Abstract
The DNase I (EC 3.1.21.1) sensitivity of transcribed yeast chromatin has been examined. We find that, in contrast to chromatin from higher eukaryotes, transcribed yeast chromatin and total yeast chromatin are equally sensitive to DNase I digestion. We interpret these results to mean that the entire yeast genome exists in a state that represents a restricted proportion of total chromatin in higher eukaryotes.
Full text
PDF
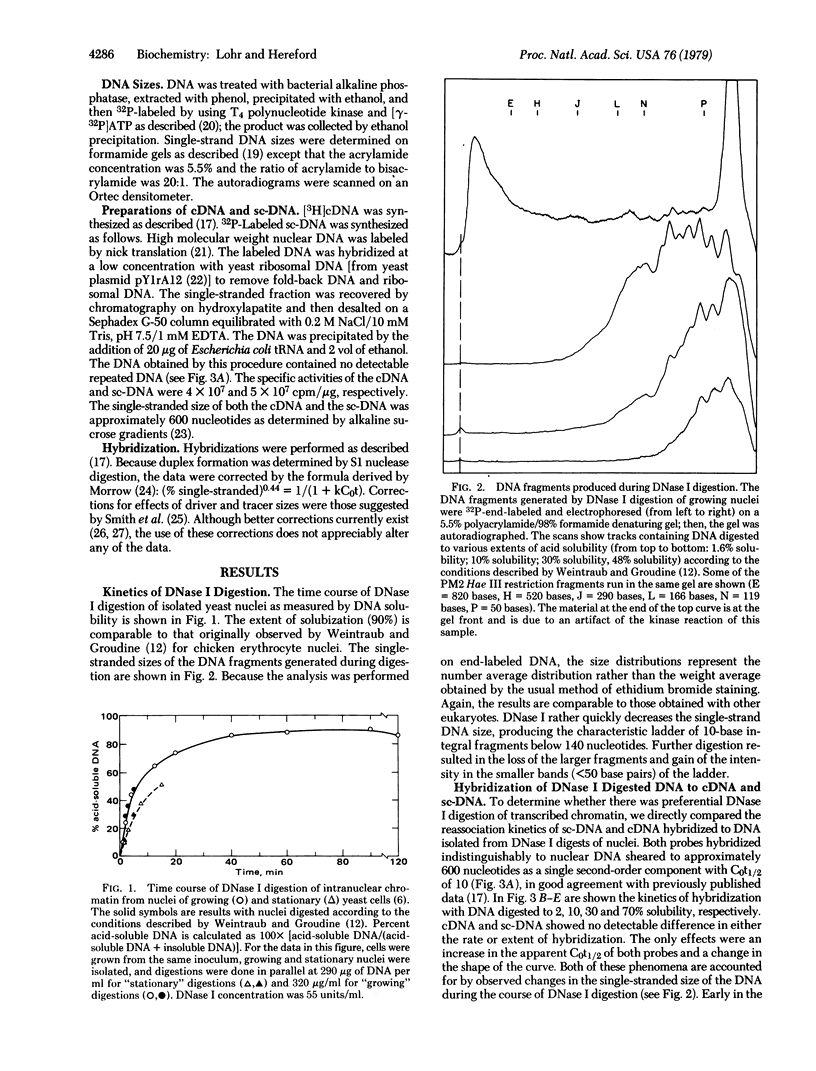
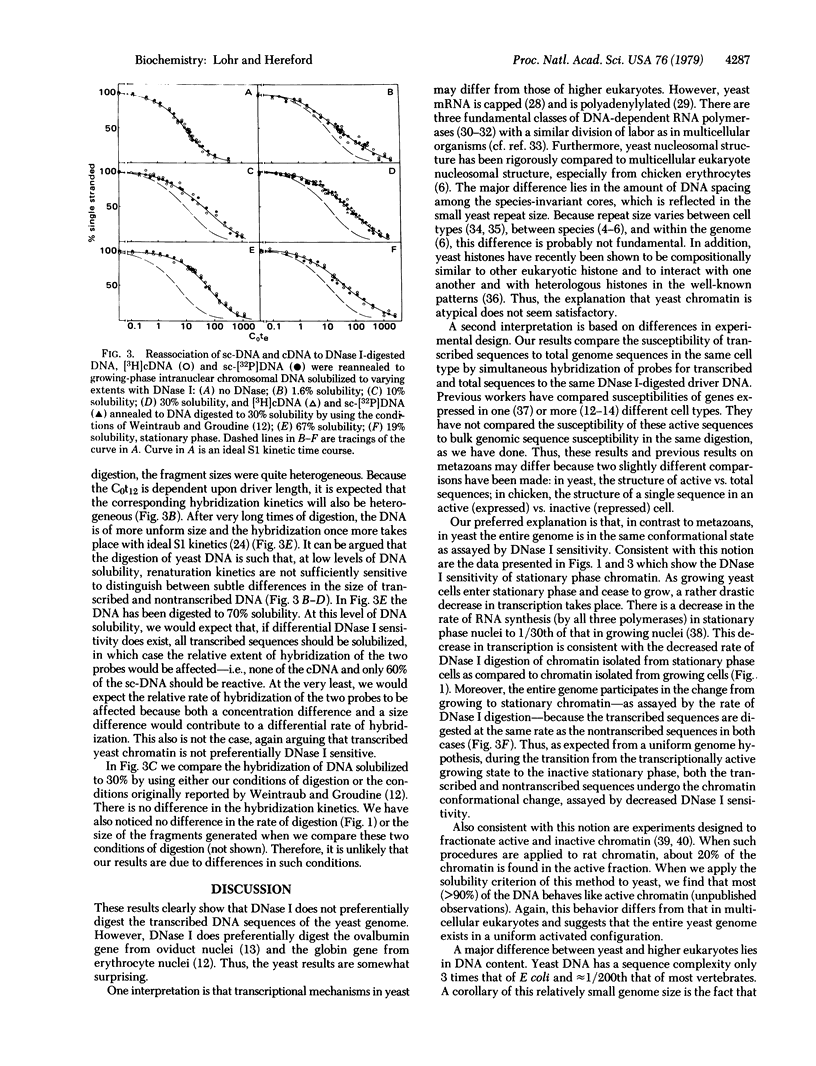

Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Axel R. Cleavage of DNA in nuclei and chromatin with staphylococcal nuclease. Biochemistry. 1975 Jul;14(13):2921–2925. doi: 10.1021/bi00684a020. [DOI] [PubMed] [Google Scholar]
- Axel R., Feigelson P., Schutz G. Analysis of the complexity and diversity of mRNA from chicken liver and oviduct. Cell. 1976 Feb;7(2):247–254. doi: 10.1016/0092-8674(76)90024-6. [DOI] [PubMed] [Google Scholar]
- Chamberlin M. E., Galau G. A., Britten R. J., Davidson E. H. Studies on nucleic acid reassociation kinetics: V. Effects of disparity in tracer and driver fragment lengths. Nucleic Acids Res. 1978 Jun;5(6):2073–2094. doi: 10.1093/nar/5.6.2073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Compton J. L., Bellard M., Chambon P. Biochemical evidence of variability in the DNA repeat length in the chromatin of higher eukaryotes. Proc Natl Acad Sci U S A. 1976 Dec;73(12):4382–4386. doi: 10.1073/pnas.73.12.4382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finch J. T., Lutter L. C., Rhodes D., Brown R. S., Rushton B., Levitt M., Klug A. Structure of nucleosome core particles of chromatin. Nature. 1977 Sep 1;269(5623):29–36. doi: 10.1038/269029a0. [DOI] [PubMed] [Google Scholar]
- Flint S. J., Weintraub H. M. An altered subunit configuration associated with the actively transcribed DNA of integrated adenovirus genes. Cell. 1977 Nov;12(3):783–794. doi: 10.1016/0092-8674(77)90277-x. [DOI] [PubMed] [Google Scholar]
- Foe V. E., Wilkinson L. E., Laird C. D. Comparative organization of active transcription units in Oncopeltus fasciatus. Cell. 1976 Sep;9(1):131–146. doi: 10.1016/0092-8674(76)90059-3. [DOI] [PubMed] [Google Scholar]
- Garel A., Axel R. Selective digestion of transcriptionally active ovalbumin genes from oviduct nuclei. Proc Natl Acad Sci U S A. 1976 Nov;73(11):3966–3970. doi: 10.1073/pnas.73.11.3966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garel A., Zolan M., Axel R. Genes transcribed at diverse rates have a similar conformation in chromatin. Proc Natl Acad Sci U S A. 1977 Nov;74(11):4867–4871. doi: 10.1073/pnas.74.11.4867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gordon C. N. Chromatin behaviour during the mitotic cell cycle of Saccharomyces cerevisiae. J Cell Sci. 1977 Apr;24:81–93. doi: 10.1242/jcs.24.1.81. [DOI] [PubMed] [Google Scholar]
- Gottesfeld J. M., Butler P. J. Structure of transcriptionally-active chromatin subunits. Nucleic Acids Res. 1977 Sep;4(9):3155–3173. doi: 10.1093/nar/4.9.3155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottesfeld J. M., Partington G. A. Distribution of messenger RNA-coding sequences in fractionated chromatin. Cell. 1977 Dec;12(4):953–962. doi: 10.1016/0092-8674(77)90160-x. [DOI] [PubMed] [Google Scholar]
- Hartwell L. H. Macromolecule synthesis in temperature-sensitive mutants of yeast. J Bacteriol. 1967 May;93(5):1662–1670. doi: 10.1128/jb.93.5.1662-1670.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hereford L. M., Rosbash M. Number and distribution of polyadenylated RNA sequences in yeast. Cell. 1977 Mar;10(3):453–462. doi: 10.1016/0092-8674(77)90032-0. [DOI] [PubMed] [Google Scholar]
- Hinnebusch A. G., Clark V. E., Klotz L. C. Length dependence in reassociation kinetics of radioactive tracer DNA. Biochemistry. 1978 Apr 18;17(8):1521–1529. doi: 10.1021/bi00601a026. [DOI] [PubMed] [Google Scholar]
- Jendrisak J. J., Burgess R. R. Studies of the subunit structure of wheat germ ribonucleic acid polymerase II. Biochemistry. 1977 May 3;16(9):1959–1964. doi: 10.1021/bi00628a032. [DOI] [PubMed] [Google Scholar]
- Lacy E., Axel R. Analysis of DNA of isolated chromatin subunits. Proc Natl Acad Sci U S A. 1975 Oct;72(10):3978–3982. doi: 10.1073/pnas.72.10.3978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levy B., Dixon G. H. Renaturation kinetics of cDNA complementary to cytoplamic polyadenylated RNA from rainbow trout testis. Accessibility of transcribed genes to pancreatic DNase. Nucleic Acids Res. 1977 Apr;4(4):883–898. doi: 10.1093/nar/4.4.883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lohr D., Corden J., Tatchell K., Kovacic R. T., Van Holde K. E. Comparative subunit structure of HeLa, yeast, and chicken erythrocyte chromatin. Proc Natl Acad Sci U S A. 1977 Jan;74(1):79–83. doi: 10.1073/pnas.74.1.79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lohr D., Ide G. Comparison on the structure and transcriptional capability of growing phase and stationary yeast chromatin: a model for reversible gene activation. Nucleic Acids Res. 1979;6(5):1909–1927. doi: 10.1093/nar/6.5.1909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lohr D., Kovacic R. T., Van Holde K. E. Quantitative analysis of the digestion of yeast chromatin by staphylococcal nuclease. Biochemistry. 1977 Feb 8;16(3):463–471. doi: 10.1021/bi00622a020. [DOI] [PubMed] [Google Scholar]
- Maniatis T., Jeffrey A., Kleid D. G. Nucleotide sequence of the rightward operator of phage lambda. Proc Natl Acad Sci U S A. 1975 Mar;72(3):1184–1188. doi: 10.1073/pnas.72.3.1184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mardian J. K., Isenberg I. Yeast inner histones and the evolutionary conservation of histone-histone interactions. Biochemistry. 1978 Sep 5;17(18):3825–3833. doi: 10.1021/bi00611a023. [DOI] [PubMed] [Google Scholar]
- McLaughlin C. S., Warner J. R., Edmonds M., Nakazato H., Vaughan M. H. Polyadenylic acid sequences in yeast messenger ribonucleic acid. J Biol Chem. 1973 Feb 25;248(4):1466–1471. [PubMed] [Google Scholar]
- Noll M. Differences and similarities in chromatin structure of Neurospora crassa and higher eucaryotes. Cell. 1976 Jul;8(3):349–355. doi: 10.1016/0092-8674(76)90146-x. [DOI] [PubMed] [Google Scholar]
- Pardon J. F., Worcester D. L., Wooley J. C., Cotter R. I., Lilley D. M., Richards R. M. The structure of the chromatin core particle in solution. Nucleic Acids Res. 1977 Sep;4(9):3199–3214. doi: 10.1093/nar/4.9.3199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petes T. D., Broach J. R., Wensink P. C., Hereford L. M., Fink G. R., Botstein D. Isolation and analysis of recombinant DNA molecules containing yeast DNA. Gene. 1978 Sep;4(1):37–49. doi: 10.1016/0378-1119(78)90013-6. [DOI] [PubMed] [Google Scholar]
- STUDIER F. W. SEDIMENTATION STUDIES OF THE SIZE AND SHAPE OF DNA. J Mol Biol. 1965 Feb;11:373–390. doi: 10.1016/s0022-2836(65)80064-x. [DOI] [PubMed] [Google Scholar]
- Schultz L. D., Hall B. D. Transcription in yeast: alpha-amanitin sensitivity and other properties which distinguish between RNA polymerases I and III. Proc Natl Acad Sci U S A. 1976 Apr;73(4):1029–1033. doi: 10.1073/pnas.73.4.1029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultz L. D. Transcriptional role of yeast deoxyribonucleic acid dependent ribonucleic acid polymerase III. Biochemistry. 1978 Feb 21;17(4):750–758. doi: 10.1021/bi00597a031. [DOI] [PubMed] [Google Scholar]
- Simpson R. T., Whitlock J. P. Mapping DNAase l-susceptible sites in nucleosomes labeled at the 5' ends. Cell. 1976 Oct;9(2):347–353. doi: 10.1016/0092-8674(76)90124-0. [DOI] [PubMed] [Google Scholar]
- Smith M. J., Britten R. J., Davidson E. H. Studies on nucleic acid reassociation kinetics: reactivity of single-stranded tails in DNA-DNA renaturation. Proc Natl Acad Sci U S A. 1975 Dec;72(12):4805–4809. doi: 10.1073/pnas.72.12.4805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sollner-Webb B., Felsenfeld G. A comparison of the digestion of nuclei and chromatin by staphylococcal nuclease. Biochemistry. 1975 Jul;14(13):2915–2920. doi: 10.1021/bi00684a019. [DOI] [PubMed] [Google Scholar]
- Sripati C. E., Groner Y., Warner J. R. Methylated, blocked 5' termini of yeast mRNA. J Biol Chem. 1976 May 25;251(10):2898–2904. [PubMed] [Google Scholar]
- Tatchell K., Van Holde K. E. Compact oligomers and nucleosome phasing. Proc Natl Acad Sci U S A. 1978 Aug;75(8):3583–3587. doi: 10.1073/pnas.75.8.3583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas J. O., Thompson R. J. Variation in chromatin structure in two cell types from the same tissue: a short DNA repeat length in cerebral cortex neurons. Cell. 1977 Apr;10(4):633–640. doi: 10.1016/0092-8674(77)90096-4. [DOI] [PubMed] [Google Scholar]
- Valenzuela P., Hager G. L., Weinberg F., Rutter W. J. Molecular structure of yeast RNA polymerase III: demonstration of the tripartite transcriptive system in lower eukaryotes. Proc Natl Acad Sci U S A. 1976 Apr;73(4):1024–1028. doi: 10.1073/pnas.73.4.1024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weintraub H., Groudine M. Chromosomal subunits in active genes have an altered conformation. Science. 1976 Sep 3;193(4256):848–856. doi: 10.1126/science.948749. [DOI] [PubMed] [Google Scholar]



