Abstract
PURPOSE OF THE REVIEW
Anaplastic Large Cell Lymphomas (ALCLs) are rare entities whose tumorigenic events have only been found in well-defined subsets. The categorization of additional molecular fingerprints is needed to advance our knowledge and to deliver successful therapies.
RECENT FINDINGS
The discovery of Anaplastic Lymphoma Kinase (ALK) fusions has provided the basis for the characterization of distinct subsets among ALCL patients. Although the oncogenic addiction of ALK signaling is proven, the tumorigenic contribution of co-activating lesions is still missing. As ALK− and ALK+ share common signatures, it is plausible that analogous mechanisms of transformation may be operating in both subsets, as confirmed by the dis-regulated activation of c-MYC, and the loss of Blimp-1 and p53/p63 axis. Nonetheless, recurrent genetic alterations for ALK− ALCL or refractory leukemic ALK+ ALCL are lacking. Moreover, although conventional chemotherapies (anthracycline-based) are most successful, i.e. in ALK+ ALCL patients, the implementation of ALK inhibitors or of anti-CD30 based treatments provides innovatites solutions, particularly in pediatric ALK+ ALCL and in chemorefractory/relapsed patients.
SUMMARY
The complete portrayal of the landscape of genetic alterations in ALCL will dictate the development of innovative chemotherapeutic and targeted therapies that will fit most with the molecular and clinical profiling of individual patients.
Keywords: Anaplastic Lymphoma Kinase, signaling pathways, kinases’ inhibitors, lymphoid differentiation, mouse models
INTRODUCTION
Peripheral T-cell lymphoma (PTCL) are a heterogeneous group of tumors derived from post-thymic elements including leukemic/disseminated, nodal and extranodal diseases [1,2,3]. As orphan diseases (12 to 15% of all non-Hodgkin’s lymphoma [NHL] in Western populations) [1,2,3,4], they include entities displaying a great variability in clinical, morphological, immunophenotypic, cytogenetic and molecular features. First described in 1985 [5], Anaplastic Large Cell Lymphoma (ALCL) of either adults (2–8% of NHL) or children (15–30% of NHL), nowadays correspond to specific subtypes of systemic peripheral T-cell lymphoma [1]. The presence of Anaplastic Lymphoma Kinase (ALK) gene fusions has provided the criteria for a new WHO classification, which contemplates a novel entity (i.e. ALCL ALK+) and proposes a provisional one, including ALCL patients, who lack ALK translocations (i.e. ALCL ALK-). Because of genetic, immunophenotypic, and clinical differences, cutaneous ALCLs (cALCL) are considered as a completely distinct subset a part from its systemic counterparts. Systemic ALCL share cytological, immunophenotypic and molecular features. However, ALK- ALCL patients have poorer performance status, more often B symptoms [6], and an overall survival (OS) rate of 36% versus 20% of PTCL-NOS patients. This suggests unique driving defects, with high oncogenic penetrance. In contrast, ALK+ ALCL have a more favorable clinical course [4], though ALK+ ALCL with an aggressive behavior could be encountered in the clinical practice [7]. It remains uncertain whether the molecular lesion(s) and/or other features determine the clinical course of ALCL patients. In fact, once patients are normalized by clinical parameters, ALK- and ALK+ ALCL display analogous prognosis (failure-free survival [FFS] and OS) [6]. Considering that ALK+ ALCL have a less complex karyotype [8,9,10,11], it plausible that ALK fusions are critical actors and that tumor progression is due to somatic mutations (minimal deletions, activating somatic mutations, etc.) disrupting the function of a limited set of genes. In contrast, the transformation of ALK- ALCL might require the consolidation/acquisition of many genetic defects that rapidly lead to systemic and more aggressive phenotype. This is in agreement suggested with their higher and heterogeneous karyotypes [8,9,10,11]. Nonetheless, the driving lesion(s) of ALK- ALCL are still to be identified, and co-drivers are lacking for both ALCL subgroups.
The lack of representative cell lines or animal models has definitively contributed in impairing our knowledge of mature T-cell lymphoma. Ultimately, this has jeopardized the design of successful therapies and the upgrade of clinical programs, particularly in patients with poor outcome (ALK- ALCL and PTCL-NOS). The recognition of the tumorigenic defects of PTCL is expected to provide patient specific "molecular fingerprints" and thus more suitable tailored therapies.
SIGNALING OF ALK FUSIONS
Chromosomal translocations of the ALK gene are documented in many ALCL, although the percentage of ALK+ ALCL varies, as a result of the inclusion criteria of the ALK- ALCL. In absence of strong classifier(s) (like ALK), the distinction between ALK- ALCL, CD30 PTCL-NOS and some enteropathy associated T-cell lymphoma represents a diagnostic challenge.
ALK gene encodes a 210kDa tyrosine kinase receptor (CD247) belonging to the insulin growth factor receptor superfamily. Its expression is largely limited to the nervous system during embryogenesis and to focal areas of the adult brain [12]. Although the physiological role of ALK in mammals is unknown, it is involved in neuronal differentiation [13] and ALK activating mutations have been found in familial and sporadic neuroblastomas [14].
The breakpoints of ALK chimera invariably occur within the intron placed between the exons 19 and 20 (NM_004304.3). Thus exons coding for the intracytoplasmic domain of ALK (exons 20–29) are then juxtaposed to different partners [15,16]. Seventy-80% of ALK+ ALCL harbor the t(2;5)(p23;q35) translocation (NPM–ALK chimera). The intracellular distribution of the fusions is due to the structure/function of ALK partners, enforcing either to nuclear/cytoplasmic, or cytoplasmic and in rare cases juxta-membranous localization. Virtually all partners (with the exclusion of MYH9-ALK) provide dimerization domains, leading to the homo/heterodimerization of the fusions and to constitutive activation of the kinase [16,17]. Conventional genomic approaches, and more recently next generation sequencing (NGS) have shown alternative ALK translocations in many types of human cancers, e.i. lung tumors [18,19]. ALCL display additional alterations involving many chromosomes [8,9,10,20], however frequent common/discrete secondary lesions are rare. Two translocations were reported in ALK-ALCL, involving the DUSP22 gene, which is juxtaposed to the FRA7H fragile site, or to the gene coding for IRF4 [21]. Boi et al. have recently shown that several ALCL display recurrent deletions affecting 17p13.3-p12 (25%) region, in which TP53 gene is located, and 6q21 (19%) encompassing PRDM1 and ATG5 genes [11]. Finally, Vaismatzis et al. have recently described a set of genomic defects in DLBCL and PTCL/ALCL encoding fusion proteins homologous to ΔNp63, a dominant-negative p63 isoform that inhibits the p53 pathway [22].
ALK chimera were originally proven to be oncogenic in vivo [23] and these data were largely confirmed in in vitro models [24] and then in genetically modified animals [25,26]. Understanding how ALK signals acts and defining the mechanisms responsible for its deregulation is critical for dissecting the mechanisms, which mediate ALK cellular transformation and provide the basis for rationale therapeutic approaches. By a large array of methods, it is now know that ALK fusions and in particular the NPM-ALK chimera interact with a plethora of molecules and elicit many pathways. These include the RAS/Erk, PLC-γ, PI3K, and Jak/Signal Transducers and Activators of Transcription (STAT), capable to control individually or in association, cell proliferation, survival, and cytoskeletal properties [19].
The activation of RAS/Erk pathway provides positive signals regulating cell growth and the inhibition of MEK (AZD6244 or shRNA) leads to cell cycle arrest, without significant changes in cell viability (Crescenzo R, personal communication). Similarly, NPM-ALK can down-modulate, via PI3K-AKT, the inhibitory action of FOXO3a, upregulating Cyclin D2 and down-regulating p27, and providing positive growth signals.
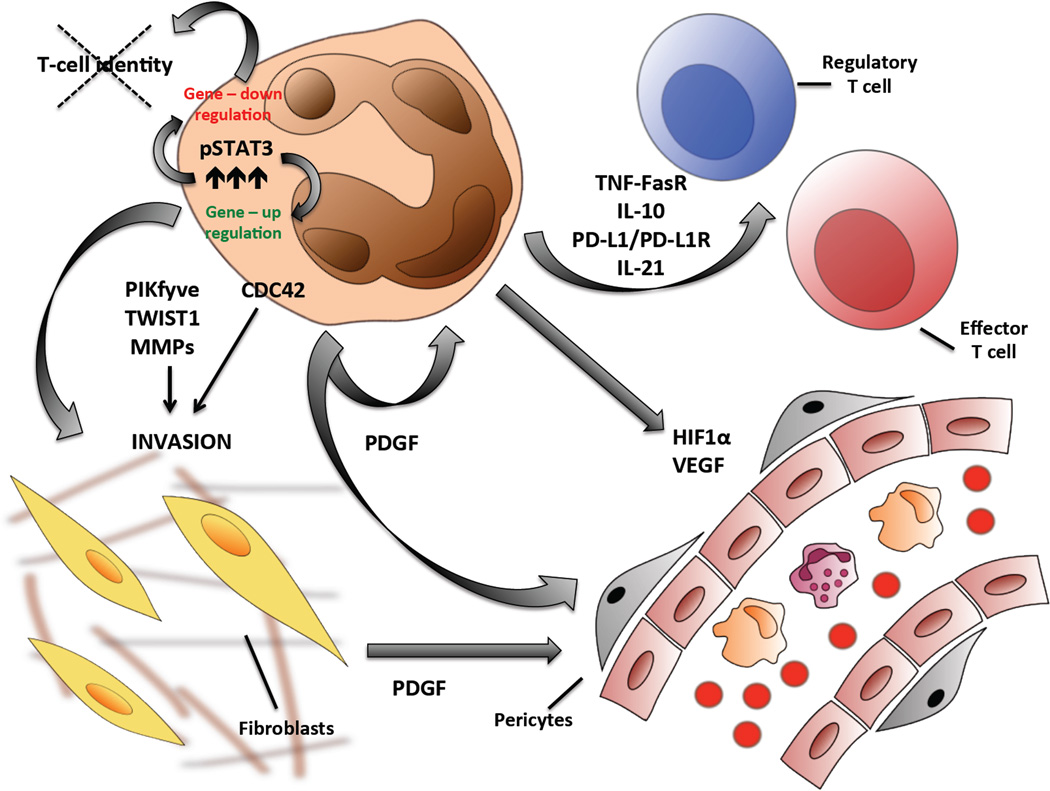
We and other groups have shown that the neoplastic phenotype of NPM-ALK is largely mediated by the STAT3. This enforces the transcription of a surplus of genes (coding and non coding), promoting cell growth and survival. In shRNA-based knockdown experiments, Piva et al. have demonstrated that several genes are directly regulated by STAT3, which display canonical STAT3 binding sites within their regulatory regions (Piva R, personal communication). Among them we mention CD30, granzyme, perforin, IL1RAP and IL2RA. From a diagnostic point of view, CD30, granzyme, perforin are known to be preferentially expressed by ALCL cells, and are commonly used in algorithms encompassing the differential diagnosis of different PTCL entities. Their transcription requires phosphorylated STAT3 complexes, which often include CEBPb and AP-1 transcription factors. Notably AP-1 members play an important role in ALK mediated transformation controlling tumor growth and positive host signals, via PDGF [27] (Fig. 1).
Figure 1. COMPLEX NETWORKING OF ALCL DISREGULATES AND CONTROLS HOST ENVIRONMENT: Tumor and host cells modulating their reciprocal functions and activities.
Oncogenic drivers in ALCL cells dirout intrinsic pathways leading to a self autonomous cell growth and overcoming host confinements; ALCL cells are also capable to modulate immunoresponsiveness changing the function of regulatory and effector T-cell through alternative mechanisms including antigen-camouflaging and producing lymphokines able to suppress immune-sorveillance (i.e. TNF/FasR, PD-L1/PD-L1R). Through cytoskeleton changes and production of pro-invasive mechanisms tumor cells have the ability to migrate locally and disseminate to distant organs. Moreover, production of pro-angiogenetic factors stimulates new vessels formation providing the necessary growth support for tumor survival. Many of these phenomena largely depend on the deregulated activation of multiple signalling pathway, and among them JAK-STAT represents a master culprit modulating gene-expression transcription (positive and negative regulation).
Legend: CDC42, Cell division control protein 42 homolog, HIF1α, Hypoxia-inducible factor 1 alpha; MMPs, Matrix metalloproteinases; PDGF, Platelet-derived growth factor; PD-L1, Programmed cell death 1 ligand 1; FasR, FAS recptor PIKfyve, FYVE finger-containing phosphoinositide kinase; IL, Interleukin; JAK, Janus Kinase; pSTAT3, phosphorylated Singnal transducer and activator of transcription 3; TNF, Tumor necrosis factor TWIST1, Twist related protein 1; VEGF, Vascular endothelial growth factor.
Zhang and coworkers have recently elucidated additional features of STAT3, demonstrating a STAT3-positive regulation of ICOS [28]. The same group had previously shown that PDL-1 expression is also regulated by STAT3 [29]. Collectively, these data demonstrate that ALCL cells engage ICOS to gain a growth advantage, and PDL-1 as novel mechanism of tumor escape, modulating the host responses. The overexpression of IL-21 [30] and deregulation of TNF/Fas/TNF [31] can also contribute to ALK tumorigenic phenotype, favoring the success of ALCL cells and overcoming host defenses. Finally, ALK signaling controls HiF1α, a factor that impacts directly in the neo-angiogenesis and provides a positive growth advantage to the lymphoma cells [32,33] (Fig. 1).
Lastly, STAT3 itself can directly or via downstream mediators down-regulate the transcription of many genes. Approximately 60% of modulated STAT3 genes are repressed after shRNA KD. Interestingly, Zhang et al have recently reported that STAT3, engaging the IL-2Rγ promoter, enhances the binding of DNA methyltransferases (DNMTs), leading ultimately to the transcriptional repression of IL-2Rγ gene. The knockdown of IL-2Rγ expression contributes to the neoplastic phenotype, as demonstrated by its forced/expression that leads to the loss of NPM-ALK protein expression, and then apoptosis. Ultimately, STAT3 down regulates T-cell associated molecules controlling T-cell identity of ALCL cells. In this context NPM-ALK provides signals capable to bypass TCR mediated activation [34,35] (Fig. 1).
STAT3 can similarly regulate the expression of several miRNA clusters (Spaccarotella E, personal communication) including the miRNA17-92 [36], known to have a role in human cancers. In ALCL, the miRNA17-92 overexpression overcomes in part the loss of STAT3 in an shRNA STAT3 inducible ALK+ ALCL model. More importantly, primary ALK+ ALCL display higher miRNA17-92 levels [36] compared to ALK- ALCL and cutaneous T-cell lymphoma and the usage of STAT3 inhibitors leading to the down-regulation of this cluster could represent an attractive strategy for the treatment of ALCL lymphoma (Lin C, personal communication).
The ability to successfully migrate and invade distant tissues contributes to the neoplastic phenotype, impairing clinical responses and long remissions. ALK signaling can efficiently module the cytoskeleton and promote invasion. The data reported by Ambrogio et al. [37] have recently been confirmed [38]. Dupuis-Coronas et al. have also shown that ALK, modulating the activity of PIKfyve, enhances the invasive capacities of NPM-ALK cells and their capacity to degrade the extracellular matrix [39]. Invasion of ALK+ ALCL cells is also modulated by the axis ALK-STAT3-Twist1 [40] (Fig. 1).
In conclusion, it is evident that the tumorigenic properties of ALK signaling are more complex that originally proposed, confirming that ALK is a powerful kinase capable to provide a complete and broad oncogenic addiction. These properties make ALK an excellent therapeutic target.
ALK+ AND ALK− ALCL: TWO SIDES OF THE SAME COIN?
The debate on distinct entities among PTCLs remains open. Novel hypotheses are emerging on the origin and relationship of different PTCL entities. The concept that ALK- ALCL should be lumped within PTCL-NOS has been recently sponsored. Alternatively, a scenario in which all ALCL are incorporated in a single group, irrespectively of the ALK expression has been contemplated. This level of uncertainty is corroborated by the fact that, once ALCL patients are stratified by stage, IPI etc. either groups display similar characteristics. In this landscape, CD30+ PTCL represent a puzzling/confounding group [41]. Their precise definition is critical and sometime questionable. Immunophenotypically, they express weak/partial CD30, and in same cases CD15 [42,43]. Cytologically display a certain monomorphism and they often have a functional TCR signaling (NFATc positive etc.) [44,45]. Clinically, CD30+ PTCL share a more aggressive clinical course, justifying their distinction and a closer relationship to PTCL-NOS. We strongly believe that these uncertainties will be solved only when distinct molecular defects will be discovered in different PTCLs.
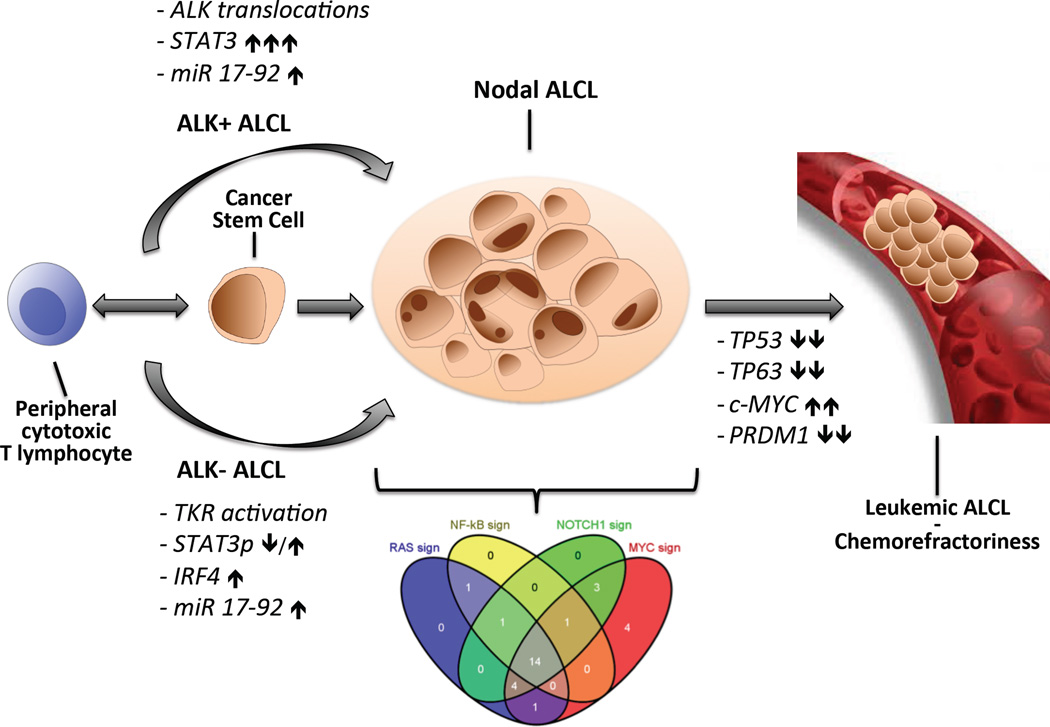
Another similar confusing topic regards the ALCL origin, and their putative normal counterpart elements. Several hypotheses have been proposed, taking in account their expression profile and unique immune-phenotype. The expression of perforin, T1A1 and granzyme has been interpreted as a specific fingerprint, supporting the idea that ALCL may derive from cytotoxic T-lymphocytes. Alternatively, we speculate that the phenotype of ALCL may rather be the result of the deregulated expression of unique pathways and/or specific defects, which impose unique/fixed profiles. It is known that transcription factors can play a critical role in T-cell differentiation and once constitutively activated can undermine physiological programs and rerouted their development. Based on this assumption, we could speculate that the constitutive activation of STAT3 might be responsible for the cytotoxic phenotype of ALK+ ALCL cells, even in cells that were committed to different lineages and/or function. This leaves the open question, why ALK- ALCL display a cytotoxic phenotype? To solve this question, we have analyzed a large cohort of ALCL samples and found that a subset of ALK- ALCL clearly shares a STAT3 expression profile and detectable nuclear pSTAT3. Moreover, both ALK+ and ALK- ALCL reveal signatures, linked to the activation of c-MYC, NOTCH-1, or NFkB, and RAS/ERK, suggesting the existence of upstream activators. Interestingly, it now evident that several ALCL co-share overlapping signatures suggesting multiple activating defects or alternatively the presence of unique lesions capable, like ALK fusions, to efficiently and concomitantly fire multiple pathways. Search of ALCL pathogenetic lesions is under evaluation and it is predicted that new information will be available soon (Fig. 2).
Figure 2. HYPOTHETICAL MODEL DEPICTING ALCL TRANSFORMATION: a common stem cell precursor and/or a normal counterpart for ALCL remain unknown?
We are still unable to understand how the recently identified dysregulation of multiple signaling pathways controls the immunophenotype and effector function of ALCL tumor cells. The discovery of leading genetic defects in both ALK+ and ALK- ALCL is critical to underline the mechanism responsible for cell transformation and tumor maintenance; this knowledge is essential to select vulnerable targets and address innovative therapies. With more successful approaches, chemorefractoriness and leukemic transformation will become more frequent and thus more appropriate strategies will be required. Indeed the discovery that the loss of several tumor-suppressor genes (TP53, TP63, BLIMP1) and the dysregulation of c-MYC identify high risk patients should provide the rational basis for the usage of new therapeutics agents (i.e. Bromodomain-inhibitors).
Legend: ALCL, Anplastic large cell lymphoma; ALK, Anaplastic lymphoma kinase; PRDM1, PR domain zinc finger protein 1 IRF4, Interferon regulatory factor 4; STAT3, Signal transducer and activator of transcription; TP, Tumor protein: TKR, Tyrosine kinase recpetor.
CAN WE USE PRECISION MEDICINE DATA TO IMPROVE THERAPUTIC COMPLIANCE?
The definition of the molecular fingerprints of neoplasms is now possible through the implementation of the impressive technologies. The NGS platforms are currently entering the clinical arena and it is plausible that, once interconnected and clinical based networks of laboratories, many patients will have individualized molecular identikits. Nonetheless, caveats on the tumorigenic contribution of individual lesions and their functional role in the maintenance of the neoplastic phenotypes remain untouched. This are a critical issues, which should be added to the overwhelming capacity of tumor cells to adapt rapidly to the environment and to stress imposed by drugs and host changes. Thus, the search the “magic bullet” may fail. Instead, the association of multiple “smart” compounds could provide higher response rates and overcome resistance. Since the cost for a novel drug is around 1billion and requires approximately 12–15 years, we need to overcome impairing inefficiencies. It is agreed that many improvements in discovery programs need to be rapidly put in place, meliorating company inefficiencies (structural and operation), selection of viable targets, defining good therapeutic biopredictors, innovative technologies, more efficient and reliable screening tests and faster and less expensive clinical tracks in molecularly defined and/or naïve patients.
While pharmaceutical companies are reshaping their pipelines, a small number of drugs is successfully introduced into clinics. This ineffective result is seemingly related to update pre-clinical models, heavily relaying on in vitro models and “xenografts mouse platforms”. Indeed, the most frequently used cell lines poorly represent human tumors [46]. This has encouraged many institutions and drug industry to acquire large library of cell lines, which can be interrogated with HTP platforms (NSG, phosphomapping etc.). The hope is to define better criteria and relationships between the genome and responses to therapies. The hope is of predicting more reliable clinical responses and dissecting responders and refractory patients. But, cell lines lack the host and its regulatory networks, have undergone ferocious in vitro selections and do not represent tumor heterogeneity.
To solve some of these issues, implants of fresh primary neoplasms are frequently introduced in severally-immunocompromised mice [47]. The generation of individualized cancer models represents an unprecedented opportunity to test battery of drugs for each individual and provide personalized oncology programs. However, these strategies need to be linked to defined genetic defects. Only combining HTP and innovative models, we can deliver a list of targetable lesions, which once validated in animals, should provide reasonable expectations. Since the successful growth of tumorgraft implants may require long period of time, new technologies interrogating the functional network in cancer and the efficacy of chemical libraries in vitro may provide alternative routes for the execution of pre-clinical trials in vivo. Our group has recently embarked in such a program and generated a battery of ALCL “Patients Derived Tumorgrafts” (PDT) [48]. These retain the immunophenotypic, genomic features of their corresponding primary tumors and display responses to conventional and innovative protocols that closely mimic those seen in donor patients. Their molecular characterization has demonstrated the presence of unique genomic defects and allowed to discover new pathogenetic translocations and activating somatic mutations. The definition of a molecular identikit in PDTs will provide not only patients’ fingerprints but also models to test the efficacy of selected drugs targeting hypothetical tumorigenic defects in each patient in vivo.
CONCLUSION
Little is known on the mechanisms leading to T-cell transformation. Nonetheless, the systematic usage of HTP platforms has recently demonstrated that recurrent defect may be present in specific subset of PTCL. Although ALK+ ALCL and ALK- ALCL display heterogeneous complex karyotypes, they share common expression signatures and disregulated signaling pathways. The usage of NGS approaches will be instrumental for the discovery of the driving defects of ALCL. New defects, even in small subgroups of patients, will provide objective diagnostic criteria and the bases for “intelligent” therapies, to be first validated in new preclinical models (i.e. PDT etc.).
KEY POINTS.
Anaplastic Large Cell Lymphoma are an heterogeneous group of lymphoma some of which carry a restricted number of genetic defects mainly involving the Anaplastic Lymphoma Kinase (ALK+ ALCL) o less frequently display alternative translocations [t(2;x)(p23;x), t(6;7)(p25.3;q32.3) and inv(3)(q26q28) leading to TBL1XR1/TP63, etc.].
The precise relationship and origin of ALCL remain unclear. Although ALK+ and ALK- share a set of genes and similar phenotypes, they are considered distinct groups with unique clinical features.
Bioinformatics algorithms have identified several ALCL signaling classifiers demonstrating the preferential expression of a restricted number of pathways. The recognition of “common hubs”, which can be targeted by selective inhibitors, represents a viable strategy for future therapeutic protocols.
Taking advantage of in vitro and in vivo models, several groups have shown that the transforming properties of ALK fusions involve a plethora of alternative modules capable to regulate intrinsic (i.e. cytoskeleton, cell growth, etc.) and/or extrinsic (cell matrix invasion, tumor-host relationships, etc.) modalities.
The identification of the driving lesion of ALCL will require the construction of international networks capable to synergize their activities and to construct large and clinically annotate tissue libraries. The collection of viable tissues will facilitate the generation of batteries of “Patient Derived Tumorgrafts”.
ACKNOWLEGMENTS
GI is supported by the Italian Association for Cancer Research (AIRC) Special Program in Clinical Molecular Oncology, Milan (5x1000 No. 10007); Regione Piemonte (ONCOPROT, CIPE 25/2005); ImmOnc (Innovative approaches to boost the immune responses, Programma Operativo Regionale, Piattaforme Innovative BIO F.E.S.R. 2007/13, Asse 1 'Ricerca e innovazione' della LR 34/2004) and the Oncology Program of Compagnia di San Paolo, Torino. RR is supported by Partnership for Cure, NIH 1 P50 MH094267-01, NIH 1 U54 CA121852-05, NIH 1R01CA164152-01. FB sponsored by the Oncosuisse KLS-02403-02-2009 (Bern, Switzerland); Anna Lisa Stiftung (Ascona, Switzerland); Nelia and Amadeo Barletta Foundation (Lausanne, Switzerland). We thank Drs Vigliani C, Fioravanti A, and Mossino M for their technical support.
REFERENCES
- 1.Swerdlow SH, Campo E, Haris NL, et al. In: WHO classification of Tumors of Haemotolopoietic and Lymphoid tissues. 4th Edition. Swerdlow SH, Campo E, Haris NL, et al., editors. Lyon: International Agency for Research on Cancer; 2008. pp. 312–317. [Google Scholar]
- 2.Inghirami G, Pileri SA. Anaplastic large-cell lymphoma. Semin Diagn Pathol. 2011;28:190–201. doi: 10.1053/j.semdp.2011.03.002. [PMID: 21850985] [DOI] [PubMed] [Google Scholar]
- 3.Rodriguez-Abreu D, Filho VB, Zucca E. Peripheral T-cell lymphomas, unspecified (or not otherwise specified): a review. Hematol Oncol. 2008;26:8–20. doi: 10.1002/hon.836. [PMID: 18050364] [DOI] [PubMed] [Google Scholar]
- 4.Vose J, Armitage J, Weisenburger D. International peripheral T-cell and natural killer/T-cell lymphoma study: pathology findings and clinical outcomes. J Clin Oncol. 2008;26:4124–4130. doi: 10.1200/JCO.2008.16.4558. [PMID: 18626005] [DOI] [PubMed] [Google Scholar]
- 5.Stein H, Mason DY, Gerdes J, et al. The expression of the Hodgkin's disease associated antigen Ki-1 in reactive and neoplastic lymphoid tissue: evidence that Reed-Sternberg cells and histiocytic malignancies are derived from activated lymphoid cells. Blood. 1985;66:848–858. [PMID: 3876124] [PubMed] [Google Scholar]
- 6.Savage KJ, Harris NL, Vose JM, et al. ALK-anaplastic large-cell lymphoma is clinically and immunophenotypically different from both ALK+ ALCL and peripheral T-cell lymphoma, not otherwise specified: report from the International Peripheral T-Cell Lymphoma Project. Blood. 2008;111:5496–5504. doi: 10.1182/blood-2008-01-134270. [PMID: 18385450] [DOI] [PubMed] [Google Scholar]
- 7.Grewal JS, Smith LB, Winegarden JD, 3rd, et al. Highly aggressive ALK-positive anaplastic large cell lymphoma with a leukemic phase and multi-organ involvement: a report of three cases and a review of the literature. Ann Hematol. 2007;86:499–508. doi: 10.1007/s00277-007-0289-3. [PMID: 17396261] [DOI] [PubMed] [Google Scholar]
- 8.Zettl A, Rüdiger T, Konrad MA, et al. Genomic profiling of peripheral T-cell lymphoma, unspecified, and anaplastic large T-cell lymphoma delineates novel recurrent chromosomal alterations. Am J Pathol. 2004;164:1837–1848. doi: 10.1016/S0002-9440(10)63742-X. [PMCID: PMC1615643] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Salaverria I, Beà S, Lopez-Guillermo A, et al. Genomic profiling reveals different genetic aberrations in systemic ALK-positive and ALK-negative anaplastic large cell lymphomas. Br J Haematol. 2008;140:516–526. doi: 10.1111/j.1365-2141.2007.06924.x. [PMID: 18275429] [DOI] [PubMed] [Google Scholar]
- 10.Boi M, Stathis A, Zucca E, et al. Genetic alterations in systemic nodal and extranodal non-cutaneous lymphomas derived from mature T cells and natural killer cells. Cancer science. 2012;103:1397–1404. doi: 10.1111/j.1349-7006.2012.02321.x. [PMID: 22568409] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Boi M, Rinaldi A, Piva R, et al. BLIMP1 Is Commonly Inactivated In Anaplastic Large T-Cell Lymphomas (ALCL) Blood (ASH Annual Meeting Abstracts) 2011;118:1131–1132. [PMID: 24004669] [Google Scholar]
- 12.Iwahara T, Fujimoto J, Wen D, et al. Molecular characterization of ALK, a receptor tyrosine kinase expressed specifically in the nervous system. Oncogene. 1997;14:439–449. doi: 10.1038/sj.onc.1200849. [PMID: 9053841] [DOI] [PubMed] [Google Scholar]
- 13.Souttou B, Carvalho NB, Raulais D, Vigny M. Activation of anaplastic lymphoma kinase receptor tyrosine kinase induces neuronal differentiation through the mitogen-activated protein kinase pathway. J Biol Chem. 2001;276:9526–9531. doi: 10.1074/jbc.M007333200. [PMID: 11121404] [DOI] [PubMed] [Google Scholar]
- 14.Mossé YP, Laudenslager M, Longo L, et al. Identification of ALK as a major familial neuroblastoma predisposition gene. Nature. 2008;455:930–935. doi: 10.1038/nature07261. [PMCID: PMC2672043] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Morris SW, Kirstein MN, Valentine MB, et al. Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin's lymphoma. Science. 1994;263:1281–1284. doi: 10.1126/science.8122112. [PMID: 8122112] [DOI] [PubMed] [Google Scholar]
- 16.Barreca A, Lasorsa E, Riera L, et al. Anaplastic lymphoma kinase in human cancer. J Mol Endocrinol. 2011;47:R11–R23. doi: 10.1530/JME-11-0004. [PMID: 21502284] [DOI] [PubMed] [Google Scholar]
- 17.Chiarle R, Voena C, Ambrogio C, et al. The anaplastic lymphoma kinase in the pathogenesis of cancer. Nat Rev Cancer. 2008;8:11–23. doi: 10.1038/nrc2291. [PMID: 18097461] [DOI] [PubMed] [Google Scholar]
- 18.Soda M, Takada S, Takeuchi K, et al. A mouse model for EML4-ALK-positive lung cancer. Proc Natl Acad Sci U S A. 2008;105:19893–19897. doi: 10.1073/pnas.0805381105. [PMCID: PMC2605003] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Tabbò F, Barreca A, Piva R, Inghirami G. ALK Signaling and Target Therapy in Anaplastic Large Cell Lymphoma. Front Oncol. 2012;2:41. doi: 10.3389/fonc.2012.00041. [PMCID: PMC3355932] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ott G, Katzenberger T, Siebert R, DeCoteau JF, et al. Chromosomal abnormalities in nodal and extranodal CD30+ anaplastic large cell lymphomas: infrequent detection of the t(2;5) in extranodal lymphomas. Genes Chromosomes Cancer. 1998;22:114–121. doi: 10.1002/(sici)1098-2264(199806)22:2<114::aid-gcc5>3.0.co;2-#. [PMID: 9598798] [DOI] [PubMed] [Google Scholar]
- 21.Feldman AL, Dogan A, Smith DI, et al. Discovery of recurrent t(6;7)(p25.3;q32.3) translocations in ALK-negative anaplastic large cell lymphomas by massively parallel genomic sequencing. Blood. 2011;117:915–919. doi: 10.1182/blood-2010-08-303305. [PMCID: PMC3035081] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Vasmatzis G, Johnson SH, Knudson RA, et al. Genome-wide analysis reveals recurrent structural abnormalities of TP63 and other p53-related genes in peripheral T-cell lymphomas. Blood. 2012;120:2280–2289. doi: 10.1182/blood-2012-03-419937. [PMID: 22855598] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kuefer MU, Look AT, Pulford K, et al. Retrovirus-mediated gene transfer of NPM-ALK causes lymphoid malignancy in mice. Blood. 1997;90:2901–2910. [PMID: 9376569] [PubMed] [Google Scholar]
- 24.Bai RY, Dieter P, Peschel C, et al. Nucleophosmin-anaplastic lymphoma kinase of large-cell anaplastic lymphoma is a constitutively active tyrosine kinase that utilizes phospholipase C-gamma to mediate its mitogenicity. Mol Cell Biol. 1998;18:6951–6961. doi: 10.1128/mcb.18.12.6951. [PMCID: PMC109278] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chiarle R, Gong JZ, Guasparri I, et al. NPM-ALK transgenic mice spontaneously develop T-cell lymphomas and plasma cell tumors. Blood. 2003;101:1919–1927. doi: 10.1182/blood-2002-05-1343. [PMID: 12424201] [DOI] [PubMed] [Google Scholar]
- 26.Turner SD, Alexander DR. What have we learnt from mouse models of NPM-ALK-induced lymphomagenesis? Leukemia. 2005;19:1128–1134. doi: 10.1038/sj.leu.2403797. [PMID: 15902287] [DOI] [PubMed] [Google Scholar]
- 27. Laimer D, Dolznig H, Kollmann K, et al. PDGFR blockade is a rational and effective therapy for NPM-ALK-driven lymphomas. Nat Med. 2012;18:1699–1704. doi: 10.1038/nm.2966. [PMID: 23064464] Taking advantage of engineered mouse modeling the authors highlight the pathogenetic role of AP-1 transcription factors and their downstream molecules (PDGF) in ALK-driven lymphomagenesis.
- 28.Zhang Q, Wang H, Kantekure K, et al. Oncogenic tyrosine kinase NPM-ALK induces expression of the growth-promoting receptor ICOS. Blood. 2011;118:3062–3071. doi: 10.1182/blood-2011-01-332916. [PMCID: PMC3175783] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Marzec M, Zhang Q, Goradia A, et al. Oncogenic kinase NPM/ALK induces through STAT3 expression of immunosuppressive protein CD274 (PD-L1, B7-H1) Proc Natl Acad Sci U S A. 2008;105:20852–20857. doi: 10.1073/pnas.0810958105. [PMCID: PMC2634900] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Dien Bard J, Gelebart P, Anand M, et al. IL-21 contributes to JAK3/STAT3 activation and promotes cell growth in ALK-positive anaplastic large cell lymphoma. Am J Pathol. 2009;175:825–834. doi: 10.2353/ajpath.2009.080982. [PMCID: PMC2716977] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wu F, Wang P, Zhang J, et al. Studies of phosphoproteomic changes induced by nucleophosmin-anaplastic lymphoma kinase (ALK) highlight deregulation of tumor necrosis factor (TNF)/Fas/TNF-related apoptosis-induced ligand signaling pathway in ALK-positive anaplastic large cell lymphoma. Mol Cell Proteomics. 2010;9:1616–1632. doi: 10.1074/mcp.M000153-MCP201. [PMCID: PMC2938097] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Marzec M, Liu X, Wong W, et al. Oncogenic kinase NPM/ALK induces expression of HIF1alpha mRNA. Oncogene. 2010;30:1372–1378. doi: 10.1038/onc.2010.505. [PMID: 21102525] [DOI] [PubMed] [Google Scholar]
- 33.Dejean E, Renalier MH, Foisseau M, et al. Hypoxia-microRNA-16 downregulation induces VEGF expression in anaplastic lymphoma kinase (ALK)-positive anaplastic large-cell lymphomas. Leukemia. 2011;25:1882–1890. doi: 10.1038/leu.2011.168. [PMID: 21778999] [DOI] [PubMed] [Google Scholar]
- 34.Bonzheim I, Geissinger E, Roth S, et al. Anaplastic large cell lymphomas lack the expression of T-cell receptor molecules or molecules of proximal T-cell receptor signaling. Blood. 2004;104:3358–3360. doi: 10.1182/blood-2004-03-1037. [PMID: 15297316] [DOI] [PubMed] [Google Scholar]
- 35.Ambrogio C, Martinengo C, Voena C, et al. NPM-ALK oncogenic tyrosine kinase controls T-cell identity by transcriptional regulation and epigenetic silencing in lymphoma cells. Cancer Res. 2009;69:8611–8619. doi: 10.1158/0008-5472.CAN-09-2655. [PMCID: PMC2784121] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Merkel O, Hamacher F, Laimer D, et al. Identification of differential and functionally active miRNAs in both anaplastic lymphoma kinase (ALK)+ and ALK-anaplastic large-cell lymphoma. Proc Natl Acad Sci U S A. 2010;107:16228–16233. doi: 10.1073/pnas.1009719107. [PMCID: PMC2941277] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Ambrogio C, Voena C, Manazza AD, et al. The anaplastic lymphoma kinase controls cell shape and growth of anaplastic large cell lymphoma through Cdc42 activation. Cancer Res. 2008;68:8899–8907. doi: 10.1158/0008-5472.CAN-08-2568. [PMCID: PMC2596920] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Colomba A, Giuriato S, Dejean E, et al. Inhibition of Rac controls NPM-ALK-dependent lymphoma development and dissemination. Blood Cancer. 2011;1(6):e2. doi: 10.1038/bcj.2011.19. [PMCID: PMC3255265] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Dupuis-Coronas S, Lagarrigue F, Ramel D, et al. The nucleophosmin-anaplastic lymphoma kinase oncogene interacts, activates, and uses the kinase PIKfyve to increase invasiveness. J Biol Chem. 2011;286:32105–32114. doi: 10.1074/jbc.M111.227512. [PMCID: PMC3173219] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhang J, Wang P, Wu F, et al. Aberrant expression of the transcriptional factor Twist1 promotes invasiveness in ALK-positive anaplastic large cell lymphoma. Cell Signal. 2012;24:852–858. doi: 10.1016/j.cellsig.2011.11.020. [PMID: 22155737] [DOI] [PubMed] [Google Scholar]
- 41.Fornari A, Piva R, Chiarle R, et al. Anaplastic large cell lymphoma: one or more entities among T-cell lymphoma? Hematol Oncol. 2009;27:161–170. doi: 10.1002/hon.897. [PMID: 19358142] [DOI] [PubMed] [Google Scholar]
- 42.Barry TS, Jaffe ES, Sorbara L, et al. Peripheral T-cell lymphomas expressing CD30 and CD15. Am J Surg Pathol. 2003;27:1513–1522. doi: 10.1097/00000478-200312000-00003. [PMID: 14657710] [DOI] [PubMed] [Google Scholar]
- 43.Gorczyca W, Tsang P, Liu Z, et al. CD30-positive T-cell lymphomas co-expressing CD15: an immunohistochemical analysis. Int J Oncol. 2003;22:319–324. [PMID: 12527929] [PubMed] [Google Scholar]
- 44.Piva R, Agnelli L, Pellegrino E, et al. Gene expression profiling uncovers molecular classifiers for the recognition of anaplastic large-cell lymphoma within peripheral T-cell neoplasms. J Clin Oncol. 2010;28:1583–1590. doi: 10.1200/JCO.2008.20.9759. [PMID: 20159827] [DOI] [PubMed] [Google Scholar]
- 45. Agnelli L, Mereu E, Pellegrino E, et al. Identification of a 3-gene model as a powerful diagnostic tool for the recognition of ALK-negative anaplastic large-cell lymphoma. Blood. 2012;120:1274–1281. doi: 10.1182/blood-2012-01-405555. [PMID: 22740451] The authors investigate, through a gene expression platform, the feasibility of a molecular stratification among T-NHL, highlighting diagnostic predictors capable to improve the correct identification of questionable cases, including CD30+ PTCL
- 46.Garber K. From human to mouse and back: 'tumorgraft' models surge in popularity. J Natl Cancer Inst. 2009;101:6–8. doi: 10.1093/jnci/djn481. [PMID: 19116380] [DOI] [PubMed] [Google Scholar]
- 47.Shultz LD, Ishikawa F, Greiner DL. Humanized mice in translational biomedical research. Nat Rev Immunol. 2007;7:118–130. doi: 10.1038/nri2017. [PMID: 17259968] [DOI] [PubMed] [Google Scholar]
- 48.Tabbò F, Barreca A, Machiorlatti R, et al. Humanized NOD/Scid/IL2g−/− tumorgrafts recapitulate primary Anaplastic Large Cell Lymphoma. AACR Meeting Abstracts. 2013 [Google Scholar]