Abstract
The correspondence between the transversion/transition ratio and the neighboring base composition in chloroplast DNA is examined. For 18 noncoding regions of the chloroplast genome, alignments between rice (Oryza sativa) and maize (Zea mays) were generated by two different methods. Difficulties of aligning noncoding DNA are discussed, and the alignments are analyzed in a manner that reduces alignment artifacts. Sequence divergence is < 10%, so multiple substitutions at a site are assumed to be rare. Observed substitutions were analyzed with respect to the A+T content of the two immediately flanking bases. It is shown that as this content increases, the proportion of transversions also increases. When both the 5'- and 3'-flanking nucleotides are G or C (A+T content of 0), only 25% of the observed substitutions are transversions. However, when both the 5'- and 3'-flanking nucleotides are A or T (A+T content of 2), 57% of the observed substitutions are transversions. Therefore, the influence of flanking base composition on substitutions, previously reported for a single noncoding region, is a general feature of the chloroplast genome.
Full text
PDF

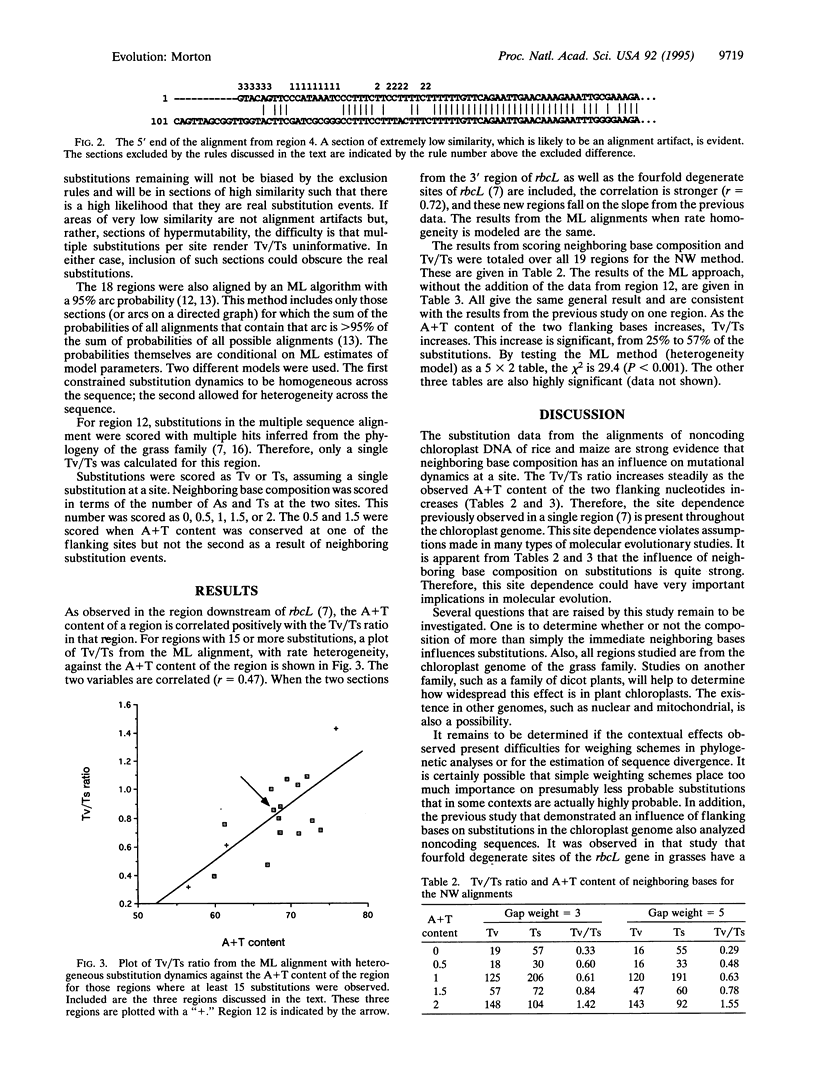


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bowman C. M., Barker R. F., Dyer T. A. In wheat ctDNA, segments of ribosomal protein genes are dispersed repeats, probably conserved by nonreciprocal recombination. Curr Genet. 1988 Aug;14(2):127–136. doi: 10.1007/BF00569336. [DOI] [PubMed] [Google Scholar]
- Brown W. M., Prager E. M., Wang A., Wilson A. C. Mitochondrial DNA sequences of primates: tempo and mode of evolution. J Mol Evol. 1982;18(4):225–239. doi: 10.1007/BF01734101. [DOI] [PubMed] [Google Scholar]
- Clegg M. T. Chloroplast gene sequences and the study of plant evolution. Proc Natl Acad Sci U S A. 1993 Jan 15;90(2):363–367. doi: 10.1073/pnas.90.2.363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gojobori T., Li W. H., Graur D. Patterns of nucleotide substitution in pseudogenes and functional genes. J Mol Evol. 1982;18(5):360–369. doi: 10.1007/BF01733904. [DOI] [PubMed] [Google Scholar]
- Golenberg E. M., Clegg M. T., Durbin M. L., Doebley J., Ma D. P. Evolution of a noncoding region of the chloroplast genome. Mol Phylogenet Evol. 1993 Mar;2(1):52–64. doi: 10.1006/mpev.1993.1006. [DOI] [PubMed] [Google Scholar]
- Hiratsuka J., Shimada H., Whittier R., Ishibashi T., Sakamoto M., Mori M., Kondo C., Honji Y., Sun C. R., Meng B. Y. The complete sequence of the rice (Oryza sativa) chloroplast genome: intermolecular recombination between distinct tRNA genes accounts for a major plastid DNA inversion during the evolution of the cereals. Mol Gen Genet. 1989 Jun;217(2-3):185–194. doi: 10.1007/BF02464880. [DOI] [PubMed] [Google Scholar]
- Jones M., Wagner R., Radman M. Repair of a mismatch is influenced by the base composition of the surrounding nucleotide sequence. Genetics. 1987 Apr;115(4):605–610. doi: 10.1093/genetics/115.4.605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li W. H., Wu C. I., Luo C. C. Nonrandomness of point mutation as reflected in nucleotide substitutions in pseudogenes and its evolutionary implications. J Mol Evol. 1984;21(1):58–71. doi: 10.1007/BF02100628. [DOI] [PubMed] [Google Scholar]
- Morton B. R., Clegg M. T. A chloroplast DNA mutational hotspot and gene conversion in a noncoding region near rbcL in the grass family (Poaceae). Curr Genet. 1993 Oct;24(4):357–365. doi: 10.1007/BF00336789. [DOI] [PubMed] [Google Scholar]
- Needleman S. B., Wunsch C. D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol. 1970 Mar;48(3):443–453. doi: 10.1016/0022-2836(70)90057-4. [DOI] [PubMed] [Google Scholar]
- Radman M., Wagner R. Mismatch repair in Escherichia coli. Annu Rev Genet. 1986;20:523–538. doi: 10.1146/annurev.ge.20.120186.002515. [DOI] [PubMed] [Google Scholar]
- Thorne J. L., Churchill G. A. Estimation and reliability of molecular sequence alignments. Biometrics. 1995 Mar;51(1):100–113. [PubMed] [Google Scholar]
- Thorne J. L., Kishino H., Felsenstein J. An evolutionary model for maximum likelihood alignment of DNA sequences. J Mol Evol. 1991 Aug;33(2):114–124. doi: 10.1007/BF02193625. [DOI] [PubMed] [Google Scholar]




