ABSTRACT
An outbreak of the fungal pathogen Cryptococcus gattii began in the Pacific Northwest (PNW) in the late 1990s. This outbreak consists of three clonal subpopulations: VGIIa/major, VGIIb/minor, and VGIIc/novel. Both VGIIa and VGIIc are unique to the PNW and exhibit increased virulence. In this study, we sequenced the genomes of isolates from these three groups, as well as global isolates, and analyzed a total of 53 isolates. We found that VGIIa/b/c populations show evidence of clonal expansion in the PNW. Whole-genome sequencing provided evidence that VGIIb originated in Australia, while VGIIa may have originated in South America, and these were likely independently introduced. Additionally, the VGIIa outbreak lineage may have arisen from a less virulent clade that contained a mutation in the MSH2 ortholog, but this appears to have reverted in the VGIIa outbreak strains, suggesting that a transient mutator phenotype may have contributed to adaptation and evolution of virulence in the PNW outbreak. PNW outbreak isolates share genomic islands, both between the clonal lineages and with global isolates, indicative of sexual recombination. This suggests that VGII C. gattii has undergone sexual reproduction, either bisexual or unisexual, in multiple locales contributing to the production of novel, virulent subtypes. We also found that the genomes of two basal VGII isolates from HIV+ patients contain an introgression tract spanning three genes. Introgression substantially contributed to intra-VGII polymorphism and likely occurred through sexual reproduction with VGI. More broadly, these findings illustrate how both microevolution and sexual reproduction play central roles in the development of infectious outbreaks from avirulent or less virulent progenitors.
IMPORTANCE
Cryptococcus gattii is the causative agent responsible for ongoing infections in the Pacific Northwest of the United States and western Canada. The incidence of these infections increased dramatically in the 1990s and remains elevated. These infections are attributable to three clonal lineages of C. gattii, VGIIa, VGIIb, and VGIIc, with only VGIIa identified once previously in the Pacific Northwest prior to the start of the outbreak, albeit in a less virulent form. This study addresses the origin and emergence of this outbreak, using whole-genome sequencing and comparison of both outbreak and global isolates. We show that VGIIa arose mitotically from a less virulent clonal group, possibly via the action of a mutator phenotype, while VGIIb was likely introduced from Australia, and VGIIc appears to have emerged in the United States or in an undersampled locale via sexual reproduction. This work shows that multiple processes can contribute to the emergence of an outbreak.
INTRODUCTION
Emergence and expansion of new pathogens are among the most important current paradigms in infectious diseases (1). Consequently, understanding the mechanisms by which these outbreaks develop is paramount. For some pathogens, emergence is enabled by changes in the host population, such as expansion of the immunocompromised population as a consequence of the HIV/AIDS epidemic and the rise in the number of organ transplant recipients. This has dramatically increased the number of annual infections caused by opportunistic pathogens like Cryptococcus neoformans, particularly among HIV/AIDS patients in sub-Saharan Africa (2). Likewise, the recent emergence of zygomycete pathogens has been driven by the expansion of a vulnerable host population (3).
Alternatively, emergence can be driven by changes in the pathogen population, often through genomic variation. In the case of the yearly outbreak of seasonal influenza, antigenic drift, whereby viral lineages undergo positive selection altering the antigenic target of the immune system, allows the virus to evade not only the immunity established in infected individuals the year before but also that of immunized individuals (4). While this process allows the small changes required for continual, seasonal reemergence, the antigenic shift that occurs by reassortment when two or more viruses exchange components while in the same host has been implicated in the emergence of large influenza outbreaks, including the 1918 pandemic and the 2009 H1N1 epidemic (5, 6). These much greater genomic reassortments (here termed macroevolution) can lead to emergence of pathogens with novel characteristics, including enhanced transmission and virulence.
In eukaryotes, recombination between lineages occurs via meiotic sexual cycles. This can have profound consequences for the development of successful pathogens. In Toxoplasma, two ancient lineages recombined to generate the modern hypervirulent clonal groups (7), and in a contemporary setting, both outcrossing and “selfing” have been implicated in outbreaks of Toxoplasma and related pathogens (8). Sexual reproduction has also recently been shown to provide de novo short-term variation in C. neoformans by dramatically increasing the incidence of aneuploidy (9). Alternatively, mitotic mutation can contribute to the emergence of pathogens. In C. neoformans, serial growth under laboratory conditions or in animals can lead to attenuation (10) or elevation of virulence (11), respectively. In both bacteria and fungi, mutation can be further enabled by development of a hypermutator state, often through mutations in DNA repair pathways. In yeast, loss of MSH2, which encodes a critical mismatch repair protein, results in instability in repeat tracts (12), which commonly occur in cell surface proteins (13), and can affect recognition by the host immune system. In C. neoformans, a hypermutator state has been shown to increase resistance to amoeba by enabling mutations in the RAM pathway and enabling pseudohyphal growth (14).
Cryptococcus gattii is a basidiomycete fungus and a pathogen of humans. Unlike its sister species C. neoformans, two of the subtypes of C. gattii, VGI and VGII, predominantly infect immunocompetent individuals. The VGII subtype is causing an ongoing outbreak in the Pacific Northwest (PNW) of the United States and in British Columbia in Canada (15–17). This outbreak originated on Vancouver Island in the late 1990s and has been expanding geographically over the ensuing decade (15, 17). Recent human and veterinary cases of C. gattii show the outbreak is continuing to expand south and east from the Pacific Northwest (18, 19). Multilocus sequence typing revealed that the outbreak consists of two subtypes of VGII, termed VGIIa and VGIIb. The VGIIb subtype is also found in other parts of the world, especially Australia, and has reduced virulence in animal models, while VGIIa is unique to the Pacific Northwest (with the exception of one isolate from Brazil that differed at one of seven multilocus sequence typing [MLST] markers examined) and exhibits increased virulence (20). Additionally, these subtypes share approximately 50% of their MLST alleles, suggesting that VGIIa may have been a more virulent progeny or sibling of VGIIb (20). Subsequent sampling identified a third, even more virulent subtype designated VGIIc, which is, so far, completely unique to the Pacific Northwest outbreak in Oregon (21). Recent work based on MLST shows that the VGII lineage itself originated ancestrally in South America (22); however, the proximal source of the outbreak strains is still unknown. Australia (20) and South America (20, 22) have both been proposed as geographical sources of origin.
Previous studies used a multilocus sequence typing (MLST) approach to test a maximum of 30 loci (20). This inherently covers only a fraction of the true diversity that is present. Dramatic reductions in the cost of whole-genome sequencing over the past several years have made whole-genome sequencing of multiple isolates tractable. However, the few studies that have incorporated this approach have primarily used it in comparison to or to supplement MLST (22, 23). In this study, we utilized whole-genome sequencing to address the clonality of individual outbreak lineages, to examine the possibility of recombination in the population, and to determine the origin of the outbreak isolates. Our findings reveal that the clonal clusters in the Pacific Northwest originated through sexual reproduction within the highly sexual VGII population but were introduced independently of each other. Furthermore, the VGIIc lineage likely developed high virulence through sexual recombination of alleles, while VGIIa underwent mitotic microevolution, potentially driven by a mutator phenotype that arose following ancestral rounds of sexual reproduction.
RESULTS
The genomes of 38 C. gattii isolates of the VGII genotype were sequenced to initiate an investigation of recombination and clonality. In addition, sequences for 17 previously published genomes from the CDC were obtained from the NCBI Sequence Read Archive (SRA) (23). The strains analyzed are summarized in Table 1. Briefly, genomic sequencing targeted three sets of strains thought to be clonal, representing the VGIIa, VGIIb, and VGIIc groups. In addition, 3 VGIIa-like, 2 VGIIb-like, and 18 other VGII isolates not from these groups were sequenced, including 3 MATa isolates. In sum, 11 VGIIa, 9 VGIIb, 10 VGIIc, and 23 other genomes sequenced in this study or previously published were compared, with two isolates sequenced by both the CDC and this study for a total set of 53 VGII whole-genome sequences. Sequences generated in this study are available in the SRA.
TABLE 1 .
C. gattii genomes sequenceda
| Population and strain | Origin | Mating type | Sequenced by |
|---|---|---|---|
| VGIIa | |||
| R265 | Vancouver Island, clinical | MATα | Broad Institute |
| RB1 | Vancouver Island, environmental | MATα | Heitman lab |
| RB59 | Vancouver Island, environmental | MATα | Heitman lab |
| EJB17 | Oregon, veterinary | MATα | Heitman lab/CDC |
| EJB21 (or B7467) | Oregon, clinical | MATα | Heitman lab |
| B7395 | Washington, clinical | MATα | Heitman lab/CDC |
| B7436 | California, veterinary | MATα | Heitman lab |
| R498 | Vancouver Island, veterinary | MATα | Heitman lab |
| R265 reseq | Vancouver Island, clinical | MATα | Heitman lab |
| B7422 | Oregon, veterinary | MATα | CDC |
| B8849 | Oregon, environmental | MATα | CDC |
| B8577 | British Columbia, environmental | MATα | CDC |
| VGIIa-like | |||
| ICB107 | Brazil, clinical | MATα | Heitman lab |
| CBS7750 | San Francisco, environmental | MATα | Heitman lab |
| NIH444 | Seattle, clinical, 1975 | MATα | Heitman lab |
| VGIIb | |||
| R272 | Vancouver Island, clinical | MATα | Heitman lab |
| NT13 | Australia, Northern Territory | MATα | Heitman lab |
| B7394 | Washington, veterinary | MATα | CDC |
| B7735 | Oregon, clinical | MATα | CDC |
| B8554 | Oregon, veterinary | MATα | CDC |
| B8828 | Washington, veterinary | MATα | CDC |
| RDH6 | Australia, Northern Territory, clinical | MATα | Heitman lab |
| V9 | Australia, veterinary | MATα | Heitman lab |
| V6 | Australia, veterinary | MATα | Heitman lab |
| V26 | Australia, veterinary | MATα | Heitman lab |
| VGIIb-like | |||
| 99/473-1 | Caribbean Islands, clinical | MATα | Heitman lab |
| B9588 | Florida, clinical | MATα | Heitman lab |
| VGIIc | |||
| B8571 | Washington, clinical | MATα | CDC |
| B8843 | Oregon, clinical | MATα | CDC |
| B8838 | Washington, clinical | MATα | CDC |
| B7466 (or EJB52) | Oregon, veterinary | MATα | CDC |
| B7737 | Oregon, clinical | MATα | CDC |
| B6863 | Oregon, clinical | MATα | CDC |
| B7390 | Idaho, clinical | MATα | CDC |
| B7432 | Oregon, clinical | MATα | CDC |
| EJB87 | Oregon, veterinary | MATα | Heitman lab |
| EJB18 | Oregon, clinical | MATα | Heitman lab |
| VGIIMATa | |||
| LA499 | Colombia, clinical | MATa | Heitman lab |
| CBS1930 | Aruba, veterinary | MATa | Heitman lab |
| VBGc11 | Puerto Rico, environmental | MATa | Heitman lab |
| VGIInt | |||
| NT3 | Australia, Northern Territory, clinical | MATα | Heitman lab |
| NT7 | Australia, Northern Territory, clinical | MATα | Heitman lab |
| NT8 | Australia, Northern Territory, clinical | MATα | Heitman lab |
| RDH2 | Australia, clinical | MATα | Heitman lab |
| RDH7 | Australia, Northern Territory, clinical | MATα | Heitman lab |
| VGII | |||
| 78-5-46 | Utah, clinical | MATα | Heitman lab |
| ICB182 | Brazil, clinical | MATα | Heitman lab |
| ICB183 | Brazil, environmental | MATα | Heitman lab |
| ICB184 | Brazil, environmental | MATα | Heitman lab |
| WA861 | Western Australia, veterinary | MATα | Heitman lab |
| WM178 | Sydney, Australia, clinical | MATα | Heitman lab |
| IP96/1120-1 | French, clinical | MATα | Heitman lab |
| 2003.125 | French, clinical | MATα | Heitman lab |
| 93.980 | French, clinical | MATα | Heitman lab |
| 98.1132 | Caribbean, clinical | MATα | Heitman lab |
The strains utilized in this study are listed. The source of the sequence is indicated, with “Heitman lab” designating isolates newly sequenced in this study.
Outbreak populations of C. gattii are highly clonal.
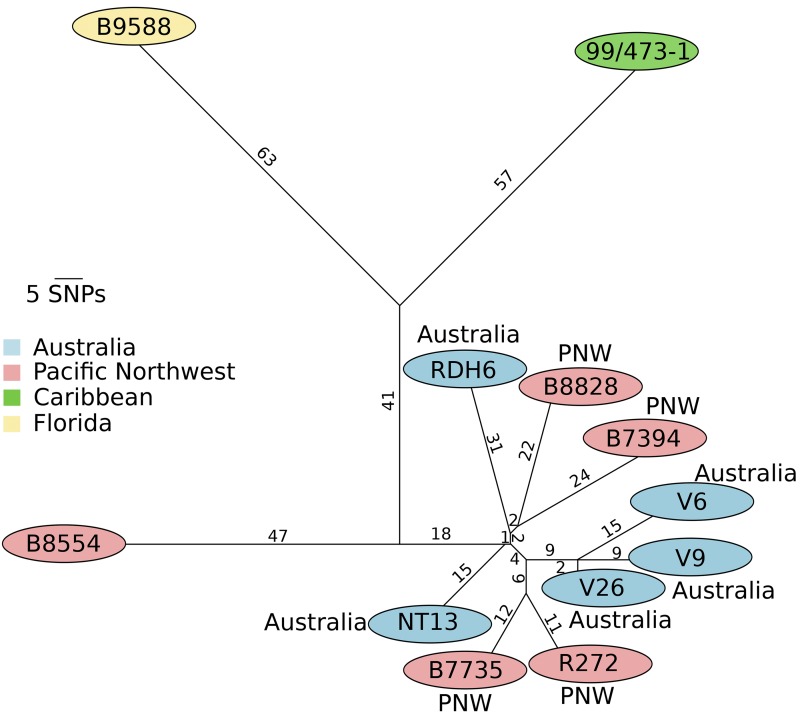
Analysis of whole-genome sequencing of C. gattii isolates revealed that the population consists of clonal subpopulations, each of which shares a mating type and is differentiated within the cluster by very limited diversity. Five clonal clusters were identified: the previously known VGIIa, VGIIb, and VGIIc groups; an additional cluster restricted to the Northern Territory of Australia comprised of isolates NT3, NT7, NT8, RDH2, and RDH7 (referred to as VGIInt [24]); and a MATa clonal cluster comprised of isolates LA499, CBS1930, and VBGc11 (referred to as VGIIMATa). After elimination of erroneous polymorphic site calls as a result of improper mapping of repetitive sequences, a total of 150 polymorphisms remained within the VGIIa group, and 288 remained within the VGIIb group, 130 within the VGIIc group, and 2,164 within the Australian VGIInt group. As most of the polymorphic alleles were private (restricted to a single isolate), any two given isolates within a group were separated by considerably fewer sites, ranging from a minimum of 14 to a maximum of 49 for VGIIa. VGIIb isolates were separated by 11 to 61 polymorphisms, excluding one outlier isolate B8554 (99 sites). Phylogenetic trees were constructed using single nucleotide polymorphisms (SNPs) derived from the whole genome. In the case of both VGIIa and VGIIb, comparison with the most closely related isolates that were still distinguishable by MLST analysis revealed substantial differences (Fig. 1A and B). For VGIIa, the South American isolate ICB107 branched outside the VGIIa cluster from the Pacific Northwest. Similarly, the Caribbean isolate 99/473-1 and the Florida isolate B9588 fell outside the VGIIb cluster otherwise shared between Australia and the Pacific Northwest. VGIIc was not closely related to any isolates from any geographic regions other than the PNW but did exhibit diversity at the whole-genome level, allowing an inference of population structure within the VGIIc clade (Fig. 1C).
FIG 1 .
Phylogenetic reconstruction shows evidence for clonal expansion in outbreak groups. Maximum likelihood phylogenies were constructed using a matrix of all available SNPs. Bootstrap values are based on 500 replicates. The scale bar indicates the number of substitutions per nucleotide position. (A) Phylogeny based on SNPs unique to VGIIa. ICB107 was also included as it differed at only one MLST marker in previous studies. (B) Phylogeny based on SNPs unique to VGIIb. 99/473-1 was also included as it was similar to VGIIb in previous studies. (C) Phylogeny based on SNPs unique to VGIIc.
Analysis of whole-genome sequencing revealed no evidence of ongoing recombination within the clonal clusters in the Pacific Northwest via a standard allele compatibility test. In the case of VGIIa, only two of the sequenced strains shared any polymorphisms (Fig. 1A). Instead the vast majority of SNPs did not show evidence of either sexual recombination or shared descent past the introduction into the Pacific Northwest. This likely indicates that sampling and sequencing of isolates has not reached saturation. VGIIb and VGIIc showed substantially more population structure, with a number of isolates sharing SNPs; however, both groups still lacked evidence for intraclade recombination via a standard allele compatibility test. The much higher frequency of shared polymorphism within the VGIIb and IIc groups suggests that sampling has been more complete in these groups or is more restricted to one subclade.
Outbreak strains have diverse proximal origins.
There has been a debate as to the geographic origin(s) of the lineages causing the Pacific Northwest outbreak. The VGIIa lineage was previously proposed to be the offspring of a cross between VGIIb and an unknown parent that may have taken place either in Australia or in the Pacific Northwest (20). Whole-genome analysis suggests that VGIIa from the Pacific Northwest is a clonal group derived from a shared common ancestor with limited diversity (150 SNPs within the entire group, with as few as 14 to 49 SNPs distinguishing any two isolates) (Fig. 2A). In addition, there is a closely related isolate from South America, ICB107. This isolate differs from the PNW VGIIa isolates by 1,174 polymorphic sites, including insertion or deletion events (indels). These sites are distributed evenly across the genome (Fig. 2B), with no substantial blocks devoid of SNPs. This suggests that these SNPs have arisen during clonal propagation since the last common ancestor of these isolates underwent a meiotic event and that VGIIa likely shares a common lineage with ICB107. While VGIIa could have entered the PNW through Australia as previously hypothesized, the generation of this lineage likely occurred much longer ago, either in South America or in a different location, after which it dispersed to both South America and the PNW. The lack of a VGIIa-like isolate from Australia suggests that South America is a more likely proximal source.
FIG 2 .
VGIIa outbreak isolates are more closely related to each other than to historical isolates or South American VGII isolates. (A) Prediction of VGIIa population structure based on haplotype network inference. Only SNPs were used to eliminate alignment artifacts. All Pacific Northwest isolates are predicted to share a relatively recent last common ancestor. The South American isolate ICB107, while the closest among isolates included in this analysis, is more distantly related. (B) SNPs distinguishing ICB107 from the VGIIa isolate R265 plotted across the genome. SNPs are distributed across the genome, suggesting ICB107 was not separated from VGIIa by a recent meiotic event.
Importantly, VGIIb isolates from the Pacific Northwest are highly similar to isolates from Australia, with some isolates clustering more closely with Australian than with other PNW VGIIb strains (Fig. 3). With one exception, a CDC-sequenced isolate from Oregon (B8554), the Australian and PNW isolates appear to share a single origin. In contrast, isolates described as VGIIb from outside the PNW or Australia (i.e., Caribbean isolate 99/473-1 and a recent Florida clinical isolate, B9588) group outside this clonal lineage. It may be more accurate to describe these isolates as VGIIb-like, as they do not appear to fit within the clonal outbreak cluster encompassing both the PNW and Australian isolates and were likely introduced to the southeast United States in a separate, unrelated event. This evidence suggests that the VGIIb/minor component of the outbreak either arrived in the PNW from Australia or arrived in both the PNW and Australia at very similar times from a common progenitor population.
FIG 3 .
VGIIb outbreak isolates cluster more closely with Australian isolates than Caribbean and Florida isolates. Shown is a prediction of VGIIb population structure based on haplotype network inference. Pacific Northwest VGIIb isolates and Australian isolates cluster together, with the exception of the Pacific Northwest isolate B8554 sequenced by the CDC.
VGIIa-like isolates have a frameshift mutation in a key DNA repair gene.
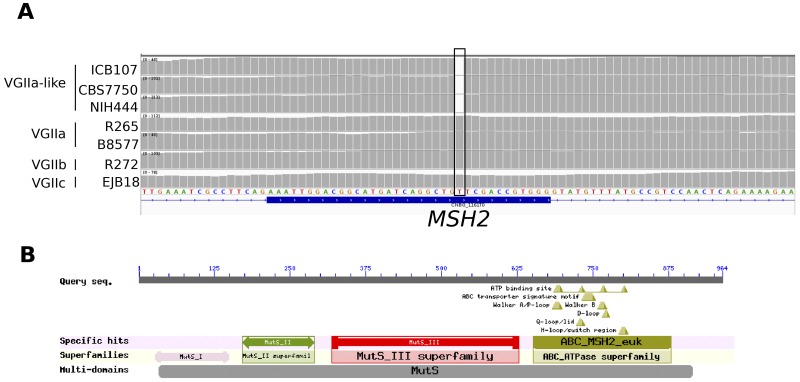
The VGIIa population consists of two distinct groups: the outbreak strains in the Pacific Northwest and a less virulent group consisting of the strains ICB107, CBS7750, and NIH444 that we call VGIIa-like (20, 21). These strains were all isolated prior to the onset of the Pacific Northwest outbreak, and all are characterized by diminished virulence. A rough estimate based on the commonly accepted estimate of 0.0081 substitution per site per million years (25) suggests that these groups diverged 91.7 years ago, although with such a small number of SNPs, this number could vary dramatically. As our results show that VGIIa and the VGIIa-like isolates do not appear to be separated by a recent meiotic event (Fig. 2B), we explored mitotic microevolution as a potential cause of the increased virulence observed in the more recently isolated VGIIa strains. SNPs and indels differentiating the VGIIa and VGIIa-like strains were examined for potential impact using SNPeff (25) (see Table S1 in the supplemental material). Twelve missense mutations were identified, but only one frameshift mutation was found (Fig. 4A). This mutant was in the 4th of 19 exons of CNBG_1161, leaving only 42 amino acids unchanged out of the 964-amino-acid length. CNBG_1161 encodes the C. gattii ortholog of Msh2, a MutS-like DNA mismatch repair protein characterized in Saccharomyces cerevisiae (12, 26) (Fig. 4B). The msh2 frameshift mutation is present in all three VGIIa-like isolates (ICB107, CBS7750, and NIH444), but the wild-type allele is present in all VGIIa outbreak isolates analyzed.
FIG 4 .
VGIIa-like low-virulence clade has a hypermutator mutation. (A) Read depth across an exon of CNBG_1161 from three VGIIa-like genomes, two VGIIa genomes, one VGIIb genome, and one VGIIc genome. All three VGIIa-like isolates have a single base deletion (denoted with a box) resulting in a frameshift mutation in the MSH2 gene. (B) Results from a conserved domain search for CNBG_1161 performed using the NCBI’s CD search program. CNBG_1161 has strong homology to the MutS or Msh2 domains involved in DNA damage repair.
Recombination is occurring between clonal clusters.
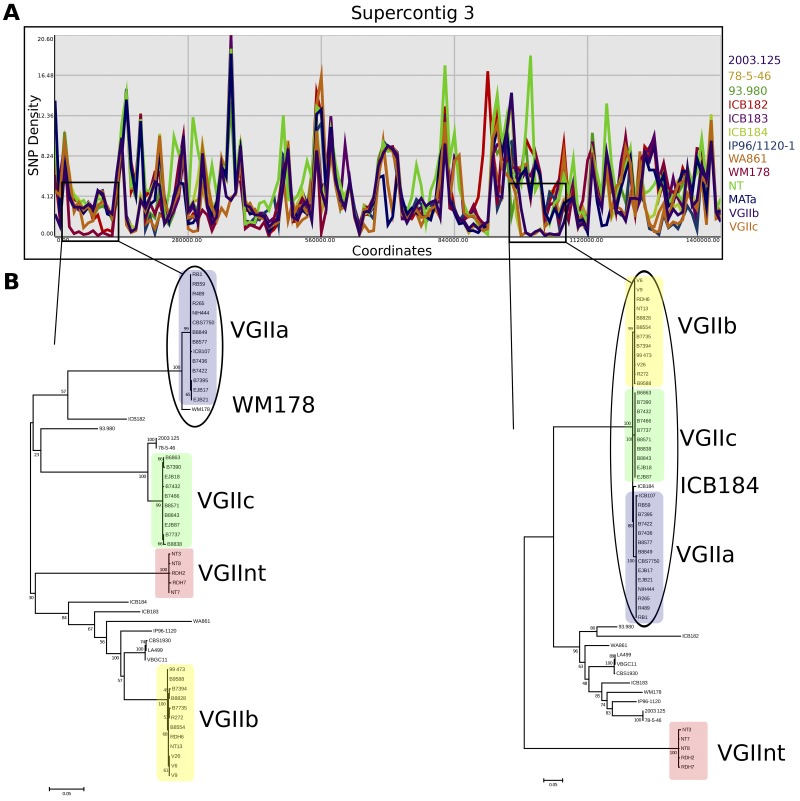
While strains within each of the five clonal clusters of C. gattii are highly related to each other, comparisons among clusters revealed substantial differences. However, the sites differentiating these clusters were not uniformly dense across the genome. This is demonstrated by incongruity between phylogenies inferred from different portions of the genome (Fig. 5), suggesting that ancestral recombination among the clonal lineages has resulted in some portions of the genome being more similar between two clonal groups than the others. Further evidence is provided by examining SNP density between isolates. This analysis demonstrates the presence of islands of identity between any two individual isolates where diversity is considerably lower than the genome-wide average (Fig. 6A). Inferring phylogenies from these regions of depressed polymorphism reveals uniquely close relationships among strains (Fig. 6B). This suggests that sexual reproduction likely contributed to the production of the ancestor of each clonal cluster, with subsequent mitotic clonal expansion of these isolates to spread throughout the environment. Notably, none of the outbreak clusters has an individual partner strain or cluster that contributes a large portion of the genome (i.e., 50% for one cross, 25% for two crosses, etc.). This suggests that either sampling has captured only a small portion of the VGII population or some of these intermediate strains may no longer exist.
FIG 5 .
VGII phylogenies are incongruent based on different loci. Shown is a pair of maximum likelihood phylogenies generated from the first 100 kb of supercontigs 6 and 7. Clonal clusters are shaded to facilitate comparison. Bootstrap values are based on 500 bootstraps. The scale bar indicates substitutions per nucleotide position. Crossing lines indicate lack of congruence.
FIG 6 .
Regions of reduced SNP density provide evidence for ancestral sexual reproduction. (A) SNP density is plotted relative to the R265 reference genome. Clonal groups are collapsed to the SNPs shared by all members of the group. Regions of unexpectedly low levels of polymorphism are boxed for emphasis. (B) Maximum likelihood phylogenies generated from regions of depressed SNP density demarcated in panel A. Bootstrap values are based on 500 bootstraps. The scale bar indicates the number of substitutions per nucleotide position. Outbreak groups are highlighted in larger font, as well as individual global isolates that contribute to the identity island.
Diploid RB59 isolate is homozygous.
A diploid VGIIa isolate RB59 was previously identified by fluorescence-activated cell sorter (FACS) analysis (20). Using MLST markers, this strain did not appear to be a hybrid of two different VGII lineages. Whole-genome sequencing further revealed that this strain is homozygous for all identified SNPs and indels. This suggests that RB59 is the result of either endoreplication or mother-daughter cell-cell fusion, both of which are possible intermediates in the unisexual cycle. This would not produce offspring with substantial polymorphism, as both parental chromosomes would be identical in both copies, but it could facilitate the production of basidiospores, which might then serve as infectious propagules, or the production of aneuploid variant progeny (9, 27).
Analysis of possible recombination/gene conversion in the MATα locus.
Recombination can occur via either bisexual or unisexual reproduction. However, the mating type locus is an exception, as this ~100-kb region is highly rearranged and divergent between MATα and MATa isolates and recombination between a and α alleles will result in acentric or dicentric chromosomes. As a result, any evidence of crossover within this locus should be the result of mating between two isolates of the same mating type. This locus has a relatively robust population structure, with the VGIIa and VGIIb clonal groups being closely related compared to the other clonal clusters (see Fig. S1A in the supplemental material).
We examined 995 SNPs within the MAT locus. Three SNPs broke apart otherwise conserved haplotypes and were suggestive of recombination via a standard allele compatibility test. Two of these lie within the coding sequence of the gene encoding Ste20 (see Fig. S1B), while the other was approximately 40 kb away in the intergenic region between the SPO14 and GEF1 genes (not shown). All three SNPs were well supported, with high-quality scores, high mapping scores, and high depth of coverage. The intergenic variant between SPO14 and GEF1 lies in a region that is colinear between the a and α alleles and so could potentially still be the result of recombination during bisexual mating. This variant also lies close to a known a-α mating gene conversion hot spot in C. neoformans (2 events out of 255 progeny tested) (28). However, the other pair lies within a gene that is diverged and rearranged between a and α alleles. Given the lack of flanking recombination tracts matching the new allele combinations, the most parsimonious explanation is that one of the alleles underwent gene conversion during α-α unisexual mating. Alternatively, it is possible that homoplasic mutations may have contributed to the signature of gene conversion. We can estimate the frequency of homoplasic mutation by measuring the frequency of multiallelic SNPs compared with biallelic SNPs in the SNP calling set. There are 1,346 multiallelic SNPs out of 223,743 total SNPs for a total frequency of 0.60%. Mutations at a biallelic site should create homoplasic mutations half as often as those at a multiallelic site, so with 995 SNPs in the MAT locus, this suggests approximately 3 homoplasic SNPs may exist in this set. As a result, we cannot confidently distinguish the two possible explanations for the origin of these SNPs.
The VGII group is recombining at the population level.
In addition to the shared genomic islands, use of a paired allele compatibility test based on SNP information (AB × ab giving rise to AB, ab, Ab, or αB) provided evidence for sexual reproduction and recombination (Fig. 7A). One hundred random loci were selected using the Genome Analysis Toolkit (GATK) SelectVariants walker out of the 262,614 polymorphic sites in the data set. These loci were collapsed into unique and informative allele combinations, leaving 46 loci, and then compared. This test showed evidence for mating and meiosis occurring between a number of different isolates at the whole-genome level, both with and without the involvement of the three sequenced MATa isolates in this study. In particular, all of the 46 loci tested showed evidence for recombination with at least 13 of the 45 other loci (29%) and as many as 40 out of 45 (89%) (Fig. 7B). Recombination provides strong evidence for the involvement of the sexual cycle in the production of these C. gattii strains, and given the paucity of a isolates in the environment, we hypothesize this involved both a-α bisexual and α-α unisexual reproduction.
FIG 7 .
Whole-genome allele compatibility test shows evidence for prolific recombination at the population level. (A) An example of a paired allele compatibility test from the VGII population. Alternative SNPs are depicted in red and the reference in white. Evidence for recombination is provided by any pairwise comparison of two loci in which strains are present where red-red, white-white, red-white, and white-red combinations are all found (AB, Ab, aB, and ab) satisfying the allele compatibility test, providing evidence for recombination. (B) One hundred random SNPs were selected from the VGII data set and collapsed into 46 unique allele patterns. The reference nucleotide is indicated by white and the variant by red. A pairwise comparison of all 46 unique loci is shown, with green shading indicating a positive result and evidence for recombination.
Genomic islands of high polymorphism in the VGII group.
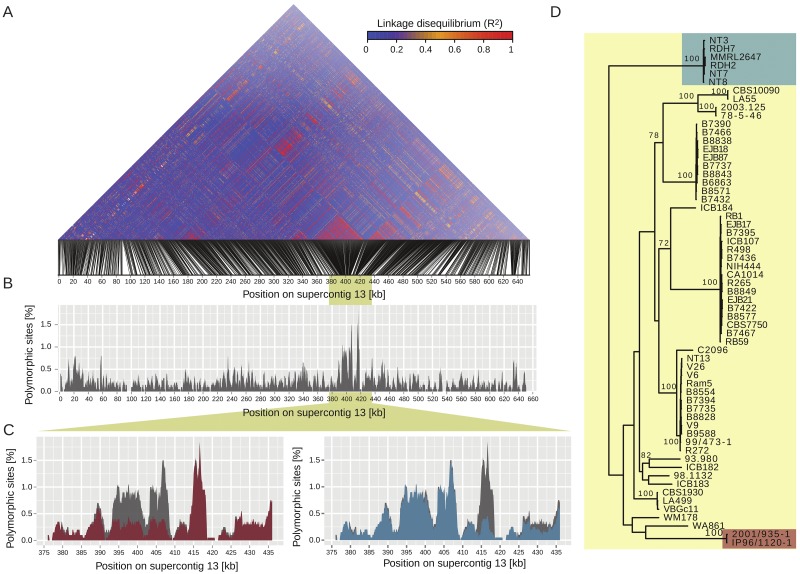
Polymorphisms among VGII isolates are not homogeneous along the chromosomes. In contrast with the islands of diminished polymorphism, we found SNP density was elevated locally, resulting in genomic islands of high polymorphism. For example, a region on R265 supercontig 13 located between 393 and 417 kb showed 3× to 4× more polymorphisms than the surrounding regions (Fig. 8B). This region coincides with two blocks of high linkage disequilibrium on supercontig 13 (Fig. 8A). We examined whether the increased polymorphisms were caused by the inclusion of particular VGII isolates and found that the high polymorphisms between positions 393 and 408 kb are caused by two outlying clonally related isolates (2001/935-1 and IP96/1120-1) (Fig. 8D). Removal of these two isolates reduced the polymorphism to levels identical to the genomic surroundings (red area in Fig. 8C). Similarly, the high level of polymorphism in the region from 414 to 417 kb is caused by the presence of six clonal isolates from the VGIInt group (NT3, NT7, NT8, RDH2, RDH7, and MMRL2647) (Fig. 8D). Removal of these six isolates from the analysis drastically reduced the level of polymorphism in this region (blue area in Fig. 8C).
FIG 8 .
Genomic islands of high polymorphism on supercontig 13 are caused by two distinct VGII clades. (A) Linkage disequilibria (R2) among SNP loci on supercontig 13. Two regions located between positions 393 and 417 kb were in high linkage disequilibria. (B) Polymorphism among VGII isolates on supercontig 13. An island of high polymorphism colocalized with high linkage disequilibria between positions 393 and 417 kb. (C) The islands of high polymorphism were caused by the presence of two distinct groups of VGII isolates. Exclusion of isolates 2001/935-1 and IP96/1120-1 reduced the polymorphism within VGII to low levels (red area) between positions 393 and 408 kb compared to polymorphism among all VGII isolates (gray area). Exclusion of isolates NT3, NT7, NT8, RDH2, RDH7, and MMRL2647 reduced the polymorphism within VGII to low levels (blue area) between positions 414 and 417 kb compared to polymorphism among all VGII isolates (gray area). (D) Maximum likelihood phylogeny of VGII isolates based on all SNP on supercontig 13. Values indicate bootstrap support among 100 replicates. Isolates 2001/935-1 and IP96/1120-1 and isolates NT3, NT7, NT8, RDH2, RDH7, and MMRL2647 each grouped into a distinct clade of VGII.
Incongruent phylogeny on supercontig 13 indicates introgression into VGII isolates.
To understand the causes of the regions of elevated polymorphism, we constructed a phylogeny based on sequence polymorphisms within the region of 393 to 408 kb on supercontig 13. We included diverse C. gattii isolates of all four VG types. The phylogeny resolved all four VG types of C. gattii as expected (Fig. 9B). However, isolates 2001/935-1 and IP96/1120-1 were placed incongruously next to VGI and VGIV instead of being clustered among other VGII isolates. Clade support for these groupings was high (95% and 73% bootstrap support, respectively). The incongruous groupings are indicative of an introgression event into isolates 2001/935-1 and IP96/1120-1 from VGI C. gattii. In contrast, in the second block from 414 to 417 kb, isolates from the VGIInt group were placed within VGII (data not shown).
FIG 9 .
Island of polymorphism on supercontig 13 shows evidence of introgression into VGII. (A) The region of high polymorphism between 393 and 408 kb on supercontig 13 contains three genes (CNBG_4871 to -4873). Variation in GC content in the region is shown in red. (B) Maximum likelihood phylogeny of C. gattii VGI, VGII, VGIII, and VGIV isolates. The tree was constructed based on 297 informative SNPs in the region of 393 to 408 kb on supercontig 13. VGII isolates 2001/935-1 and IP96/1120-1 most closely grouped with VGI. Values indicate bootstrap support among 100 replicates.
The affected region from 393 to 408 kb comprises three genes: CNBG_4871, CNBG_4872, and CNBG_4873 (Fig. 9A). All of these genes showed considerable amino acid polymorphism among C. gattii isolates (see Fig. S2 in the supplemental material). Isolates 2001/935-1 and IP96/1120-1 share amino acid polymorphism with different VG groups at different positions in the protein, suggesting that the introgressed region was affected by recombination or the donor is an unknown VG type. In CNBG_4873, the two isolates also have an amino acid at position 44 that was not found in any other C. gattii isolate. The three genes in the affected region from 393 to 408 kb are not yet functionally characterized in C. gattii. A homology search based on a translated nucleotide sequences identified CNBG_4871 as being a likely ortholog of AVO3 in Saccharomyces cerevisiae. Avo3 is part of the TORC2 complex that contains the Tor2 protein and is involved in polarization of the actin cytoskeleton. CNBG_4872 is predicted to encode the ATP-dependent DNA helicase Mph1 that dissociates Rad51 D-loops. CNBG_4873 is predicted to encode the metal homeostasis protein Bsd2 with roles in metal ion transport and vacuolar protein targeting.
Introgressed isolate from an HIV+ patient is aneuploid.
Both isolates 2001/935-1 and IP96/1120-1 do not belong to an outbreak VGII subtype but do group within the VGII clade (Fig. 8D). Both isolates are of clinical origin and were isolated from HIV-positive patients from Africa. Isolate IP96/1120-1 was isolated in France from a patient who moved from Africa and isolate 2001/935-1 was isolated from a bronchoalveolar lavage sample from a Senegalese patient (20). Despite the near clonal nature of the two isolates 2001/935-1 and IP96/1120-1, we found that IP96/1120-1 likely had multiple partial and complete chromosomal duplications not found in 2001/935-1. Read depth on supercontigs 6 and 13 indicated partially duplicated regions and read depth on supercontig 14 indicated duplication of the entire region covered by this supercontig (Fig. 10). Interestingly, the introgressed region on supercontig 13 is found within a duplicated region of this supercontig. All three duplications contain only homozygous variants, suggesting the extra copies arose by duplicating the original chromosomal region represented by the supercontig, rather than through mating with a different strain, although α-α mating or autodiploidization followed by sequential chromosome loss cannot be ruled out. Duplication of chromosomes readily occurs in Cryptococcus following exposure to antifungal compounds and during sexual reproduction (9, 29). Chromosomal disomy in IP96/1120-1 isolated from an HIV+ patient may have been a direct consequence of antifungal treatment in the patient.
FIG 10 .
Aligned read depth on supercontigs of IP96/1120-1. Read depth was approximately 2-fold higher than the genome-wide mean coverage on supercontigs 6, 13, and 14. Variation in read depth on supercontigs 6 and 13 suggests partial duplications across the region of the chromosome mapping to these supercontigs. Read depth on supercontig 14 suggests duplication of the entire region represented by this supercontig.
In addition to the aneuploidy observed in IP96/1120-1, two additional isolates were found to be aneuploid (see Fig. S3 in the supplemental material). The VGIIb isolate B8828 is disomic across supercontig 13, while the French clinical isolate 93.980 is disomic across supercontig 14 and parts of the region aligning to supercontig 6 occur in 1, 2, 3, 4, 5, or 6 copies (see Fig. S4 in the supplemental material). These isolates are of clinical and veterinary origin, respectively, which may indicate that aneuploidy could be an adaptation to growth in an animal host.
DISCUSSION
Understanding the evolution of virulence in pathogens is key to understanding the development of emerging diseases. The sudden emergence of an outbreak of the previously primarily tropical pathogen C. gattii in the temperate Pacific Northwest in the late 1990s raised a number of questions about the emergence of this outbreak, namely, the genetic and geographic origin of this pathogen.
Adaptation and emergence of virulence can take place through both small changes introduced through random mutation, as well as through reassortment of preexisting mutations in the population. The VGII C. gattii population shows examples of both types of adaptation. The VGIIa lineage is a result of small-scale mitotic adaptation from a shared last common ancestor with the strains we have termed VGIIa-like. These lineages display different virulence potentials, and the VGIIa outbreak strains have increased virulence relative to the VGIIa-like isolates ICB107, CBS7750, and NIH444 (20, 21). Both CBS7750 and NIH444 are closely related to the VGIIa clonal outbreak cluster, and thus comparison of the genomes may allow identification of the critical changes impacting virulence that enabled the emergence of the VGIIa subclade of the Pacific Northwest outbreak. In fact, the first VGIIa-like isolate, NIH444, was isolated in Seattle in the 1970s. This raises the obvious question of why, if VGIIa has been present in the Pacific Northwest for so long, did the outbreak not begin until the late 1990s? Our whole-genome sequencing analysis suggests that microevolution may have led to heightened virulence and expansion of these isolates as an outbreak and likely occurred over a relatively short time (from ~1970 to 1999) in the Pacific Northwest, although the ancestral origin of these isolates may lie in South America, along with that of the VGIIa-like isolate ICB107. There is precedent for dramatic changes in virulence through mitotic growth in the serially passaged H99 laboratory isolates (10). An alternative hypothesis is that both VGIIa and VGIIa-like strains arose somewhere else and were both introduced to the Pacific Northwest, potentially at different times. We favor the explanation that VGIIa arose from the VGIIa-like lineage via mitotic mutation. The recent development of a congenic VGIIa mating pair (30) will allow experimental crosses between R265a and the VGIIa-like isolates to test these and other hypotheses.
Furthermore, our results suggest that the microevolution of VGIIa may have been mediated by a hypermutator phenotype as a result of an early frameshift mutation in the ortholog of MSH2, a key mismatch repair component. This may be a transient mutation that was repaired in the VGIIa lineage, after allowing short-term adaptation for higher virulence. In C. neoformans, there is evidence that hypermutators aid in adaptation to stress caused by amoeba (14); however, in this case the hypermutator strains themselves have diminished virulence in mice. This is counter to experiments done with signature-tagged mutants in C. neoformans, where both the msh2Δ and mlh1Δ mismatch repair mutants appear to proliferate faster in mouse lungs in a signature-tagged mutant experiment (31). Further experiments will be necessary to test whether Msh2 contributes to virulence in VGII C. gattii. In this case, we suspect the main contribution of the mutation in MSH2 is not a direct effect on virulence, but instead it may have potentiated changes in the progenitor of the VGIIa group, resulting in a more fit lineage that gave rise to the VGIIa outbreak strains. The msh2 frameshift mutation is only present in the VGIIa-like isolates that we hypothesize are the progenitors of the VGIIa outbreak lineage in which the MSH2 gene has been restored to wild type. Further experiments will be necessary to test these hypotheses.
In contrast, the novel VGIIc type is, as a whole, restricted to the Pacific Northwest, possibly only to the state of Oregon, and has dramatically enhanced virulence (21). This clonal lineage, unlike VGIIa, appears to have been generated as a new, highly virulent subtype via mating. The virulence phenotype of VGIIc exceeds that of any other VGII subtype tested (21), and it appears to be a clonal expansion of a single original strain in the PNW. An alternative is that VGIIc exists but has not been sampled in another geographic region. Both models are possible, but we favor a recent origin for two reasons: (i) the high virulence potential of VGIIc suggests that we would likely observe human or veterinary cases if it were present in another highly populated locale, and (ii) ongoing sampling since the start of the PNW outbreak has been unable to identify a single VGIIc, or even VGIIc-like, isolate outside the PNW. Additional sampling may reveal other VGIIc isolates in other parts of the world, but this lineage appears to be the result of a meiotic event resulting in a more fit progeny.
The proximal geographic source of the VGII outbreak has been debated. Hypotheses have included an origin in Australia (20) as well as in South America (20, 22). Data from whole-genome analysis suggest that both hypotheses may be at least partially correct. In fact, it appears that VGIIa arose in its most recent, virulent form in the Pacific Northwest, although it likely originated ancestrally in South America, and VGIIb came to the Pacific Northwest by way of Australia, although VGIIb-like isolates exist in the Caribbean and now in Florida (32) within the United States. In contrast, VGIIc arose in either the Pacific Northwest or some unsampled location elsewhere in the world. An origin other than the PNW seems less likely given the high virulence potential of VGIIc, unless it lies in a pristine niche to which the human population and other animals have not yet been exposed. Notably, whole-genome data suggest that each of the molecular types of the PNW population may have been introduced independently and possibly at different times. Sequencing of additional South American VGIIa-like isolates, as well as additional global VGIIb-like isolates, will help increase confidence in these hypotheses.
As a whole, the VGII group is robustly recombining at the population level based on a standard allele compatibility test. Within our set of samples, multiple loci from the randomly selected set show evidence for sexual recombination that rely on involvement of the clonal outbreak groups from the Pacific Northwest to complete the allele compatibility test. We cannot be certain whether these groups were the parents in such a putative cross or the offspring. Likewise, we cannot be certain that additional sampling will not reveal additional strains that may have these allele pairs. However, in combination with the presence of islands of identity within the genome, this provides strong evidence that the clonal outbreak lineages are the result of sexual reproduction, at least part of which resulted in shared genomic portions between these clonal outbreak groups and their ancestral progenitors.
Additionally, while we cannot rule out a-α opposite sex mating, it is likely that α-α unisexual mating is contributing to recombination within the VGII group globally. Outside of South America, there arze very few MATa isolates, and there are multiple genotypes that have not been isolated from South America, despite relatively extensive sampling. In this study, we sequenced three MATa isolates representing two MLST types. These isolates were more similar than predicted by the MLST approach and likely represent a single progeny from a cross that has mitotically expanded over time, analogous to the VGIIa group. Notably, these isolates have previously been shown to be fertile in an a-α laboratory cross (21), but recombination appears to be rare in the environment.
The population genomic and phylogenetic analyses of supercontig 13 strongly suggest that the genomes of isolates 2001/935-1 and IP96/1120-1 contain introgressed sequences. A very recent introgression event is expected to leave a mosaic structure of multiple introgressed sequences in the genome, although introgression events across a species boundary can result in single introgressed tracts. The highly localized nature of the discordant phylogenetic signals and evidence for recombination among haplotypes in the introgressed tract suggests that the introgression event did not occur in very recent generations, or occurred across a species boundary. The source of introgression is most likely an isolate from the VGI group. Introgression from related VG groups into VGII is likely as at least VGII and VGIII can produce viable progeny and transmit hypervirulence traits (33). The strong linkage disequilibrium and restricted introgression tract suggest that selection favored the fixation of the introgressed sequences (34).
Genomes from the VGII C. gattii isolates suggest that it is a population heavily characterized by ancestral sexual recombination, including evidence for introgression from other VG types, but also by more recent clonal expansions. We found that neither of the two original components of the VGII outbreak in the PNW appears to have been the immediate product of sexual reproduction. Instead, VGIIa may have arisen in South America and undergone expansion, driven by a transient mutator, to become more virulent in the PNW, and VGIIb appears to have arrived in the PNW largely unchanged from Australia. In contrast, VGIIc, as a result of its uniqueness to the PNW and its novel high virulence, may have arisen through a recent mating in the Pacific Northwest. This provides evidence that mitosis-driven mutations, migration, and sexual reproduction can all contribute to the development of an outbreak.
MATERIALS AND METHODS
Genome sequencing.
DNA was prepared using the cetyltrimethylammonium bromide (CTAB) isolation method as previously described (28). Library construction and genome sequencing were performed at the University of North Carolina Next Generation Sequencing Facility using a Hiseq2500. Isolates were multiplexed and run with 16 samples in each lane. Paired-end sequence libraries with approximately 300-base inserts were constructed and sequenced to a length of 100 bp per end. The Illumina Pipeline (v.1.8.2) was employed for initial processing.
Reference-based assembly.
Sequences were mapped to the R265 reference genome (35) using the short read component of the BWA aligner (36). Further refinement was performed using the Genome Analysis Toolkit (GATK) (37) pipeline version 2.4-9, including SAMtools (38) and Picard to convert from sam to bam format and to clean, sort, fix read groups, and mark duplicates, respectively. Indels were also realigned using GATK’s IndelRealigner. SNPs and indels were called with the GATK Unified Genotyper, using the haploid ploidy setting. SNP filtering was performed using VCFtools (39) to remove all SNP calls with missing data in at least one sample, a call quality below a PHRED score of 30, or a mean depth below 10. This eliminated about 7% of the initial calls, almost entirely as a result of the missing data parameter. This data set was analyzed using SnpEff (46). For some analysis, including the MAT locus, the original unfiltered set was used because MATa isolates contributed missing sites across much of the MAT locus. In addition, to examine recombination within the clonal groups, SNPS were manually filtered by visually examining the BAM files using IGV for clear evidence of misalignment of reads to eliminate obviously erroneous calls. For the introgression analyses, additional outgroup isolate data sets from VGI, VGII, VGIII, and VGIV were downloaded from the Short Read Archive and analyzed as described above (see Table S2 in the supplemental material).
Inference of phylogeny.
Maximum likelihood phylogenies were constructed using MEGA5 (40) either after alignment using Kalign (41) on artificial fasta files using GATK’s FastaAlternateReferenceMaker or from SNP matrices extracted from the VCF format using a custom Perl script, included in Text S1 in the supplemental material. Bootstrap analysis was based on a total of 500 replicates. MEGA trees were midpoint rooted. Support was calculated using 500 bootstraps. Phylogenetic analyses of introgression tracts were performed using RAxML 8.0.20 (42) using the GTRCAT nucleotide substitution model and rapid bootstrap analysis for 500 replicates.
Haplotype network inference
Haplotype networks were constructed using TCS 1.21 (43) from SNP matrices within the clonal lineages. This relies on the assumption that the strains were not separated by meiotic events, which we found evidence for from paired allele comparisons, and essentially treats the entire genome of each clonal group as a single haplotype. Indels were not used for this analysis to eliminate alignment errors.
Linkage disequilibrium analyses.
Chromosome-wide linkage disequilibria (R2) were calculated for all SNP markers with a minor allele frequency of at least 0.03 using VCFtools version 0.0.12a (39) and visualized with the R statistical software (44). Script has been included in Text S1 in the supplemental material.
Whole-genome allele compatibility tests.
One hundred random SNPs out of the 262,614 SNPs in the call set were selected using the random selection component of GATK’s SelectVariants walker. These SNPs were then collapsed into 46 unique allele patterns. Each pattern was compared pairwise with each other pattern to test for the presence of all four allele combinations. Formally, this is a test that all four products of meiosis are present: AB, Ab, aB, and ab. This analysis assumes that homoplasy is rare, an assumption that is supported by the low frequency of multiallelic SNPs in the data set, which should be more frequent than homoplasic mutations.
Nucleotide sequence accession number.
Sequences have been deposited into the Sequence Read Archive under project accession no. SRP044232.
SUPPLEMENTAL MATERIAL
MAT locus shows evidence of gene conversion. (A) Maximum likelihood phylogeny was derived from alignment of the MAT locus. This region corresponds to R265 supercontig 18, from nucleotides 26430 to 121589. Five hundred bootstraps were used to calculate support. The scale bar indicates substitutions per nucleotide site. (B) Allele compatibility test of region that is suggestive of recombination/gene conversion within the MAT locus. Reference alleles are indicated in green and alternate alleles in red. Each locus depicted is biallelic. In the paired allele compatibility diagram below, evidence for recombination is indicated by the presence of all four alleles and an hourglass shape, which is colored red for emphasis. Stars indicate loci containing allele combinations found only once within MAT, which break apart otherwise conserved haplotypes. Download
Amino acid polymorphisms in proteins encoded by genes CNBG_4871 to -4873 located in the introgressed region of the VGII isolates 2001/935-1 and IP96/1120-1. Variable amino acid positions are shown for distinct protein haplotypes grouped by the VG type of C. gattii. The protein haplotype found in introgressed isolates 2001/935-1 and IP96/1120-1 is highlighted in yellow. Download
Supercontig copy number variation among VGII isolates. The aligned read depth of resequenced isolates was normalized to the genome-wide mean coverage. Orange represents supercontigs with a read depth identical to the genome-wide average (normalized to 1). Red shows supercontigs with a 2-fold-increased read depth compared to the genome-wide average. Shades of color between orange and red indicate supercontigs with putative partial duplications. Download
French clinical isolate 93.980 has highly aneuploid supercontig 6. Depth of coverage across supercontig 6 of 93.980 was determined using the IGVtools count program and visualized using IGV (45). The genome average depth of coverage is indicated by a solid black horizontal line. Regions of the supercontig 6 range include 1N, 2N, 3N, 4N, 5N, and 6N. Download
Predicted impact of VGIIa-like SNPs and indels. SNPeff predictions of SNP and indel effects on genes that divide the VGIIa and VGIIa-like lineages.
Outgroup isolates for introgression analyses. Additional VGI, VGII, VGIII, and VGIV isolates were used as outgroups in the phylogenetic introgression analyses.
Scripts. Download
ACKNOWLEDGMENTS
We thank Rodrigo Olarte for helpful comments about analysis of allele compatibility data and Sheng Sun for helpful comments on the manuscript. We also thank Teun Boekhout, the CBS Fungal Biodiversity Center, and Shawn Lockhart, CDC, for providing strains.
This study was supported by NIH/NIAID R37 grant AI39115-17 and R01 grant AI50113-10 to J.H. D.C. was supported by grant PA00P3_145360 from the Swiss National Science Foundation and by a grant from the Canadian Institutes of Health Research to J.W.K. D.A.C. was supported by the Australian National Health and Medical Research Council, grant #571354.
Footnotes
Citation Billmyre RB, Croll D, Li W, Mieczkowski P, Carter DA, Cuomo CA, Kronstad JW, Heitman J. 2014. Highly recombinant VGII Cryptococcus gattii population develops clonal outbreak clusters through both sexual macroevolution and asexual microevolution. mBio 5(4):e01494-14. doi:10.1128/mBio.01494-14.
REFERENCES
- 1. Morens DM, Fauci AS. 2013. Emerging infectious diseases: threats to human health and global stability. PLoS Pathog. 9:e1003467. 10.1371/journal.ppat.1003467 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Park BJ, Wannemuehler KA, Marston BJ, Govender N, Pappas PG, Chiller TM. 2009. Estimation of the current global burden of cryptococcal meningitis among persons living with HIV/AIDS. AIDS 23:525–530. 10.1097/QAD.0b013e328322ffac [DOI] [PubMed] [Google Scholar]
- 3. Marr KA, Carter RA, Crippa F, Wald A, Corey L. 2002. Epidemiology and outcome of mould infections in hematopoietic stem cell transplant recipients. Clin. Infect. Dis. 34:909–917. 10.1086/339202 [DOI] [PubMed] [Google Scholar]
- 4. Bush RM, Bender CA, Subbarao K, Cox NJ, Fitch WM. 1999. Predicting the evolution of human influenza A. Science 286:1921–1925. 10.1126/science.286.5446.1921 [DOI] [PubMed] [Google Scholar]
- 5. Smith GJ, Vijaykrishna D, Bahl J, Lycett SJ, Worobey M, Pybus OG, Ma SK, Cheung CL, Raghwani J, Bhatt S, Peiris JS, Guan Y, Rambaut A. 2009. Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459:1122–1125. 10.1038/nature08182 [DOI] [PubMed] [Google Scholar]
- 6. Smith GJ, Bahl J, Vijaykrishna D, Zhang J, Poon LL, Chen H, Webster RG, Peiris JS, Guan Y. 2009. Dating the emergence of pandemic influenza viruses. Proc. Natl. Acad. Sci. U. S. A. 106:11709–11712. 10.1073/pnas.0904991106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Grigg ME, Bonnefoy S, Hehl AB, Suzuki Y, Boothroyd JC. 2001. Success and virulence in Toxoplasma as the result of sexual recombination between two distinct ancestries. Science 294:161–165. 10.1126/science.1061888 [DOI] [PubMed] [Google Scholar]
- 8. Wendte JM, Miller MA, Lambourn DM, Magargal SL, Jessup DA, Grigg ME. 2010. Self-mating in the definitive host potentiates clonal outbreaks of the apicomplexan parasites Sarcocystis neurona and Toxoplasma gondii. PLoS Genet. 6:e1001261. 10.1371/journal.pgen.1001261 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Ni M, Feretzaki M, Li W, Floyd-Averette A, Mieczkowski P, Dietrich FS, Heitman J. 2013. Unisexual and heterosexual meiotic reproduction generate aneuploidy and phenotypic diversity de novo in the yeast Cryptococcus neoformans. PLoS Biol. 11:e1001653. 10.1371/journal.pbio.1001653 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Janbon G, Ormerod KL, Paulet D, Byrnes EJ, Yadav V, Chatterjee G, Mullapudi N, Hon C-C, Billmyre RB, Brunel F, Bahn Y-S, Chen W, Chen Y, Chow EWL, Coppée J-Y, Floyd-Averette A, Gaillardin C, Gerik KJ, Goldberg J, Gonzalez-Hilarion S, Gujja S, Hamlin JL, Hsueh Y-P, Ianiri G, Jones S, Kodira CD, Kozubowski L, Lam W, Marra M, Mesner LD, Mieczkowski PA, Moyrand F, Nielsen K, Proux C, Rossignol T, Schein JE, Sun S, Wollschlaeger C, Wood IA, Zeng Q, Neuvéglise C, Newlon CS, Perfect JR, Lodge JK, Idnurm A, Stajich JE, Kronstad JW, Sanyal K, Heitman J, Fraser JA, Cuomo CA, Dietrich FS. 2014. Analysis of the genome and transcriptome of Cryptococcus neoformans var. grubii reveals complex RNA expression and microevolution leading to virulence attenuation. PLoS Genet. 10:e1004261. 10.1371/journal.pgen.1004261 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Hu G, Chen SH, Qiu J, Bennett JE, Myers TG, Williamson PR. 2014. Microevolution during serial mouse passage demonstrates FRE3 as a virulence adaptation gene in Cryptococcus neoformans. mBio 5:e00941-14. 10.1128/mBio.00941-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Strand M, Prolla TA, Liskay RM, Petes TD. 1993. Destabilization of tracts of simple repetitive DNA in yeast by mutations affecting DNA mismatch repair. Nature 365:274–276. 10.1038/365274a0 [DOI] [PubMed] [Google Scholar]
- 13. Verstrepen KJ, Jansen A, Lewitter F, Fink GR. 2005. Intragenic tandem repeats generate functional variability. Nat. Genet. 37:986–990. 10.1038/ng1618 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Magditch DA, Liu TB, Xue C, Idnurm A. 2012. DNA mutations mediate microevolution between host-adapted forms of the pathogenic fungus Cryptococcus neoformans. PLoS Pathog. 8:e1002936. 10.1371/journal.ppat.1002936 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Byrnes EJ, Bildfell RJ, Frank SA, Mitchell TG, Marr KA, Heitman J. 2009. Molecular evidence that the range of the Vancouver Island outbreak of Cryptococcus gattii infection has expanded into the Pacific Northwest in the United States. J. Infect. Dis. 199:1081–1086. 10.1086/597306 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Kidd SE, Hagen F, Tscharke RL, Huynh M, Bartlett KH, Fyfe M, Macdougall L, Boekhout T, Kwon-Chung KJ, Meyer W. 2004. A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on Vancouver Island (British Columbia, Canada). Proc. Natl. Acad. Sci. U. S. A. 101:17258–17263. 10.1073/pnas.0402981101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Hoang LM, Maguire JA, Doyle P, Fyfe M, Roscoe DL. 2004. Cryptococcus neoformans infections at Vancouver Hospital and Health Sciences Centre (1997-2002): epidemiology, microbiology and histopathology. J. Med. Microbiol. 53:935–940. 10.1099/jmm.0.05427-0 [DOI] [PubMed] [Google Scholar]
- 18. Singer LM, Meyer W, Firacative C, Thompson GR, Samitz E, Sykes JE. 2014. Antifungal drug susceptibility and phylogenetic diversity among Cryptococcus isolates from dogs and cats in North America. J. Clin. Microbiol. 52:2061–2070. 10.1128/JCM.03392-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Lockhart SR, Iqbal N, Harris JR, Grossman NT, DeBess E, Wohrle R, Marsden-Haug N, Vugia DJ. 2013. Cryptococcus gattii in the United States: genotypic diversity of human and veterinary isolates. PLoS One 8:e74737. 10.1371/journal.pone.0074737 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Fraser JA, Giles SS, Wenink EC, Geunes-Boyer SG, Wright JR, Diezmann S, Allen A, Stajich JE, Dietrich FS, Perfect JR, Heitman J. 2005. Same-sex mating and the origin of the Vancouver Island Cryptococcus gattii outbreak. Nature 437:1360–1364. 10.1038/nature04220 [DOI] [PubMed] [Google Scholar]
- 21. Byrnes EJ, Li W, Lewit Y, Ma H, Voelz K, Ren P, Carter DA, Chaturvedi V, Bildfell RJ, May RC, Heitman J. 2010. Emergence and pathogenicity of highly virulent Cryptococcus gattii genotypes in the northwest United States. PLoS Pathog. 6:e1000850. 10.1371/journal.ppat.1000850 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Hagen F, Ceresini PC, Polacheck I, Ma H, van Nieuwerburgh F, Gabaldón T, Kagan S, Pursall ER, Hoogveld HL, van Iersel LJ, Klau GW, Kelk SM, Stougie L, Bartlett KH, Voelz K, Pryszcz LP, Castañeda E, Lazera M, Meyer W, Deforce D, Meis JF, May RC, Klaassen CH, Boekhout T. 2013. Ancient dispersal of the human fungal pathogen Cryptococcus gattii from the Amazon rainforest. PLoS One 8:e71148. 10.1371/journal.pone.0071148 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Gillece JD, Schupp JM, Balajee SA, Harris J, Pearson T, Yan Y, Keim P, DeBess E, Marsden-Haug N, Wohrle R, Engelthaler DM, Lockhart SR. 2011. Whole genome sequence analysis of Cryptococcus gattii from the Pacific Northwest reveals unexpected diversity. PLoS One 6:e28550. 10.1371/journal.pone.0028550 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Carter D, Campbell L, Saul N, Krockenberger M. 2011. Sexual reproduction of Cryptococcus gattii: a population genetics perspective, p 299–311 In Heitman J, Kozel TR, Kwon-Chung KJ, Perfect JR, Casadevall A. (ed), Cryptococcus—from human pathogen to model yeast. ASM Press, Washington, DC [Google Scholar]
- 25. Rosenblum EB, James TY, Zamudio KR, Poorten TJ, Ilut D, Rodriguez D, Eastman JM, Richards-Hrdlicka K, Joneson S, Jenkinson TS, Longcore JE, Parra Olea G, Toledo LF, Arellano ML, Medina EM, Restrepo S, Flechas SV, Berger L, Briggs CJ, Stajich JE. 2013. Complex history of the amphibian-killing chytrid fungus revealed with genome resequencing data. Proc. Natl. Acad. Sci. U. S. A. 110:9385–9390. 10.1073/pnas.1300130110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Reenan RA, Kolodner RD. 1992. Isolation and characterization of two Saccharomyces cerevisiae genes encoding homologs of the bacterial HexA and MutS mismatch repair proteins. Genetics 132:963–973 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Velagapudi R, Hsueh YP, Geunes-Boyer S, Wright JR, Heitman J. 2009. Spores as infectious propagules of Cryptococcus neoformans. Infect. Immun. 77:4345–4355. 10.1128/IAI.00542-09 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Sun S, Hsueh YP, Heitman J. 2012. Gene conversion occurs within the mating-type locus of Cryptococcus neoformans during sexual reproduction. PLoS Genet. 8:e1002810. 10.1371/journal.pgen.1002810 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Sionov E, Lee H, Chang YC, Kwon-Chung KJ. 2010. Cryptococcus neoformans overcomes stress of azole drugs by formation of disomy in specific multiple chromosomes. PLoS Pathog. 6:e1000848. 10.1371/journal.ppat.1000848 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Zhu P, Zhai B, Lin X, Idnurm A. 2013. Congenic strains for genetic analysis of virulence traits in Cryptococcus gattii. Infect. Immun. 81:2616–2625. 10.1128/IAI.00018-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Liu OW, Chun CD, Chow ED, Chen C, Madhani HD, Noble SM. 2008. Systematic genetic analysis of virulence in the human fungal pathogen Cryptococcus neoformans. Cell 135:174–188. 10.1016/j.cell.2008.07.046 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Kunadharaju R, Choe U, Harris JR, Lockhart SR, Greene JN. 2013. Cryptococcus gattii, Florida, USA, 2011. Emerg. Infect. Dis. 19:519–521. 10.3201/eid1903.121399 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Voelz K, Ma H, Phadke S, Byrnes EJ, Zhu P, Mueller O, Farrer RA, Henk DA, Lewit Y, Hsueh Y-P, Fisher MC, Idnurm A, Heitman J, May RC. 2013. Transmission of hypervirulence traits via sexual reproduction within and between lineages of the human fungal pathogen Cryptococcus gattii. PLoS Genet. 9:e1003771. 10.1371/journal.pgen.1003771 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Hedrick PW. 2013. Adaptive introgression in animals: examples and comparison to new mutation and standing variation as sources of adaptive variation. Mol. Ecol. 22:4606–4618. 10.1111/mec.12415 [DOI] [PubMed] [Google Scholar]
- 35. D’Souza CA, Kronstad JW, Taylor G, Warren R, Yuen M, Hu G, Jung WH, Sham A, Kidd SE, Tangen K, Lee N, Zeilmaker T, Sawkins J, McVicker G, Shah S, Gnerre S, Griggs A, Zeng Q, Bartlett K, Li W, Wang X, Heitman J, Stajich JE, Fraser JA, Meyer W, Carter D, Schein J, Krzywinski M, Kwon-Chung KJ, Varma A, Wang J, Brunham R, Fyfe M, Ouellette BFF, Siddiqui A, Marra M, Jones S, Holt R, Birren BW, Galagan JE, Cuomo CA. 2011. Genome variation in Cryptococcus gattii, an emerging pathogen of immunocompetent hosts. mBio 2:e00342-10. 10.1128/mBio.00342-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Li H, Durbin R. 2009. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. 10.1093/bioinformatics/btp324 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA. 2010. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20:1297–1303. 10.1101/gr.107524.110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, 1000 Genome Project Data Processing Subgroup 2009. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25:2078–2079. 10.1093/bioinformatics/btp352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST, McVean G, Durbin R, 1000 Genomes Project Analysis Group 2011. The variant call format and VCFtools. Bioinformatics 27:2156–2158. 10.1093/bioinformatics/btr330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. 2011. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28:2731–2739. 10.1093/molbev/msr121 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Lassmann T, Sonnhammer EL. 2005. Kalign—an accurate and fast multiple sequence alignment algorithm. BMC Bioinformatics 6:298. 10.1186/1471-2105-6-298 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Stamatakis A. 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. 10.1093/bioinformatics/btu033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Clement M, Posada D, Crandall KA. 2000. TCS: a computer program to estimate gene genealogies. Mol. Ecol. 9:1657–1659. 10.1046/j.1365-294x.2000.01020.x [DOI] [PubMed] [Google Scholar]
- 44. R Development Core Team 2005. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria [Google Scholar]
- 45. Thorvaldsdóttir H, Robinson JT, Mesirov JP. 2013. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14:178–192. 10.1093/bib/bbs017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM. 2012. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6:80–92. 10.4161/fly.19695 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
MAT locus shows evidence of gene conversion. (A) Maximum likelihood phylogeny was derived from alignment of the MAT locus. This region corresponds to R265 supercontig 18, from nucleotides 26430 to 121589. Five hundred bootstraps were used to calculate support. The scale bar indicates substitutions per nucleotide site. (B) Allele compatibility test of region that is suggestive of recombination/gene conversion within the MAT locus. Reference alleles are indicated in green and alternate alleles in red. Each locus depicted is biallelic. In the paired allele compatibility diagram below, evidence for recombination is indicated by the presence of all four alleles and an hourglass shape, which is colored red for emphasis. Stars indicate loci containing allele combinations found only once within MAT, which break apart otherwise conserved haplotypes. Download
Amino acid polymorphisms in proteins encoded by genes CNBG_4871 to -4873 located in the introgressed region of the VGII isolates 2001/935-1 and IP96/1120-1. Variable amino acid positions are shown for distinct protein haplotypes grouped by the VG type of C. gattii. The protein haplotype found in introgressed isolates 2001/935-1 and IP96/1120-1 is highlighted in yellow. Download
Supercontig copy number variation among VGII isolates. The aligned read depth of resequenced isolates was normalized to the genome-wide mean coverage. Orange represents supercontigs with a read depth identical to the genome-wide average (normalized to 1). Red shows supercontigs with a 2-fold-increased read depth compared to the genome-wide average. Shades of color between orange and red indicate supercontigs with putative partial duplications. Download
French clinical isolate 93.980 has highly aneuploid supercontig 6. Depth of coverage across supercontig 6 of 93.980 was determined using the IGVtools count program and visualized using IGV (45). The genome average depth of coverage is indicated by a solid black horizontal line. Regions of the supercontig 6 range include 1N, 2N, 3N, 4N, 5N, and 6N. Download
Predicted impact of VGIIa-like SNPs and indels. SNPeff predictions of SNP and indel effects on genes that divide the VGIIa and VGIIa-like lineages.
Outgroup isolates for introgression analyses. Additional VGI, VGII, VGIII, and VGIV isolates were used as outgroups in the phylogenetic introgression analyses.
Scripts. Download