Abstract
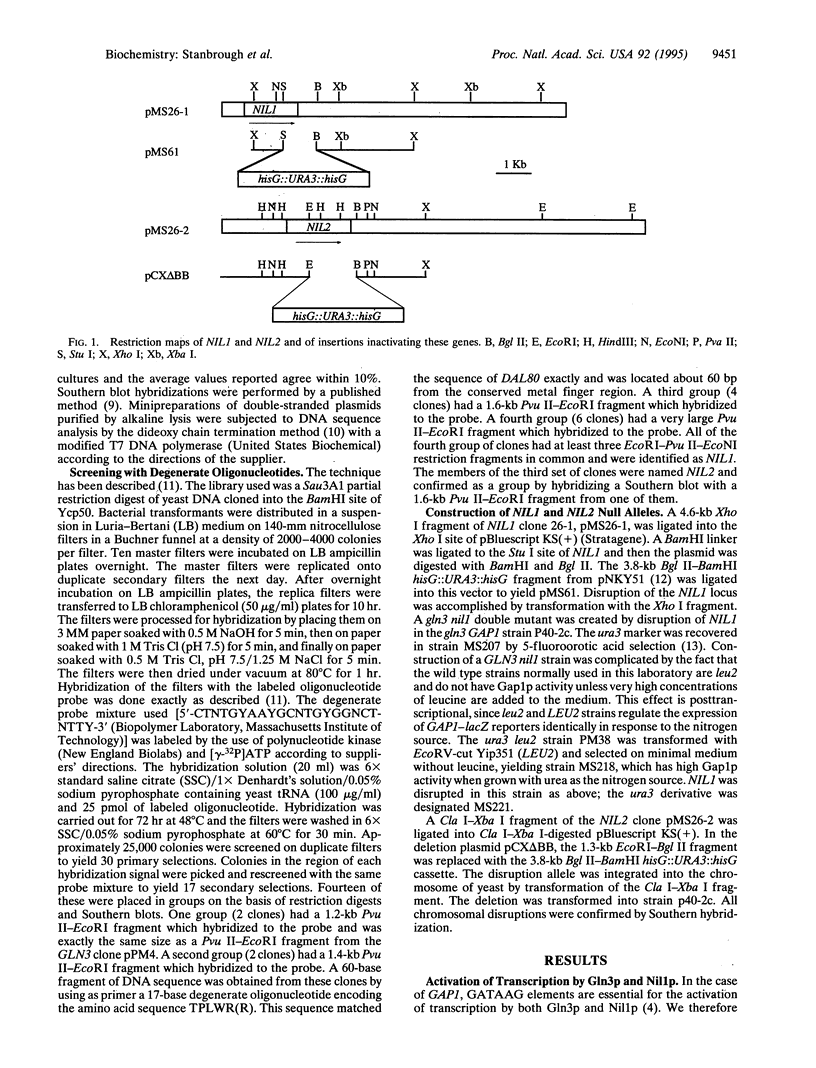
We have isolated the NIL1 gene, whose product is an activator of the transcription of nitrogen-regulated genes, by virtue of the homology of its zinc-finger domain to that of the previously identified activator, the product of GLN3. Disruption of the chromosomal NIL1 gene enabled us to compare the effects of Gln3p and of Nil1p on the expression of the nitrogen-regulated genes GLN1, GDH2, and GAP1, coding respectively for glutamine synthetase, NAD-linked glutamate dehydrogenase, and general amino acid permease. Our results show that the nature of GATAAG sequence that serve as the upstream activation sequence elements for these genes determines their abilities to respond to Gln3p and Nil1p. The results further indicate that Gln3p is inactivated by an increase in the intracellular concentration of glutamine and that Nil1p is inactivated by an increase in intracellular glutamate.
Full text
PDF



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alani E., Cao L., Kleckner N. A method for gene disruption that allows repeated use of URA3 selection in the construction of multiply disrupted yeast strains. Genetics. 1987 Aug;116(4):541–545. doi: 10.1534/genetics.112.541.test. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blinder D., Magasanik B. Recognition of nitrogen-responsive upstream activation sequences of Saccharomyces cerevisiae by the product of the GLN3 gene. J Bacteriol. 1995 Jul;177(14):4190–4193. doi: 10.1128/jb.177.14.4190-4193.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boeke J. D., Trueheart J., Natsoulis G., Fink G. R. 5-Fluoroorotic acid as a selective agent in yeast molecular genetics. Methods Enzymol. 1987;154:164–175. doi: 10.1016/0076-6879(87)54076-9. [DOI] [PubMed] [Google Scholar]
- Coschigano P. W., Magasanik B. The URE2 gene product of Saccharomyces cerevisiae plays an important role in the cellular response to the nitrogen source and has homology to glutathione s-transferases. Mol Cell Biol. 1991 Feb;11(2):822–832. doi: 10.1128/mcb.11.2.822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Courchesne W. E., Magasanik B. Ammonia regulation of amino acid permeases in Saccharomyces cerevisiae. Mol Cell Biol. 1983 Apr;3(4):672–683. doi: 10.1128/mcb.3.4.672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunningham T. S., Cooper T. G. Expression of the DAL80 gene, whose product is homologous to the GATA factors and is a negative regulator of multiple nitrogen catabolic genes in Saccharomyces cerevisiae, is sensitive to nitrogen catabolite repression. Mol Cell Biol. 1991 Dec;11(12):6205–6215. doi: 10.1128/mcb.11.12.6205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller S. M., Magasanik B. Role of NAD-linked glutamate dehydrogenase in nitrogen metabolism in Saccharomyces cerevisiae. J Bacteriol. 1990 Sep;172(9):4927–4935. doi: 10.1128/jb.172.9.4927-4935.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller S. M., Magasanik B. Role of the complex upstream region of the GDH2 gene in nitrogen regulation of the NAD-linked glutamate dehydrogenase in Saccharomyces cerevisiae. Mol Cell Biol. 1991 Dec;11(12):6229–6247. doi: 10.1128/mcb.11.12.6229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minehart P. L., Magasanik B. Sequence and expression of GLN3, a positive nitrogen regulatory gene of Saccharomyces cerevisiae encoding a protein with a putative zinc finger DNA-binding domain. Mol Cell Biol. 1991 Dec;11(12):6216–6228. doi: 10.1128/mcb.11.12.6216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minehart P. L., Magasanik B. Sequence of the GLN1 gene of Saccharomyces cerevisiae: role of the upstream region in regulation of glutamine synthetase expression. J Bacteriol. 1992 Mar;174(6):1828–1836. doi: 10.1128/jb.174.6.1828-1836.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitchell A. P., Magasanik B. Regulation of glutamine-repressible gene products by the GLN3 function in Saccharomyces cerevisiae. Mol Cell Biol. 1984 Dec;4(12):2758–2766. doi: 10.1128/mcb.4.12.2758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murakami Y., Naitou M., Hagiwara H., Shibata T., Ozawa M., Sasanuma S., Sasanuma M., Tsuchiya Y., Soeda E., Yokoyama K. Analysis of the nucleotide sequence of chromosome VI from Saccharomyces cerevisiae. Nat Genet. 1995 Jul;10(3):261–268. doi: 10.1038/ng0795-261. [DOI] [PubMed] [Google Scholar]
- Ptashne M. How eukaryotic transcriptional activators work. Nature. 1988 Oct 20;335(6192):683–689. doi: 10.1038/335683a0. [DOI] [PubMed] [Google Scholar]
- Rasmussen S. W. A 37.5 kb region of yeast chromosome X includes the SME1, MEF2, GSH1 and CSD3 genes, a TCP-1-related gene, an open reading frame similar to the DAL80 gene, and a tRNA(Arg). Yeast. 1995 Jul;11(9):873–883. doi: 10.1002/yea.320110909. [DOI] [PubMed] [Google Scholar]
- Roeder G. S., Fink G. R. DNA rearrangements associated with a transposable element in yeast. Cell. 1980 Aug;21(1):239–249. doi: 10.1016/0092-8674(80)90131-2. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stanbrough M., Magasanik B. Transcriptional and posttranslational regulation of the general amino acid permease of Saccharomyces cerevisiae. J Bacteriol. 1995 Jan;177(1):94–102. doi: 10.1128/jb.177.1.94-102.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]



