA
- D3G
delphinidin 3-glucoside
- D3R
delphinidin 3-rutinoside
- SAM
S-adenosyl-l-Met
- VR
V23 x R51
- OGB
cultivar Old Glory Blue
- UTR
untranslated region
- SNP
single nucleotide polymorphism
- CCoA-MT
caffeoyl-Coenzyme A O-methyltransferase
- Ms
Medicago sativa
- RT
reverse transcription
- LC
liquid chromatography
- MS/MS
tandem mass spectrometry
- m/z
mass-to-charge ratio
- MS(+)
positive mode mass spectroscopy
- MS(−)
negative mode mass spectroscopy
- MS/MS(+)
positive mode tandem mass spectroscopy
Footnotes
This work was supported by a European Cooperation in Science and Technology fellowship (COST–STSM–858–4287 to S.P.).
The online version of this article contains Web-only data.
References
- Acevedo De la Cruz A, Hilbert G, Rivière C, Mengin V, Ollat N, Bordenave L, Decroocq S, Delaunay JC, Delrot S, Mérillon JM, et al. (2012) Anthocyanin identification and composition of wild Vitis spp. accessions by using LC-MS and LC-NMR. Anal Chim Acta 732: 145–152 [DOI] [PubMed] [Google Scholar]
- Akita Y, Kitamura S, Hase Y, Narumi I, Ishizaka H, Kondo E, Kameari N, Nakayama M, Tanikawa N, Morita Y, et al. (2011) Isolation and characterization of the fragrant cyclamen O-methyltransferase involved in flower coloration. Planta 234: 1127–1136 [DOI] [PubMed] [Google Scholar]
- Alfenito MR, Souer E, Goodman CD, Buell R, Mol J, Koes R, Walbot V. (1998) Functional complementation of anthocyanin sequestration in the vacuole by widely divergent glutathione S-transferases. Plant Cell 10: 1135–1149 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ando T, Saito N, Tatsuzawa F, Kakefuda T, Yamakage K, Ohtani E, Koshi-ishi M, Matsusake Y, Kokubun H, Watanabe H, et al. (1999) Floral anthocyanins in wild taxa of Petunia (Solanaceae). Biochem Syst Ecol 27: 623–650 [Google Scholar]
- Brouillard R, Chassaing S, Fougerousse A. (2003) Why are grape/fresh wine anthocyanins so simple and why is it that red wine color lasts so long? Phytochemistry 64: 1179–1186 [DOI] [PubMed] [Google Scholar]
- Brugliera F, Holton TA, Stevenson TW, Farcy E, Lu CY, Cornish EC. (1994) Isolation and characterization of a cDNA clone corresponding to the Rt locus of Petunia hybrida. Plant J 5: 81–92 [DOI] [PubMed] [Google Scholar]
- Buer CS, Imin N, Djordjevic MA. (2010) Flavonoids: new roles for old molecules. J Integr Plant Biol 52: 98–111 [DOI] [PubMed] [Google Scholar]
- Comai L, Moran P, Maslyar D. (1990) Novel and useful properties of a chimeric plant promoter combining CaMV 35S and MAS elements. Plant Mol Biol 15: 373–381 [DOI] [PubMed] [Google Scholar]
- de Vetten N, Quattrocchio F, Mol J, Koes R. (1997) The an11 locus controlling flower pigmentation in petunia encodes a novel WD-repeat protein conserved in yeast, plants, and animals. Genes Dev 11: 1422–1434 [DOI] [PubMed] [Google Scholar]
- de Vetten N, ter Horst J, van Schaik HP, de Boer A, Mol J, Koes R. (1999) A cytochrome b5 is required for full activity of flavonoid 3′,5′-hydroxylase, a cytochrome P450 involved in the formation of blue flower colors. Proc Natl Acad Sci USA 96: 778–783 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Vlaming P, Cornu A, Farcy E, Gerats AGM, Maizonnier D, Wiering H, Wijsman HJW. (1984) Petunia hybrida: a short description of the action of 91 genes, their origin and their map location. Plant Mol Biol Rep 2: 21–42 [Google Scholar]
- Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, Dufayard JF, Guindon S, Lefort V, Lescot M, et al. (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36: W465–W469 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferrer JL, Zubieta C, Dixon RA, Noel JP. (2005) Crystal structures of alfalfa caffeoyl coenzyme A 3-O-methyltransferase. Plant Physiol 137: 1009–1017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gang DR, Lavid N, Zubieta C, Chen F, Beuerle T, Lewinsohn E, Noel JP, Pichersky E. (2002) Characterization of phenylpropene O-methyltransferases from sweet basil: facile change of substrate specificity and convergent evolution within a plant O-methyltransferase family. Plant Cell 14: 505–519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grotewold E. (2006) The genetics and biochemistry of floral pigments. Annu Rev Plant Biol 57: 761–780 [DOI] [PubMed] [Google Scholar]
- Hoballah ME, Gübitz T, Stuurman J, Broger L, Barone M, Mandel T, Dell’Olivo A, Arnold M, Kuhlemeier C. (2007) Single gene-mediated shift in pollinator attraction in Petunia. Plant Cell 19: 779–790 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hugueney P, Provenzano S, Verriès C, Ferrandino A, Meudec E, Batelli G, Merdinoglu D, Cheynier V, Schubert A, Ageorges A. (2009) A novel cation-dependent O-methyltransferase involved in anthocyanin methylation in grapevine. Plant Physiol 150: 2057–2070 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huits HSM, Gerats AGM, Kreike MM, Mol JNM, Koes RE. (1994) Genetic control of dihydroflavonol 4-reductase gene expression in Petunia hybrida. Plant J 6: 295–310 [DOI] [PubMed] [Google Scholar]
- Ibdah M, Zhang XH, Schmidt J, Vogt T. (2003) A novel Mg2+-dependent O-methyltransferase in the phenylpropanoid metabolism of Mesembryanthemum crystallinum. J Biol Chem 278: 43961–43972 [DOI] [PubMed] [Google Scholar]
- Jonsson LMV, Aarsman MEG, de Vlaming P, Schram AW. (1984a) On the origin of anthocyanin methyltransferase isozymes of Petunia hybrida and their role in regulation of anthocyanin methylation. Theor Appl Genet 68: 459–466 [DOI] [PubMed] [Google Scholar]
- Jonsson LMV, Aarsman MEG, Poulton JE, Schram AW. (1984b) Properties and genetic control of four methyltransferases involved in methylation of anthocyanins in flowers of Petunia hybrida. Planta 160: 174–179 [DOI] [PubMed] [Google Scholar]
- Jonsson LMV, Aarsman MEG, Schram AW, Bennink GJH. (1982) Methylation of anthocyanins by cell-free extracts of flower buds of Petunia hybrida. Phytochemistry 21: 2457–2459 [Google Scholar]
- Jonsson LMV, Aarsman MEG, van Diepen J, de Vlaming P, Smit N, Schram AW. (1984c) Properties and genetic control of anthocyanin 5-O-glucosyltransferase in flowers of Petunia hybrida. Planta 160: 341–347 [DOI] [PubMed] [Google Scholar]
- Jonsson LMV, de Vlaming P, Wiering H, Aarsman MEG, Schram AW. (1983) Genetic control of anthocyanin-O-methyltransferase activity in flowers of Petunia hybrida. Theor Appl Genet 66: 349–355 [DOI] [PubMed] [Google Scholar]
- Kamsteeg J, van Brederode J, van Nigtevecht G. (1979) Genetics of anthocyanin formation in petals of the red campion (Silene dioica (L) ClairV.). Genetica 51: 5–13 [Google Scholar]
- Karimi M, Inzé D, Depicker A. (2002) GATEWAY vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 7: 193–195 [DOI] [PubMed] [Google Scholar]
- Kim BG, Lee HJ, Park Y, Lim Y, Ahn JH. (2006) Characterization of an O-methyltransferase from soybean. Plant Physiol Biochem 44: 236–241 [DOI] [PubMed] [Google Scholar]
- Koes R, Verweij W, Quattrocchio F. (2005) Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci 10: 236–242 [DOI] [PubMed] [Google Scholar]
- Kroon J, Souer E, de Graaff A, Xue Y, Mol J, Koes R. (1994) Cloning and structural analysis of the anthocyanin pigmentation locus Rt of Petunia hybrida: characterization of insertion sequences in two mutant alleles. Plant J 5: 69–80 [DOI] [PubMed] [Google Scholar]
- Lam KC, Ibrahim RK, Behdad B, Dayanandan S. (2007) Structure, function, and evolution of plant O-methyltransferases. Genome 50: 1001–1013 [DOI] [PubMed] [Google Scholar]
- Lücker J, Martens S, Lund ST. (2010) Characterization of a Vitis vinifera cv. Cabernet Sauvignon 3′,5′-O-methyltransferase showing strong preference for anthocyanins and glycosylated flavonols. Phytochemistry 71: 1474–1484 [DOI] [PubMed] [Google Scholar]
- Pichersky E, Gang DR. (2000) Genetics and biochemistry of secondary metabolites in plants: an evolutionary perspective. Trends Plant Sci 5: 439–445 [DOI] [PubMed] [Google Scholar]
- Provenzano S (2011) The genetics of anthocyanin production, accumulation and display: a comparative study in different species. PhD thesis. VU University, Amsterdam, The Netherlands [Google Scholar]
- Quattrocchio F, Wing J, van der Woude K, Souer E, de Vetten N, Mol J, Koes R. (1999) Molecular analysis of the anthocyanin2 gene of petunia and its role in the evolution of flower color. Plant Cell 11: 1433–1444 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quattrocchio F, Wing JF, Leppen H, Mol J, Koes RE. (1993) Regulatory genes controlling anthocyanin pigmentation are functionally conserved among plant species and have distinct sets of target genes. Plant Cell 5: 1497–1512 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saito K, Yonekura-Sakakibara K, Nakabayashi R, Higashi Y, Yamazaki M, Tohge T, Fernie AR. (2013) The flavonoid biosynthetic pathway in Arabidopsis: structural and genetic diversity. Plant Physiol Biochem 72: 21–34 [DOI] [PubMed] [Google Scholar]
- Slimestad R, Aaberg A, Andersen OM. (1999) Acylated anthocyanins from petunia flowers. Phytochemistry 50: 1081–1086 [Google Scholar]
- Souer E, Rebocho AB, Bliek M, Kusters E, de Bruin RA, Koes R. (2008) Patterning of inflorescences and flowers by the F-box protein DOUBLE TOP and the LEAFY homolog ABERRANT LEAF AND FLOWER of petunia. Plant Cell 20: 2033–2048 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spelt C, Quattrocchio F, Mol JN, Koes R. (2000) anthocyanin1 of petunia encodes a basic helix-loop-helix protein that directly activates transcription of structural anthocyanin genes. Plant Cell 12: 1619–1632 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Styles ED, Ceska O. (1972) Flavonoid pigments in genetic strains of maize. Phytochemistry 11: 3019–3021 [Google Scholar]
- van Houwelingen A, Souer E, Spelt K, Kloos D, Mol J, Koes R. (1998) Analysis of flower pigmentation mutants generated by random transposon mutagenesis in Petunia hybrida. Plant J 13: 39–50 [DOI] [PubMed] [Google Scholar]
- van Tunen AJ, Koes RE, Spelt CE, van der Krol AR, Stuitje AR, Mol JNM. (1988) Cloning of the two chalcone flavanone isomerase genes from Petunia hybrida: coordinate, light-regulated and differential expression of flavonoid genes. EMBO J 7: 1257–1263 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verweij W, Spelt C, Di Sansebastiano GP, Vermeer J, Reale L, Ferranti F, Koes R, Quattrocchio F. (2008) An H+ P-ATPase on the tonoplast determines vacuolar pH and flower colour. Nat Cell Biol 10: 1456–1462 [DOI] [PubMed] [Google Scholar]
- Wang J, Pichersky E. (1999) Identification of specific residues involved in substrate discrimination in two plant O-methyltransferases. Arch Biochem Biophys 368: 172–180 [DOI] [PubMed] [Google Scholar]
- Wiering H. (1974) Genetics of flower colour in Petunia hybrida Hort. Genen Phaenen 17: 117–134 [Google Scholar]
- Wiering H, De Vlaming P. (1973) Glycosylation and methylation patterns of anthocyanins in Petunia hybrida. I. The gene Gf. Genen Phaenen 16: 35–50 [Google Scholar]
- Wiering H, De Vlaming P. (1977) Glycosylation and methylation patterns of anthocyanins in Petunia hybrida. II. The genes Mf1 and Mf2. Z. Pfanzenzüchtung 78: 113–123 [Google Scholar]
- Wils CR, Brandt W, Manke K, Vogt T. (2013) A single amino acid determines position specificity of an Arabidopsis thaliana CCoAOMT-like O-methyltransferase. FEBS Lett 587: 683–689 [DOI] [PubMed] [Google Scholar]
- Winkel-Shirley B. (2001) Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126: 485–493 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamazaki M, Yamagishi E, Gong Z, Fukuchi-Mizutani M, Fukui Y, Tanaka Y, Kusumi T, Yamaguchi M, Saito K. (2002) Two flavonoid glucosyltransferases from Petunia hybrida: molecular cloning, biochemical properties and developmentally regulated expression. Plant Mol Biol 48: 401–411 [DOI] [PubMed] [Google Scholar]
- Yoshimura Y, Zaima N, Moriyama T, Kawamura Y. (2012) Different localization patterns of anthocyanin species in the pericarp of black rice revealed by imaging mass spectrometry. PLoS ONE 7: e31285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zubieta C, He XZ, Dixon RA, Noel JP. (2001) Structures of two natural product methyltransferases reveal the basis for substrate specificity in plant O-methyltransferases. Nat Struct Biol 8: 271–279 [DOI] [PubMed] [Google Scholar]
- Zubieta C, Ross JR, Koscheski P, Yang Y, Pichersky E, Noel JP. (2003) Structural basis for substrate recognition in the salicylic acid carboxyl methyltransferase family. Plant Cell 15: 1704–1716 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.