Abstract
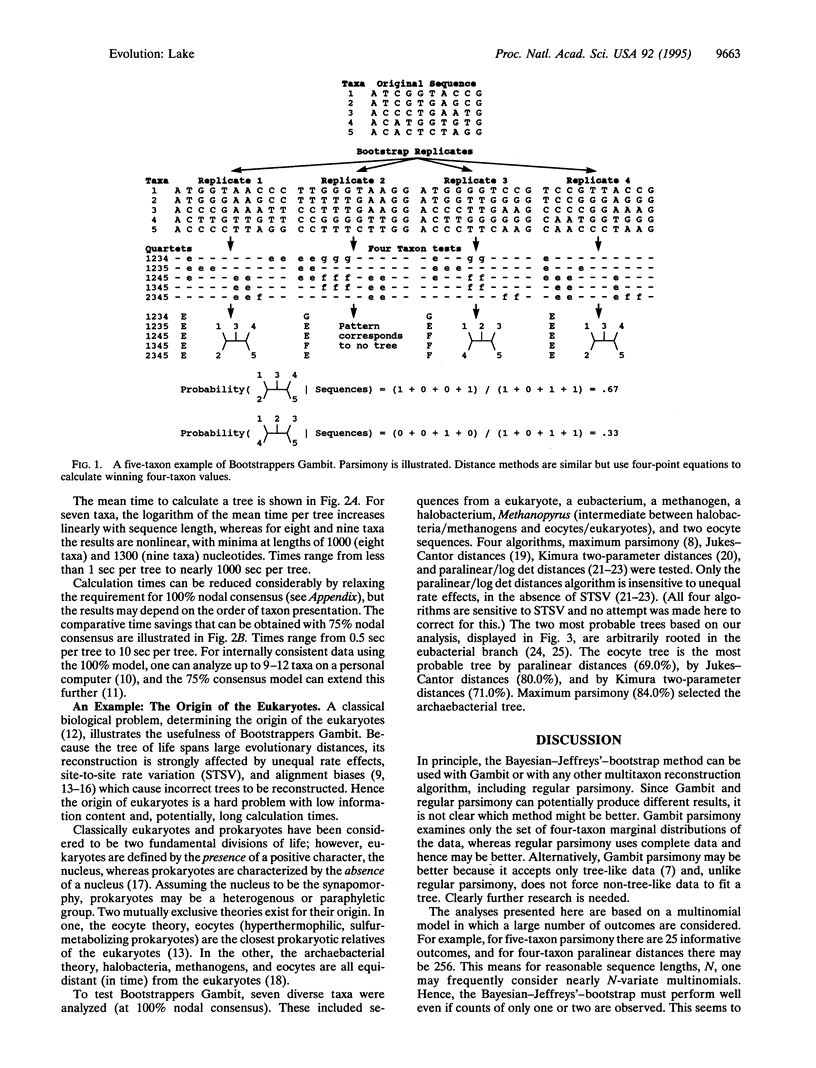
The reconstruction of multitaxon trees from molecular sequences is confounded by the variety of algorithms and criteria used to evaluate trees, making it difficult to compare the results of different analyses. A global method of multitaxon phylogenetic reconstruction described here, Bootstrappers Gambit, can be used with any four-taxon algorithm, including distance, maximum likelihood, and parsimony methods. It incorporates a Bayesian-Jeffreys'-bootstrap analysis to provide a uniform probability-based criterion for comparing the results from diverse algorithms. To examine the usefulness of the method, the origin of the eukaryotes has been investigated by the analysis of ribosomal small subunit RNA sequences. Three common algorithms (paralinear distances, Jukes-Cantor distances, and Kimura distances) support the eocyte topology, whereas one (maximum parsimony) supports the archaebacterial topology, suggesting that the eocyte prokaryotes are the closest prokaryotic relatives of the eukaryotes.
Full text
PDF



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Cavalli-Sforza L. L., Edwards A. W. Phylogenetic analysis. Models and estimation procedures. Am J Hum Genet. 1967 May;19(3 Pt 1):233–257. [PMC free article] [PubMed] [Google Scholar]
- Churchill G. A., von Haeseler A., Navidi W. C. Sample size for a phylogenetic inference. Mol Biol Evol. 1992 Jul;9(4):753–769. doi: 10.1093/oxfordjournals.molbev.a040757. [DOI] [PubMed] [Google Scholar]
- Cousineau B., Cerpa C., Lefebvre J., Cedergren R. The sequence of the gene encoding elongation factor Tu from Chlamydia trachomatis compared with those of other organisms. Gene. 1992 Oct 12;120(1):33–41. doi: 10.1016/0378-1119(92)90006-b. [DOI] [PubMed] [Google Scholar]
- Creti R., Citarella F., Tiboni O., Sanangelantoni A., Palm P., Cammarano P. Nucleotide sequence of a DNA region comprising the gene for elongation factor 1 alpha (EF-1 alpha) from the ultrathermophilic archaeote Pyrococcus woesei: phylogenetic implications. J Mol Evol. 1991 Oct;33(4):332–342. doi: 10.1007/BF02102864. [DOI] [PubMed] [Google Scholar]
- Felsenstein J. Estimating effective population size from samples of sequences: a bootstrap Monte Carlo integration method. Genet Res. 1992 Dec;60(3):209–220. doi: 10.1017/s0016672300030962. [DOI] [PubMed] [Google Scholar]
- Gogarten J. P., Kibak H., Dittrich P., Taiz L., Bowman E. J., Bowman B. J., Manolson M. F., Poole R. J., Date T., Oshima T. Evolution of the vacuolar H+-ATPase: implications for the origin of eukaryotes. Proc Natl Acad Sci U S A. 1989 Sep;86(17):6661–6665. doi: 10.1073/pnas.86.17.6661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halanych K. M., Bacheller J. D., Aguinaldo A. M., Liva S. M., Hillis D. M., Lake J. A. Evidence from 18S ribosomal DNA that the lophophorates are protostome animals. Science. 1995 Mar 17;267(5204):1641–1643. doi: 10.1126/science.7886451. [DOI] [PubMed] [Google Scholar]
- Iwabe N., Kuma K., Hasegawa M., Osawa S., Miyata T. Evolutionary relationship of archaebacteria, eubacteria, and eukaryotes inferred from phylogenetic trees of duplicated genes. Proc Natl Acad Sci U S A. 1989 Dec;86(23):9355–9359. doi: 10.1073/pnas.86.23.9355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lake J. A. Origin of the eukaryotic nucleus determined by rate-invariant analysis of rRNA sequences. Nature. 1988 Jan 14;331(6152):184–186. doi: 10.1038/331184a0. [DOI] [PubMed] [Google Scholar]
- Lake J. A. Reconstructing evolutionary trees from DNA and protein sequences: paralinear distances. Proc Natl Acad Sci U S A. 1994 Feb 15;91(4):1455–1459. doi: 10.1073/pnas.91.4.1455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lake J. A. The order of sequence alignment can bias the selection of tree topology. Mol Biol Evol. 1991 May;8(3):378–385. doi: 10.1093/oxfordjournals.molbev.a040654. [DOI] [PubMed] [Google Scholar]
- Maslov D. A., Avila H. A., Lake J. A., Simpson L. Evolution of RNA editing in kinetoplastid protozoa. Nature. 1994 Mar 24;368(6469):345–348. doi: 10.1038/368345a0. [DOI] [PubMed] [Google Scholar]
- Miller G. Animal model for Epstein-Barr lymphoma. Nature. 1986 Feb 20;319(6055):626–626. doi: 10.1038/319626c0. [DOI] [PubMed] [Google Scholar]
- Rivera M. C., Lake J. A. Evidence that eukaryotes and eocyte prokaryotes are immediate relatives. Science. 1992 Jul 3;257(5066):74–76. doi: 10.1126/science.1621096. [DOI] [PubMed] [Google Scholar]
- Steel M. A., Lockhart P. J., Penny D. Confidence in evolutionary trees from biological sequence data. Nature. 1993 Jul 29;364(6436):440–442. doi: 10.1038/364440a0. [DOI] [PubMed] [Google Scholar]
- Woese C. R. Bacterial evolution. Microbiol Rev. 1987 Jun;51(2):221–271. doi: 10.1128/mr.51.2.221-271.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]



