Abstract
Acute promyelocytic leukemia (APL) is characterized by the t(15;17)(q22;q21), which results in the fusion of the promyelocytic leukemia (PML) gene at 15q22 with the retinoic acid α-receptor (RARA) gene at 17q21. The current study presents the case of a 54-year-old female with APL carrying the atypical PML/RARA fusion signal due to a novel complex variant translocation t(15;16;17)(q22;q24;q21), as well as the classical PML/RARA fusion signal. Subsequent array comparative genomic hybridization revealed somatic, cryptic deletions on 3p25.3, 8q23.1 and 12p13.2-p13.1, and a duplication on 8q11.2; however, no genetic material loss or gain was observed in the breakpoint regions of chromosomes 15, 16 or 17. To the best of our knowledge, this is the first report of the coexistence of two abnormal clones, one classical and one variant, presenting simultaneously in addition to cryptic chromosome segmental imbalances in an adult APL patient.
Keywords: promyelocytic leukemia/retinoic acid α-receptor, array comparative genomic hybridization, variant translocation, acute promyelocytic leukemia, fluorescence in situ hybridization
Introduction
Acute promyelocytic leukemia (APL) is characterized by particular clinical features that are important in differentiating it from other acute myeloid leukemias (AMLs) and determining an accurate diagnosis. These clinical features include unique hemorrhagic syndrome, disseminated intravascular coagulation (DIC) and association with the translocation between chromosomes 15 and 17, resulting in the formation of two reciprocal fusion genes; promyelocytic leukemia (PML)/retinoic acid α-receptor (RARA) on chromosome 15 and RARA/PML on chromosome 17. These fusion genes are sensitive to retinoid differentiating agents, such as all-trans retinoic acid (ATRA), and novel antiapoptotic agents, including arsenic trioxide (1,2).
Although t(15;17) has been found in ~90% of APL patients, variant translocations have been reported in a few APL patients, which are described as simple translocations involving chromosome 15 or 17 with any other chromosomes [t(15;v) or t(17;v)] or complex translocations characterized by the involvement of additional chromosome(s) in addition to chromosomes 15 and 17 [ct(15;17;v)] (3–5). In the last few years, a number of studies have focused on the alternate translocation in APL, for example t(5;17)(q35;q21) forming NPM/RARA, t(11;17)(q23;q21) producing PLF/RARA fusion and t(11;17)(q13;q21) generating NUMA/RARA, providing advanced insights into the pathogenesis of APL (3,6,7). However, little is known concerning the complex variant translocations in APL. The current study reports a patient who presented with the classical t(15;17) and complex variant t(15;16;17)(q22;q24;q21), which were demonstrated by traditional cytogenetic analysis, including G-banding karyotype and fluorescence in situ hybridization (FISH). In addition, cryptic losses on 3p25.3, 8q23.1 and 12p13.2-p13.1, and a gain of chromosome 8q11.2, were revealed in the level of array comparative genomic hybridization (CGH). This study was approved by the Institutional Review Board (IRB) at the University of Oklahoma Health Sciences Center (IRB no. 13100; Oklahoma City, OK, USA). The patient provided consent.
Case report
Clinical presentation
The current study presents a 54-year-old female with APL who was admitted to the University of Oklahoma Health Sciences Center due to fever and bleeding gums. The peripheral blood examination showed a hemoglobin count of 7.1 g/dl (normal range, 12–15.5 g/dl) and platelet count of 35×103/μl (normal range, 150–450×103/μl), as well as a white blood cell count of 11.95×103/μl (normal range, 3.5–10.5×103/μl) with 86% blasts, characterized by small to large cells with irregular, lobulated and bilobed nuclei with prominent nucleoli. The blast cells in the peripheral blood exhibited scant to moderate rare blue granulated cytoplasm and the bone marrow aspirate revealed no cellular particles. The touch preparations showed suboptimal cellular morphology, but revealed numerous blasts that were more frequently granulated than those present in the peripheral smear. The core biopsy of the bone marrow revealed a cellularity of >95% consisting of sheets of immature cells. A diagnosis of APL, M3 variant was determined according to the French-American-British Cooperative Group criteria (8).
Cytogenetics, FISH and array CGH analyses
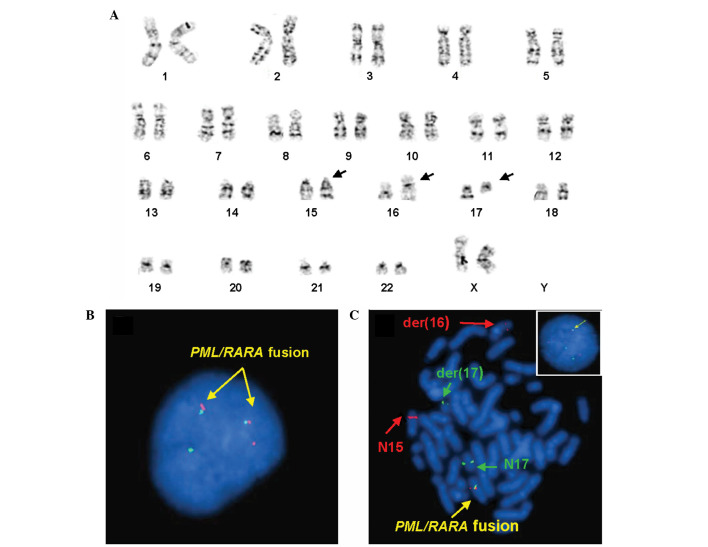
The karyotype analysis at diagnosis revealed that the 20 metaphases analyzed exhibited a variant translocation among three chromosome; chromosomes 15, 16 and 17. The karyotype was designated as 46,XX,t(15;16;17)(q24;q24;q21)[20] (Fig. 1A). FISH analyses using the LSI PML/RARA dual color and dual fusion probe (Abbott Molecular, Inc., Des Plaines, IL, USA) was applied to the uncultured and cultured cells. On uncultured interphase cells, 12 out of 200 cells (6%) exhibited two classical PML/RARA fusion signals, one SpectrumOrange and one SpectrumGreen signal, which were the result of the classical t(15;17) (Fig. 1B) (http://www.abbottmolecular.com/us/siteMap.html#sthash.483IvlrV.dpuf). While 168 out of 200 cells (84%) exhibited one fusion signal, two SpectrumOrange signals and two SpectrumGreen signals, which were the result of the complex variant translocation, t(15;16;17) (Fig. 1C) (9). By combining the karyotyping and FISH results, it was demonstrated on metaphase cells that a part of the PML gene labeled with SpectrumOrange was present on the derivative chromosome 16 and a part of the RARA gene labeled with SpectrumGreen signal was present on the derivative chromosome 17 (Fig. 1C). Subsequent cohybridization of whole chromosome painting probes (WCP) 15 and 16 (Cytocell, Ltd., Cambridge, UK) and cohybridization of WCP17 (Abbott Molecular, Inc.), as well as the subtelomeric probes for chromosome 16p and q (Abbott Molecular, Inc.) on metaphase cells, confirmed the complex translocations among chromosomes 15, 16 and 17 (Fig. 2A and B) (http://www.abbottmolecular.com/us/siteMap.html#sthash.483IvlrV.dpuf) (10).
Figure 1.
(A) Karyotype analysis revealed 46,XX,t(15;16;17)(q24;q24;q21) and the arrows indicate the t(15;16;17)(q24;q24;q21). Fluorescence in situ hybridization analyses using LSI PML (15q22; Spectrum Orange)/RARA (17q21; SpectrumGreen) dual color/dual fusion probe revealed a (B) typical PML/RARA dual color dual fusion signal and (C) variant t(15;17)(q22;q21) in the metaphase and interphase cells. PML/RARA, promyelocytic leukemia/retinoic acid α-receptor; N, normal; der, derivative.
Figure 2.
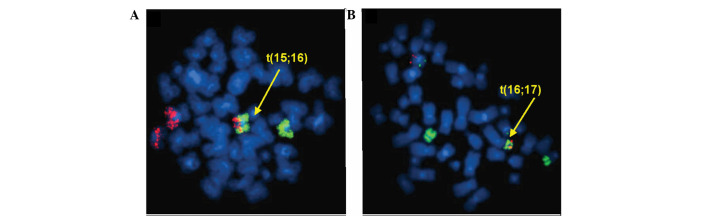
Fluorescence in situ hybridization analyses using cohybridization of (A) WCP15 (SpectrumOrange) with WCP16 (SpectrumGreen) showed t(15;16) and (B) WCP17 (SpectrumGreen) with subtelomeric probe for chromosome 16p (SpectrumGreen) and q (SpectrumOrange) showed t(16;17). WCP, whole chromosome painting probes.
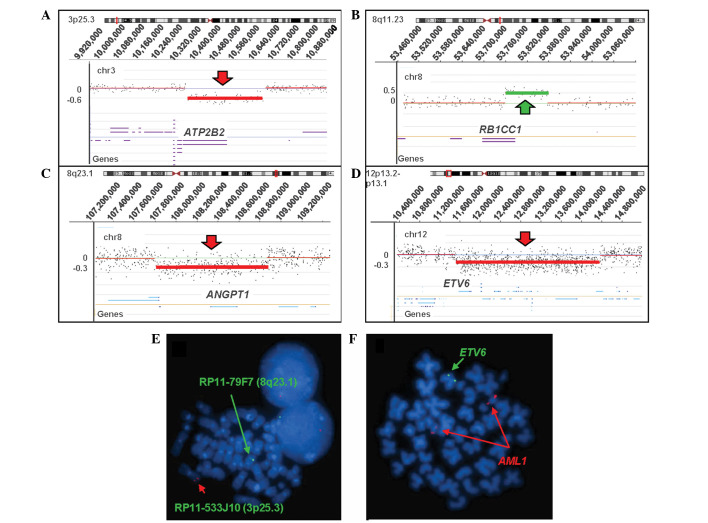
Array CGH analysis showed the acquired genomic aberrations. No loss or gain was identified on chromosomes 15, 16, or 17, but cryptic losses of genomic material on 3p25.3, 8q23.1 and 12p13.1-p13.2, and a cryptic gain of 8q11.23 were detected (Fig. 3A–D). Several cancer-related genes were located in these genomic imbalanced regions. A loss of 3p25.3 (10,364,050–10,670,236 bp hg18; ~0.3 Mb) encompasses the plasma membrane Ca2+-ATPase 2 (ATP2B2) gene, a loss of 8q23.1 (107,811,331–108,868,110 bp hg18; ~1.1 Mb) contains the angiopoietin 1 (ANGPT1) gene, a loss of 12p13.2-p13.1 (11,619,439–14,472,130 bp hg18; ~2.9 Mb) involves ~23 genes, including the ETS variant 6 (ETV6) gene which is highly associated with t(12;21) in acute lymphoblastic leukemia, and a gain of 8q11.23 (53,762,025–53,876,752 bp hg18; ~0.1 Mb) contains the tumor suppressor gene retinoblastoma 1 inducible coiled-coil 1 (RB1CC1). Subsequent FISH confirmed the array CGH observations. The cultured cells were hybridized with home-brew probes using bacterial artificial chromosome clones, RP11-533J10 labeled with SpectrumOrange on 3p25.3 and RP11-79F7 labeled with SpectrumGreen on 8q23.1 (Invitrogen Life Technologies, Carlsbad, CA, USA), and 194 out of 200 cells (97%) presented with one SpectrumOrange signal and 104 out of 200 cells (52%) showed one SpectrumGreen signal, demonstrating the deletions of 3p25.3 and 8q23.1, respectively (Fig. 3E). The percentage of the cells with deletions was close to the percentage of cells with t(15;17) combined with t(15;16;17). These deletions are likely to be somatically acquired and not constitutional changes. In addition, hybridization with the LSI ETV6/AML1 dual color ES probe (Abbott Molecular, Inc.) confirmed the deletion of 12p13.1-p13.2 involving the ETV6 genes (Fig. 3F) and 116 out of 200 (58%) cells were observed with deletion of the ETV6 gene. However, gain of 8q11.23 was not be confirmed by FISH due to the small size (only 100 bp) of the duplication.
Figure 3.
Results of the oligoarray CGH using NimbleGen SegMNT and the RefSeq genes in the abnormal region (University of California, Santa Cruz genome browser hg18). The Y-axis indicates a gain or loss of genetic material, while the X-axis indicates the genomic position of each feature on the chromosome. The red arrow indicates loss and the green arrow indicates gain and the significant gene in the lost or gained region is listed. (A) Loss of 3p25.3 (10,364,050–10,670,236 bp, hg18; ~0.3 Mb). (B) Gain of 8q11.23 (53,762,025–53,876,752bp, hg18; ~0.1 Mb), which is potentially constitutional rather than a disease-related duplication. (C) Loss of 8q23.1 (107,811,331–108,868,110 bp, hg 18; ~1.1 Mb). (D) Loss of 12p13.2-p13.1 (11,619,439–14,472,130 bp, hg18; ~2.9 Mb). Fluorescence in situ hybridization analyses using bacterial artificial chromosome RP11-533J10 (3p25.3; SpectrumOrange), RP11-79F7 (8p23.1; SpectrumGreen) and LSI ETV6 (SpectrumGreen) confirmed the array CGH results of the losses of (E) 3p25.3 and 8q23.1, and (F) ETV6 gene. CGH, comparative genomic hybridization; ETV6, ETS variant 6; AML1, acute myeloid leukemia-1.
Discussion
It is rare that complex variant translocations between chromosomes 15 and 17 [ct(15;17;v)] involving more than three chromosomes in APL result in the PML/RARA fusion gene. To the best of our knowledge, 45 cases have been reported (5,6,9,11–40), which account for 10% of APLs lacking classical t(15;17) (6). The clinical characterizations of the reported cases of APL with ct(15;17;v), including the present study, are summarized in Table I. No distinct clinical features have been observed in APL with ct(15;17;v) compared with APL with classical t(15;17). Generally, the ratio between males and females is 1:1, the median age at diagnosis is 45 years and 77% of ct(15;17;v) cases are accompanied with DIC. This is similar to the DIC ratio of cases with classical t(15;17). All observed ct(15;17;v) cases exhibited good responses to ATRA. Among these ct(15;17;v), nine recurrent breakpoints on the third or fourth chromosome consisted of the ct(15;17;v), including 1p36, 2q21, 3p21, 4q21, 11q13, 18q12, 20p13, 22q11.2 and Xq13. The current study presents the third case of APL with ct(15;17;v) involving chromosome 16. The first was described in a human immunodeficiency virus patient with secondary APL and the second was described in an APL patient with four chromosome translocations involving chromosomes 5, 15, 16 and 17 (23,29). However, the breakpoint of chromosome 16 in the current case (16q24) was different from those identified in the two previously reported cases. The breakpoint 16q24 has been described in AML with t(16;21)(q24;q22), which leads to the fusion gene RUNX1/MTG16 and is predominantly associated with therapy-related AML (41). Whether the MTG16 gene is located at the breakpoint and forms a fusion gene in the present case requires clarification.
Table I.
Clinical characterization of reported acute promyelocytic leukemia cases with variant t(15;17).
| Author (ref.) | Cytogenetic abnormality | Gender | Age, years | FAB | DIC | Cell clones | Treatment | Relapse | Survival, months |
|---|---|---|---|---|---|---|---|---|---|
| Berger et al (11) | t(15;17;4)(q22;q12;q21) | M | NA | M3 | NA | 46,XY, t(15;17;4)(q22;q12;q21) | CNA | NA | |
| Bernstein et al (12) | t(2;15;17)(q21?;q25?6?;q21?2?) | F | 23 | M3v | ± | 46,XX 46,XX,t(2;15;17)( q21?;q25?6?;q21?2?) 9p+ |
C + | 6.5 | |
| Bernstein et al (13) | t(3;15;17)(p21;q25?6?;q21?2?) | M | 28 | M3 | + | 46,XY 46,XY,t(3;15;17)( p21;q25?6?;q21?2?) 4p+ |
CNR | 0.2 | |
| Ohyashiki et al (14) | t(1;5;15;17)(p36;q31;q22;q12) | F | 34 | M3 | + | 46,XX 47,XX,+X, t(1;5;15;17)(p36;q31;q22;q12) |
CNR | 1 | |
| Callen et al (15) | t(X;15;17)(q13;q21;q21) | F | 48 | M3v? | − | 46,XX/46,XX, t(X;15;17)(q13;q21;q21) Relapse: 46,XX, t(X;15;17)(q13;q21;q21) |
C + | 13 | |
| Bjerrum et al (16) | t(2;17;15)(q21;q21;q22or23or24) | M | 10 | M3 | ± | 46,XY, t(2;17;15)(q21;q21;q22or23or24) 46,XY,t(15;17)(q22or23or24;q21) |
C + | 34 | |
| 46,XY, t(2;17;15)(q21;q21;q22or23or24),8p+ | A − | 10+ | |||||||
| Osella et al (17) | t(1;15;17)(p36;q22;q21.1) | M | 46 | M3 | + | 46,XY 46,XY, t(1;15;17)(p36;q22;q21.1) 46,XY, t(1;15;17)(p36;q22;q21.1),+8 |
|||
| Saitoh et al (18) | t(15;19;17)(q22;p13;q12) | F | 16 | M3 | NA | 46,XX 46,XX, t(15;19;17)(q22;p13;q12) |
A − | 7+ | |
| Wan et al (19) | t(X;17;15)(q13;q12;q21) | F | 30 | M3 | + | 46,XX, t(X;17;15)(q13;q12;q21) | C − | 2+ | |
| Fujishima et al (20) | t(2;15;17)(q21;q22;q21) | M | 77 | M3 | + | 46,XY, t(2;15;17)(q21;q22;q21) | C+A | − | 12+ |
| Liu et al (21) | t(4;15;17)(q21;q22;q21) | M | 37 | M3 | + | 46,XY 46,XY, t(4;15;17)(q21;q22;q21) |
As2O3 | − | 10+ |
| Misawa et al (22) | t(7;17;15)(p22;q21.1orq12;q22) | M | 35 | M3 | + | 46,XY, t(7;17;15)(p22;q21.1orq12;q22) | C − | 10+ | |
| Zaccaria et al (23) | t(13;15;17;20)(q22;q22;q12;p13) | F | 74 | M3 | + | 46,XX 47,XX,+8, t(1;5;15;17)(p36;q31;q22;q12) |
ANR | 0.2 | |
| t(5;15;16;17)(q22;q22;p13;q12) | M | 45 | M3 | + | 46,XY 47,XY, t(5;15;16;17)(q22;q22;p13;q12) |
C+A | + | 24 | |
| Ogawa et al (24) | t(15;17;18)(q21q22;q12;q12) | M | 53 | M3 | + | 46,XY,del(15)(q21q22),der(17)t(17;18)(q12;q12), der(18)t(17;18)(q12;q12)ins(18;15)(q12;q21q22) | C − | 3+ | |
| Chen et al (25) | t(11;15;17)(q13;q22;q12) | F | 15 | M3 | NA | 46,XX, t(11;15;17)(q13;q22;q12) | NA | NA | NA |
| t(11;15;17)(q13;q22;q12) | M | 41 | M3 | NA | 46,XY 46,XY, t(11;15;17)(q13;q22;q12) |
NA | NA | NA | |
| t(5;15;17)(q13;q22;q12) | M | 50 | M3 | NA | 46,XY 46,XY, t(5;15;17)(q13;q22;q12) |
NA | NA | NA | |
| Calabrese et al (26) | t(15;17;21)(q22;q12;q22) | M | 38 | M3 | NA | 46,XY, t(15;17;21)(q22;q12;q22) | C+BMT | + | 3 |
| der(4)t(4;17)(q11;p11),der(15)t(15;17)(q22;q21),der(17)t(15;17)(q22;q21)t(4;17)(q11;p11) | F | 22 | M3 | NA | 46,XX,der(4)t(4;17)(q11;p11),der(15)t(15;17)(q22;q21),der(17)t(15;17)(q22;q21)t(4;17)(q11;p11) | C+A | − | 44+ | |
| Park et al (27) | t(1;15;17)(p31;q22;q21) | M | 46 | M3 | + | 46,XY | |||
| 46,XY,del(1)(p22),del(3)(p25),der(17)t(1;15;17)(17pter→17q21::15q21→15q22::1p36→1p31::1 5q21→15q22::17q21→17pter) |
C+A | + | 9 | ||||||
| Yamamoto et al (28) | t(15;20;17)(q22;p13;q21) | M | 39 | M3 | + | 46,XY 46,XY, t(15;20;17)(q22;p13;q21) |
C+A | − | 12+ |
| Kudva et al (29) | t(15;16;17)(q22;q13;q21) | M | 27 | M3v | − | 46,XY 46,XY, t(15;16;17)(q22;q13;q21) |
C+A | − | 40+ |
| Eclache et al (30) | t(6;15;17)(q25;q22;q21) | F | 56 | M3 | NA | 46,XX 46,XX, t(6;15;17)(q25;q22;q21) |
C+A | − | 5+ |
| Xu et al (5) | t(15;17) relapsed with t(4;15;17)(q27-28;q22;q21) | F | 49 | M3 M5a |
NA | 46,XX,dup(8)(q?),t(4;15;17)(q27-28;q22;q21) | C+A | + | 10 |
| García-Casado et al (31) | t(15;17(q22;q21),t(17;20)(q21;q12) | F | 31 | M3 | − | 46,XX | |||
| 46,XX,der(15) t(15;17(q22;q21),t(17;20)(q21;q12) | C+A | − | 60+ | ||||||
| Yoo et al (32) | t(10;17;15;22)(q22;q21;q22;q11.2) | F | 47 | M3 | NA | 46,XX,t(10;17;15;22)(q22;q21;q22;q11.2) | C+A | − | 1+ |
| Miyazaki et al (33) | t(15;17) involved 8q21 and 14p? | M | 78 | M3v | − | 46,XY 46,XY,t(15;17)(q22;q21) 46,XY,der(15)t(15;17)(q22;q21),der(17)(17pter→1 7q21::15q22→15qter::8q21→8qter) 46,XY,der(15)t(15;17)(q22;q21),der(17)(17pter→17q21::15q22→15qter::8q21→8qter::14p?→14qter) |
C+A | − | 14+ |
| Abe et al (34) | t(5;17;15)(q11;q21;q22) | F | 22 | M3v | + | 46,XX 46,XX, t(5;17;15)(q11;q21;q22) |
C+A | − | 12+ |
| Stavropoulou et al (35) | t(15;17) ,t(17;18) | F | 46 | M3 | NA | 46,XX,der(15)t(15;17)(q22;q21),der(17)t(15;17)(q 22;q21)t(17;18)(q21;q12),der(18)t(15;17)(q22;q21)t(17;18)(q21;q12) | C+A | − | 2 |
| Kato et al (36) | t(15;22;17)(q22;q11.2;q21) | M | 44 | M3 | + | 46,XY 46,XY,t(15;22;17)(q22;q11.2;q21) |
C+A | − | 1+ |
| Grimwade et al (6) | t(1;17;15)(p32;q21;q22) | M | NA | M3 | NA | 46,XY 46,XY, t(1;17;15)(p32;q21;q22) 46,idem,add(21)(p13) |
NA | NA | NA |
| Grimwade et al (6) | t(7;17;15)(q22;q21;q22) | F | NA | M3v | NA | 46,XX, t(7;17;15)(q22;q21;q22) | NA | NA | NA |
| t(6;17;15)(p21;q21;q22) | M | NA | M3v | NA | 46,XY 46,XY, t(6;17;15)(p21;q21;q22) |
NA | NA | NA | |
| t(8;17;15)(q22;q21;q22) | F | NA | M3 and M3r | NA | 46,XX 46,XX, t(8;17;15)(q22;q21;q22)47,idem,+8 |
NA | NA | NA | |
| t(13;17;15)(p13;q21;q22) | F | NA | M3v | NA | 46,XX 46,XX, t(13;17;15)(p13;q21;q22) |
NA | NA | NA | |
| t(5;17;15)(q14;q21;q22) | F | NA | M3 | NA | 46,XX,t(5;17;15)(q14;q21;q22 48,idem,+8,+21 | NA | NA | NA | |
| Tirado et al (36) | t(15;17;17)(q22;q23;q21) | F | 52 | M3 | − | 46,XX, t(15;17;17)(q22;q23;q21) | NA | − | 12+ |
| Freeman et al (37) | t(3;17;15)(q27;q21;q22) | M | 78 | M3 | − | 46,XY 46,XY, t(3;17;15)(q27;q21;q22) |
C+A | − | NA |
| t(8;17;15)(q24.3;q12;q22) | F | 49 | M3 | + | 46,XX 46,XX, t(8;17;15)(q24.3;q12;q22) |
C+A | − | NA | |
| Huret et al (38) | t(4;15;17)(q21;q26;q22) relapsed | M | 67 | M3 | + | 46,XY, 46,XY, t(4;15;17)(q21;q26;q22) 46,XY, add(1)(p?),t(1;18)(p36;p10),add(10)(p?), t(15;17)(q26;q22) |
C | + | 25+ |
| McKinney et al (39) | t(3;17;15)(p21;q21;q22) | F | 51 | M3 | + | 46,XX,t(3;17;15)(p21;q21;q22) | C | + | 33 |
| t(3;17;15)(p12;q21;q22) | M | 63 | M3 | + | 46,XY 46,XY, t(3;17;15)(q12;q21;q22) |
C | + | 14 | |
| Galieni et al (40) | t(1;17;15)(q23;q23;q22) | M | 41 | M3 | NA | 46,XY | |||
| 46,XY, t(1;17;15)(q23;q23;q22)47,idem, +10 | C+A | − | 8+ |
FAB, French-American-British classification of acute leukemia; DIC: disseminated intravascular coagulation; NA, not available; NR, not remission; C, chemotherapy; A, all-trans retinoic acid; As2O3, arsenic trioxide; BMT, bone marrow transplantation; M, male; F, female.
In addition, the association between classical t(15;17) and ct(15;17;v) clones remains unclear. Several previous studies have assumed that the complicated rearrangements originate from the single standard translocation and then quickly outgrow this stem line (16,26,33). The current study is the first to report the presentation of two abnormal clones, classical t(15;17) and ct(15;17;v), simultaneously in an adult APL patient, providing indirect evidence that the variant translocation possibly evolves from the classical t(15;17).
It is well known that chromosomal imbalances, including the deletion(s) or amplification(s) of key driver gene(s), may promote the malignant transformation of leukemia (42–44). In the reported 45 APL cases of ct(15;17;v), eight cases presented with trisomy 8 or partial trisomy, including 8p+, 4p+, 9p+, +10 and +X. However, deletions, particularly small or cryptic deletions, have not yet been reported. In the present study, array CGH revealed the cryptic chromosome aberrations, including the deletions of ATP2B2, ANGPT1 and ETV6, and a gain of RB1CC1 genes, but no imbalances of the breakpoints of chromosomes 15, 16 or 17. FISH analysis not only confirmed the deletions of ATP2B2, ANGPT1 and ETV6, but also demonstrated that these deletions were somatically acquired events since normal cells coexist with abnormal cells, which may potentially be involved in the leukemogenesis. However, the gain of 8q11.23 has not been confirmed due to the one size of duplication, which is too small to confirm by FISH. Furthermore, the possibility of a constitutional gain rather than disease-related gain cannot be ruled out. A divergence of ratio has also been identified between the deletions of ATP2B2 (97%), ANGPT1 (52%), ETV6 (90%) and the PML/RARA fusion (classical and variant combined; 90%) by FISH, suggesting that these unbalanced genetic events may occur asynchronously in leukemogenesis.
In conclusion, the current study was the first to identify the classical t(15;17) and complex variant t(15;16;17) in an adult patient with APL using FISH. Furthermore, cryptic genomic alterations involving leukemia-related genes, such as ATP2B2, ANGPT1, ETV6 and RB1CC1, were inferred in the level of array CGH and confirmed by FISH. It may be proposed that the malignant transformation of APL with complex variant translocation presents the following multi-step progression: i) Classical translocation; ii) formation of complex variant translocation by third chromosome involvement; and iii) further genomic changes. The further genomic changes, with the exception of the fusion of PML/RARA in APL, may be implicated in the heterogenicity of therapy outcome. Advanced study on these genes is likely to aid the elucidation of the oncogene or tumor suppressor gene candidates that potentially affect the prognosis of APL.
References
- 1.Degos L, Dombret H, Chomienne C, et al. All-trans-retinoic acid as a differentiating agent in the treatment of acute promyelocytic leukemia. Blood. 1995;85:2643–2653. [PubMed] [Google Scholar]
- 2.Shen ZX, Chen GQ, Ni JH, et al. Use of arsenic trioxide (As2O3) in the treatment of acute promyelocytic leukemia (APL): II. Clinical efficacy and pharmacokinetics in relapsed patients. Blood. 1997;89:3354–3360. [PubMed] [Google Scholar]
- 3.Redner RL. Variations on a theme: the alternate translocations in APL. Leukemia. 2002;16:1927–1932. doi: 10.1038/sj.leu.2402720. [DOI] [PubMed] [Google Scholar]
- 4.Brunel V, Lafage-Pochitaloff M, Alcalay M, Pelicci PG, Birg F. Variant and masked translocations in acute promyelocytic leukemia. Leuk Lymphoma. 1996;22:221–228. doi: 10.3109/10428199609051752. [DOI] [PubMed] [Google Scholar]
- 5.Xu L, Zhao WL, Xiong SM, et al. Molecular cytogenetic characterization and clinical relevance of additional, complex and/or variant chromosome abnormalities in acute promyelocytic leukemia. Leukemia. 2001;15:1359–1368. doi: 10.1038/sj.leu.2402205. [DOI] [PubMed] [Google Scholar]
- 6.Grimwade D, Biondi A, Mozziconacci MJ, et al. Characterization of acute promyelocytic leukemia cases lacking the classic t(15;17): results of the European Working Party. Groupe Français de Cytogénétique Hématologique, Groupe de Français d’Hematologie Cellulaire, UK Cancer Cytogenetics Group and BIOMED 1 European Community-Concerted Action ‘Molecular Cytogenetic Diagnosis in Haematological Malignancies’. Blood. 2000;96:1297–1308. [PubMed] [Google Scholar]
- 7.Rohr SS, Pelloso LA, Borgo A, et al. Acute promyelocytic leukemia associated with the PLZF-RARA fusion gene: two additional cases with clinical and laboratorial peculiar presentations. Med Oncol. 2012;29:2345–2347. doi: 10.1007/s12032-011-0147-y. [DOI] [PubMed] [Google Scholar]
- 8.Bennett JM, Catovsky D, Daniel MT, et al. Proposals for the classification of the acute leukaemias. French-American-British (FAB) co-operative group. Br J Haematol. 1976;33:451–458. doi: 10.1111/j.1365-2141.1976.tb03563.x. [DOI] [PubMed] [Google Scholar]
- 9.Kato T, Hangaishi A, Ichikawa M, Motokura T, Takahashi T, Kurokawa M. A new three-way variant t(15;22;17)(q22;q11.2;q21) in acute promyelocytic leukemia. Int J Hematol. 2009;89:204–208. doi: 10.1007/s12185-008-0253-6. [DOI] [PubMed] [Google Scholar]
- 10.Ried T, Schröck E, Ning Y, Wienberg J. Chromosome painting: a useful art. Hum Mol Genet. 1998;7:1619–1626. doi: 10.1093/hmg/7.10.1619. [DOI] [PubMed] [Google Scholar]
- 11.Berger R, Flandrin G, Bernheim A, et al. Cytogenetic studies on 519 consecutive de novo acute nonlymphocytic leukemias. Cancer Genet Cytogenet. 1987;29:9–21. doi: 10.1016/0165-4608(87)90026-4. [DOI] [PubMed] [Google Scholar]
- 12.Bernstein R, Mendelow B, Pinto MR, Morcom G, Bezwoda W. Complex translocations involving chromosomes 15 and 17 in acute promyelocytic leukaemia. Br J Haematol. 1980;46:311–314. doi: 10.1111/j.1365-2141.1980.tb05972.x. [DOI] [PubMed] [Google Scholar]
- 13.Bernstein R, Pinto MR, Morcom G, et al. Karyotype analysis in acute nonlymphocytic leukemia (ANLL): comparison with ethnic group, age, morphology, and survival. Cancer Genet Cytogenet. 1982;6:187–199. doi: 10.1016/0165-4608(82)90056-5. [DOI] [PubMed] [Google Scholar]
- 14.Ohyashiki K, Oshimura M, Uchida H, et al. Cytogenetic and ultrastructural studies on ten patients with acute promyelocytic leukemia, including one case with a complex translocation. Cancer Genet Cytogenet. 1985;14:247–255. doi: 10.1016/0165-4608(85)90190-6. [DOI] [PubMed] [Google Scholar]
- 15.Callen DF, Dale BM, Sage RE, Ford JH. A complex translocation in acute promyelocytic leukemia. Cancer Genet Cytogenet. 1985;16:45–48. doi: 10.1016/0165-4608(85)90076-7. [DOI] [PubMed] [Google Scholar]
- 16.Bjerrum OW, Philip P, Pressler T, Tygstrup I. Acute promyelocytic leukemia with t(15;17) and t(2;17;15) Cancer Genet Cytogenet. 1987;28:107–111. doi: 10.1016/0165-4608(87)90359-1. [DOI] [PubMed] [Google Scholar]
- 17.Osella P, Wyandt H, Vosburgh E, Milunsky A. Report of a variant t(1;15;17)(p36;q22;q21.1) in a patient with acute promyelocytic leukemia. Cancer Genet Cytogenet. 1991;57:201–207. doi: 10.1016/0165-4608(91)90153-l. [DOI] [PubMed] [Google Scholar]
- 18.Saitoh K, Miura I, Kobayashi Y, et al. A new variant translocation of t(15;17) in a patient with acute promyelocytic leukemia (M3): t(15;19;17)(q22;p13;q12) Cancer Genet Cytogenet. 1998;102:15–18. doi: 10.1016/s0165-4608(97)00299-9. [DOI] [PubMed] [Google Scholar]
- 19.Wan TS, Chim CS, So CK, Chan LC, Ma SK. Complex variant 15;17 translocations in acute promyelocytic leukemia. A case report and review of three-way translocations. Cancer Genet Cytogenet. 1999;111:139–143. doi: 10.1016/s0165-4608(98)00230-1. [DOI] [PubMed] [Google Scholar]
- 20.Fujishima M, Takahashi N, Miura I, et al. A PML/RARA chimeric gene on chromosome 2 in a patient with acute promyelocytic leukemia (M3) associated with a new variant translocation: t(2;15;17)(q21;q22;q21) Cancer Genet Cytogenet. 2000;120:80–82. doi: 10.1016/s0165-4608(99)00238-1. [DOI] [PubMed] [Google Scholar]
- 21.Liu S, Li Q, Pang W, et al. A new complex variant t(4;15;17) in acute promyelocytic leukemia: fluorescence in situ hybridization confirmation and literature review. Cancer Genet Cytogenet. 2001;130:33–37. doi: 10.1016/s0165-4608(01)00464-2. [DOI] [PubMed] [Google Scholar]
- 22.Misawa S, Lee E, Schiffer CA, Liu Z, Testa JR. Association of the translocation (15;17) with malignant proliferation of promyelocytes in acute leukemia and chronic myelogenous leukemia at blastic crisis. Blood. 1986;67:270–274. [PubMed] [Google Scholar]
- 23.Zaccaria A, Testoni M, Martinelli G, et al. Four-chromosomes complex translocations in acute promyelocytic leukemia: description of two cases. Eur J Haematol. 1994;52:129–133. doi: 10.1111/j.1600-0609.1994.tb01302.x. [DOI] [PubMed] [Google Scholar]
- 24.Ogawa S, Mitani K, Sato Y, et al. Detection of the PML/RAR alpha fusion gene in acute promyelocytic leukemia with a complex translocation involving chromosomes 15, 17, and 18. Cancer Genet Cytogenet. 1993;69:113–117. doi: 10.1016/0165-4608(93)90084-y. [DOI] [PubMed] [Google Scholar]
- 25.Chen Z, Morgan R, Stone JF, Sandberg AA. Identification of complex t(15;17) in APL by FISH. Cancer Genet Cytogenet. 1994;72:73–74. doi: 10.1016/0165-4608(94)90114-7. [DOI] [PubMed] [Google Scholar]
- 26.Calabrese G, Min T, Stuppia L, et al. Complex chromosome translocations of standard t(8;21) and t(15;17) arise from a two-step mechanism as evidenced by fluorescence in situ hybridization analysis. Cancer Genet Cytogenet. 1996;91:40–45. doi: 10.1016/s0165-4608(96)00096-9. [DOI] [PubMed] [Google Scholar]
- 27.Park JP, Fairweather RB. Complex t(1;15;17) in acute promyelocytic leukemia with duplication of RAR alpha and PML sequences. Cancer Genet Cytogenet. 1996;89:52–56. doi: 10.1016/0165-4608(95)00365-7. [DOI] [PubMed] [Google Scholar]
- 28.Yamamoto K, Hamaguchi H, Nagata K, Kobayashi M, Takashima T, Taniwaki M. A new complex translocation (15;20;17)(q22;p13;q21) in acute promyelocytic leukemia. Cancer Genet Cytogenet. 1998;101:89–94. doi: 10.1016/s0165-4608(97)00251-3. [DOI] [PubMed] [Google Scholar]
- 29.Kudva GC, Maliekel K, Richart JM, et al. Acute promyelocytic leukemia and HIV-1 infection: case report and review of the literature. Am J Hematol. 2004;77:287–290. doi: 10.1002/ajh.20192. [DOI] [PubMed] [Google Scholar]
- 30.Eclache V, Viguie F, Frocrain C, et al. A new variant t(6;15;17)(q25;q22;q21) in acute promyelocytic leukemia: fluorescence in situ hybridization confirmation. Cancer Genet Cytogenet. 2005;159:69–73. doi: 10.1016/j.cancergencyto.2004.09.013. [DOI] [PubMed] [Google Scholar]
- 31.García-Casado Z, Cervera J, Valencia A, et al. A t(17;20)(q21;q12) masking a variant t(15;17)(q22;q21) in a patient with acute promyelocytic leukemia. Cancer Genet Cytogenet. 2006;168:73–76. doi: 10.1016/j.cancergencyto.2005.12.014. [DOI] [PubMed] [Google Scholar]
- 32.Yoo SJ, Seo EJ, Lee JH, Seo YH, Park PW, Ahn JY. A complex, four-way variant t(15;17) in acute promyelocytic leukemia. Cancer Genet Cytogenet. 2006;167:168–171. doi: 10.1016/j.cancergencyto.2005.12.006. [DOI] [PubMed] [Google Scholar]
- 33.Miyazaki K, Kikukawa M, Kiuchi A, Shin K, Iwamoto T, Ohyashiki K. Complex translocations derived stepwise from standard t(15;17) in a patient with variant acute promyelocytic leukemia. Cancer Genet Cytogenet. 2007;176:127–130. doi: 10.1016/j.cancergencyto.2007.03.013. [DOI] [PubMed] [Google Scholar]
- 34.Abe S, Ishikawa I, Harigae H, Sugawara T. A new complex translocation t(5;17;15)(q11;q21;q22) in acute promyelocytic leukemia. Cancer Genet Cytogenet. 2008;184:44–47. doi: 10.1016/j.cancergencyto.2008.03.006. [DOI] [PubMed] [Google Scholar]
- 35.Stavropoulou C, Georgakakos VN, Manola KN, et al. 5′RARA submicroscopic deletion from new variant translocation involving chromosomes 15, 17, and 18, in a case of acute promyelocytic leukemia. Cancer Genet Cytogenet. 2008;182:50–55. doi: 10.1016/j.cancergencyto.2007.12.011. [DOI] [PubMed] [Google Scholar]
- 36.Tirado CA, Golembiewski-Ruiz V, Horvatinovich J, et al. Cytogenetic and molecular analysis of an unusual case of acute promyelocytic leukemia with a t(15;17;17)(q22;q23;q21) Cancer Genet Cytogenet. 2003;145:31–37. doi: 10.1016/s0165-4608(03)00027-x. [DOI] [PubMed] [Google Scholar]
- 37.Freeman CE, Mercer DD, Ye Y, Van Brunt J, 3rd, Li MM. Cytogenetic and molecular characterization of complex three-way translocations in acute promyelocytic leukemia. Beijing Da Xue Xue Bao. 2009;41:477–479. (In Chinese) [PubMed] [Google Scholar]
- 38.Huret JL, Couet D, Guilhot F, Brizard A, Tanzer J. A two-step t(4;der(15)) t(15;17) complex translocation in an acute promyelocytic leukaemia and review of the literature. Leuk Res. 1987;11:761–765. doi: 10.1016/0145-2126(87)90014-2. [DOI] [PubMed] [Google Scholar]
- 39.McKinney CD, Golden WL, Gemma NW, Swerdlow SH, Williams ME. RARA and PML gene rearrangements in acute promyelocytic leukemia with complex translocations and atypical features. Genes Chromosomes Cancer. 1994;9:49–56. doi: 10.1002/gcc.2870090109. [DOI] [PubMed] [Google Scholar]
- 40.Galieni P, Marotta G, Vessichelli F, et al. Variant t(1;15;17)(q23;q22;q23) in a case of acute promyelocytic leukemia. Leukemia. 1996;10:1658–1661. [PubMed] [Google Scholar]
- 41.De Braekeleer E, Douet-Guilbert N, Le Bris MJ, Morel F, Férec C, De Braekeleer M. RUNX1-MTG16 fusion gene in acute myeloblastic leukemia with t(16;21)(q24;q22): case report and review of the literature. Cancer Genet Cytogenet. 2008;185:47–50. doi: 10.1016/j.cancergencyto.2008.04.011. [DOI] [PubMed] [Google Scholar]
- 42.Tybäkinoja A, Elonen E, Piippo K, Porkka K, Knuutila S. Oligonucleotide array-CGH reveals cryptic gene copy number alterations in karyotypically normal acute myeloid leukemia. Leukemia. 2007;21:571–574. doi: 10.1038/sj.leu.2404543. [DOI] [PubMed] [Google Scholar]
- 43.Zagaria A, Anelli L, Albano F, Vicari L, Schiavone EM, et al. Molecular cytogenetic characterization of deletions on der(9) in chronic myelocytic leukemia. Cancer Genet Cytogenet. 2006;167:97–102. doi: 10.1016/j.cancergencyto.2006.01.011. [DOI] [PubMed] [Google Scholar]
- 44.Paulsson K, Cazier JB, Macdougall F, et al. Microdeletions are a general feature of adult and adolescent acute lymphoblastic leukemia: Unexpected similarities with pediatric disease. Proc Natl Acad Sci USA. 2008;105:6708–6713. doi: 10.1073/pnas.0800408105. [DOI] [PMC free article] [PubMed] [Google Scholar]