Summary
In recent years, long noncoding RNAs (lncRNAs) have emerged as an important class of regulators of gene expression. Notably, they exhibit several distinctive features that confer these functions, including exquisite cell- and tissue-specific expression and the capacity to transduce higher order regulatory networks. Here we review evidence showing that lncRNAs exert critical functions in adult tissue stem cells, including skin, brain, and muscle, as well as developmental patterning and pluripotency. We highlight new approaches for ascribing lncRNA functions and discuss mammalian dosage compensation as a classic example of a lncRNA network that provides broader insights into the class overall.
Introduction
Efforts to understand how tissues are patterned during development and maintained by stem cells throughout life have traditionally focused on the protein-coding genome. Over the past decade, however, our understanding of the non-coding genome and its impact on cell fate has dramatically expanded. Contrary to previous notions of genome organization and function, the identification of thousands of long and short noncoding RNAs (ncRNAs) has revealed that much of the genome is in fact transcribed. Long noncoding RNAs (lncRNAs) are operationally defined as transcripts of greater than 200 nucleotides that function by means other than coding for proteins; lncRNAs are typically transcribed by RNA polymerase II and are frequently spliced and polyadenylated (reviewed by (Rinn and Chang, 2012). As a class, lncRNAs tend to be expressed at lower levels and are predominantly localized in the nucleus, in contrast to messenger RNAs, which are abundant and enriched in the cytoplasm (Derrien et al., 2012). Notwithstanding these generalizations, lncRNAs exhibit a wide range of expression levels and distinct cytotopic localizations, reflecting a large and diverse class of regulators (reviewed by (Batista and Chang, 2013)). Several well-studied examples of lncRNAs suggest that they can operate through distinct modes, including as signals, scaffolds for protein-protein interactions, molecular decoys, and guides to target elements in the genome or transcriptome (Wang and Chang, 2011). The discovery of novel lncRNAs has historically outpaced their functional annotation, however efforts to more specifically ascribe function to either previously identified or novel lncRNAs have increased in recent years. Stem cells offer an attractive system for studying lncRNA function since previous findings have suggested that lncRNA expression is more cell type specific than mRNA expression (Cabili et al., 2011), leading to the possibility that lncRNAs may be key regulators of cell fate.
Here we review recent developments that illuminate the roles of lncRNAs in stem cell biology. We explore efforts to characterize the functions of lncRNAs in the development and patterning of several somatic tissues, including skin, brain, and musculature. Additionally, we examine how lncRNAs contribute to the pluripotent state and can be used to assess reprogramming status.
LncRNAs in Adult Tissue Stem Cells
Skin: an ideal model
Studying the biology of tissues at the molecular level necessitates robust model systems. While there are few systems that are suitable for detailed molecular characterization, well-developed human models exist for the skin based on ex vivo tissue regeneration that can also be grafted in vivo (Sen et al., 2010; Truong et al., 2006). Such models provide cellular material for molecular and biochemical studies that would be otherwise inaccessible and offer a system for testing the function of lncRNAs. Surveying the pattern of gene expression during epidermal differentiation, Khavari and colleagues discovered two key lncRNAs, ANCR and TINCR, that are expressed in epidermal stem cells and their terminally differentiated progeny, respectively (Kretz et al., 2012; 2013) (Figure 1). ANCR (Anti-differentiation noncoding RNA) provides a prime example of a lncRNA that controls the differentiation state of a somatic stem cell (Kretz et al., 2012). Specifically, ANCR depletion results in ectopic differentiation of epidermal stem cells, implying that ANCR’s role is to suppress the differentiation pathway in the epidermis and maintain the stem cell compartment.
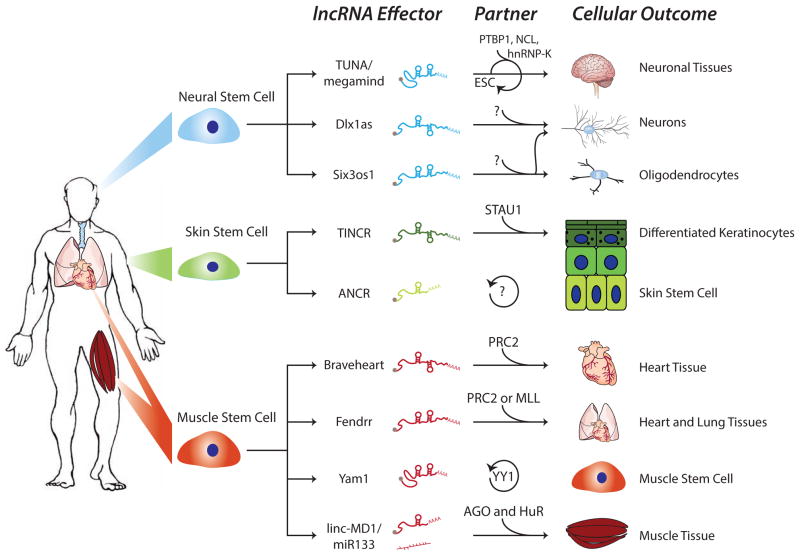
Figure 1. lncRNAs control differentiation and self-renewal.
Several lncRNAs that regulate specific somatic tissue stem cell renewal or differentiation and their protein partners are depicted. Some lncRNAs maintain the stem cell state while others promote a differentiation program. Their functions are often facilitated by protein partners that impart the ability to activate or repress gene expression or post-transcriptionally regulate other RNAs.
While ANCR appears to inhibit differentiation, a different lncRNA termed terminal differentiation induced noncoding RNA (TINCR) promotes epidermal differentiation (Kretz et al., 2013). TINCR is kept at very low levels in epidermal stem cells, but it is dramatically induced upon differentiation. Mechanistic studies of TINCR revealed that TINCR is a cytoplasmic lncRNA that interacts with the RNA-binding protein (RBP) STAU1 and converts STAU1 into an mRNA stability factor (Figure 1). Together, TINCR and STAU1 bind to and functionally stabilize mRNAs that encode structural and regulatory proteins critical for terminally differentiated keratinocytes. Additionally, TINCR expression is down regulated in human squamous cell carcinoma providing evidence that lncRNAs can functionally regulate healthy and disease tissues.
The development of two techniques made these insights possible: (i) RNA interactome analysis (RIA), which allows the retrieval and unbiased discovery of RNAs interacting with a lncRNA of interest, and (ii) protein microarray hybridization, which allows rapid discovery of direct RBP partners of a lncRNA (Kretz et al., 2013). Moreover, both ANCR and TINCR were identified from large-scale expression profiling studies, suggesting that many additional lncRNAs may be identified and characterized using this system. Indeed, the differentiation of the skin is a multistep and highly regulated process that could benefit from the diverse set of lncRNAs hiding in the genome. The development of techniques such as RIA and the implementation of protein microarrays facilitated the functional characterization of TINCR, but are applicable to uncovering mechanisms of other lncRNAs. Within the skin, the regulated and sequential expression of lncRNAs is clearly essential for their function, thus understanding what controls the spatiotemporal expression of lncRNAs, such as ANCR and TINCR, should be the focus of future studies.
Regulation in the brain
Transcription and alternative splicing in the brain appear to be the most complex among all organs (Mehler and Mattick, 2007; Mercer et al., 2008). An early example of lncRNAs controlling neural cell fates involves the Evf2 lncRNA and the Dlx5/6 genomic locus (Bond et al., 2009). Evf2 is transcribed antisense to Dlx6, which encodes a transcription factor, and is located immediately downstream of the Dlx5 genomic locus. The act of transcribing Evf2 can control the levels of Dlx6 in cis, and after disengaging the polymerase, Evf2 acts in trans to modulate the methylation of the Dlx5/6 enhancer and transcription of Dlx5. Therefore, by regulating the cellular levels of the Dlx5 and Dlx6 transcription factors, Efv2 controls GABAergic interneuron activity (Berghoff et al., 2013; Bond et al., 2009). A different study characterizing another lncRNA important for neural differentiation found that an enhancer region of the gene encoding the Neurogenin 1 transcription factor was transcribed and produced a lncRNA that positively regulated Neurogenin 1 expression (Onoguchi et al., 2012). These few examples begin to build the case that lncRNAs play an important role in neural biology.
The starting point of many lncRNA studies is unbiased gene expression analysis, which can reveal novel lncRNAs and their expression pattern in a developmental context. Recent large-scale efforts have employed next generation sequencing (“-seq”) technologies, from RNA-seq to chromatin immunoprecipitation (ChIP-seq), to identify transcripts and define their genomic positions (reviewed by (Rinn and Chang, 2012)). In the mouse brain, Lim and colleauges isolated three separate regions: subventricular zone (SVZ), olfactory bulb (OB) and the dentate gyrus (DG), and subjected these samples to short read RNA-seq and ChIP-seq (Ramos et al., 2013). Over 3,600 novel lncRNAs were identified and clustering of the lncRNAs and mRNAs by their expression patterns revealed that the lncRNAs were more tissue-specific than mRNAs, consistent with previous reports (Cabili et al., 2011). Application of CaptureSeq, a technique that circumvents some drawbacks of short-read sequencing (Mercer et al., 2011), to further characterize the transcriptome of adult SVC tissue doubled the number (to ~7000) of novel lncRNAs identified. To functionally validate the cataloging effort, two lncRNAs were identified by selecting loci marked by H3K4me3, which is associated with expressed genes, in NPC-SVC cells. This search identified Six3os and Dlx1as for further testing. Notably, Six3os has been previously reported to control retinal development (Rapicavoli et al., 2011). To characterize the neural role of Six3os and Dlx1as, SVZ neural progenitor cells were challenged in a 7-day differentiation assay with short hairpin RNAs targeting the two lncRNAs. Depletion of Six3os lncRNA lead to fewer Tuj1 (neuron marker) and OLIG2 (oligodentrocyte marker) positive cells, whereas depletion of Dlx1as specifically affected the number of Tuj1 positive cells (Figure 1). While the molecular mechanisms of these lncRNAs were not explored, Six3os has been shown to physically interact with Ezh2, a component of the Polycomb Repressive Complex 2 (PRC2), to repress specific genes in retinal cells (Rapicavoli et al., 2011). These examples illustrate that mapping spatiotemporal patterns of lncRNAs can highlight functional transcripts. Larger scale validation efforts will be required to fully realize the extent of lncRNA regulation in the different regions of the brain.
A complementary approach identifies potential lncRNA regulators based on their loss of function phenotypes in large-scale depletion studies (Guttman et al., 2012; Lin et al., 2014). Rana and colleagues targeted 1,280 mouse lncRNAs and identified 20 lncRNAs that were required for the maintenance of mESC pluripotency. One lncRNA, named TUNA, was previously identified as megamind in zebrafish. TUNA/Megamind depletion in zebrafish led to impaired locomotor response (Ulitsky et al., 2011). TUNA is highly conserved in human and fish, is required for the maintenance of pluripotency, and is also expressed in the brain, spinal cord, and eyes in adult tissues, Indeed, TUNA expression was increased when mESCs differentiated towards the neural lineages, and TUNA depletion inhibited neural differentiation of ESCs (Figure 1). Purifying proteins that associate with in vitro transcribed TUNA identified hnRNP-K, Nucleolin (NCL), and PTBP1 as interaction partners. Importantly, depletion of several of these proteins phenocopied TUNA depletion (Lin et al., 2014). An important caveat to consider is that while the candidate approach characterized TUNA, Six3os, and Dlx1as lncRNAs as successful validation of genome-wide screens, such approaches leave the function of thousands of other transcripts, many of which may play important roles, unaddressed.
Many lncRNAs have been implicated in the regulation of chromatin states (Rinn and Chang, 2012) but direct evidence for their association has only recently been possible through the development Chromatin Isolation by RNA Purification (ChIRP, and others methods discussed below)(Chu et al., 2011). ChIRP uses DNA capture probes to retrieve a specific lncRNA with its associated genomic DNA targets, and together with deep sequencing can generate a genome-wide map of lncRNA-chromatin interactions. Careful optimization of in vivo crosslinking, both of the chemical crosslinking agent and duration, and selection of proper oligonucleotide probes are important to obtain reliable measurement. This process often includes multiple but distinct DNA capture probe sets, probes targeting irrelevant RNAs as negative controls, and positive control regions to assay during pilot experiments (Chu et al., 2011). Successful implementation of ChIRP has revealed the lncRNA TUNA occupies promoter regions of NANOG, Sox2, and Fgf4, genes that are important for pluripotency and neural lineage commitment (Lin et al., 2014). Together with its protein partners and its chromatin localization, TUNA may regulate gene expression at both the transcriptional and post-transcriptional level. Thus, TUNA represents a lncRNA that is important for at least two cell states (ESC pluripotency and neural differentiation) and likely operates through multiple molecular mechanisms. This example highlights the concept that a single lncRNA can, under different cellular context and protein partners, function to control multiple molecular pathways.
lncRNAs and muscle
LncRNAs also control development of mesodermal tissues and have similarly benefited from large-scale sequencing efforts to identify functionally important transcripts. One example of a heart specific lncRNA named Braveheart was first functionally characterized as a key factor involved in cardiac lineage commitment because its depletion resulted in a severe reduction in the number of spontaneous beating cardiomyocytes formed during embryoid body differentiation (Klattenhoff et al., 2013) (Figure 1). Further characterization of Braveheart found that it interacts with Suz12, a subunit of PRC2, and acts as in trans to regulate heart-specific differentiation genes such as MesP1. The regulation of master drivers of cardiac differentiation such as MesP1 by Braveheart, offers new tools towards the goal of achieving highly efficient and reproducible in vitro reprograming (Burridge et al., 2012). Producing cardiomyocytes from induced pluripotency stem cells (iPSCs) or directly from other differentiated cell types, may benefit from engineering specific lncRNA expression during in vitro production.
While small interfering RNA (siRNA) knockdown of lncRNAs (used in most of the discussed work) often provides a great deal of insight into function, off-target effects and incomplete depletion must always been considered. As with protein coding genes, knockout (KO) strategies offer potential remedies to these siRNA-related issues, but the specific strategy employed is critical (discussed below: Developmental Patterning by lncRNAs). Utilizing this concept Herrmann and colleagues inserted a pre-mature polyadenylation (polyA) signal into the lncRNA Fendrr’s locus to promote accumulation of the full-length Fendrr RNA (Grote et al., 2013). Initial characterization of Fendrr found it expressed in the caudal end of the lateral plate mesoderm (LPM), which develops into the structures like the heart and body wall. Fendrr KO resulted in embryonic lethality at E13.75, abdominal wall defects, and pooling of blood in the right atrium. By partnering with both activating (Mixed Lineage Leukemia (MLL), WDR5) and silencing (PRC2) chromatin complexes Fendrr was proposed to modulate the epigenetic landscape during development (Figure 1). More recently ChIRP was used to show that Fendrr physically associates with the promoters of FoxF1 and Pitx2 mRNAs, two genes repressed by Fendrr (Grote and Herrmann, 2013; Grote et al., 2013). Fendrr therefore represents a dual-function lncRNA that may control both positive and negative chromatin modifying complexes to guide development.
Long RNAs controlling small RNAs
The differentiation of a myoblast progenitor cell (MB) to a fully differentiated muscle cell is a highly regulated process that relies on Ying Yang 1 (YY1), a multifunctioning transcription factor (Deng et al., 2010; Lu et al., 2012). Examination of YY1’s chromatin binding pattern in MBs revealed that it bound the promoter of many ncRNA loci, and these target noncoding genes were named YY1 associated muscle lncRNAs (Yam)(Lu et al., 2013). Characterization of one of these lncRNAs, Yam1, identified it as a key regulator of myogenesis, as it was able to repress key muscle differentiation genes including myogenin, Tnni2, and α-actin, (Figure 1). Furthermore, Yam1 increased levels of microRNA-715 (miR-715), which targets Wnt7b, a protein that normally promotes muscle differentiation (Lu et al., 2013). Yam1 thus provides evidence that in muscle lncRNAs can modulate the levels of both mRNAs and other ncRNAs, such as miRNAs, providing additional network control to cells.
The regulation of miRNA networks reveals an additional mechanism through which lncRNAs exert control. Recently, multiple lncRNAs have been shown to act as competing endogenous RNAs (ceRNAs), where the lncRNAs are proposed to bind to and compete miRNAs away from cognate mRNA targets (Tay et al., 2014). Pseudogene lncRNAs are prime candidates for the ceRNA mechanism because they may share multiple miRNA binding sites, allowing more effective competition with cognate mRNAs. The ceRNA hypothesis requires that ceRNAs are expressed highly enough and have sufficient numbers of miRNA binding sites to substantially affect the pool of cellular miRNAs. Recent work exploring the dynamics of miRNA-regulated gene repression have shown that it is highly susceptible to thresholds. In certain contexts, small concentration changes of miRNA-mRNA or miRNA-ceRNA pairs can substantially modulate the gene expression network (Bosia et al., 2013; Mukherji et al., 2011). Moreover, one example of a ceRNA, linc-MD1, has been previously show to regulate muscle differentiation through its ability to sponge miR-133 and miR-135 away from the mRNAs MAML1 and MEF2C (Cesana et al., 2011). These two mRNAs are important transcriptional activators of the muscle differentiation program. Linc-MD1 itself contains a miR-133b, which represses muscle differentiation when processed. Recent molecular characterization of this network revealed the RBP HuR bound to linc-MD1 and the levels of linc-MD1 positively correlated with HuR protein abundance (Legnini et al., 2014) (Figure 1)., HuR controlled the fate of linc-MD1, as cellular depletion of HuR favored the processing of linc-MD1 into miR-133b, tipping the balance in favor of the miRNA over the ceRNA.. HuR has known roles in myogenesis and its interaction with linc-MD1 fine-tunes the levels of miRNAs important in the muscle differentiation program. Together, these studies explore lncRNA functions in muscle tissue and help to expand the possible modes of lncRNA functions within the already complex system of miRNA-mediated gene regulation.
Developmental Patterning by lncRNAs
LncRNAs also orchestrate the patterning of cells into tissues and organs during development. HOTAIR lncRNA was one of the first characterized lncRNAs that acts at distance (in trans) to modulate Hox gene expression (Rinn et al., 2007). HOTAIR is a repressive lncRNA and serves a scaffold between two distinct chromatin modification complexes (Rinn et al., 2007; Tsai et al., 2010). Other Hox encoded lncRNAs such as, HOTTIP, Mistral, and HOTAIRm1 were shown to regulate different members of HoxA genes (Bertani et al., 2011; Wang et al., 2011; Zhang et al., 2009). For example, HOTTIP is expressed in distal anatomic structures and activates the expression of HOXA9-HOXA13 genes to promote distal limb development (Wang et al., 2011). Characterization of these lncRNAs has often occurred through overexpression or siRNA knockdown studies. While these strategies often yield relevant results, transcriptional modulation is often not complete, especially using siRNA (or even short hairpin RNA), necessitating alternative methods.
Recently there have been a number of studies utilizing gene KO to understand lncRNA biology (Grote et al., 2013; Li et al., 2013; Sauvageau et al., 2013). At least three KO strategies have been reported: 1) Insertion of a polyA signal near the transcription start sites; 2) Insertion of a reporter gene under the control of the endogenous promoter; 3) Complete deletion of the lncRNA locus. The latter is the most dramatic and may, in addition to removing the lncRNA exons/intron structure, remove unknown regulatory elements. Insertion of a reporter gene has the advantage of being able to monitor expression of the lncRNA throughout development, however depending on which sequences are replaced, it may also carry similar drawbacks as the deletion strategy. Finally, insertion of a polyA signal near the transcription start sites likely has the least off target effects, however by not removing downstream sequences, cryptic start sites or inefficient polyA and cleavage events could cause background expression from the lncRNA locus.
Elucidating lncRNA tissue patterning by KO models
Recent efforts have begun to utilize full KO strategies to characterize additional lncRNAs including Hox encoded ones. The developmental functions of mouse Hotair were investigated by full lncRNA locus deletion in the mouse (Li et al., 2013). Loss of Hotair resulted in aberrant patterning of the skeletal system during development as was evident in abnormalities in the wrist and spine, including a switch of vertebral segment identity called homeotic transformation. Further, genome-wide characterization of the Hotair KO mouse confirmed that murine Hotair acted similarly to human HOTAIR, namely as a trans acting lncRNA controlling histone modification at specific genomic loci (Li et al., 2013). More recently, in an effort to dramatically expand the number of lncRNA KOs, Rinn and colleagues used the reporter gene approach to generate 18 separate lncRNA knockout mice (Sauvageau et al., 2013). By replacing lncRNA exonic regions with a LacZ construct, both KO and tagging was achieved. Three of the 18 lncRNAs (Fendrr, Peril, and Mdgt) showed variable penetrance and lethality. The Mdgt and Pint KO lead to abnormally low body weight and slower growth. The detailed characterization of the lncRNA Brn1b revealed its role in cortical development; specifically this lncRNA was important for the embryonic patterning in certain areas of projection neurons. By creating a large number of lncRNA KO mice and characterizing many of their functions in vivo, this study helped solidify the functional importance of lncRNAs. While thousands of lncRNAs remain to be genetically manipulated, new and more facile genome-editing tools should speed future characterization (Mali et al., 2013).
Sauvageu et al. also generated a new Fendrr KO mouse (Sauvageau et al., 2013). Under these conditions Fendrr was expressed much more widely than previously observed and most highly in the developing lung. Fendrr KO resulted in perinatal lethality, as Fendrr −/− embryoseither failed to initiate breathing or stop breathing within 5 hours of birth, neither of which was observed in WT pups. While the most striking phenotype of this KO was pulmonary, heart septal defects were also apparent even though their LacZ construct did not stain the heart for expression. This discrepancy is an important example of the possible phenotypic difference achieved by differential KO strategies such as reporter construct replacement or early polyA termination (Grote et al., 2013; Sauvageau et al., 2013). Specifically, addition of the polyA sites resulted in minimal disruption of the endogenous Fendrr locus but extremely low levels of Fendrr were still detectable (Grote and Herrmann, 2013). On the other hand, the LacZ construct replaced ~20kb of the genome resulting in a complete lack of Fendrr transcripts, however this large replacement may have removed other functional elements from the genome responsible for regulating other genes. Therefore, while both approaches confirmed loss of the lncRNA transcript, additional investigation is necessary and careful consideration of the cellular outcomes from any particular targeting strategy must be including in the experimental design.
Single cell analysis of lncRNA function
Most transcript profiling experiments of lncRNAs have employed bulk measurements, reporting results from an average of thousands or millions of cells. Recent work at the single cell level has revealed how much heterogeneity exists even within a “clonal” population of cells (Buganim et al., 2012; Shalek et al., 2013). Thus, it follows that examination of the non-coding genome and its function at the single cell level could also reveal novel modes of action. Additionally, while some studies have successfully elucidated the role lncRNAs present at a low copy number (Wang et al., 2011), the accuracy of such reports remains challenging when working with bulk populations.
Recent characterization of a lncRNA, named lincHOXA1, located in the 3′ end of the HoxA cluster by Raj and colleagues brought to light the importance of carefully examining, at the single cell level, the function of lowly expressed lncRNAs (Maamar et al., 2013). Initial analysis, at the bulk cell level, ascribed a positive correlation to the expression of lincHOXA1 and a nearby mRNA HOXA1. Surprisingly however, single cell analysis revealed an anti-correlation, and specifically a switch-like relationship was observed such that if a cell had above 10 copies of lincHOXA1, HOXA1 was repressed. Knockdown studies used targeting with both siRNA and antisense oligonucleotides (ASO, via RNase H-mediated cleavage of the target RNA). The two depletion methods differ in their capacity to reduce lincHOXA1 levels on the chromatin versus total levels, with siRNA treatment unable to efficiently lower chromatin-associated transcripts. Functionally, lincHOXA1 was found to partner with Purine-Rich Element-Binding Protein B (PURB) and exert transcriptional silencing of HOXA1. Importantly, this study highlights two key and common methodological decision points: context of cellular measurements and transcriptional knockdown strategies. In this case, bulk measurements would have masked the anti-correlated relationship between lincHOXA1 and HOXA1, which could have led to key misinterpretations. Additionally, use of siRNAs, which were effective in reducing total cellular levels of lincHOXA1, were not efficient at depleting the functional lincHOXA1 transcripts. Future work examining the molecular roles of both coding and noncoding transcripts should choose carefully the methods and context in which experiments are performed. As single cell analysis and ASO technology become more robust and widely adopted, it is likely that many unknown features of known lncRNAs may be revealed.
LncRNAs Regulation of Pluripotency
The richness of the lncRNA regulatory landscape is perhaps best exemplified in ESCs, where the noncoding transcriptome has been under intense study. The expansive number of genomic datasets, both RNA- and chromatin-based, now available in ESCs provides a rich database to characterize lncRNA function. Recent progress in understanding lncRNA control of pluripotency and dosage compensation mechanisms have revealed intimate connections between lncRNAs and chromatin state (Figure 2). Some of the most studied lncRNA binding proteins belong to chromatin modification complexes, including PRC2 and MLL, which act to suppress and activate, respectively, transcription through methylation of histone protein.
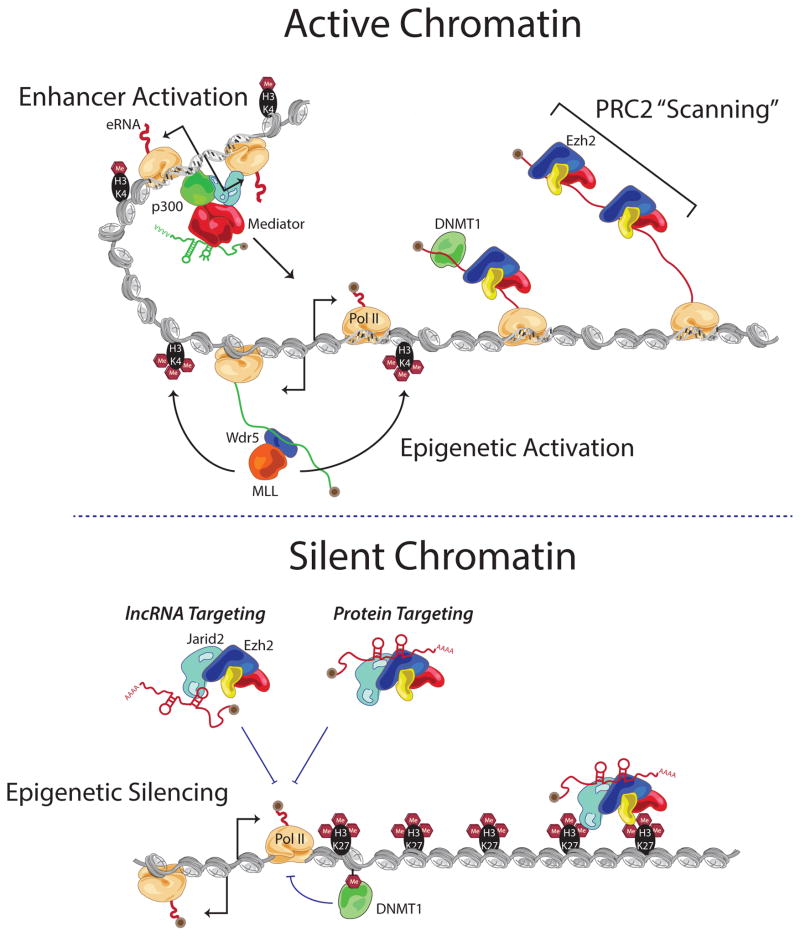
Figure 2. lncRNAs program active and silent chromatin states.
(Top) In ESCs active chromatin is achieved and maintained through multiple mechanisms. Cis acting lncRNAs can recruit the MLL/WDR5 complex to deposit H3K4me3 at promoters. Enhancer regions can transcribe enhancer RNAs (eRNAs); some enhancer-like RNAs bring Mediator to promoters to contribute to gene activation. Additionally, through interactions with the nascent transcribed RNA, canonical silencing factors such as PRC2 and DNMT1 are titrated away from active chromatin. (Bottom) Chromatin also employs many lncRNA-based mechanisms to stay silent. Ezh2 and JAIRD2 (subunits of PRC2) may bind lncRNAs to facilitate specific chromatin targeting or to enhance PRC2 complex assembly and stability. Additionally, when nascent RNA production is low, DNMT1 can interact with the chromatin and act to silence through DNA methylation.
Transition between cell states
Characterization of the transcriptome of ESCs has revealed many lncRNAs that participate in the regulation of the pluripotent state (Guttman et al., 2012; 2009; Lin et al., 2014; Ng et al., 2012; Sheik Mohamed et al., 2010). Through a comprehensive “perturb-and-measure” strategy, Guttman et al. showed that dozens of lncRNAs are required for the setting the gene expression patterns of mouse ESCs or the first step of differentiation toward different germ layers (Guttman et al., 2012). A subset of these lncRNAs bound one or more chromatin modification complexes, including readers, writers, or erasers of repressive histone modifications.
In contrast, the “regulator of reprogramming” lncRNA, lincRNA-RoR was identified as an important factor for the reprogramming process as its depletion or over expression lead to a lower or higher efficiency of reprogramming fibroblasts to iPSCs, respectively (Loewer et al., 2010). However only recently was the molecular mechanism investigated (Wang et al., 2013). Pull down experiments with lincRNA-RoR specifically isolated miR-145-5p, 181a-5p, and 99b-3p, as well as the miR targeting protein Argonaute2 (Ago2). These miRs have been previously shown to regulate core pluripotency factors such as Pou5f1, Sox2, and NANOG, suggesting that lincRNA-RoR might act as a ceRNA. Indeed, functional assays revealed that lincRNA-RoR regulated the mature form of miR-145, characteristic of a ceRNA. Loss of lncRNA-RoR caused human ESCs (hESCs) to differentiate towards mesoderm and ectoderm, while overexpression conferred a differentiation defect. Additionally, in the context of cancer, a rapidly proliferative state similar to ESCs, lincRNA-RoR was recently shown to act in a regulatory loop suppressing the expression of the tumor suppressor p53 (Zhang et al., 2013). Together, this characterization of lincRNA-RoR further advances the idea that each lncRNA may control many pathways in different cellular contexts including tumor growth and core pluripotency gene network utilizing a ceRNA mechanism.
Activation of the epigenome with lncRNAs
To date, the vast majority of lncRNAs have annotated functions in repressive complexes, with only a few examples of activating or enhancing lncRNAs (Wang et al., 2011; Zhang et al., 2009). HOTTIP, named due to its location at the distal “tip” of the HOXA gene cluster enforces an active chromatin state by recruiting the WDR5 subunit of the MLL complex (Wang et al., 2011) (Figure 2). The HOTTIP locus comes into spatial proximity with its target genes, and all the while the expression level of HOTTIP remains near one copy per cell (Wang et al., 2011). The low copy number of HOTTIP ensures that HOTTIP acts precisely in cis on target genes defined by proximity in three dimensional nuclear space, but not broadly on other genes. More recently, biochemical characterization of the interaction between WDR5 and HOTTIP revealed a specific RNA binding pocket of WDR5 and that RNA binding could stabilize chromatin-associated WDR5(Yang et al., 2014). This finding suggested that in vivo, not only the localization, but also the half-life of WDR5 could be modulated by HOTTIP. Given that WDR5/MLL acts at many genomic loci, RNA immunoprecipitation-seq (RIP-seq) was used to identify over 1400 WDR5 target RNAs, including many coding and noncoding RNAs. A lncRNA binding pocket on WDR5 was discovered, and a specific mutation of the RNA binding pocket selectively abrogated RNA binding but no other functions of the WDR5-MLL complex (Yang et al., 2014). This selective WDR5 mutant revealed that RNA binding is important for the temporal stability of the active chromatin mark H3K4me3 over time and maintenance of ESC pluripotency. These studies suggest a generalizable mechanism for functional MLL/WDR5-RNA interaction. Specifically, HOTTIP acts in cis, and is expressed at far too low levels per cell to globally modulate the MLL/WDR5 chromatin localization. The RIP-seq of WDR5 in mESCs (which do not express HOTTIP) revealed that more than one thousand cellular RNAs could interact with and may modulate the chromatin modification complex. Because WDR5 targets over 10,000 genomic sites (Ang et al., 2011), whether the three-dimensional organization of the genome facilitates lncRNA co-regulation of the mESC self-renewal program remains to be addressed in future studies.
Epigenetic repression through lncRNA-PRC2 interactions
Unlike activating chromatin complexes, chromatin-modifying complexes that repress transcription have been more extensively studied in the context of lncRNA interactions, resulting in a richer set of known interactions. The focus of many of these studies has been the PRC2 complex, responsible for depositing H3K27me3, which plays roles pluripotency, differentiation, XCI, and diseases such as cancer (Margueron and Reinberg, 2011). An initial survey of the RNA-interactome of Ezh2 yielded more than 9,000 target RNAs using RIP-seq in mESCs (Zhao et al., 2010). Recently, two studies have revisited this observation to further clarify the interplay between RNA and PRC2 (Davidovich et al., 2013; Kaneko et al., 2013b) (Figure 2). Biochemical interaction and photoactivated RNA-crosslinking experiments suggest that Ezh2 can interact with numerous RNAs, including the 5′ end of nascent RNAs that are actively transcribed. The apparently specific interactions of PRC2 with several lncRNAs in lysate and in vivo are not recapitulated in vitro by the core PRC2 complex alone. The promiscuous RNA binding of Ezh2 may be modulated by additional proteins, such as Jarid2 and others, to facilitate higher degrees of specificity in vivo ((Davidovich et al., 2013) and see below). Moreover, Ezh2 may scan the genome surveying the transcriptional status of its targets. Actively transcribed regions may continually push Ezh2 away via their elongating mRNAs, while silent regions or those stably bound by lncRNAs (generated in trans) can be silenced. This proposed mechanism reinforces the status quo of gene transcription and silencing, and is consistent with the known genetic role of Polycomb group proteins in chromatin state maintenance.
A similar RNA surveillance mechanism is also employed by the DNA methylase DNMT1 that interacts with many cellular transcripts, including the nonpolyadenylated extra codingCEBPA (ecCEBPA) lncRNA. The ecCEBPA lncRNA adopts a characteristic stem-loop structure critical for interaction with DNMT1 and, when transcribed, acts to shield the CEBPA locus from DNA methylation (Di Ruscio et al., 2013) (Figure 2). These two examples provide evidence that cells employ RNAs to modulate the deposition of repressive epigenetic marks on a genome-wide manner. Nonetheless, recognition of the potentially broad interactions between RNA and PRC2 highlights the need for high-quality in vivo controls and validation of RNA-protein interactions. Methodological choice is critical as each assay type has its own strengths and weaknesses, which will impact the results obtained and conclusions drawn.
While PCR2 operates in a wide range of cell types, certain subunits such as JARID2, are specifically expressed and partner with PCR2 in ESCs and certain dividing cells, including cancer cells (Pasini et al., 2010; Peng et al., 2009; Shen et al., 2009). These initial studies established JARID2’s capacity to regulate the stability of the PRC2 complex as well as its enzymatic activity (Figure 2). Further expanding the cellular functions of JARID2, in vitro RNA binding assays and in vivo PAR-CLIP suggest that JARID2 directly interacts with cellular RNAs (Kaneko et al., 2013a). JARID2 and Ezh2 reproducibly crosslinked to 106 and 165 lncRNAs, respectively, and 53 lncRNAs were commonly bound. The MEG3 lncRNA was bound by both subunits of PRC2, however the RNA binding region (RBR) of JARID2 provided the largest contribution of MEG3 binding to PRC2. Additionally, cellular levels of MEG3 contribute to PRC2’s chromatin association, as low expression of MEG3 resulted in the loss of PRC2 subunits from specific loci leading to derepression of the nearby genes. Finally, the in vitro interaction between JARID2 and Ezh2 was facilitated by HOTAIR and MEG3, and Ezh2’s chromatin association was shown to be partially dependent on JARID2’s RNA-binding domain. Thus, JARID2, an ESC-specific subunit of PRC2, appears to modulate the localization of PRC2, and thus the chromatin state, in an RNA-dependent manner. While this study offers an additional layer of regulation with respect to the Polycomb complex, little is known about the other RNA targets of JARID2, which may significantly contribute to its cellular function. Additionally, studies to rigorously interrogate the enzymatic properties of the PRC2 complex inside cells with and without its RNA partners will be very informative.
A lncRNA Network to Control Dosage Compensation
Dosage compensation of genes encoded on the X chromosome is accomplished by divergent strategies in different species; however, the use of lncRNAs is a common feature. In Drosophila, dosage compensation is achieved by precisely upgregulating the X-chromosome in males by 2-fold (Lucchesi et al., 2005). A desire to understand how dosage compensation operates fueled the development of ChIRP and CHART, genomic tools that map the chromatin-association of lncRNAs (Chu et al., 2011; Simon et al., 2011). Initially, ChIRP and CHART were applied to the Drosophila roX2 lncRNA. which provided evidence that roX2 co-occupies genomic loci with the known dosage compensation protein factors on the X chromosome. Importantly, they proved that mapping the genomic locations of lncRNAs can generate novel hypotheses for functions of lncRNAs. While studies in Drosophila and other model systems have provided key insights into mechanisms of dosage compensation, we will focus on recent investigations conducted in mammalian cells.
Xist Spreading
In mammals, the strategy for dosage compensation is reverse from Drosophila: female cells selectively repress one entire chromosome by upregulating the repressive lncRNA Xist (Lee, 2012). Xist is transcribed from the X-inactivation Center (XIC) and is responsible for physically coating and silencing the X-chromosome targeted for the Barr body (the Inactive X, Xi). Another lncRNA, Tsix, is transcribed from the active X chromosome (Xa) and enforces silencing of Xist (Lee, 2012). These two lncRNAs, together with others described below, form a complex RNA-protein regulatory network that controls X chromosome dosage compensation in mammals.
Traditional techniques such as immunofluorescence (IF) and RNA fluorescence in situ hybridization (FISH) have been widely applied to study X-chromosome Inactivation (XCI) and have arrived at a consensus mechanism: elevated Xist expression from the future Xi leads to a cloud-like coating of Xist on Xi and finally epigenetic silencing and chromatin compaction. While informative, IF and RNA-FISH studies had resolution limitations, and as was true for the roX2 RNA, specifically mapping the genomic locations of Xist held the promise of answering mechanistic features of its function. Recently, application of CHART and the development and application of RAP (a method with similar principals to ChIRP and CHART) to the Xist lncRNA defined its precise chromatin-association (Engreitz et al., 2013; Simon et al., 2014). Together the studies revealed that the initiation of Xist spreading occurs from the Xist locus to distinct sites across the X-chromosome that are not directly adjacent to its locus. These regions are highly accessible by DNaseI footprinting and contain many genes that are actively transcribed prior to silencing. Once Xist is deposited on these early sites, it proceeds to spread and coat the rest of the chromosome to fully silence all but a few genes that escape XCI. It is proposed that the initial disposition process is mediated through higher-order chromatin architecture (Engreitz et al., 2013), however experimental design differences between the two studies described above make it difficult to directly compare the chromatin conformation results measured. While further investigation is clearly needed to solidify and refine these results, using high-resolution genomic tools (ChIRP, CHART, or RAP) can provide critical insight into lncRNA-controlled systems previously hidden from view.
Mechanisms of Xist Regulation
Intense study of the Xist regulatory network has uncovered many novel lncRNAs in and around the XIC, often illuminating novel mechanistic concepts for how lncRNAs function. Within the lncRNA network that controls Xist, Tsix and Jpx oppose each other’s function by repressing or activating, respectively, the transcription of Xist (Lee, 2000; Tian et al., 2010). Recently, additional characterization of the Jpx pathway revealed an unexpected interplay between the lncRNA Jpx and CCCTC-binding factor (CTCF), a major DNA binding protein involved in higher order chromosomal folding and interactions (Sun et al., 2013). During female mESC differentiation CTCF is lost from the Xist locus, therefore allowing allele specific Xist upregulation. Molecular characterization of this regulatory loop revealed that CTCF directly binds Jpx and this interaction can titrate CTCF from its DNA targets. Within the conceptual framework of dosage compensation, this puts Jpx and CTCF as central players in the balance between activation and silencing of the X chromosome. Cellular levels of Jpx, as partially determined by the number of X chromosomes, would control the ability of CTCF to bind and inhibit transcription at the Xist locus only under conditions when XCI is required. Another recent study more globally characterized the RNA-binding capacity of CTCF and found a multitude of RNA targets, including Wrap53, a lncRNA that controls the induction of the tumor suppressor p53 upon DNA-damage (Saldaña-Meyer et al., 2014). Interestingly, biochemical characterization of CTCF’s protein domains revealed that the RBR and RNA promoted multimerization of CTCF.
While Xist is modulated by CTCF localization and the spatiotemporal deposition of Xist has been initially defined through CHART and RAP, how Xist interacts with protein effectors of XCI remains poorly understood. The repeat A (RepA) domain of Xist has been reported to mediate the interaction with the PRC2 complex (Zhao et al., 2008). Recent characterization of the JARID2 subunit of PRC2 also implicates it in functionally interacting with Xist (da Rocha et al., 2014). Specifically, the authors observed JARID2 and other PCR2 subunits co-occupying genomic regions on the Xi, and a requirement for JARID2 for the deposition of H3K27me3. Further, Xist deletion experiments defined the RepB and RepF regions within the RNA as responsible for JARID2 targeting to the Xi. Interesting, this function was not depended on its previously identified RBR (Kaneko et al., 2013a) suggesting that JARID2 is a multifunctioning RNA binding protein, that mediates the association of PRC2 to the Xi through Xist. These examples suggest that within the context of XCI, as well as during other critical cellular decisions, lncRNAs (such as Xist) can act to modulate chromosome architecture and chromatin modification patterns.
XCI as a marker of reprogramming
The ability to transform differentiated cells back into pluripotent cells holds tremendous possibilities for regenerative medicine, but many hurdles still remain before this technology is fully matured (Alvarado and Yamanaka, 2014). Because biallelic X activation is a key epigenetic marker of pluripotency, the status of Xist and Xist-mediated gene silencing (or lack thereof) can be exploited to phenotype ESCs and iPSCs (Figure 3). Careful analysis of human iPSCs derived from female cells revealed that many carried an Xi, failing to undergo X-chromosome reactivation (XCR) and are epigenetically dynamic, suggesting that the derivation of hiPSCs may not result in pristinely pluripotent cells as desired (Tchieu et al., 2010). A subsequent study used X-inactivation markers to segregate populations of hiPSCs and found that female derived iPSCs are likely to be less stable in culture than male derived cells (Anguera et al., 2012). Indeed, erosion of dosage compensation has been observed in female hiPSCs over time in culture, significantly impacting the potential use of these cells for modeling x-linked disease (Mekhoubad et al., 2012). More recent work characterized XCR in the context of iPSC reprogramming and found PRDM14, involved in the ESC pluripotency network, controls Xist silencing (Payer et al., 2013). With the help of Tsix, PRDM14 represses Xist activators (Rnf12 and Jpx) and the Xist locus itself by recruiting PRC2, placing PRDM14 expression as a marker for XCR. Work from Heard and colleagues also explored how Xist status can directly regulate ESC differentiation, notably within the framework of the primed/metastable and ground/naïve states, with the latter representing a more primordial state of pluripotency. Schultz et al. reported that an X-linked inhibitor of MAPK signaling couples the status of X chromosomes to ESC differentiation. In the ground state where both X chromosomes are active, MAPK is inhibited concomitantly with other molecular changes that block ESC differentiation (Schulz et al., 2014). Upon X chromosome inactivation in the primed state, the relief of MAPK inhibition leads to high MAPK signaling and the capacity to proceed with differentiation. Therefore the characteristic expression of Xist and X-silencing genes provides new ways to evaluate the efficiency and ultimately control of reprograming during iPSC generation. Combining traditional pluripotency markers with new markers like X-inactivation will be critical to achieve the standardization and consistency necessary for clinical application of iPSC technologies.
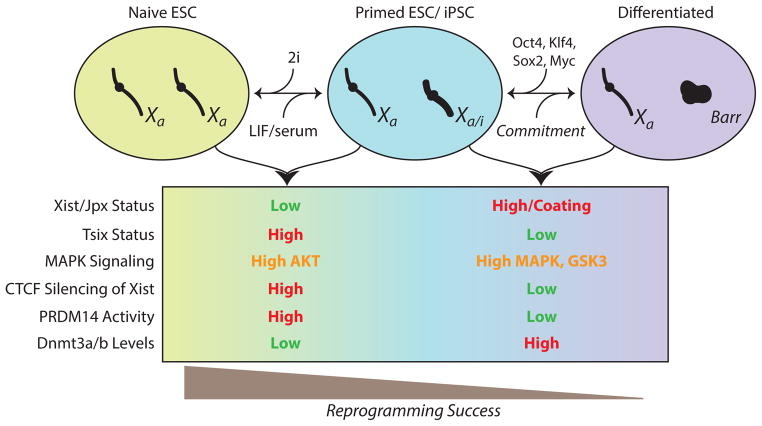
Figure 3. lncRNAs mark ESC state and reprogramming success.
X-chromosome inactivation (XCI) is a key step in the commitment of ESCs to differentiated cell types. The network on lncRNAs, signaling pathways, and protein effectors that control XCI are depicted. These features can distinguish the stemness of different ESC states and iPSC quality.
Lessons and Future Prospects
While the myriad examples to date highlight the functions of a small fraction of known lncRNAs, they illustrate the principle that lncRNAs are intimately involved in the specification, self-renewal, differentiation, and patterning of stem cells and their differentiated progenies. It is reasonable to anticipate that similar principles will be uncovered in many additional organ systems and cell types. A frequently asked question is “Why RNA?” LncRNAs exhibit exquisite cell type and organ-specific expression patterns, in fact, to a greater extent than mRNAs. Evolution has likely taken advantage of this fertile soil of cell-type and state-specific transcription to evolve regulatory functions. Thus, one area of future investigation should focus on the regulation of lncRNA expression—what exactly makes them different and endows them with such state-specific expression? A second challenge for the field is the need to predict the functions of lncRNAs from primary sequence. Finally, understanding how the structure of lncRNAs guides their function remains largely unexplored. As has been true for protein biochemistry, understanding the physical conformations lncRNAs adopt inside cells will undoubtedly uncover novel functional domains and structural elements responsible for their cellular activities.
Acknowledgments
We apologize to colleagues whose work was not discussed due to space constraints. We thank M. Wernig, E. Heard, R.C. Spitale, A.J. Rubin and member of the Chang lab for comments. Supported by NIH Medical Scientist Training Program (R.A.F.), NIH and California Institute for Regenerative Medicine (H.Y.C.), H.Y.C. is an Early Career Scientist of the Howard Hughes Medical Institute.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Alvarado AS, Yamanaka S. Rethinking Differentiation: Stem Cells, Regeneration, and Plasticity. Cell. 2014;157:110–119. doi: 10.1016/j.cell.2014.02.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ang YS, Tsai SY, Lee DF, Monk J, Su J, Ratnakumar K, Ding J, Ge Y, Darr H, Chang B, et al. Wdr5 Mediates Self-Renewal and Reprogramming via the Embryonic Stem Cell Core Transcriptional Network. Cell. 2011;145:183–197. doi: 10.1016/j.cell.2011.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anguera MC, Sadreyev R, Zhang Z, Szanto A, Payer B, Sheridan SD, Kwok S, Haggarty SJ, Sur M, Alvarez J, et al. Molecular Signatures of Human Induced Pluripotent Stem Cells Highlight Sex Differences and Cancer Genes. Stem Cell. 2012;11:75–90. doi: 10.1016/j.stem.2012.03.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Batista PJ, Chang HY. Long noncoding RNAs: cellular address codes in development and disease. Cell. 2013;152:1298–1307. doi: 10.1016/j.cell.2013.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berghoff EG, Clark MF, Chen S, Cajigas I, Leib DE, Kohtz JD. Evf2 (Dlx6as) lncRNA regulates ultraconserved enhancer methylation and the differential transcriptional control of adjacent genes. Development. 2013;140:4407–4416. doi: 10.1242/dev.099390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bertani S, Sauer S, Bolotin E, Sauer F. Short Article. Molecular Cell. 2011;43:1040–1046. doi: 10.1016/j.molcel.2011.08.019. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- Bond AM, VanGompel MJW, Sametsky EA, Clark MF, Savage JC, Disterhoft JF, Kohtz JD. Balanced gene regulation by an embryonic brain ncRNA is critical for adult hippocampal GABA circuitry. Nat Neurosci. 2009;12:1020–1027. doi: 10.1038/nn.2371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bosia C, Pagnani A, Zecchina R. Modelling Competing Endogenous RNA Networks. PLoS ONE. 2013;8:e66609. doi: 10.1371/journal.pone.0066609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buganim Y, Faddah DA, Cheng AW, Itskovich E, Markoulaki S, Ganz K, Klemm SL, van Oudenaarden A, Jaenisch R. Single-Cell Expression Analyses during Cellular Reprogramming Reveal an Early Stochastic and a Late Hierarchic Phase. Cell. 2012;150:1209–1222. doi: 10.1016/j.cell.2012.08.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burridge PW, Keller G, Gold JD, Wu JC. Production of De Novo Cardiomyocytes: Human Pluripotent Stem Cell Differentiation and Direct Reprogramming. Cell Stem Cell. 2012;10:16–28. doi: 10.1016/j.stem.2011.12.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JL. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes & Development. 2011;25:1915–1927. doi: 10.1101/gad.17446611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cesana M, Cacchiarelli D, Legnini I, Santini T, Sthandier O, Chinappi M, Tramontano A, Bozzoni I. A Long Noncoding RNA Controls Muscle Differentiation by Functioning as a Competing Endogenous RNA. Cell. 2011;147:358–369. doi: 10.1016/j.cell.2011.09.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu C, Qu K, Zhong FL, Artandi SE, Chang HY. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Molecular Cell. 2011;44:667–678. doi: 10.1016/j.molcel.2011.08.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- da Rocha ST, Boeva V, Escamilla-Del-Arenal M, Ancelin K, Granier C, Matias NR, Sanulli S, Chow J, Schulz E, Picard C, et al. Jarid2 Is Implicated in the Initial Xist-Induced Targeting of PRC2 to the Inactive X Chromosome. Molecular Cell. 2014;53:301–316. doi: 10.1016/j.molcel.2014.01.002. [DOI] [PubMed] [Google Scholar]
- Davidovich C, Zheng L, Goodrich KJ, Cech TR. Promiscuous RNA binding by Polycomb repressive complex 2. Nature Structural & Molecular Biology. 2013;20:1250–1257. doi: 10.1038/nsmb.2679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng Z, Cao P, Wan MM, Sui G. Yin Yang 1: a multifaceted protein beyond a transcription factor. Transcription. 2010;1:81–84. doi: 10.4161/trns.1.2.12375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Research. 2012;22:1775–1789. doi: 10.1101/gr.132159.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Ruscio A, Ebralidze AK, Benoukraf T, Amabile G, Goff LA, Terragni J, Figueroa ME, De Figueiredo Pontes LL, Alberich-Jorda M, Zhang P, et al. DNMT1-interacting RNAs block gene-specific DNA methylation. Nature. 2013;503:371–376. doi: 10.1038/nature12598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engreitz JM, Pandya-Jones A, McDonel P, Shishkin A, Sirokman K, Surka C, Kadri S, Xing J, Goren A, Lander ES, et al. The Xist lncRNA Exploits Three-Dimensional Genome Architecture to Spread Across the X Chromosome. Science. 2013;341:1237973–1237973. doi: 10.1126/science.1237973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grote P, Herrmann BG. The long non-coding RNA Fendrr links epigenetic control mechanisms to gene regulatory networks in mammalian embryogenesis. RNA Biol. 2013 doi: 10.4161/rna.26165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grote P, Wittler L, Hendrix D, Koch F, Währisch S, Beisaw A, Macura K, Bläss G, Kellis M, Werber M, et al. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev Cell. 2013;24:206–214. doi: 10.1016/j.devcel.2012.12.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;457:223–227. doi: 10.1038/nature07672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guttman M, Donaghey J, Carey BW, Garber M, Grenier JK, Munson G, Young G, Lucas AB, Ach R, Bruhn L, et al. linc RNAs act in the circuitry controlling pluripotency and differentiation. Nature. 2012;477:295–300. doi: 10.1038/nature10398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaneko S, Bonasio R, Saldaña-Meyer R, Yoshida T, Son J, Nishino K, Umezawa A, Reinberg D. Interactions between JARID2 and Noncoding RNAs Regulate PRC2 Recruitment to Chromatin. Molecular Cell. 2013a:1–11. doi: 10.1016/j.molcel.2013.11.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaneko S, Son J, Shen SS, Reinberg D, Bonasio R. PRC2 binds active promoters and contacts nascent RNAs in embryonic stem cells. Nature Structural & Molecular Biology. 2013b;20:1258–1264. doi: 10.1038/nsmb.2700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klattenhoff CA, Scheuermann JC, Surface LE, Bradley RK, Fields PA, Steinhauser ML, Ding H, Butty VL, Torrey L, Haas S, et al. Braveheart, a Long Noncoding RNA Required for Cardiovascular Lineage Commitment. Cell. 2013;152:570–583. doi: 10.1016/j.cell.2013.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kretz M, Webster DE, Flockhart RJ, Lee CS, Zehnder A, Lopez-Pajares V, Qu K, Zheng GXY, Chow J, Kim GE, et al. Suppression of progenitor differentiation requires the long noncoding RNA ANCR. Genes & Development. 2012;26:338–343. doi: 10.1101/gad.182121.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kretz M, Siprashvili Z, Chu C, Webster DE, Zehnder A, Qu K, Lee CS, Flockhart RJ, Groff AF, Chow J, et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature. 2013;493:231–235. doi: 10.1038/nature11661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee JT. Disruption of imprinted X inactivation by parent-of-origin effects at Tsix. Cell. 2000;103:17–27. doi: 10.1016/s0092-8674(00)00101-x. [DOI] [PubMed] [Google Scholar]
- Lee JT. Epigenetic regulation by long noncoding RNAs. Science. 2012;338:1435–1439. doi: 10.1126/science.1231776. [DOI] [PubMed] [Google Scholar]
- Legnini I, Morlando M, Mangiavacchi A, Fatica A, Bozzoni I. Short Article. Molecular Cell. 2014;53:506–514. doi: 10.1016/j.molcel.2013.12.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li L, Liu B, Wapinski OL, Tsai MC, Qu K, Zhang J, Carlson JC, Lin M, Fang F, Gupta RA, et al. Targeted disruption of Hotair leads to homeotic transformation and gene derepression. Cell Reports. 2013;5:3–12. doi: 10.1016/j.celrep.2013.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin N, Chang K-Y, Li Z, Gates K, Rana ZA, Dang J, Zhang D, Han T, Yang C-S, Cunningham TJ, et al. An Evolutionarily Conserved Long Noncoding RNA TUNA Controls Pluripotency and Neural Lineage Commitment. Molecular Cell. 2014 doi: 10.1016/j.molcel.2014.01.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loewer S, Cabili MN, Guttman M, Loh YH, Thomas K, Park IH, Garber M, Curran M, Onder T, Agarwal S, et al. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nature Genetics. 2010;42:1113–1117. doi: 10.1038/ng.710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu L, Sun K, Chen X, Zhao Y, Wang L, Zhou L, Sun H, Wang H. Genome-wide survey by ChIP-seq reveals YY1 regulation of lincRNAs in skeletal myogenesis. Embo J. 2013;32:2575–2588. doi: 10.1038/emboj.2013.182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu L, Zhou L, Chen EZ, Sun K, Jiang P, Wang L, Su X, Sun H, Wang H. A Novel YY1-miR-1 regulatory circuit in skeletal myogenesis revealed by genome-wide prediction of YY1-miRNA network. PLoS ONE. 2012;7:e27596–e27596. doi: 10.1371/journal.pone.0027596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lucchesi JC, Kelly WG, Panning B. Chromatin remodeling in dosage compensation. Annu Rev Genet. 2005 doi: 10.1146/annurev.genet.39.073003.094210. [DOI] [PubMed] [Google Scholar]
- Maamar H, Cabili MN, Rinn J, Raj A. linc-HOXA1 is a noncoding RNA that represses Hoxa1 transcription in cis. Genes & Development. 2013;27:1260–1271. doi: 10.1101/gad.217018.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mali P, Esvelt KM, Church GM. Cas9 as a versatile tool for engineering biology. Nature Methods. 2013 doi: 10.1038/nmeth.2649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Margueron R, Reinberg D. The Polycomb complex PRC2 and its mark in life. Nature. 2011;469:343–349. doi: 10.1038/nature09784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehler MF, Mattick JS. Noncoding RNAs and RNA editing in brain development, functional diversification, and neurological disease. Physiol Rev. 2007;87:799–823. doi: 10.1152/physrev.00036.2006. [DOI] [PubMed] [Google Scholar]
- Mekhoubad S, Bock C, de Boer AS, Kiskinis E, Meissner A, Eggan K. Erosion of Dosage Compensation Impacts Human iPSC Disease Modeling. Stem Cell. 2012;10:595–609. doi: 10.1016/j.stem.2012.02.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mercer TR, Dinger ME, Mariani J, Kosik KS, Mehler MF, Mattick JS. Noncoding RNAs in Long-Term Memory Formation. Neuroscientist. 2008;14:434–445. doi: 10.1177/1073858408319187. [DOI] [PubMed] [Google Scholar]
- Mercer TR, Gerhardt DJ, Dinger ME, Crawford J, Trapnell C, Jeddeloh JA, Mattick JS, Rinn JL. targeted rNA sequencing reveals the deep complexity of the human transcriptome. Nat Biotechnol. 2011;30:99–104. doi: 10.1038/nbt.2024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mukherji S, Ebert MS, Zheng GXY, Tsang JS, Sharp PA, van Oudenaarden A. MicroRNAs can generate thresholds in target gene expression. Nature Genetics. 2011;43:854–859. doi: 10.1038/ng.905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ng SY, Johnson R, Stanton LW. Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. Embo J. 2012;31:522–533. doi: 10.1038/emboj.2011.459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Onoguchi M, Hirabayashi Y, Koseki H, Gotoh Y. A noncoding RNA regulates the neurogenin1 gene locus during mouse neocortical development. 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pasini D, Cloos PAC, Walfridsson J, Olsson L, Bukowski JP, Johansen JV, Bak M, Tommerup N, Rappsilber J, Helin K. JARID2 regulates binding of the Polycomb repressive complex 2 to target genes in ES cells. Nature. 2010;464:306–310. doi: 10.1038/nature08788. [DOI] [PubMed] [Google Scholar]
- Payer B, Rosenberg M, Yamaji M, Yabuta Y, Koyanagi-Aoi M, Hayashi K, Yamanaka S, Saitou M, Lee JT. Tsix RNA and the Germline Factor, PRDM14,Link X Reactivation and Stem Cell Reprogramming. Molecular Cell. 2013;52:805–818. doi: 10.1016/j.molcel.2013.10.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peng JC, Valouev A, Swigut T, Zhang J, Zhao Y, Sidow A, Wysocka J. Jarid2/Jumonji coordinates control of PRC2 enzymatic activity and target gene occupancy in pluripotent cells. Cell. 2009;139:1290–1302. doi: 10.1016/j.cell.2009.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramos AD, Diaz A, Nellore A, Delgado RN, Park KY, Gonzales-Roybal G, Oldham MC, Song JS, Lim DA. Integration of genome-wide approaches identifies lncRNAs of adult neural stem cells and their progeny in vivo. Cell Stem Cell. 2013;12:616–628. doi: 10.1016/j.stem.2013.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rapicavoli NA, Poth EM, Zhu H, Blackshaw S. The long noncoding RNA Six3OS acts in trans to regulate retinal development by modulating Six3 activity. Neural Dev. 2011;6:32–32. doi: 10.1186/1749-8104-6-32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinn JL, Chang HY. Genome Regulation by Long Noncoding RNAs. Annu Rev Biochem. 2012;81:145–166. doi: 10.1146/annurev-biochem-051410-092902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007;129:1311–1323. doi: 10.1016/j.cell.2007.05.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saldaña-Meyer R, González-Buendía E, Guerrero G, Narendra V, Bonasio R, Recillas-Targa F, Reinberg D. CTCF regulates the human p53 gene through direct interaction with its natural antisense transcript, Wrap53. Genes & Development. 2014;28:723–734. doi: 10.1101/gad.236869.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sauvageau M, Goff LA, Lodato S, Bonev B, Groff AF, Gerhardinger C, Sanchez-Gomez DB, Hacisuleyman E, Li E, Spence M, et al. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. Elife. 2013;2:e01749–e01749. doi: 10.7554/eLife.01749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schulz EG, Meisig J, Nakamura T, Okamoto I, Sieber A, Picard C, Borensztein M, Saitou M, Blüthgen N, Heard E. The Two Active X Chromosomes in Female ESCs Block Exit from the Pluripotent Stateby Modulating the ESC Signaling Network. Stem Cell. 2014;14:203–216. doi: 10.1016/j.stem.2013.11.022. [DOI] [PubMed] [Google Scholar]
- Sen GL, Reuter JA, Webster DE, Zhu L, Khavari PA. DNMT1 maintains progenitor function in self-renewing somatic tissue. Nature. 2010;463:563–567. doi: 10.1038/nature08683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shalek AK, Satija R, Adiconis X, Gertner RS, Gaublomme JT, Raychowdhury R, Schwartz S, Yosef N, Malboeuf C, Lu D, et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature. 2013;498:236–240. doi: 10.1038/nature12172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheik Mohamed J, Gaughwin PM, Lim B, Robson P, Lipovich L. Conserved long noncoding RNAs transcriptionally regulated by Oct4 and Nanog modulate pluripotency in mouse embryonic stem cells. Rna. 2010;16:324–337. doi: 10.1261/rna.1441510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen X, Kim W, Fujiwara Y, Simon MD, Liu Y, Mysliwiec MR, Yuan GC, Lee Y, Orkin SH. Jumonji modulates polycomb activity and self-renewal versus differentiation of stem cells. Cell. 2009;139:1303–1314. doi: 10.1016/j.cell.2009.12.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simon MD, Pinter SF, Fang R, Sarma K, Rutenberg-Schoenberg M, Bowman SK, Kesner BA, Maier VK, Kingston RE, Lee JT. High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation. Nature. 2014;504:465–469. doi: 10.1038/nature12719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simon MD, Wang CI, Kharchenko PV, West JA, Chapman BA, Alekseyenko AA, Borowsky ML, Kuroda MI, Kingston RE. The genomic binding sites of a noncoding RNA. Proc Natl Acad Sci USa. 2011;108:20497–20502. doi: 10.1073/pnas.1113536108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun S, Del Rosario BC, Szanto A, Ogawa Y, Jeon Y, Lee JT. Jpx RNA Activates Xist by Evicting CTCF. Cell. 2013;153:1537–1551. doi: 10.1016/j.cell.2013.05.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505:344–352. doi: 10.1038/nature12986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tchieu J, Kuoy E, Chin MH, Trinh H, Patterson M, Sherman SP, Aimiuwu O, Lindgren A, Hakimian S, Zack JA, et al. Female Human iPSCs Retain an Inactive X Chromosome. Stem Cell. 2010;7:329–342. doi: 10.1016/j.stem.2010.06.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian D, Sun S, Lee JT. The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell. 2010;143:390–403. doi: 10.1016/j.cell.2010.09.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Truong AB, Kretz M, Ridky TW, Kimmel R, Khavari PA. p63 regulates proliferation and differentiation of developmentally mature keratinocytes. Genes & Development. 2006;20:3185–3197. doi: 10.1101/gad.1463206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, Shi Y, Segal E, Chang HY. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329:689–693. doi: 10.1126/science.1192002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ulitsky I, Shkumatava A, Jan CH, Sive H, Bartel DP. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell. 2011;147:1537–1550. doi: 10.1016/j.cell.2011.11.055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang KC, Chang HY. Molecular Mechanisms of Long Noncoding RNAs. Molecular Cell. 2011;43:904–914. doi: 10.1016/j.molcel.2011.08.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, Lajoie BR, Protacio A, Flynn RA, Gupta RA, et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature. 2011;472:120–124. doi: 10.1038/nature09819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Xu Z, Jiang J, Xu C, Kang J, Xiao L, Wu M, Xiong J, Guo X, Liu H. Endogenous miRNA Sponge lincRNA-RoR Regulates Oct4, Nanog, and Sox2in Human Embryonic Stem Cell Self-Renewal. Dev Cell. 2013;25:69–80. doi: 10.1016/j.devcel.2013.03.002. [DOI] [PubMed] [Google Scholar]
- Yang YW, Flynn RA, Chen Y, Qu K, Wan B, Wang KC, Lei M, Chang HY. Essential role of lncRNA binding for WDR5 maintenance of active chromatin and embryonic stem cell pluripotency. Elife. 2014;3:e02046–e02046. doi: 10.7554/eLife.02046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang A, Zhou N, Huang J, Liu Q, Fukuda K, Ma D, Lu Z, Bai C, Watabe K, Mo YY. The human long non-coding RNA-RoR is a p53 repressor in response to DNA damage. Cell Res. 2013;23:340–350. doi: 10.1038/cr.2012.164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Lian Z, Padden C, Gerstein MB, Rozowsky J, Snyder M, Gingeras TR, Kapranov P, Weissman SM, Newburger PE. A myelopoiesis-associated regulatory intergenic noncoding RNA transcript within the human HOXA cluster. Blood. 2009;113:2526–2534. doi: 10.1182/blood-2008-06-162164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science. 2008 doi: 10.1126/science.1163045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao J, Ohsumi TK, Kung JT, Ogawa Y, Grau DJ, Sarma K, Song JJ, Kingston RE, Borowsky M, Lee JT. Genome-wide identification of polycomb-associated RNAs by RIP-seq. Molecular Cell. 2010;40:939–953. doi: 10.1016/j.molcel.2010.12.011. [DOI] [PMC free article] [PubMed] [Google Scholar]