Abstract
Myotonic dystrophy disorders are caused by expanded CUG repeats in non-coding regions. To reveal mechanisms of CUG repeat pathogenesis we used C. elegans expressing CUG repeats to identify gene inactivations that modulate CUG repeat toxicity. We identified 15 conserved genes that function as suppressors or enhancers of CUG repeat-induced toxicity and modulate formation of nuclear RNA foci by CUG repeats. These genes regulated CUG repeat-induced toxicity through distinct mechanisms including RNA export and RNA clearance, suggesting that CUG repeat toxicity is mediated by multiple pathways. A subset is shared with other degenerative disorders. The nonsense-mediated mRNA decay (NMD) pathway plays a conserved role regulating CUG repeat RNA transcript levels and toxicity, and NMD recognition of toxic RNAs depends on 3′UTR GC nucleotide content. Our studies suggest a broader surveillance role for NMD where variations in this pathway influence multiple degenerative diseases.
Keywords: CUG repeats, myotonic dystrophy, DM1, RNA toxicity, repeat disorders, nonsense-mediated decay, Caenorhabditis elegans
Introduction
Expansions in nucleotide repeat sequences cause many neuromuscular degenerative disorders1 and can occur in noncoding as well as coding regions of genes. Expansions of CTG repeats in the 3′ untranslated region (3′UTR) of the DMPK protein kinase gene causes myotonic dystrophy 1 (DM1), an autosomal dominant degenerative disease2. DM1 CTG expansions range up to >2,000 repeats; normal CTG lengths range from 5–36 repeats. RNA toxicity is the cause of DM1 pathology, where transcripts containing expanded CUG repeats accumulate in the nucleus as discrete RNA foci3. The length of repeat expansion correlates with DM1 disease onset and severity4,5. Expanded CUG repeat RNA transcripts disrupt alternative RNA splicing mediated by muscleblind-like (MBNL)6 and the CUG binding protein 1 (CUG-BP1)7 RNA binding protein families, causing toxicity. However, disruption of these splicing factors, in particular of MBNL1, does not explain the many phenotypes observed in DM disorders. The identification of new factors that function as modifiers of DM1-associated RNA toxicity8–10, and the demonstration that in DM1 mouse models changes are detected in mRNA transcript levels and in transcript splicing not observed in mbnl1 knockout mice11,12, suggest the involvement of additional unknown factors and mechanisms in expanded CUG repeat pathogenesis. Here we find that many of the mammalian toxic features of non-coding expanded CUG repeats can be recapitulated in C. elegans muscle. Analysis of C. elegans muscle function defects caused by expanded CUG repeats, together with cell biological analysis of these aberrant RNAs in wild type and in strains with a library of genes inactivated, identified gene inactivations that modify expanded CUG repeat toxicity and CUG repeat foci accumulation, the hallmark of DM disorders. These modifiers of expanded CUG repeat toxicity include the nonsense-mediated mRNA decay (NMD) pathway, which targets CUG repeat-containing transcripts for degradation. NMD regulation of CUG repeat foci accumulation is a conserved mechanism present in both C. elegans and human cells. Recognition of these CUG repeat-containing transcripts for degradation by NMD is dependent on repeat-sequence composition.
Results
Expanded CUG repeats cause C. elegans muscle defects
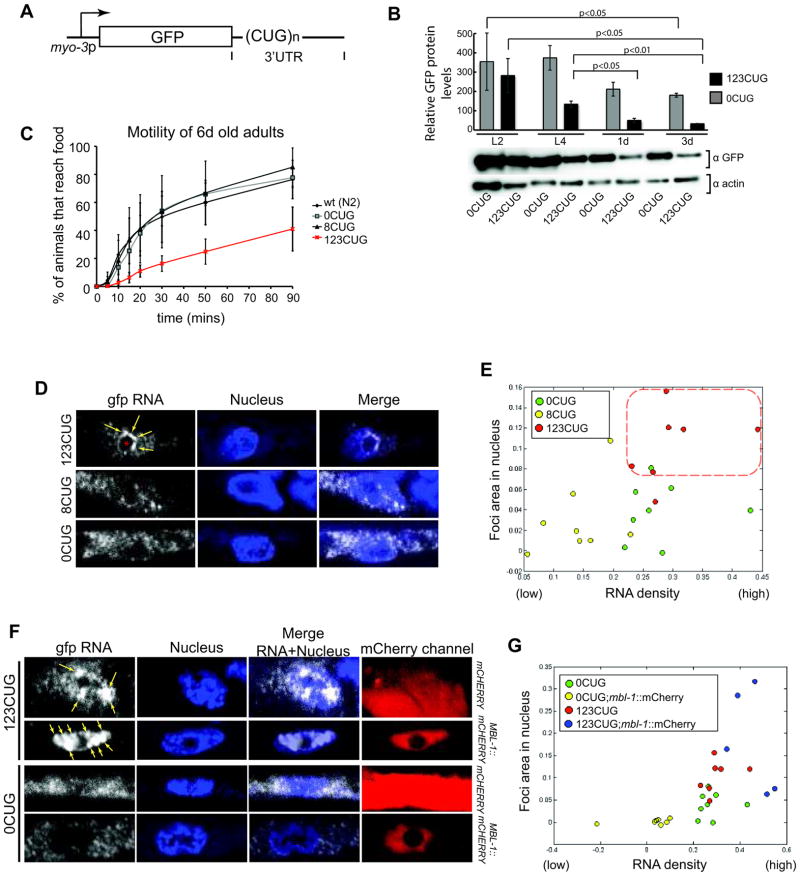
We generated a set of C. elegans reporter genes expressing GFP with 3′UTR containing various lengths of CTG repeats in body wall muscle cells, using the myo-3 muscle-specific promoter (Fig. 1A). Reporter constructs without any CUG repeats in the 384-nt 3′UTR from the let-858 gene (0CUG) displayed strong GFP fluorescence at all developmental stages, with a modest decline during adulthood. Analogous constructs with eight CUG repeats showed similar results with mild changes in GFP fluorescence (data not shown). In contrast, the presence of 123 CUG repeats in the 3′UTR (123CUG, a pathogenic repeat length in mammalian myocytes) resulted in a sharp decline in GFP fluorescence as animals developed to adults. Western blotting analyses revealed a sharp decrease in GFP protein levels in 3 day (3d) old adult stage animals of the 123CUG strain (12% compared to protein levels at the L2 larval stage). The 3d adult stage animals of control 0CUG strain showed 50% of the GFP levels in L2) (Fig. 1B). We used the decline in adult stage GFP fluorescence in123CUG transgenic animals for RNAi screens to identify genes that influence toxicity of expanded CUG repeats.
Figure 1.
Expanded CUG-dependent C. elegans muscle phenotypes
(A) Diagram of CUG-containing plasmids for expression in C. elegans muscle cells, under the myo-3 promoter. n indicates number of CUG repeats. (B) Quantification of GFP expression levels from reporter genes with 123CUG repeats or 0 CUG repeats in the 3′UTR, relative to actin. Graph shows mean and s.d. for 3 independent experiments, p was determined by Student’s t test. Bottom, western blots using GFP and actin antibodies, actin was used for sample normalization. (C) Motility assays for 6d adults. Data plotted corresponds to average percentage of population to reach food at each time point. Error bars represent SD from at least 3 independent experiments; in each experiment, 3–5 replicas of ca. 100–150 animals were analyzed. (D) Confocal single molecule RNA fluorescence in situ hybridization (SM-FISH) images of C. elegans muscle cells for GFP RNA transcripts (right, white); nucleus are stained with DAPI (blue). Yellow arrows indicate expanded CUG nuclear foci, and the red asterisk (●) indicates the nucleolus. (E) Computational analysis of SM-FISH muscle cell images of 0CUG, 8CUG and 123CUG animals. Each dot corresponds to an analyzed SM-FISH image. The red dotted square indicates the region of clustering of the 123CUG images (red dots). (F) confocal SM-FISH images of C. elegans muscle cells for GFP RNA transcripts (right, white); nucleus are stained with DAPI and mCherry fluorescence is shown on the right. The strains express GFP with 123CUG or 0CUG in a mCHERRY or MBL-1::mCHERRY backgrounds. Yellow arrows indicate expanded CUG nuclear foci. MBL-1::mCHERRY localizes to the nucleus. (G) Computational analysis of SM-FISH images of 0CUG, 0CUG;mbl-1::mCherry, 123CUG and 123CUG;mbl-1::mCherry animals.
We investigated the function of C. elegans muscle expressing CUG repeats by assessing locomotion phenotypes of these animals. We quantified motor defects by determining the percentage of animals that reached an attractant E. coli food ring (2cm radius) on an agar plate in 90min (Fig 1C and Supplementary Fig. 1A). The 123CUG strains exhibited severe motility deterioration at 6d adulthood, moving about five fold slower than wild type or control transgenic animals carrying 8CUGs or 0CUG constructs, which were similar to wild type. We also observed earlier locomotion defects in synchronized populations of 123CUG animals at the 2d adult stage (Supplementary Fig. 1B) and at the L4 stage (data not shown), whereas strains’ bearing 8CUG or 0CUG repeats showed no motility defects. Thus, expanded CUG repeats cause progressive muscle dysfunction as C. elegans ages, as in other organisms including mammals13–15.
Because nuclear inclusions of expanded CUG repeat RNAs are characteristic of DM, we investigated whether 123CUG RNA transcripts formed nuclear foci in C. elegans muscle cells. We used single molecule RNA fluorescence in situ hybridization (SM-FISH) that has higher sensitivity and specificity than traditional FISH16 (see Supplementary Notes). The repeat-containing region of the expanded RNA transcript is known to interact inappropriately with RNA-binding proteins17; therefore we chose RNA probes complementary to the GFP sequence expected to be accessible in SM-FISH. SM-FISH detected the accumulation of expanded mRNA transcripts in foci as ‘large’, often amorphous, bright fluorescent structures, with 123CUG repeats mRNAs causing the accumulation of 2 to 5 nuclear foci per cell (Fig. 1D). Many individual fluorescence spots, likely corresponding to individual mRNAs, were also observed in the nucleus in the 123CUG strain (Fig. 1D). In contrast, animals expressing 0CUG or 8CUG repeat RNA transcripts lacked multiple bright nuclear foci, and exhibited a predominantly cytoplasmic distribution of RNA ‘single’ transcripts (Fig. 1D).
For a systematic analysis of all SM-FISH data to quantify foci formation and nuclear versus cytoplasmic RNA distribution for 123CUG repeats vs controls, we developed an algorithm that analyzed pixel intensity and cellular distribution in SM-FISH images (Supplementary Fig. 1C–E and Supplementary Notes). We examined the SM-FISH images collected for the nuclear versus cytoplasmic distribution of CUG repeat RNA transcripts as foci or as ‘concentrated single transcripts’ (high RNA density areas) (Supplementary Fig. 1C–E). Consistent with the SM-FISH images (Fig. 1D), the analysis of multiple 123CUG images showed a higher nuclear fluorescence intensity, corresponding to nuclear foci and ‘single’ RNA transcripts (Fig. 1E), clearly distinct from the control 0CUG samples. The quantitative analysis also distinguished the 8CUG from the 0CUG samples, indicating that there are fewer RNA transcripts in the nucleus of 8CUG animals compared to 0CUG strains.
The mammalian splicing protein MBNL1 binds to RNA transcripts containing expanded CUG repeats6, and in myotonic dystrophy is sequestered by expanded CUG foci18. We examined by SM-FISH and by mosaic analysis in vivo (see Supplementary Notes) whether the C. elegans MBNL1 orthologue, MBL-119, bound the 123CUG foci detected in muscle cells. Expression of mbl-1 in a 123CUG background caused a marked increase in foci size relative to the 123CUG strain alone (Fig. 1F, G, Supplementary Fig. 2A–D). Mosaic analysis showed that MBL-1 caused the retention of expanded CUG repeat RNA transcripts in large nuclear foci disrupting transport to the cytoplasm and GFP translation (Supplementary Fig. 2E and Supplementary Notes). These effects were not observed with GFP mRNAs with 0CUG in a strain expressing MBL-1. Thus, as in other organisms, MBL-1 interacts in vivo with expanded CUG transcripts in C. elegans, and MBL-1 association with expanded CUG repeat transcripts decreases mRNA export to the cytoplasm and translation. Down-regulation of mbl-1 by RNAi did not disrupt or enhance 123CUG transcript foci accumulation (Supplementary Fig. 2F). MBL-1 down-regulation, as we show below, can affect the levels of expanded CUG transcript available for translation. These data suggested that additional regulatory factors contribute to expanded CUG foci accumulation and toxicity. Taken together our results support the hypothesis that the RNA aggregated transcripts identified by SM-FISH correspond to the key foci characteristic of DM.
Screen for modifiers of expanded CUG-mediated toxicity
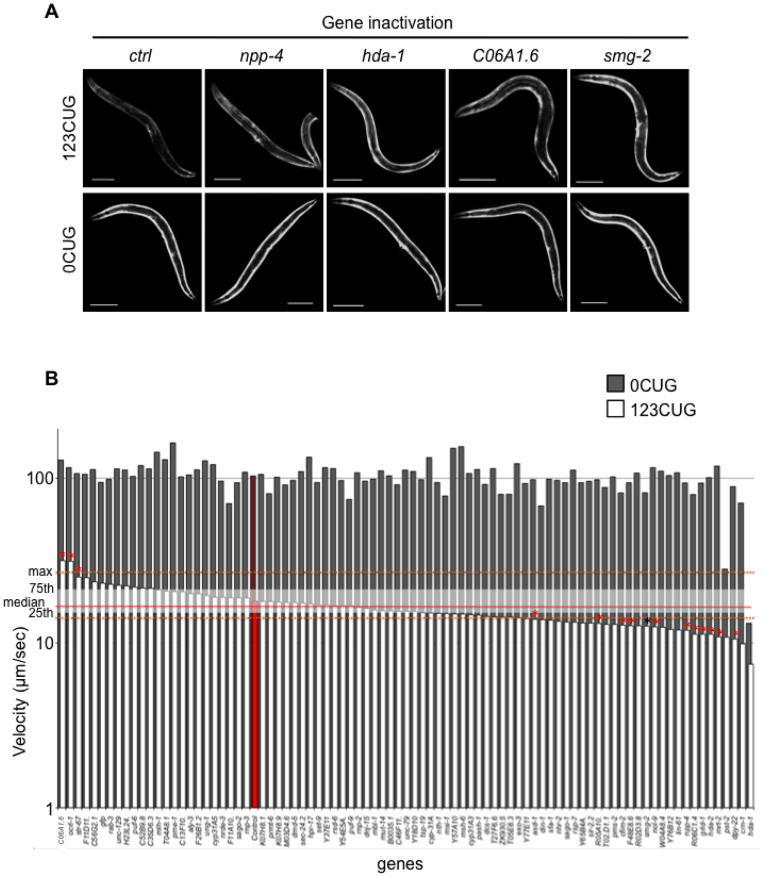
To identify genes that mediate expanded CUG repeat RNA pathogenesis, we used RNAi to reveal gene inactivations that can modify expanded CUG repeat RNA toxicity. A two-step screen was performed, with an initial fluorescent-based RNAi screen, followed by a secondary motility-based screen on hits from the primary screen (Supplementary Fig. 3A). For the fluorescent-based screen, we assayed for gene inactivations that disrupt the late stage down-regulation of GFP fluorescence specific to the 123CUG strain. We screened an RNAi library of 403 clones targeting genes that encode RNA-binding proteins and factors implicated in small RNA pathways20. This type of sub-library was expected to have a high representation of genes involved in expanded CUG repeat toxicity. Of the 403 genes tested, after re-screening in triplicate, 84 gene inactivations were selected that induced an increase in late developmental stage GFP fluorescence specifically in the 123CUG strain without affecting the control 0CUG strain (Figure 2A, Supplementary Fig. 3B, Supplementary Table 1). We tested each of the 84 gene inactivations identified for their ability to modulate the motility defect observed in 123CUG animals (see Online Methods). The 123CUG animals on the control RNAi showed a severe loss in motility, with a median velocity of ≈17μm/sec, compared to the 0CUG strain on the same control RNAi at ≈100μm/sec (Fig. 2B) similar to wild type animals. We identified 14 gene inactivations that significantly (p<0.01 using the two-sample Kolmogorov-Smirnov test) increased or decreased the velocity of 123CUG animals without affecting the control (0CUG) animals (Fig. 2B, Table 1).
Figure 2.
Identification of gene inactivation that modulate expanded CUG repeat toxicity (A) Gene inactivations that disrupt the late stage down-regulation of GFP fluorescence mediated by 123 CUG repeats in the 3′ UTR. Fluorescent microscopy images of the strains 123CUG and the control 0CUG, on different RNAi gene inactivations: empty vector control (ctrl), npp-4, hda-1, C06A1.6 and smg-2. Images were taken at the 3d old adult stage. Bar, 200μm. (B) Genetic suppressors and enhancers of expanded CUG repeat toxicity. Graph of velocity measurements of 0CUG (grey) and 123CUG (white) animals fed on different gene inactivations. The plotted velocities (μm/sec) correspond to the median of at least two experiments, where the red bars correspond to strains fed on control vector. Red line indicates the median velocity, and white shading represents the 25th and 75th percentile for the 123CUG animals fed on control vector. The dotted orange line represents the maximum and minimum of the median velocity for 123CUG animals fed on control vector. Indicated by red asterisk (*) are the significant gene inactivations, where significance was determined by Kolmogorov-Smirnov p-value. The black asterisk indicates the gene smg-2.
Table 1.
Modifiers of expanded CUG toxicity
| Gene inactivation | Gene | Molecular Function | Class | Human ortholog | Motility | Relative Velocity as a percentage of 123CUG on ctrl vector | RNA foci relative to 123CUG alone | |
|---|---|---|---|---|---|---|---|---|
|
|
||||||||
| Toxicity Enhancer | K10C9.6 | str-67 | G-protein coupled receptor | Signaling | OR4F5 | improved | 148 | decrease |
| C16C2.3 | ocrl-1 | inositol-1,4,5-triphosphate 5-phosphatase | Signaling | OCRL | improved | 184 | mild decrease | |
| C06A1.6 | uncharacterized | Cytoskeleton homology | KRTAP5-7 | improved | 187 | decrease | ||
|
|
||||||||
| Toxicity Suppressor | R05A10.1 | uncharacterized | Signaling | ADCY4 | worsened | 78 | increase | |
| K09B11.2 | nol-9 | polynucleotide 5′hydroxyl-kinase (nucleolar protein) | RNA Processing | NOL9 | worsened | 74 | increase | |
| Y48G8AL.6 | smg-2 | helicase | RNA Processing and Degradation | UPF1 | worsened | 75 | increase | |
| Y54E5A.4 | npp-4 | nuclear pore complex protein | RNA Transport | NUPL1 | worsened | 70 | increase | |
| R74.5 | asd-1 | alternative splicing family member | RNA Processing | FOX2 | worsened | 82 | mild increase | |
| F47A4.2 | dpy-22 | mediator complex subunit transcriptional mediator of RNA | Transcription | MED12L | worsened | 62 | mild increase | |
| C08B11.2 | hda-2 | histone deacetylase | Transcription | HDAC1 | worsened | 66 | decrease | |
| Y41C4A.14 | mrt-2 | conserved DNA-damage checkpoint protein | DNA Repair and Recombination | RAD1 | worsened | 65 | decrease | |
| F29C4.7 | grld-1 | RNA-binding protein (splicing) | RNA Processing | RBM15B | worsened | 66 | mild decrease | |
| R06C1.4 | uncharacterized | RNA Processing and Degradation; Translation | CSTF2T | worsened | 66 | mild decrease | ||
| D1046.1 | cfim-2 | cleavage and polyadenylation factor | RNA Processing and Degradation | CPFS7 | worsened | 76 | no change | |
| F48E8.6 | ribonuclease | RNA processing and Degradation | DIS3L2 | worsened | 75 | no change | ||
The list of genetic modifiers of expanded CUG toxicity identified can be categorized into the following three major classes: genes involved in transcription, signaling, and RNA processing and degradation (Table 1). Some of the genes identified had been previously implicated in polyglutamine (polyQ) repeat disorders: the hda-2, mrt-2 and smg-2 genes21–23, corresponding to a histone deacetylase, a RAD1 911 complex DNA damage checkpoint protein, and a RNA helicase part of the nonsense-mediated decay pathway, respectively. smg-2 was included in the final list as an additional gene inactivation that affected both the 123CUG repeat transgene and the 0CUG control transgene; smg-2 gene inactivation caused a mild decrease in motility of the 0CUG strain, but caused a much stronger loss of motility for 123CUG repeat strain and was the strongest hit from the fluorescent screen for suppression of the 123CUG-specific decline in GFP fluorescence. The identification in our screen of common regulators of expanded repeat diseases supports the view that repeat-associated disorders, where repeats occur in either coding or non-coding regions, share several protein cofactors.
CUG toxicity modulators affect nuclear foci accumulation
We examined whether any of the 15 gene inactivations that modulated expanded CUG repeat toxicity changed RNA foci accumulation of 123CUG transcripts. One prediction is that gene inactivations that improve the motility of animals expressing 123CUG RNAs would also cause a decrease in foci size or number and similarly, gene inactivations that caused further motility impairment would lead to an increase in foci size or number (Table 1). Of the 15 genes identified, inactivation of ocrl-1/inositol-1,4,5-triphosphate 5-phosphatase, str-67/GPCR chemoreceptor and C06A1.6, led to an improvement of motility in strains expressing 123CUG repeats in muscle (Table 1). Examination of GFP mRNA localization by SM-FISH in 123CUG muscles revealed a significant reduction in the number of nuclear foci when these three genes are inactivated (Fig. 3A, B, Supplementary Fig. 4, 5). The suppression of 123CUG foci was particularly striking for C06A1.6 gene inactivation, where 123CUG foci were now few and small, with SM-FISH signals close to 0CUG control levels (Fig. 3A, Supplementary Fig. 4). However, distribution of expanded RNA ‘single’ transcripts was still observed preferentially in the nucleus versus the cytoplasm for all 3 gene inactivations, suggesting a role for ocrl-1, str-67 and C06A1.6 in foci formation rather than in cellular distribution of RNA. No significant changes in RNA localization, and no foci accumulation, were found in the control 0CUG strain, when these 3 genes were inactivated (Fig. 3A, Supplementary Fig. 4, 5). Together, these data support a model in which ocrl-1, str-67 and C06A1.6 gene activities normally enhance the toxicity of expanded CUG repeats by contributing to 123CUG foci formation, and inactivation of these genes results in decreased toxicity.
Figure 3.
Suppressors and enhancers of expanded CUG toxicity affect nuclear foci. (A) Confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue) and merge of C. elegans muscle cells. Shown are 123CUG and the 0CUG control in different RNAi gene inactivations: empty vector control (ctrl), C06A1.6 and npp-4. Yellow arrows indicate expanded CUG nuclear foci. (B) Computational analysis of SM-FISH images of 123CUG animals with different gene inactivations and control (ctrl). Results are plotted as bar graphs were gene inactivations corresponding to bars on the right of the control exhibit an increase in detected foci area (in red), and conversely bars on the left of the control exhibit a decrease in foci area (in green), relative to the control. The cfim-2 and F48E8.6 gene inactivations are similar to ctrl. (C) C. elegans muscle cells confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue), merge of GFP RNA and nucleus images, and mCherry translational fusion protein. Strains imaged are 123CUG and 0CUG animals, in a mCHERRY (control) or NPP-4::mCHERRY backgrounds. Yellow arrows indicate expanded CUG nuclear foci.
For the 12 gene inactivations that further reduced motility in 123CUG animals, six gene inactivations caused an increase in foci size present in the nucleus of 123CUG body wall muscle cells. These genes are npp-4/nuclear pore complex protein, asd-1/alternative splicing regulator, smg-2/nonsense-mediated decay (NMD) factor, nol-9/polynucleotide 5′-hydroxyl-kinase, dpy-22/transcriptional mediator protein and R05A10.1 (Fig. 3, Table 1, Supplementary Fig. 4, 5). For some genes, such as npp-4, a change in RNA localization was observed, with transcript enrichment in the nucleus relative to the cytoplasm (Supplementary Fig. 4, 5). For all these genes, except smg-2, no significant changes in transcript distribution were observed for the control 0CUG mRNA. smg-2 gene inactivation in the control 0CUG led to a slight increase in transcript signal, in both the nucleus and cytoplasm, without affecting nuclear to cytoplasm RNA distribution or leading to foci formation. Inactivation of the other 6 genes either caused a reduction in foci sizes or did not cause a significant change in aggregate size or number (Table 1). The reduction of foci number associated with an increase in toxicity suggested that, in certain conditions, the accumulation of non-aggregated CUG-expanded RNAs can be a major contributor of cellular dysfunction. Similar to what was previously suggested24, these ‘free’ toxic RNAs would have the potential to affect the activity of a wider range of RNA-binding proteins than when in an ‘aggregated’ state.
To further establish that the genes identified were involved in the regulation of expanded CUG-mediated toxicity, we overexpressed npp-4/nuclear pore complex component and asd-1/alternative splicing regulator as mCherry fusion proteins in body wall muscle cells in C. elegans. Down-regulation of npp-4 and asd-1 by RNAi caused an increase in nuclear expanded CUG RNA foci sizes (Table 1). C. elegans expressing these proteins fused to the fluorophore mCherry in either 123CUG or control 0CUG backgrounds, were analyzed by SM-FISH for a change in accumulation of 123CUG RNA in nuclear foci. Overexpressing either of these genes led to a decrease in foci number in a 123CUG background relative to the 123CUG parental strain (Supplementary Fig. 6A). In contrast, overexpression of these proteins in the 0CUG strain had no effect on GFP mRNA transcript distribution. Expression of mCherry alone (Fig. 1F), or a different protein, such as RNP-2, had no effect on 123CUG foci size or number (Supplementary Fig. 6A). Thus some of the genes identified are dosage sensitive components of the CUG repeat toxicity pathway.
Nonsense-mediated decay targets 3′ UTRs with CUG repeats
smg-2 RNAi in 123CUG animals caused an increase in nuclear RNA foci sizes, an increase in muscle cell toxicity with loss of motility and increase in GFP fluorescence signal relative to the control. smg-2 gene inactivation on control 0CUG strains had no effect on nuclear foci, and the mild increase in toxicity detected was not comparable to that observed in the 123CUG strain. In addition, smg-2 acts as a common regulator of expanded repeat-containing disorders by also suppressing protein aggregation caused by expanded CAG repeats in the coding regions of the Huntingtin gene, associated to Huntington’s disease21.
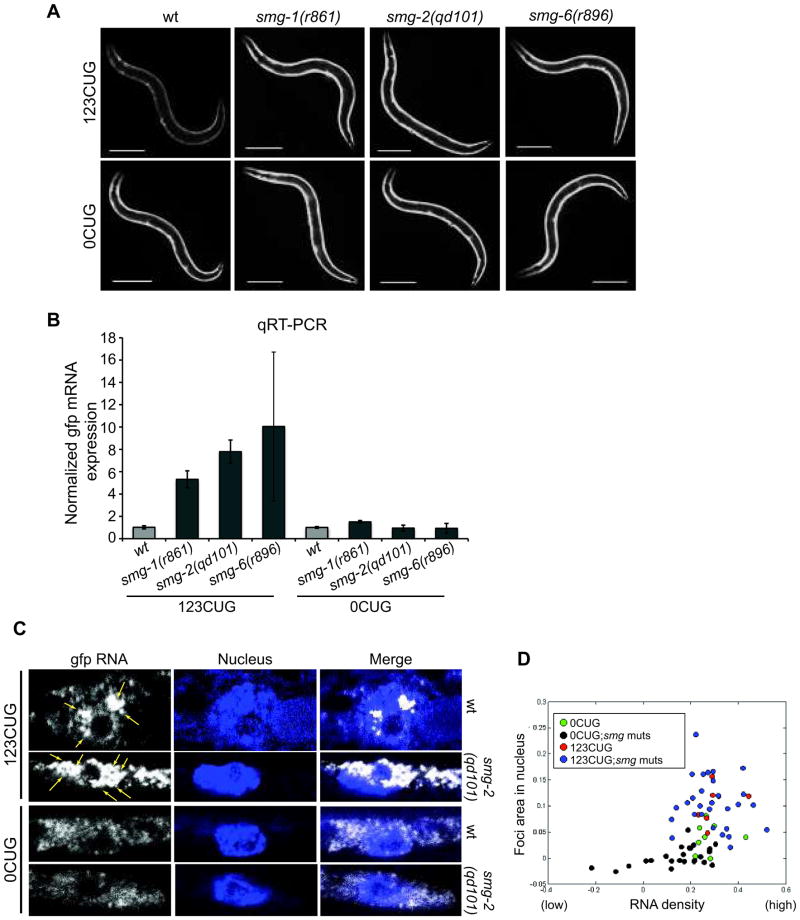
smg-2 encodes an RNA helicase and is a conserved component of the nonsense-mediated mRNA decay (NMD) pathway. The NMD pathway is an evolutionary conserved surveillance mechanism that detects mRNAs containing premature stop codons, preventing toxic expression of truncated proteins25. The identification of smg-2 as a modulator of expanded CUG toxicity suggested that the NMD pathway may recognize and target for degradation RNA transcripts with expanded CUG repeats, even in the 3′ UTRs of non-truncated open reading frames. We analyzed the effects of mutations in NMD components on GFP transcripts bearing 123CUG repeats or control 0CUG in muscle cells using smg-1(r861), smg-2(qd101) and smg-6(r896) mutants. We observed that 123CUG animals in the background of any of the smg mutants showed a strong increase in GFP fluorescence signal relative to the parental strain (Fig. 4A). No such change in fluorescence was observed for the control 0CUG animals (Fig. 4A). Quantitative RT-PCR showed that, mRNA levels of gfp bearing 123CUG repeats were increased by several fold: ≈5.3 fold in smg-1(r861), ≈7.8 fold in smg-2(qd101) and ≈10.1 fold in smg-6(r896) backgrounds, compared to wild type (Fig. 4B). However, no significant change was observed in the levels of gfp mRNA without any CUG repeats in the 3′UTR in the different smg mutant backgrounds compared to the wild type (Fig. 4B). Thus the NMD pathway targets the mRNA transcripts containing the expanded CUG repeats for degradation.
Figure 4.
The NMD pathway modulates expanded CUG transcripts degradation and nuclear foci accumulation. (A) Fluorescent microscopy images of 2d old adult animals expressing either 123CUG repeats or 0CUG in the backgrounds: wild type (wt), smg-2(qd101), smg-1(r861) and smg-6(r896). Scale bars correspond to 200μm. (B) qRT-PCR assay for gfp levels in animals expressing either 123CUG repeats or the control GFP in different backgrounds: wild type (wt), smg-2(qd101), smg-1(r861) and smg-6(r896). Wild type=1.0. Error bars represent SEM for three biological replicates. (C) Confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue) and merge of C. elegans muscle cells. The strains imaged are 123CUG and 0CUG animals, in wild type (wt) and smg-2(qd101). Yellow arrows indicate expanded CUG nuclear foci. (D) Computational analysis of SM-FISH images of 0CUG (green dots), 0CUG in smg mutant backgrounds (blue dots), 123CUG (red dots) and 123CUG in smg mutant backgrounds (black dots).
We analyzed by SM-FISH and computational image analysis the gfp RNA transcript accumulation in 123CUG and the control 0CUG strains in the different smg mutant backgrounds. Disruption of NMD pathway in 123CUG animals caused an increase in foci size and number in the nucleus (Fig. 4C, Supplementary Fig. 6B) and in most cells the accumulation of foci-like structures in the cytoplasm as well (Fig. 4C, D, Supplementary Fig. 6B). Conversely, in the smg mutant animals expressing the control 0CUG we observed a uniform distribution of RNA transcripts with a large number present preferentially in the cytoplasm (Fig. 4C, and Supplementary Fig. 6B). Thus the NMD pathway recognizes RNA transcripts containing expanded CUG repeats and disruptions in NMD cause the accumulation of expanded CUG toxic RNA species in the nucleus, leading to cellular dysfunction (Supplementary Fig. 8A).
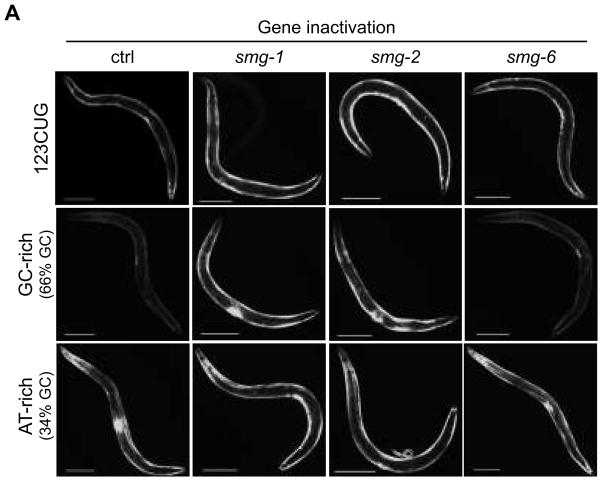
To examine whether the skewed sequence composition of expanded CUG repeat sequences targets them for the NMD pathway, we explored the influence of GC composition on NMD. 3′UTRs are typically A/U rich (≈65–70% AT-rich), exhibiting a nucleotide composition distinct from coding (≈50–55% AT-rich) or intergenic regions26,27. The let-858 3′ UTR to which the 123 CUG repeat was added is 384 nucleotides and 30% GC. The added CUG repeat elements are rich in G and C nucleotides (≈66%) that may contribute to the recognition by the NMD pathway. We generated expression plasmids in which the 3′UTR (CTG)n sequence was substituted by a non-repeat sequence with either a 66% or 34% GC nucleotide content (Supplementary Fig. 7A, B). The DNA sequences used were cloned from non-C. elegans organisms or from entirely synthetic nucleotide sequences bearing similar GC percentages to avoid a possible recognition of endogenous signal sequences (Supplementary Notes). GFP reporter genes bearing GC-rich 3′ UTR elements from non-C. elegans organisms exhibited weaker GFP fluorescence, or no fluorescence at all in the case of synthetic sequences, compared to those bearing the corresponding AT-rich elements (Fig. 5, Supplementary Fig. 7B). Strains expressing GC-rich elements from a non-C. elegans genome placed in the 3′UTR of the GFP reporter gene showed a significant increase in fluorescence when either smg-1 or smg-2 were inactivated by RNAi, whereas no change in GFP intensity was detected for AT-rich (Fig. 5, Supplementary Fig. 7A, B). Fusion genes engineered with synthetic, random high GC percentage sequences showed a stronger increase in fluorescence in the smg-2 background relative to two regulators of smg-2 phosphorylation smg-1 or smg-6 (Supplementary Fig. 7A, B). These data demonstrate that the results observed for the GC-rich versus AT-rich sequences were not due to a sequence-specific endogenous 3′ UTR identity signal present in the sequence used. These results further establish that the increase in distance between the stop codon and the polyA signal due to the addition of the CUG repeat sequence does not contribute to NMD recognition, since no repression was observed for AT-rich transcripts. These data support a model in which mRNAs, containing CUG repeats in their 3′UTR, are NMD substrates. Furthermore, it reveals that the NMD recognition of CUG-containing mRNA is dependent on nucleotide composition, either due to the presence of a GC-rich sequence in a region usually A/U-rich, or due to the formation of specific secondary structures associated to the presence of these nucleotides. While both the GC-rich 3′ UTR element and the 123CUG repeat element reporter genes are responsive to disruption of the NMD pathway, none of the 15 gene inactivations that strongly disable 123CUG repeat repression in muscle disrupt the repression conferred by GC-rich element. Thus, the detection and localization to foci of 123CUG repeats by these genes is distinct from the detection and degradation of GC rich elements by the NMD system.
Figure 5.
3′UTR CUG repeat sequence composition triggers NMD recognition for degradation. Fluorescent microscopy images of the strains 123CUG, GC-rich and AT-rich, in different RNAi gene inactivations: empty vector control (ctrl), smg-1, smg-2, and smg-6. Images of 3d old adult animals. Bar, 200μm.
To establish whether NMD recognition of expanded CUG repeats is a conserved cellular mechanism, we analyzed the nuclear RNA foci phenotype of NMD gene inactivations in human DM1 patient fibroblast cells expressing 2000 CUG repeats in the DMPK1 mRNA, as well as in control fibroblasts expressing a DMPK1 mRNA with 7 to 35 such CUG repeats. We tested for changes in foci number when the human orthologue of smg-2, UPF1, was inactivated by RNAi. We used SM-FISH for RNA foci detection, with 5 probes complementary to the CUG repeat region and 23 probes complementary to the last three exons of DMPK1 which are not composed of CUG repeats. UPF1 was down-regulated using siRNAs in DM1 and in normal fibroblasts and these cells were analyzed by SM-FISH 24h post siRNA-transfection. For both control fibroblasts and fibroblasts isolated from DM1 patients, UPF1 siRNAs decreased UPF1 protein levels by 35%–40% compared to scrambled siRNAs (Supplementary Fig. 7C, D). There was lower cell recovery after UPF1 knockdown, suggesting that knockdown of NMD components may cause a loss of cell viability, deflating the measured level of UPF1 knockdown. But even with the modest UPF1 knockdown, SM-FISH analysis revealed an increase in the number of nuclear foci in DM1 cells treated with UPF1 siRNAs compared to untreated DM1 cells or DM1 cells treated with mock siRNAs (Fig. 6A). In contrast, normal fibroblast cells bearing just a few CUG repeats in the DMPK gene exhibited no nuclear foci in both untreated or treated with UPF1 siRNAs (Fig. 6A). The number of foci present in the DM1 cells was quantified (see Online Methods) and UPF1 down-regulation caused a significant increase in the percentage of cells containing a higher number of foci (Fig. 6B). Our data supports a conserved role for NMD in the identification of transcripts bearing GC-rich sequences in their 3′UTR. Furthermore, our results support the function of NMD as an important element in the toxicity of expanded CUG repeat transcripts in myotonic dystrophy 1.
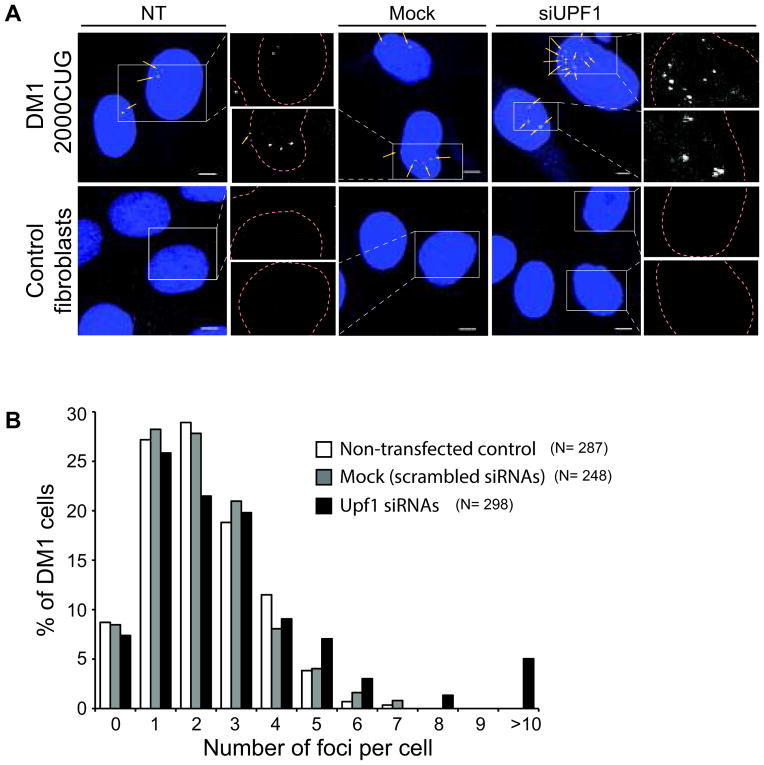
Figure 6.
NMD downregulation causes an increase in CUG repeat mRNA foci number in myotonic dystrophy 1 patient fibroblast cells. (A) SM-FISH of DM1-affected or normal human fibroblast cells in which UPF1 was downregulated relative to control non-transfected or transfected with scrambled siRNAs (mock) cells. The DM1 human fibroblast cell line used expressed the gene dmpk bearing 2000CUG in its 3′UTR. (B) Histogram represents the distribution of the number of foci in DM1 cells that were downregulated for UPF1, mock and non-transfected controls. UPF1 downregulation led to a significant increase in the number of nuclear foci present relative to mock (p<0.0001) and non-transfected cells (p<0.00003), using t-student test. N indicates the total number of cells analyzed. Two independent experiments were performed. Bar, 5μm.
Discussion
The gene inactivations that modulate phenotypes of expanded CUG RNA repeats comprise multiple pathways, beyond splicing dysregulation. We demonstrate the involvement of a number of previously unknown genes as modulators of expanded CUG toxicity and expanded CUG repeat foci formation. The demonstration that different gene inactivations, all expanded CUG repeat toxicity suppressors, have opposing effects on foci accumulation (Table 1, Supplementary Fig. 8B), supports the hypothesis that these genes act in distinct pathways. Genes where a direct correlation exists between expanded CUG repeat toxicity and foci accumulation (Supplementary Fig. 8B) include genes where modulation of expanded RNA toxicity can occur by: clearance of CUG-containing RNA transcripts, binding of expanded CUG RNA preventing foci formation or promotion of mRNA transport from the nucleus. Inactivation of these genes causes an increase in the toxic expanded CUG species present in the nucleus. One example is smg-2/NMD helicase inactivation. Another class of suppressor gene inactivations do not correlate with an increase in foci formation (Supplementary Fig. 8B); these proteins may detect cellular damage or bind to expanded CUG repeats.
The identification in our screen of additional splicing factors, such as the asd-1 and grld-1 genes, that when inactivated caused an increase in expanded CUG toxicity was reasonable (Table 1). Unlike MBL1 overexpression (Fig. 1F, Supplementary Fig. 2D), ASD-1 overexpression led to a decrease in expanded CUG nuclear foci accumulation (Supplementary Fig. 6). ASD-1 is an alternative splicing factor and belongs to the Fox-1 splicing family. In vertebrates, MBNL genes are silenced by Fox-1/2 splicing factors28. Two mechanisms for ASD-1 suppression of expanded CUG repeat toxicity emerge: 1) ASD-1 regulates functional MBNL1 levels available by modulating splicing variants; 2) ASD-1 may bind directly or indirectly to expanded CUG repeats and affect toxicity.
Most of the gene inactivations identified make the response to expanded CUG repeats more toxic and promote the accumulation of larger RNA foci in the nuclei, suggesting that these genes constitute a CUG repeat detoxification pathway that blunts their toxicity.
Commonalities have been suggested in degenerative pathways between repeat-based RNA-mediated disorders, and protein-mediated disorders1. RNA toxicity has been implicated in polyQ expansion disorders, and MBNL1 functions as a modulator of polyQ toxicity through its interaction with CAG-containing RNA transcripts29. A subset of the genes identified in our screen as modifiers of expanded CUG toxicity are modulators of polyQ aggregation or toxicity, hda-2, mrt-2 and smg-2 genes21,23. npp-4, although not previously linked to repeat expansion disorders, is part of the nuclear pore complex together with npp-8, and npp-8 had been identified as a modulator of polyQ aggregation30. The identification of pathways that function as common regulators to a broad class of triplet nucleotide pathogenic expansions supports the model of common toxic mechanisms for coding and non-coding triplet repeat disorders.
The NMD pathway is a conserved mechanism of mRNA surveillance that regulates the expression of 5–10% of the human, D. melanogaster and yeast transcriptomes31,32. We find that in addition to its expected target transcripts, NMD modulates the abundance of transcripts containing CUG repeats in their 3′UTR, reducing the accumulation and nuclear foci formation of these toxic RNA species (Supplementary Fig. 8) in both C. elegans and human cells (Fig. 4, 6 and Supplementary Fig. 6B). Sequence composition is key in the recognition by NMD of RNA transcripts containing 3′UTR CUGs; a similar G/C-rich (≈66%) sequence, when present in the 3′UTR, is also recognized by NMD, whereas an A/T-rich sequence is not.
With the identification of NMD genes as modulators of expanded CAG repeat protein-based disorders21, our results suggest broader surveillance roles for the NMD pathway. RNA transcripts containing expanded CAG repeats, also GC-rich, are likely to form secondary structures that may directly or indirectly trigger the NMD pathway33. Additionally, NMD has been mapped to nuclear surveillance leading to nuclear RNA degradation as well as cytoplasmic degradation. Our data showing a striking accumulation of nuclear RNA foci and cytoplasmic RNA foci in NMD mutants suggests a role for NMD not only in the cytoplasm but also in nuclear clearance of expanded RNA repeat transcripts.
Modulation of the NMD pathway may offer a therapeutic approach for myotonic dystrophy patients as well as other repeat-based degenerative disorders. Pharmacological compounds that increase NMD pathway activity may clear CUG-containing RNA toxic species, with the potential to significantly ameliorate DM-related symptoms. A comparable approach, applied to distinct disorders, is currently being tested in clinical trials using compounds that promote NMD read-through34. NMD efficiency varies across tissues35 and between individuals36,37, with significant clinical implications38. These variations in NMD efficiency may have significant implications for trinucleotide repeat disease onset or progression.
Online Methods
Plasmids and Constructs
Mammalian CTG repeat sequences were amplified from plasmids pR26eGFP+100 and pR26eGFP+20039 using Extended High Fidelity from Roche in 6% DMSO and 1M betaine (Sigma). CTG repeats were cloned into the C. elegans pPD118.20 vector bearing the myo-3 body wall muscle-specific promoter, GFP, and the let-858 3′ UTR. The mbl-1 and rnp-2 genes were amplified from C. elegans N2 genomic DNA, and asd-1 and npp-4 from cDNA, using Phusion polymerase (Finnzymes). These genes were cloned into the C. elegans vectors pPD49.26 and pPD30.38 (Addgene) bearing the unc-54 body wall muscle-specific promoter. The GC-rich and AT-rich nucleotide sequences were cloned from the coding region of the 1,4-alpha-glucan branching enzyme gene of Pseudomonas aeruginosa (glgB) and the 3′utr region of the Arabidopsis thaliana myb domain protein 51 gene (myb51), respectively. The synthetic GC-rich and AT rich sequences were synthesized (GenScript). The GC-rich and AT-rich sequences were amplified and cloned into the C. elegans pPD118.20 (Addgene) vector bearing the myo-3 body wall muscle-specific promoter. Detailed cloning information, including primers used, is indicated in Supplementary Notes.
C. elegans Strains
Nematodes were handled using standard methods40 and experiments were performed at 20°C, unless otherwise indicated. The C. elegans N2 Bristol strain was used as wild-type strain. Strains generated for this study are indicated in Supplementary Table 2. Transgenes containing gfp fused to different CTG lengths were integrated by exposing animals to UV irradiation and strains were outcrossed 5 times. Several independent strains were obtained carrying the different GFP transgenes and the different strains generated exhibited similar length-dependent phenotypes, as described in the Results section. The remaining transgenic strains expressed their transgenes as extrachromosomal arrays.
RNA Fluorescence In Situ Hybridization (RNA FISH)
Oligonucleotide probes were designed and SM-FISH was performed as described in Raj et al41. SM-FISH was performed in 3d adult animals, and in human fibroblast cells 24h post siRNA transfection, using probes synthesized by BioSearch Technologies. Two probe sets were used for C. elegans samples, each with thirty-four probes complementary to gfp. One set of probes used was labeled with the dye CAL Fluor Red 590, and the other set with Quasar 670. A distinct probe set was used for the fibroblast cell samples, comprised of twenty-eight probes, labeled with the CAL Fluor Red 590 dye and targeting the CUG repeat region and the 3′ region of the dmpk mammalian gene (see Supplementary Notes). DAPI was used for nuclear staining and SM-FISH images were collected with an Olympus FV-1000 confocal microscope with an Olympus PlanApo 60 3 Oil 1.45 NA objective at 4 zoom, and a 559 nm (mCherry/CALFluor probe), 635nm (Quasar probe) and 405nm (DAPI) diode laser.
SM-FISH computational image analysis
To analyze SM-FISH images, an algorithm was developed to quantify the RNA intensity pixel by pixel in the image. Based on its intensity, each pixel was categorized into one of three RNA populations present in the cell: ‘single’ RNAs (low RNA density), several RNA transcripts (high RNA density), and RNA foci structures (Supplementary Fig. 1E). Pixel intensity corresponding to fluorescence intensity correlates with the number of RNA transcripts present41. DAPI staining was used to identify the nucleus in each cell. Because the accumulation of foci in DM is characterized by its nuclear localization (asymmetric cellular foci distribution), we used the cytoplasmic region in each image to normalize for variations in staining. This approach would allow also the detection of changes in nuclear foci accumulation. This algorithm allowed us to calculate for each nucleus the percent of foci (pixels) and of “high density RNA” (pixels) from the total pixel population. The data was plotted where each ‘dot’ represents a nucleus, with the Y axis representing the percentage of foci pixels and the X axis indicating the percentage of pixels with ‘high density’ RNA. For an example, see Supplementary Notes.
C. elegans Fluorescence Imaging
For in vivo imaging, animals were mounted on a 2% agar pad on a glass slide and immobilized in 1mg/ml levamisole (Sigma). Fluorescence imaging was done on a Zeiss AxioImager.Z1 Microscope.
RNAi screens
RNAi-mediated gene inactivation was by feeding42 in a 12-well plate RNAi bacterial culture 2x concentrated. Animals were synchronized by NaOCl bleaching and overnight hatching in M9. Twenty to thirty L1 larval stage animals (≈24h after synchronization) were aliquoted onto agar plates containing a 48h culture of RNAi bacteria expressing double-stranded RNA, and allowed to develop to adulthood. The drug 5-fluorodexoyuridine was added at the L4-larval stage to a final concentration of 0.1mg/ml, to inhibit progeny production. Each 12-well plate contained the empty L4440 control vector as a negative control. Animals were analyzed either as 3d and as 4d old adults for the GFP fluorescence screen, or at 2d old adults for the locomotion-based toxicity screen. The RNAi clones identified as positives from the screen were verified by sequencing of the insert. Additional information on the RNAi screens is provided in the Supplementary Notes.
C. elegans locomotion assays
The locomotion assay on plates with a ring of OP50 food attractant was performed as previously described43. The percentage of age-synchronized animals that reached the OP50 food in 90min was determined. The second locomotion assay, with analysis of animal velocity, was performed at room temperature and off food. Each experiment performed contained a control corresponding to 123CUG and 0CUG animals fed on control vector (L4440). The locomotion behavior was recorded on a Zeiss Discovery Stereomicroscope using Axiovision software. The center of mass was recorded for each animal on each video frame using object-tracking software in Axiovision. Imaging began 30min after animals were removed from food and recordings were 30sec long. For each assay, 20–45 2d old age-synchronized animals were recorded. The motility data was analyzed using the two-sample Kolmogorov-Smirnov test to compare the distributions of the values in the two data vectors x1 and x2. The null hypothesis is that x1 and x2 are from the same continuous distribution. This test was applied in two different ways 1) using the median velocities of all experiments obtained from all the 123CUG or 0CUG animals fed on control vector and 2) using the experimental internal control corresponding to the median velocity of the 123CUG or 0CUG on control vector. RNAi clones were only considered positive if strongly significant on both analyzes.
qRT-PCR
Total RNA was isolated from synchronized 2d old C. elegans adults using Trizol (Invitrogen) followed by chloroform extraction and isopropanol precipitation. Samples were DNase treated with Turbo DNA-free (Invitrogen) and cDNA was synthesized from 1μg total RNA using Retroscript (Invitrogen). Quantitative RT-PCR assays of mRNA (SYBR Green, Bio-Rad) levels were done according to Bio-Rad recommendations. Three independent biological samples were used for all strains analyzed for gfp levels, and we used rpl-32 levels for normalization across samples. The 2−ΔΔct method was used for comparing relative levels of mRNAs. Primers are listed as Supplementary information.
Protein blot assays
Proteins were extracted from synchronized animals and actin levels were used for normalization across samples. Three independent biological samples were used for all strains analyzed. Harvested C. elegans samples were boiled for 10min in Laemmli buffer, spun and the supernatant collected. Proteins were resolved on 4–12% Bis-Tris SDS polyacrilamide gels, transferred to nitrocellulose membranes and probed with GFP and actin antibodies (Roche, Cat#11814460001; Abcam, ab3280). Protein levels were quantified on a Typhoon phosphoimager using the ImageQuant TL software (GE Healthcare Life Sciences). p values were calculated using Student’s t test.
Mammalian Cell Culture
Human lymphoblast cell lines were obtained from the Coriell Cell repository corresponding to cells from unaffected individuals (GM07492) and fibroblast from DM1-affected individuals (GM03989). Cells were maintained in high glucose EMEM (Lonza) supplemented with 15% fetal bovine serum, 1x antibiotic-antimycotic (Gibco) and 1x non-essential amino acids solution (Sigma), at 37°C, 5% CO2.
siRNA knockdown of UPF1 in human cells
Fibroblast cells were transfected with UPF1 ON-TARGETplus SMARTpool siRNA (Thermo Scientific, cat. No. J-011763-05), or nontargeting siRNA as control (Thermo Scientific, cat. No. D-001810-01) for 24h, using Lipofectamine RNAiMAX (Invitrogen) according to the manufacturer’s protocol. The final siRNA concentration used was 100nM. Cells were fixed after transfection for analysis by FISH as described in Raj et al41 (see Supplementary Notes). Knockdown efficiency was monitored by Western Blotting with a UPF1 (kindly provided by Dr. Lykke-Andersen) and GAPDH specific antibodies.
Foci quantification in human fibroblasts
Nuclear foci in DM1-affeceted fibroblasts were quantified using the CellProfiler software (http://www.cellprofiler.org), and specifically a script in CellProfiler, “Speckle Counting’ that allows the identification of individual cells, their nuclei, together with the number of foci present (see Supplementary Notes). The percentage of DM1 cells containing different numbers of nuclear foci was plotted and the p value calculated using two sample t-test function in the Matlab package.
Supplementary Material
Supplementary Fig. 1. C. elegans expressing expanded CUG repeats exhibit locomotion defects. (A) Representation of motility assays performed using agar plates containing an E. coli food ring. The food ring had a 2cm radius. (B) Motility assays for 2d adults. Data plotted corresponds to the average percentage of population to reach the food at each time point. Error bars represent SD from at least 3 independent experiments; in each experiment, 3–5 replicas of ca. 100–150 animals were analyzed. (C–E) Computational analysis of SM-FISH images. (C) Analysis starts with computational identification of the nuclear region based on DAPI staining in an SM-FISH image of a 123CUG animal. Following nucleus identification, (D) there is computational delineation of cytoplasmic versus nuclear spaces in the SM-FISH image corresponding to the GFP RNA transcript probes. (E) Analysis of pixel intensities for each SM-FISH image, corresponding to low RNA, high RNA densities and RNA foci in both the nucleus and cytoplasm.
Supplementary Fig. 2. Expression of MBL-1::mCherry in C. elegans muscle cells increases expanded CUG transcript recruitment and mutant transcript nuclear foci accumulation. Schematic drawing of the MBL-1::mCHERRY construct (A), and C. elegans body wall muscle cells (B). (C) MBL-1::mCHERRY exhibits a diffuse cellular distribution with nuclear accumulation. (D) C. elegans muscle cells confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue), merge of GFP RNA and nucleus images, and mCherry translational fusion protein. The muscle cells imaged correspond to animals expressing 123CUG repeats and 0CUG, in a mCHERRY (control) or MBL-1::mCHERRY backgrounds. Yellow arrows indicate expanded CUG nuclear foci. MBL-1::mCHERRY localizes to the nucleus. (E) Genetic mosaic analysis of GFP intensity shows that GFP fluorescence, from 123CUG mRNA transcripts, absent in cells expressing mbl-1::mCherry, relative to neighboring cells that fail to express mbl-1::mCherry. GFP fluorescence is not affected in the 0CUG control animals expressing mbl-1::mCherry. (F) Confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue) and merge of C. elegans muscle cells. The strains imaged were 123CUG and 0CUG in: empty vector control (ctrl) and mbl-1 gene inactivations. Yellow arrows indicate expanded CUG nuclear foci.
Supplementary Fig. 3. Screen approach for the identification of modulators of expanded CUG toxicity. (A) Representation of RNAi screen steps in the identification of modulators of expanded CUG repeat pathogenesis. (B) Fluorescent microscopy images of the strains 123CUG and the control 0CUG, on different RNAi gene inactivations: empty vector control (ctrl), mbl-1 and aly-3. Images were taken at the 3d old adult stage. Bar, 200μm.
Supplementary Fig. 4. Suppressors and enhancers of expanded CUG toxicity have distinct effects on expanded CUG nuclear foci accumulation. Confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue) and merge of C. elegans muscle cells. The strains imaged were 123CUG and the control 0CUG, in different RNAi gene inactivations: empty vector control (ctrl), C06A1.6, str-67, mrt-2, npp-4 and smg-2. Yellow arrows indicate expanded CUG nuclear foci.
Supplementary Fig. 5. Gene inactivations have different effects on foci accumulation in the nucleus. Computational analysis of SM-FISH images of 0CUG animals (yellow dots), control (green dots), and 123CUG animals fed different gene inactivations (blue dots) and control vector (red dots). Each ‘dot’ shown in the graph represents one analyzed SM-FISH image, corresponding to a single imaged cell. The red dotted square indicates the region of clustering of the samples corresponding to 123CUG animals on control vector (red dots). Labeled on the graph on the left, above the red box, are the gene inactivations (blue dots) that cause an increase in bright pixel intensity, corresponding to an increase in foci size or number, relative to the 123CUG on control. The ‘grouping’ of 123CUG npp-4 inactivations in the upper right corner of the graph indicates both an increase in nuclear foci and in nuclear ‘single’ transcript localization relative to the 0CUG npp-4 controls that localize further to the left in the graph. The inset section displayed shows gene inactivations (blue dots) that cause a decrease in bright pixel intensity, relative to the 123CUG on control vector, corresponding to a decrease in foci size or number.
Supplementary Fig. 6. (A) Over-expression of expanded CUG repeat suppressors causes a decrease in expanded CUG nuclear foci accumulation. C. elegans muscle cells confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue), merge of GFP RNA and nucleus images, and mCherry translational fusion protein. The strains imaged are animals expressing 123CUG repeats and 0CUG in the following transgenic backgrounds: mCHERRY, NPP-4::mCHERRY, ASD-1::mCHERRY and RNP-2::mCHERRY. RNP-2 corresponds to the U1 small nuclear ribonucleoprotein A, and RNP-2::mCherry exhibits nuclear localization in C. elegans muscle cells. (B) Mutants in the NMD pathway cause an increase in expanded CUG nuclear foci accumulation. Confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue) and merge of C. elegans muscle cells. The strains imaged were animals expressing 123CUG repeats and 0CUG, in the following backgrounds: wild type (wt), smg-1(r861) and smg-6(r896). Yellow arrows indicate expanded CUG nuclear foci.
Supplementary Fig. 7. (A–B) Sequence composition of CUG repeat sequences in the 3′UTR contributes to NMD transcript recognition for degradation. (A) Schematic drawing of the GC-rich or AT-rich plasmids for expressions in C. elegans muscle cells. (B) Fluorescent microscopy images of strains expressing a GFP with a 300bp ‘artificial’ insert in their 3′UTR containing the following GC percentages: 31%, 32%, 60% and 70%. Also included are the control strains containing 3′UTR inserts cloned from A. thaliana (34%GC) and P. aeruginosa (66%GC). These strains are shown in a wt background and in the background of the following smg mutants: smg-1(5861), smg-2(qd101) and smg-6(r896). The ‘fluorescence’ observed in the 60%GC and 70%GC strains in a wild type background corresponded to the characteristic gut autofluorescence, and no GFP signal was observed in the body wall muscle cells of these animals. Images were taken of animals at the L4 stage. Bar, 100μm. (C–D) Western blot analysis of UPF1 down-regulation (24h post-transfection) by siRNA pool of unaffected (C) and DM1 (D) fibroblast cells, using UPF1-specific antibody. Fibroblasts showed a decrease of 40% in UPF1 levels relative to cells transfected with scrambled siRNAs (mock cells) in both unaffected (C) as well as DM1 (D) cells. GAPDH levels were used for normalization across samples.
Supplementary Fig. 8. Model of regulation of expanded RNAs. (A) Model for regulation of expanded RNA toxicity by the NMD pathway: NMD targets expanded CUG repeat transcripts for degradation reducing the levels of toxic RNAs present in the cells. A decrease in NMD function results in accumulation of toxic transcripts with increase in nuclear RNA foci and increase in toxicity with loss of motility. (B) Model for regulation of expanded RNA foci accumulation by the modulators of RNA toxicity identified: different pathways regulate expanded CUG repeat toxicity; an increase in foci causes a decrease in locomotion however, a decrease in foci doesn’t necessarily correlate with a decrease in muscle toxicity.
Supplementary Table 1. Modifiers of expanded CUG-specific decrease in GFP fluorescence.
Acknowledgments
We thank Prof. M. Mahadevan for providing plasmids bearing the CUG repeats, S. Fischer and J. Kim for the RNAi library used in the screen and D. Kim for the smg-2(qd101) strain, and Lykke-Andersen for the UPF1 mammalian antibody. We are grateful to the J. Kaplan laboratory for the use of the Olympus FV-1000 confocal microscope, and the B. Seed lab for the use of their tissue culture facilities. We are also thankful to S. Djonovic and S. Choi for reagents and technical assistance; J. Bai for reagents, technical assistance and helpful discussions; A. Connery for help with the CellProfiler software and J. Urbach for help editing the manuscript. We thank the Caenorhabditis elegans Genetics Center for strains; the American Heart Association for funding S.M.D.A.G and NIH AG043184 for funding GR.
References
- 1.Todd PK, Paulson HL. RNA-mediated neurodegeneration in repeat expansion disorders. Ann Neurol. 2010;67:291–300. doi: 10.1002/ana.21948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Brook JD, et al. Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell. 1992;69:385. doi: 10.1016/0092-8674(92)90418-c. [DOI] [PubMed] [Google Scholar]
- 3.Taneja KL, McCurrach M, Schalling M, Housman D, Singer RH. Foci of trinucleotide repeat transcripts in nuclei of myotonic dystrophy cells and tissues. J Cell Biol. 1995;128:995–1002. doi: 10.1083/jcb.128.6.995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Tokgozoglu LS, et al. Cardiac involvement in a large kindred with myotonic dystrophy. Quantitative assessment and relation to size of CTG repeat expansion. JAMA. 1995;274:813–9. [PubMed] [Google Scholar]
- 5.Groh WJ, Lowe MR, Simmons Z, Bhakta D, Pascuzzi RM. Familial clustering of muscular and cardiac involvement in myotonic dystrophy type 1. Muscle Nerve. 2005;31:719–24. doi: 10.1002/mus.20310. [DOI] [PubMed] [Google Scholar]
- 6.Miller JW, et al. Recruitment of human muscleblind proteins to (CUG)(n) expansions associated with myotonic dystrophy. EMBO J. 2000;19:4439–48. doi: 10.1093/emboj/19.17.4439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Timchenko NA, et al. RNA CUG repeats sequester CUGBP1 and alter protein levels and activity of CUGBP1. J Biol Chem. 2001;276:7820–6. doi: 10.1074/jbc.M005960200. [DOI] [PubMed] [Google Scholar]
- 8.Yadava RS, et al. RNA toxicity in myotonic muscular dystrophy induces NKX2-5 expression. Nat Genet. 2008;40:61–8. doi: 10.1038/ng.2007.28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kim DH, et al. HnRNP H inhibits nuclear export of mRNA containing expanded CUG repeats and a distal branch point sequence. Nucleic Acids Res. 2005;33:3866–74. doi: 10.1093/nar/gki698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Garcia-Lopez A, et al. Genetic and chemical modifiers of a CUG toxicity model in Drosophila. PLoS One. 2008;3:e1595. doi: 10.1371/journal.pone.0001595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Osborne RJ, et al. Transcriptional and post-transcriptional impact of toxic RNA in myotonic dystrophy. Hum Mol Genet. 2009;18:1471–81. doi: 10.1093/hmg/ddp058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Du H, et al. Aberrant alternative splicing and extracellular matrix gene expression in mouse models of myotonic dystrophy. Nat Struct Mol Biol. 2010;17:187–93. doi: 10.1038/nsmb.1720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.de Haro M, et al. MBNL1 and CUGBP1 modify expanded CUG-induced toxicity in a Drosophila model of myotonic dystrophy type 1. Hum Mol Genet. 2006;15:2138–45. doi: 10.1093/hmg/ddl137. [DOI] [PubMed] [Google Scholar]
- 14.Mankodi A, et al. Myotonic dystrophy in transgenic mice expressing an expanded CUG repeat. Science. 2000;289:1769–73. doi: 10.1126/science.289.5485.1769. [DOI] [PubMed] [Google Scholar]
- 15.Chen KY, et al. Length-dependent toxicity of untranslated CUG repeats on Caenorhabditis elegans. Biochem Biophys Res Commun. 2007;352:774–9. doi: 10.1016/j.bbrc.2006.11.102. [DOI] [PubMed] [Google Scholar]
- 16.Raj A, van den Bogaard P, Rifkin SA, van Oudenaarden A, Tyagi S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat Methods. 2008;5:877–9. doi: 10.1038/nmeth.1253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Yuan Y, et al. Muscleblind-like 1 interacts with RNA hairpins in splicing target and pathogenic RNAs. Nucleic Acids Res. 2007;35:5474–86. doi: 10.1093/nar/gkm601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jiang H, Mankodi A, Swanson MS, Moxley RT, Thornton CA. Myotonic dystrophy type 1 is associated with nuclear foci of mutant RNA, sequestration of muscleblind proteins and deregulated alternative splicing in neurons. Hum Mol Genet. 2004;13:3079–88. doi: 10.1093/hmg/ddh327. [DOI] [PubMed] [Google Scholar]
- 19.Sasagawa N, Ohno E, Kino Y, Watanabe Y, Ishiura S. Identification of Caenorhabditis elegans K02H8.1 (CeMBL), a functional ortholog of mammalian MBNL proteins. J Neurosci Res. 2009;87:1090–7. doi: 10.1002/jnr.21942. [DOI] [PubMed] [Google Scholar]
- 20.Kim JK, et al. Functional genomic analysis of RNA interference in C. elegans. Science. 2005;308:1164–7. doi: 10.1126/science.1109267. [DOI] [PubMed] [Google Scholar]
- 21.Zhang S, Binari R, Zhou R, Perrimon N. A genomewide RNA interference screen for modifiers of aggregates formation by mutant Huntingtin in Drosophila. Genetics. 2010;184:1165–79. doi: 10.1534/genetics.109.112516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Illuzzi J, Yerkes S, Parekh-Olmedo H, Kmiec EB. DNA breakage and induction of DNA damage response proteins precede the appearance of visible mutant huntingtin aggregates. J Neurosci Res. 2009;87:733–47. doi: 10.1002/jnr.21881. [DOI] [PubMed] [Google Scholar]
- 23.Bates EA, Victor M, Jones AK, Shi Y, Hart AC. Differential contributions of Caenorhabditis elegans histone deacetylases to huntingtin polyglutamine toxicity. J Neurosci. 2006;26:2830–8. doi: 10.1523/JNEUROSCI.3344-05.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ho TH, et al. Colocalization of muscleblind with RNA foci is separable from mis-regulation of alternative splicing in myotonic dystrophy. J Cell Sci. 2005;118:2923–33. doi: 10.1242/jcs.02404. [DOI] [PubMed] [Google Scholar]
- 25.Zhang S, Ruiz-Echevarria MJ, Quan Y, Peltz SW. Identification and characterization of a sequence motif involved in nonsense-mediated mRNA decay. Mol Cell Biol. 1995;15:2231–44. doi: 10.1128/mcb.15.4.2231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Jan CH, Friedman RC, Ruby JG, Bartel DP. Formation, regulation and evolution of Caenorhabditis elegans 3′UTRs. Nature. 2011;469:97–101. doi: 10.1038/nature09616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Mangone M, et al. The landscape of C. elegans 3′UTRs. Science. 2010;329:432–5. doi: 10.1126/science.1191244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhang C, et al. Defining the regulatory network of the tissue-specific splicing factors Fox-1 and Fox-2. Genes Dev. 2008;22:2550–63. doi: 10.1101/gad.1703108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Li LB, Yu Z, Teng X, Bonini NM. RNA toxicity is a component of ataxin-3 degeneration in Drosophila. Nature. 2008;453:1107–11. doi: 10.1038/nature06909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Nollen EA, et al. Genome-wide RNA interference screen identifies previously undescribed regulators of polyglutamine aggregation. Proc Natl Acad Sci U S A. 2004;101:6403–8. doi: 10.1073/pnas.0307697101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Rehwinkel J, Letunic I, Raes J, Bork P, Izaurralde E. Nonsense-mediated mRNA decay factors act in concert to regulate common mRNA targets. RNA. 2005;11:1530–44. doi: 10.1261/rna.2160905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wittmann J, Hol EM, Jack HM. hUPF2 silencing identifies physiologic substrates of mammalian nonsense-mediated mRNA decay. Mol Cell Biol. 2006;26:1272–87. doi: 10.1128/MCB.26.4.1272-1287.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lemm I, Ross J. Regulation of c-myc mRNA decay by translational pausing in a coding region instability determinant. Mol Cell Biol. 2002;22:3959–69. doi: 10.1128/MCB.22.12.3959-3969.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Finkel RS. Read-through strategies for suppression of nonsense mutations in Duchenne/Becker muscular dystrophy: aminoglycosides and ataluren (PTC124) J Child Neurol. 2010;25:1158–64. doi: 10.1177/0883073810371129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zetoune AB, et al. Comparison of nonsense-mediated mRNA decay efficiency in various murine tissues. BMC Genet. 2008;9:83. doi: 10.1186/1471-2156-9-83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Resta N, et al. A homozygous frameshift mutation in the ESCO2 gene: evidence of intertissue and interindividual variation in Nmd efficiency. J Cell Physiol. 2006;209:67–73. doi: 10.1002/jcp.20708. [DOI] [PubMed] [Google Scholar]
- 37.Linde L, Boelz S, Neu-Yilik G, Kulozik AE, Kerem B. The efficiency of nonsense-mediated mRNA decay is an inherent character and varies among different cells. Eur J Hum Genet. 2007;15:1156–62. doi: 10.1038/sj.ejhg.5201889. [DOI] [PubMed] [Google Scholar]
- 38.Holbrook JA, Neu-Yilik G, Hentze MW, Kulozik AE. Nonsense-mediated decay approaches the clinic. Nat Genet. 2004;36:801–8. doi: 10.1038/ng1403. [DOI] [PubMed] [Google Scholar]
- 39.Amack JD, Mahadevan MS. The myotonic dystrophy expanded CUG repeat tract is necessary but not sufficient to disrupt C2C12 myoblast differentiation. Hum Mol Genet. 2001;10:1879–87. doi: 10.1093/hmg/10.18.1879. [DOI] [PubMed] [Google Scholar]
- 40.Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974;77:71–94. doi: 10.1093/genetics/77.1.71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Raj A, van den Bogaard P, Rifkin SA, van Oudenaarden A, Tyagi S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat Methods. 2008;5:877–9. doi: 10.1038/nmeth.1253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kamath RS, Martinez-Campos M, Zipperlen P, Fraser AG, Ahringer J. Effectiveness of specific RNA-mediated interference through ingested double-stranded RNA in Caenorhabditis elegans. Genome Biol. 2001;2:RESEARCH0002. doi: 10.1186/gb-2000-2-1-research0002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Gidalevitz T, Ben-Zvi A, Ho KH, Brignull HR, Morimoto RI. Progressive disruption of cellular protein folding in models of polyglutamine diseases. Science. 2006;311:1471–4. doi: 10.1126/science.1124514. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Fig. 1. C. elegans expressing expanded CUG repeats exhibit locomotion defects. (A) Representation of motility assays performed using agar plates containing an E. coli food ring. The food ring had a 2cm radius. (B) Motility assays for 2d adults. Data plotted corresponds to the average percentage of population to reach the food at each time point. Error bars represent SD from at least 3 independent experiments; in each experiment, 3–5 replicas of ca. 100–150 animals were analyzed. (C–E) Computational analysis of SM-FISH images. (C) Analysis starts with computational identification of the nuclear region based on DAPI staining in an SM-FISH image of a 123CUG animal. Following nucleus identification, (D) there is computational delineation of cytoplasmic versus nuclear spaces in the SM-FISH image corresponding to the GFP RNA transcript probes. (E) Analysis of pixel intensities for each SM-FISH image, corresponding to low RNA, high RNA densities and RNA foci in both the nucleus and cytoplasm.
Supplementary Fig. 2. Expression of MBL-1::mCherry in C. elegans muscle cells increases expanded CUG transcript recruitment and mutant transcript nuclear foci accumulation. Schematic drawing of the MBL-1::mCHERRY construct (A), and C. elegans body wall muscle cells (B). (C) MBL-1::mCHERRY exhibits a diffuse cellular distribution with nuclear accumulation. (D) C. elegans muscle cells confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue), merge of GFP RNA and nucleus images, and mCherry translational fusion protein. The muscle cells imaged correspond to animals expressing 123CUG repeats and 0CUG, in a mCHERRY (control) or MBL-1::mCHERRY backgrounds. Yellow arrows indicate expanded CUG nuclear foci. MBL-1::mCHERRY localizes to the nucleus. (E) Genetic mosaic analysis of GFP intensity shows that GFP fluorescence, from 123CUG mRNA transcripts, absent in cells expressing mbl-1::mCherry, relative to neighboring cells that fail to express mbl-1::mCherry. GFP fluorescence is not affected in the 0CUG control animals expressing mbl-1::mCherry. (F) Confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue) and merge of C. elegans muscle cells. The strains imaged were 123CUG and 0CUG in: empty vector control (ctrl) and mbl-1 gene inactivations. Yellow arrows indicate expanded CUG nuclear foci.
Supplementary Fig. 3. Screen approach for the identification of modulators of expanded CUG toxicity. (A) Representation of RNAi screen steps in the identification of modulators of expanded CUG repeat pathogenesis. (B) Fluorescent microscopy images of the strains 123CUG and the control 0CUG, on different RNAi gene inactivations: empty vector control (ctrl), mbl-1 and aly-3. Images were taken at the 3d old adult stage. Bar, 200μm.
Supplementary Fig. 4. Suppressors and enhancers of expanded CUG toxicity have distinct effects on expanded CUG nuclear foci accumulation. Confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue) and merge of C. elegans muscle cells. The strains imaged were 123CUG and the control 0CUG, in different RNAi gene inactivations: empty vector control (ctrl), C06A1.6, str-67, mrt-2, npp-4 and smg-2. Yellow arrows indicate expanded CUG nuclear foci.
Supplementary Fig. 5. Gene inactivations have different effects on foci accumulation in the nucleus. Computational analysis of SM-FISH images of 0CUG animals (yellow dots), control (green dots), and 123CUG animals fed different gene inactivations (blue dots) and control vector (red dots). Each ‘dot’ shown in the graph represents one analyzed SM-FISH image, corresponding to a single imaged cell. The red dotted square indicates the region of clustering of the samples corresponding to 123CUG animals on control vector (red dots). Labeled on the graph on the left, above the red box, are the gene inactivations (blue dots) that cause an increase in bright pixel intensity, corresponding to an increase in foci size or number, relative to the 123CUG on control. The ‘grouping’ of 123CUG npp-4 inactivations in the upper right corner of the graph indicates both an increase in nuclear foci and in nuclear ‘single’ transcript localization relative to the 0CUG npp-4 controls that localize further to the left in the graph. The inset section displayed shows gene inactivations (blue dots) that cause a decrease in bright pixel intensity, relative to the 123CUG on control vector, corresponding to a decrease in foci size or number.
Supplementary Fig. 6. (A) Over-expression of expanded CUG repeat suppressors causes a decrease in expanded CUG nuclear foci accumulation. C. elegans muscle cells confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue), merge of GFP RNA and nucleus images, and mCherry translational fusion protein. The strains imaged are animals expressing 123CUG repeats and 0CUG in the following transgenic backgrounds: mCHERRY, NPP-4::mCHERRY, ASD-1::mCHERRY and RNP-2::mCHERRY. RNP-2 corresponds to the U1 small nuclear ribonucleoprotein A, and RNP-2::mCherry exhibits nuclear localization in C. elegans muscle cells. (B) Mutants in the NMD pathway cause an increase in expanded CUG nuclear foci accumulation. Confocal SM-FISH images of GFP RNA transcripts (white), DAPI stained nucleus (blue) and merge of C. elegans muscle cells. The strains imaged were animals expressing 123CUG repeats and 0CUG, in the following backgrounds: wild type (wt), smg-1(r861) and smg-6(r896). Yellow arrows indicate expanded CUG nuclear foci.
Supplementary Fig. 7. (A–B) Sequence composition of CUG repeat sequences in the 3′UTR contributes to NMD transcript recognition for degradation. (A) Schematic drawing of the GC-rich or AT-rich plasmids for expressions in C. elegans muscle cells. (B) Fluorescent microscopy images of strains expressing a GFP with a 300bp ‘artificial’ insert in their 3′UTR containing the following GC percentages: 31%, 32%, 60% and 70%. Also included are the control strains containing 3′UTR inserts cloned from A. thaliana (34%GC) and P. aeruginosa (66%GC). These strains are shown in a wt background and in the background of the following smg mutants: smg-1(5861), smg-2(qd101) and smg-6(r896). The ‘fluorescence’ observed in the 60%GC and 70%GC strains in a wild type background corresponded to the characteristic gut autofluorescence, and no GFP signal was observed in the body wall muscle cells of these animals. Images were taken of animals at the L4 stage. Bar, 100μm. (C–D) Western blot analysis of UPF1 down-regulation (24h post-transfection) by siRNA pool of unaffected (C) and DM1 (D) fibroblast cells, using UPF1-specific antibody. Fibroblasts showed a decrease of 40% in UPF1 levels relative to cells transfected with scrambled siRNAs (mock cells) in both unaffected (C) as well as DM1 (D) cells. GAPDH levels were used for normalization across samples.
Supplementary Fig. 8. Model of regulation of expanded RNAs. (A) Model for regulation of expanded RNA toxicity by the NMD pathway: NMD targets expanded CUG repeat transcripts for degradation reducing the levels of toxic RNAs present in the cells. A decrease in NMD function results in accumulation of toxic transcripts with increase in nuclear RNA foci and increase in toxicity with loss of motility. (B) Model for regulation of expanded RNA foci accumulation by the modulators of RNA toxicity identified: different pathways regulate expanded CUG repeat toxicity; an increase in foci causes a decrease in locomotion however, a decrease in foci doesn’t necessarily correlate with a decrease in muscle toxicity.
Supplementary Table 1. Modifiers of expanded CUG-specific decrease in GFP fluorescence.