Abstract
Pneumonia is one of the major health care problems in developing and industrialized countries and is associated with considerable morbidity and mortality. Despite advances in knowledge of this illness, the availability of intensive care units (ICU), and the use of potent antimicrobial agents and effective vaccines, the mortality rates remain high1. Streptococcus pneumoniae is the leading pathogen of community-acquired pneumonia (CAP) and one of the most common causes of bacteremia in humans. This pathogen is equipped with an armamentarium of surface-exposed adhesins and virulence factors contributing to pneumonia and invasive pneumococcal disease (IPD). The assessment of the in vivo role of bacterial fitness or virulence factors is of utmost importance to unravel S. pneumoniae pathogenicity mechanisms. Murine models of pneumonia, bacteremia, and meningitis are being used to determine the impact of pneumococcal factors at different stages of the infection. Here we describe a protocol to monitor in real-time pneumococcal dissemination in mice after intranasal or intraperitoneal infections with bioluminescent bacteria. The results show the multiplication and dissemination of pneumococci in the lower respiratory tract and blood, which can be visualized and evaluated using an imaging system and the accompanying analysis software.
Keywords: Infection, Issue 84, Gram-Positive Bacteria, Streptococcus pneumoniae, Pneumonia, Bacterial, Respiratory Tract Infections, animal models, community-acquired pneumonia, invasive pneumococcal diseases, Pneumococci, bioimaging, virulence factor, dissemination, bioluminescence, IVIS Spectrum
Introduction
Respiratory tract infections caused by viruses or bacteria remain one of the most common community-acquired or clinical problems worldwide causing approximately one third of all death worldwide. The key bacterial species are Haemophilus influenzae and Streptococcus pneumoniae2. However, these bacterial species are normally common constituents of the natural respiratory tract flora. Thus bacterial carriage is also of certain risk for invasive disease and depending on the immune status or predispositions of the individuals. The asymptomatic colonization is triggered to invasive infections. Streptococcus pneumoniae is the leading pathogen of community-acquired pneumonia (CAP) and one of the most common causes of bacteremia in humans. In healthy individuals S. pneumoniae (pneumococci) are often asymptomatic and harmless colonizers of the upper respiratory tract, where they are confronted with nonpathogenic bacteria of the resident flora but also with pathogens such as Haemophilus spp. or Staphylococcus aureus and the first line of the human immune defense system. Carriage rates are highest in young children (37%) and even higher within crowded day care centers (58%)3-5. The youngest population and the elderly, receiving the pneumococcus via aerosol transmission from carriers and nasopharyngeal secretions6, belong to the high risk groups and vaccination using one of the pneumococcal conjugate vaccines (PCV10 or PCV13 in children and 23-valent polysaccharide PPSV23 in adults) is recommended in the United States (US) and many European countries4. The PPSV23 covers serotypes responsible for ~90% of the bacteremic pneumococcal diseases in the US and Europe, preventing thus efficiently invasive pneumococcal diseases (IPD) in adults, while the PCVs cover the most prevalent serotypes in children. Consequently, IPD due to vaccine types (VT) are reduced but nonvaccine serotypes displaying a high virulence potential and antibiotic resistance have emerged4,7-12. The nasopharynx as the reservoir is the starting point for pneumococci to spread to the sinuses or middle ears initiating harmful local infections. More important, pneumococci spread directly via the airway to the bronchia and lung resulting in life-threatening CAP4,13. Lung infections are often accompanied with tissue and barrier destruction, thus enabling the pathogen to spread into the blood and causing IPD. Incidences of CAP and IPD are highest in immunocompromised persons or at the extremes of age4,13. The circumstances responsible for the conversion from a commensal to a pathogen with high virulence are still under debate. However, besides changes in the host susceptibility and evolutionary adaptation accompanied with higher virulence and the increase in antibiotic resistances have been suggested to have a crucial impact on pneumococcal infections14-16.
The pathogen is endowed with a multiplicity of adhesins mediating intimate contact to mucosal epithelial cells. After surmounting the airway mucus, pneumococcal adherence to host cells is facilitated via direct interactions of surface-exposed adhesins with cellular receptors and by exploiting extracellular matrix components or serum proteins as bridging molecules4,17,18. As versatile pathogens pneumococci are also equipped with factors involved in evasion of host immune defense mechanisms. Moreover, they have the capacity to adapt to various host milieus such as the lung, blood, and cerebrospinal fluid (CSF), respectively5,17,19,20.
The impact of bacterial factors on pathogenesis and inflammatory host responses is investigated in experimental animal models of pneumonia, bacteremia, or meningitis21-25. Despite being a human pathogen, these models are well-established to decipher pneumococcal tissue tropism, virulence mechanisms, or protectivity of pneumococcal vaccine candidates. The genetic background of inbred mouse strains determines the susceptibility to pneumococci. BALB/c mice intranasally infected with pneumococci were found to be resistant, while CBA/Ca and SJL mice were more susceptible against pneumococcal infections22. This implies that, similar to humans, the genetic background and the host defense mechanisms determine the outcome of the infection. Hence, further efforts are necessary to unravel resistance loci in the genome of mice less susceptible to pneumococcal infections. The findings have led to changes in in vivo virulence protocols. Instead of the inbred BALB/c mice often used in the past, the highly susceptible CD-1/MF1 outbred mouse strains are nowadays often used to study the effect of loss-of-function pneumococcal virulence or fitness factors26-28. Moreover, the availability of bioluminescent pneumococci and optical imaging techniques allows the real-time bioluminescence bioimaging of infections. In pneumococci the optimized luxABCDE gene cassette (plasmid pAUL-A Tn4001 luxABCDE Kmr) has been inserted into a single integration site of the chromosome by transposon mutagenesis. Bioluminescent pneumococci have been employed to assess the attenuation of pneumococcal mutants deficient in virulence or fitness factors and their translocation from one anatomical site to another26,28-31.
Here we provide a protocol for the bioimaging of pneumococcal infections in a murine pneumonia or sepsis model. Amplification and dissemination of bioluminescent pneumococci in intranasally or intraperitoneally infected mice can easily be monitored over time using an optical imaging system and the same animal at different time points.
Protocol
The animal infection experiments described here must be performed in strict accordance with the local and international (e.g. European Health Law of the Federation of Laboratory Animal Science Associations (FELASA)) guidelines and regulations for the use of vertebrate animals. The experiments have to be approved by the local ethical board and Institutional Animal Care Committee. All experiments with S. pneumoniae in the laboratory or the animal infections are conducted in a Class II Biosafety Cabinet.
1. Preparation of the Infectious Stock of Bioluminescent Pneumocococi
Prepare stock pneumococcal cultures in Todd-Hewitt broth supplemented with 1% (w/v) yeast extract and a final concentration of 20% glycerol for maintenance at 80 °C.
Plate a small amount of deep-frozen stock pneumococcal culture onto a blood agar plate and incubate the bacteria for 10 hr at 37 °C in 5% CO2. The blood agar plate contains the appropriate antibiotics such as kanamycin (150 µg/ml) for the transposon insertion of the luxABCDE gene cassette into the genome.
Sub-cultivate a fresh colony onto a new blood agar plate for a maximum of 10 hr.
Inoculate THY medium supplemented with 10% (v/v) heat inactivated (30 min at 56 °C) fetal bovine serum (FBS) with pneumococci and start the culture at an OD600 of 0.05-0.07.
Incubate pneumococci without agitation at 37 °C and 5% CO2 until the culture reaches an OD600 of 0.35-0.40. Growth will take between 3-4 hr.
Harvest pneumococci by centrifugation at 3,750 x g for 10 min and resuspend bioluminescent pneumococci in phosphate buffered saline pH 7.4 (PBS) supplemented with 0.5% FBS.
Adjust the inoculum to the desired concentration in PBS/10% FBS. The infection dose depends on the pneumococcal strain used and the genetic background of the mice. To infect outbred CD-1 mice intranasally with S. pneumoniae D39lux the infection dose should be adjusted to 1 x 107 CFU in a suspension of 20 µl supplemented with 90 Units hyaluronidase. Do not shake or vortex pneumococci, since this will trigger autolysis.
Verify the inoculum concentration by plating 10 fold serial dilutions onto blood agar and enumeration of the colonies after overnight growth at 37 °C in 5% CO2.
Verify the bioluminescence of the pneumococcal inoculum by measuring the bioluminescence using an optical imaging system.
2. Intranasal and Intraperitoneal Infection of Mice with Bioluminescent Pneumococci
Use female outbred mice in the age of 7-10 weeks for the acute pneumonia and sepsis model. The weight of these mice should be between 20-30 g.
Anesthetize mice by intraperitoneal injection of ketamine and xylazine. Prepare a mixture 1.0 mg ketamine and 0.1 mg xylazine in 100 µl sterile 0.9 % (w/v) sodium chloride for mice with a body weight of 20 g. This is consistent with a dose of 50 mg/kg body weight for ketamine and 5 mg/kg body weight for xylazine. To be accurate, weigh mice before the injection of the anesthetics.
Inject the mixture into the intraperitoneal cavity and place the animals back into the cage until the anesthetics act and the animals are narcotized. Breathing of mice will become very slow and regular. Mice have to be fully narcotized, so that the intranasal inoculation and inhalation of the bacterial suspension will not result in sneezing and hence, loss of inoculum. This can be assessed by pinching the mouse at the end of their tail; a fully narcotized mouse will lack responsiveness.
Gently pick up a narcotized mouse and hold the mouse between the fingers and thumb with the nose upright (Figure 1).
Use a pipette with long narrow tips (Gel Loader Tips) and drop bioluminescent pneumococci in multiple small droplets onto the nares (10 µl/nostril) of the mouse26-29. The mouse will involuntarily inhale the bacteria. Hold the mouse 1-2 min upright and observe whether the mouse sneezes or keeps the inoculum.
Infect 7-10 week old CD-1 mice intranasally with an infection dose of 1 x 107 or 7.5 x 107 pneumococci of strain D39 and TIGR4, respectively, to monitor the infection in real time. The detection limit for the system without any absorption by the host tissue is approximately 1 x 106 bioluminescent bacteria.
Infect mice intraperitoneally (I.P.) to assess and bioimage virulence of pneumococci in a sepsis mouse infection model. Use 5 x 103 CFU in 100 µl of strain D39 and 1 x 104 of strain TIGR4. Mice are not anesthetized when using the I.P. route.
Inoculate control mice with PBS.
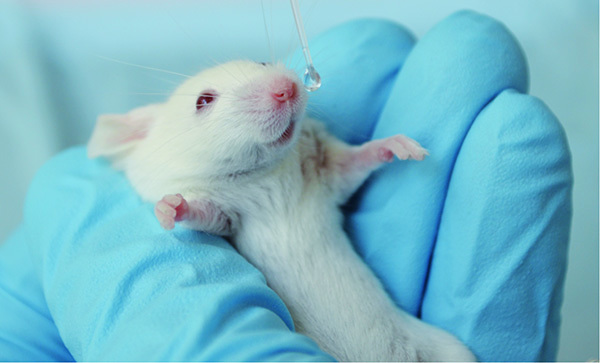 Figure 1.
Intranasal infection of CD-1 mice with S. pneumoniae. Click here to view larger image.
Figure 1.
Intranasal infection of CD-1 mice with S. pneumoniae. Click here to view larger image.
3. Visualizing Pneumococcal Dissemination into the Lung and Blood Using the Imaging System
Image dissemination of pneumococci after intranasal infection of female outbred CD1 mice in real-time using an in vivo imaging system.
Switch on the imaging system and start the image analysis software located on the preconfigurated computer.
The imaging system initializes automatically and the CCD camera is thermoelectrically cooled to 90 °C before being active.
The settings and binning depend on the number of mice measured simultaneously and on the strength of the bioluminescent signal. The field of view (FOV) to monitor the bioluminescence of the animals is variable and the magnification ranges between an FOV of 22.5 cm to measure five mice, and an FOV of 3.9 cm to measure a single mouse or parts of the animal like the chest or head.
Use routinely an exposure time of 1 min, a medium degree of binning, and an FOV of 22.5 cm (set D). Firstly, select the luminescent imaging mode and then set the imaging parameters.
The emission of the signal may depend on the bacterial strain and mouse strain used, and the pigment of the mouse. Shave the chest of mice when using e.g. C57BL/6 mice, which can be advantageous when imaging the development of pneumonia in these mice.
Image infected mice at prechosen time intervals. Measure the bioluminescence of mice in the acute pneumonia model at time intervals of maximum 8-12 hr. Monitor also the welfare of infected mice by observing their appearance, behavior, and determination of the loss of weight.
Anesthetize mice in the narcotic chamber of the anesthesia system of the imaging equipment prior to the measurements in the chamber. Start inhalation of a mixture of isoflurane and oxygen and wait until the mice breathe slowly and regularly.
Place anesthetized animals into the imaging chamber of the system and keep mice in the supine position. The CCD camera on the top of the chamber will then image the murine respiratory tract.
Place the animals with their nose into to tubes to allow inhalation of the anesthetics.
Isoflurane entry into the imaging chamber via a gas tubing is allowed to maintain anesthesia.
To activate the CCD camera on top of the chamber and to start bioluminescent optical imaging, press “Acquire” in the imaging software. Immediately a photograph of the mice in the closed chamber appears on the screen. After one minute of measurement an overlay of the bioluminescence data and the photo is shown.
Stop anesthesia after the image overlay appears in the window of the software and put the mice back into their cages.
Mice are monitored for recovery.
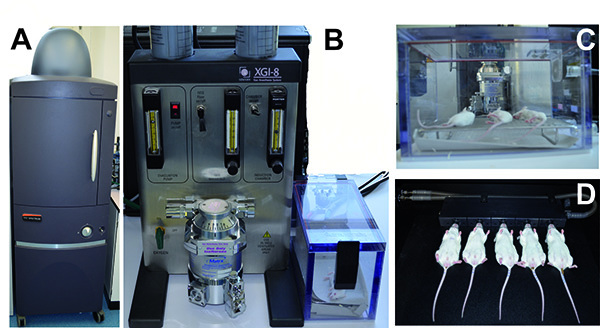 Figure 2.
Anesthesia and monitoring of mice infected with bioluminescent pneumococci using the IVIS anesthesia and IVIS Spectrum system, respectively. A) The IVIS Spectrum imaging system. B) The IVIS anesthesia system with the incubation chamber. C) Mice anesthetized in the incubation chamber. D) Mice located within the IVIS Spectrum Imaging chamber with their nose inhaling the anesthetic. Click here to view larger image.
Figure 2.
Anesthesia and monitoring of mice infected with bioluminescent pneumococci using the IVIS anesthesia and IVIS Spectrum system, respectively. A) The IVIS Spectrum imaging system. B) The IVIS anesthesia system with the incubation chamber. C) Mice anesthetized in the incubation chamber. D) Mice located within the IVIS Spectrum Imaging chamber with their nose inhaling the anesthetic. Click here to view larger image.
4. Quantification and Evaluation of the Bioluminescence of Mice Infected with Bioluminescent S. pneumoniae
Determine the bioluminescence intensities of the mice or of a selected region of the mice by quantifying the total photon emission (photons/sec) using the image analysis software.
Use the values of the photon emission that are provided in an Excel data sheet to prepare a graph. Since the data variation is often high within the grouped mice, generate a box whisker graph to display the differences in mice infected with the different pneumococcal strains.
Provide the photons per mice or alternatively the results as the average of radiance (photons/sec/cm2/sr) of a selected area.
Measurement data of ROI (region-of-interest) in photon mode can be quantitatively compared across different in vivo imaging systems with different camera settings, since the measurements in units of radiance automatically take into account camera settings (e.g. integration time, binning, f/stop, and field of view).
Representative Results
The acquisition and uptake of methionine is of central importance for pneumococci to maintain fitness in their host niche32,33. The methionine ABC transporter lipoprotein is encoded in D39 by the spd_0151 gene (TIGR4: sp_0149) and named MetQ 32. Pneumococci further produce methionine biosynthesis enzymes (D39: Spd_0510 – Spd_0511; TIGR4 Sp_0585 – Sp_0586, MetE and MetF). The lack of methionine in a chemically defined medium affects growth of pneumococci and similar, the lack of the methionine binding lipoprotein MetQ impaired uptake of methionine32,33. This defect could be restored by the addition of high concentrations of methionine. In the mouse colonization experiments the mutant deficient in MetQ was not attenuated in virulence as shown in coinfection experiments with the isogenic wild-type, while the lack of MetQ attenuated pneumococci in mouse models of sepsis and pneumonia as shown by determining the bacterial load32. Real-time optical bioimaging using the IVIS Spectrum used here allows monitoring the multiplication and dissemination of bacteria in a single infected animal at different time points.
In the example shown here, we have assessed the impact of the methionine binding lipoprotein MetQ on colonization and virulence by applying the acute pneumonia infection mouse model. CD-1 outbred mice (n=10) were infected intranasally with 1 x 107 bacteria of wild-type strain S. pneumoniae D39lux (PN149) or its isogenic mutant D39luxΔmetQ (Table 1). The metQ mutant (PN252) was constructed by insertion-deletion mutagenesis of the metQ gene in bioluminescent D39 pneumococci. Therefore, the metQ gene, 5´ sequence and 3´sequence were amplified by PCR using primers 382 and 385. The PCR product was cloned into the vector pGEM-T-easy, which resulted in plasmid p559 (Table 1). By inverse PCR using primers 383 and 384 (Table 2) the metQ sequence (nt 58 to nt 777) was deleted and replaced by an erythromycin resistance gene cassette (ermB) amplified by PCR using plasmid pE89 (Table 1) as DNA-template and the primer pair ermB_105/ermB_106 (Table 2). The resulting plasmid p56334 was transformed into pneumococci as described previously using competence-stimulating peptide-1. Mutants deficient in the lipoprotein MetQ (Spd_0151) were cultivated in THY or on blood agar plates supplemented with erythromycin (5 µg/ml).
The health status of the mice was continuously monitored. Animals were sacrificed when showing no proper response or signs of morbidity. Moreover, pneumococcal dissemination from the nasopharynx into the lungs and blood, leading to severe pneumonia and sepsis, was monitored by measuring the bioluminescence every eight hours. In each group of infected mice 7 out of 10 mice developed severe signs of illness, while the other 3 mice showed no bioluminescence and survived (Figure 3A). Mice successfully infected with the bioluminescent wild-type D39lux developed 32 hours post-infection severe lung infections, which progresses within 8-16 hr to sepsis (Figure 3A). At time point 96 hr all mice succumbed that had developed pneumonia and sepsis post-infection with D39lux. The loss-of-function of the methionine binding lipoprotein significantly impaired the virulence of pneumococci. After 32 hr only one mouse infected with the metQ mutant showed a weak lung infection, while the others showed no obvious signs of illness and pneumonia. However, after 48 hr severe lung infections became also apparent in mice infected with the mutant. As a result, these mice developed sepsis 72 hr post-infection and became moribund later than 72 hr post-infection, demonstrating that in the acute pneumonia infection model the deficiency of MetQ attenuates pneumococci.
Table 1. Strain and plasmid list.
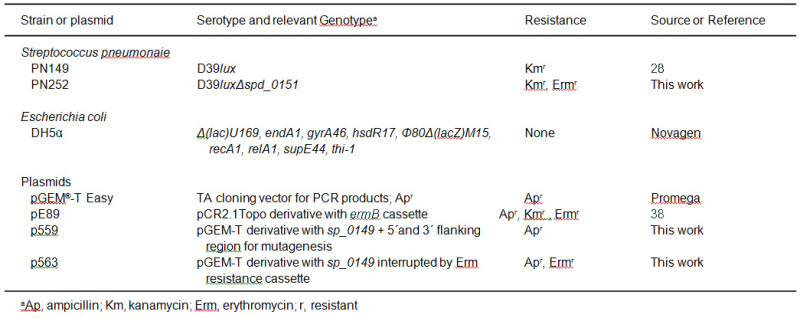
Table 2.Primer list. Restriction sites are underlined.
| Primer Intended Use | Primer Name | Sequence (5'-3') |
| Insertion-deletion mutagenesis | ||
| Amplification of sp_0149 + 5' and 3' flanking region | 382 385 | 5'-CTACTACTAGAATTCATGCTGAACACACGGACAAC-3' 5'-AACCTTCCAAGCTGCAGCCGCTCCCTCCATGATAAAG-3' |
| Inverse PCR of sp_0149 + 5' and 3' flanking region (pGEMTeasy) | 383 384 | 5'-ATCATCATCATCGAAGCTTAGCCAAACCT GCGACTGTAG-3' 5'-ACTCACTCACTGAAGCTTATCGCAGCTTA CCACACAGA-3' |
| Antibiotic cassette amplification | ||
| erythromycin (ermB) | ermB_105 ermB_106 | 5'-GATGATGATGATCCCGGGTACCAAGCTTGA ATTCACGGTTCGTGTTCGTGCTG-3' 5'-AGTGAGTGAGTCCCGGGCTCGAGAAGCTT GAATTCGTAGGCGCTAGGGACCTC-3' |
The bioluminescence measured using the imaging software was also calculated. Similar to the visualization of the infection, the quantification of the bioluminescent flux showed significant differences between wild-type infected and metQ infected mice 32 and 40 hr after infection. Thus, the loss-of-function of MetQ attenuates pneumococci and results in a significant delay of invasive infection, but does not result in avirulent bacteria.
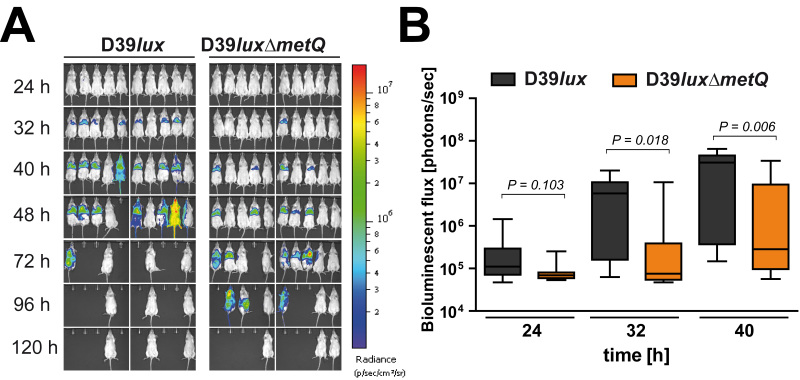 Figure 3.Monitoring of pneumococcal dissemination in mice after intranasal infection. A) Bioluminescent optical imaging of mice infected intranasally with D39lux or the isogenic mutant D39luxΔmetQ. B) The values of the bioluminescence intensities of grouped mice (n = 10) are shown for indicated time points in a box whisker graph. Click here to view larger image.
Figure 3.Monitoring of pneumococcal dissemination in mice after intranasal infection. A) Bioluminescent optical imaging of mice infected intranasally with D39lux or the isogenic mutant D39luxΔmetQ. B) The values of the bioluminescence intensities of grouped mice (n = 10) are shown for indicated time points in a box whisker graph. Click here to view larger image.
Discussion
All experiments conducted in animals have to be approved by local authorities and ethics commissions. In in vivo infection experiments the bacterial load in the various host niches of infected animals has to be determined at various time points post-infection. Under these experimental conditions the animals have to be sacrificed prior to the isolation of the bacteria from the blood, nasopharynx, bronchoalvelar lavage, or organs such as the lungs, spleen, and brain. To calculate the number of bacteria per host niche and assess the effect of bacterial factors on the virulence, the blood or organs have to be isolated followed by bacterial recovering and plating on solid media. The bacterial load is then quantified by enumeration of the CFU. To reach statistically significant data each time point needs a group of mice, which under certain experimental conditions leads to difficulties in handling the experiment due to the high number of mice.
In contrast, the real-time bioimaging method is a less time-demanding approach and each individual bacteria-infected mouse can be monitored several times over a period of time. The imaging technique allows visualization of the bacteria within the animal at different time points post-infection or even for an extended time period. Moreover, the bioluminescence measured with the optical image analysis software further enables the quantification of photons emitted by the infected animal. This can be limited to a defined area of the mouse or the whole animal can be considered. In the in vivo infection experiments pneumococci-infected mice have to be monitored in intervals of 8-12 hr, since the spread of pneumococci can progress rapidly in outbred CD-1 mice. The real time imaging can be employed to visualize transmigration of pneumococci from the nasopharynx into the lung and blood, and in addition, the multiplication of pneumococci in the blood after intraperitoneal infection can be monitored by the dramatic increase in bioluminescence intensity26,28,29.
When comparing the virulence potential of wild-type pneumococci and isogenic mutants it is essential to test first the effect of the loss-of-gene under in vitro culture conditions. Growth differences between the wild-type bacteria and mutants, in particular in chemically defined medium, already indicate a decreased bacterial fitness26. Thus, attenuation observed under in vivo conditions is associated with an altered physiology leading to less robust pneumococci during colonization and invasive infections.
However, it has to be mentioned that this imaging technique is often not sufficient to demonstrate the effect of gene knockouts on pneumococcal virulence. Similar to other pathogenic bacteria pneumococci are versatile microorganisms and have evolved multifaceted mechanisms to encounter the host and evade the immune system. Consequently, pneumococci produce on the one hand proteins with manifold functions such as the adhesin PspC and, on the other hand, they are endowed with several proteins exhibiting similar functions which thus can compensate the defect of a protein due to mutations4,17,18,35. In these scenarios other in vivo infection experiments such as the coinfection approach has to be employed to decipher the effect of the loss-of-protein function in the mutant for colonization or invasive infections27,28.
Pneumococci colonize and infect preferentially humans but were also isolated from pets or zoo animals4,36,37. To decipher the impact of virulence or fitness factors mouse infection models are used. However, not all pneumococcal strains are mouse virulent, and more importantly, the susceptibility of mice depends on the genetic background of the animals22. Pneumococcal virulence studies and protection studies are nowadays conducted with CD-1 outbred mice. However, knockout mice or transgenic mice are also of high interest to decipher the role of host factors for invasive pneumococcal diseases. The knockout or transgenic mice are often generated in mouse strains such as C57/BL6 or Balb/c. These mouse strains are less susceptible against pneumococcal infections and high infection doses are required to obtain a lethal dose, 50% (LD50). Depending on the infection dose this resistance of mice may impair real time bioimaging of pneumococcal dissemination to the lungs or blood, since pneumococci are limited to multiply in a manner needed for detection by the imaging system. To overcome this limitation and to be able to image the dissemination in real-time, high infection doses have to be utilized. The risk of this approach could be that differences in the host susceptibility cannot be explored as dissemination progresses too rapidly.
In conclusion, the use of the imaging system represents an excellent approach to monitor, compare and characterize pneumococcal infections in mice infected via the intranasal or intraperitoneal route. This approach is in particular suitable to compare the virulence potential between wild-type and mutants of S. pneumoniae.
Disclosures
The authors have nothing to disclose.
Acknowledgments
Research in the lab was supported by Grants from the Deutsche Forschungsgemeinschaft (DFG HA 3125/3-2, DFG HA 3125/4-2) and the Federal Ministry of Education and Research (BMBF) Medical Infection Genomics (FKZ 0315828A) to SH.
References
- Niederman MS, et al. Guidelines for the management of adults with community-acquired pneumonia. Diagnosis, assessment of severity, antimicrobial therapy, and prevention. Am. J. Respir. Crit. Care Med. 2001;163:1730–1754. doi: 10.1164/ajrccm.163.7.at1010. [DOI] [PubMed] [Google Scholar]
- WHO The global burden of disease: 2004 update. World Health Organization. 2008.
- Bogaert D, et al. Colonisation by Streptococcus pneumoniae and Staphylococcus aureus in healthy children. Lancet. 2004;363:1871–1872. doi: 10.1016/S0140-6736(04)16357-5. [DOI] [PubMed] [Google Scholar]
- Gamez G, Hammerschmidt S. Combat pneumococcal infections: adhesins as candidates for protein-based vaccine development. Curr. Drug Targets. 2012;13:323–337. doi: 10.2174/138945012799424697. [DOI] [PubMed] [Google Scholar]
- Mook-Kanamori BB, Geldhoff M, vander Poll T, Dvan de Beek D. Pathogenesis and pathophysiology of pneumococcal meningitis. Clin. Microbiol. Rev. 2011;24:557–591. doi: 10.1128/CMR.00008-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Musher DM. How contagious are common respiratory tract infections. N. Engl. J. Med. 2003;348:1256–1266. doi: 10.1056/NEJMra021771. [DOI] [PubMed] [Google Scholar]
- Brueggemann AB, Pai R, Crook DW, Beall B. Vaccine escape recombinants emerge after pneumococcal vaccination in the United States. PLoS Pathog. 2007;3 doi: 10.1371/journal.ppat.0030168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munoz-Almagro C, et al. Emergence of invasive pneumococcal disease caused by nonvaccine serotypes in the era of 7-valent conjugate vaccine. Clin. Infect. Dis. 2008;46:174–182. doi: 10.1086/524660. [DOI] [PubMed] [Google Scholar]
- Whitney CG. Impact of conjugate pneumococcal vaccines. Pediatr. Infect. Dis. J. 2005;24:729–730. doi: 10.1097/01.inf.0000174138.25465.ec. [DOI] [PubMed] [Google Scholar]
- Whitney CG, et al. Decline in invasive pneumococcal disease after the introduction of protein-polysaccharide conjugate vaccine. N. Engl. J. Med. 2003;348:1737–1746. doi: 10.1056/NEJMoa022823. [DOI] [PubMed] [Google Scholar]
- Lynch JP, 3rd, Zhanel GG. Streptococcus pneumoniae: epidemiology and risk factors, evolution of antimicrobial resistance, and impact of vaccines. Curr. Opin. Pulm. Med. 2010;16:217–225. doi: 10.1097/MCP.0b013e3283385653. [DOI] [PubMed] [Google Scholar]
- Singleton RJ, et al. Invasive pneumococcal disease caused by nonvaccine serotypes among Alaska native children with high levels of 7-valent pneumococcal conjugate vaccine coverage. JAMA. 2007;297:1784–1792. doi: 10.1001/jama.297.16.1784. [DOI] [PubMed] [Google Scholar]
- Dockrell DH, Whyte MK, Mitchell TJ. Pneumococcal pneumonia: mechanisms of infection and resolution. Chest. 2012;142:482–491. doi: 10.1378/chest.12-0210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lieberman TD, et al. Parallel bacterial evolution within multiple patients identifies candidate pathogenicity genes. Nat. Genet. 2011;43:1275–1280. doi: 10.1038/ng.997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang J, Tauschek M, Robins-Browne RM. Control of bacterial virulence by AraC-like regulators that respond to chemical signals. Trends Microbiol. 2011;19:128–135. doi: 10.1016/j.tim.2010.12.001. [DOI] [PubMed] [Google Scholar]
- Young BC, et al. Evolutionary dynamics of Staphylococcus aureus during progression from carriage to disease. Proc. Natl. Acad. Sci. U.S.A. 2012;109:4550–4555. doi: 10.1073/pnas.1113219109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kadioglu A, Weiser JN, Paton JC, Andrew PW. The role of Streptococcus pneumoniae virulence factors in host respiratory colonization and disease. Nat. Rev. Microbiol. 2008;6:288–301. doi: 10.1038/nrmicro1871. [DOI] [PubMed] [Google Scholar]
- Voss S, Gamez G, Hammerschmidt S. Impact of pneumococcal microbial surface components recognizing adhesive matrix molecules on colonization. Mol. Oral Microbiol. 2012;27:246–256. doi: 10.1111/j.2041-1014.2012.00654.x. [DOI] [PubMed] [Google Scholar]
- Koppe U, Suttorp N, Opitz B. Recognition of Streptococcus pneumoniae by the innate immune system. Cell. Microbiol. 2012;14:460–466. doi: 10.1111/j.1462-5822.2011.01746.x. [DOI] [PubMed] [Google Scholar]
- Paterson GK, Mitchell TJ. Innate immunity and the pneumococcus. Microbiology. 2006;152:285–293. doi: 10.1099/mic.0.28551-0. [DOI] [PubMed] [Google Scholar]
- Gerber J, et al. A mouse model of Streptococcus pneumoniae meningitis mimicking several features of human disease. Acta Neuropathol. 2001;101:499–508. doi: 10.1007/s004010000326. [DOI] [PubMed] [Google Scholar]
- Gingles NA, et al. Role of genetic resistance in invasive pneumococcal infection: identification and study of susceptibility and resistance in inbred mouse strains. Infect. Immun. 2001;69:426–434. doi: 10.1128/IAI.69.1.426-434.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holmes AR, et al. The pavA gene of Streptococcus pneumoniae encodes a fibronectin-binding protein that is essential for virulence. Mol. Microbiol. 2001;41:1395–1408. doi: 10.1046/j.1365-2958.2001.02610.x. [DOI] [PubMed] [Google Scholar]
- Koedel U, Klein M, Pfister HW. New understandings on the pathophysiology of bacterial meningitis. Curr. Opin. Infect. Dis. 2010;23:217–223. doi: 10.1097/QCO.0b013e328337f49e. [DOI] [PubMed] [Google Scholar]
- Medina E. Murine model of pneumococcal pneumonia. Methods Mol. Biol. 2010;602:405–410. doi: 10.1007/978-1-60761-058-8_22. [DOI] [PubMed] [Google Scholar]
- Hartel T, et al. Impact of glutamine transporters on pneumococcal fitness under infection-related conditions. Infect. Immun. 2011;79:44–58. doi: 10.1128/IAI.00855-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hermans PW, et al. The streptococcal lipoprotein rotamase A (SlrA) is a functional peptidyl-prolyl isomerase involved in pneumococcal colonization. J. Biol. Chem. 2006;281:968–976. doi: 10.1074/jbc.M510014200. [DOI] [PubMed] [Google Scholar]
- Jensch I, et al. PavB is a surface-exposed adhesin of Streptococcus pneumoniae contributing to nasopharyngeal colonization and airways infections. Mol. Microbiol. 2010;77:22–43. doi: 10.1111/j.1365-2958.2010.07189.x. [DOI] [PubMed] [Google Scholar]
- Kadioglu A, et al. Pneumococcal protein PavA is important for nasopharyngeal carriage and development of sepsis. Mol. Oral Microbiol. 2010;25:50–60. doi: 10.1111/j.2041-1014.2009.00561.x. [DOI] [PubMed] [Google Scholar]
- Orihuela CJ, Gao G, Francis KP, Yu J, Tuomanen EI. Tissue-specific contributions of pneumococcal virulence factors to pathogenesis. J. Infect. Dis. 2004;190:1661–1669. doi: 10.1086/424596. [DOI] [PubMed] [Google Scholar]
- Francis KP, et al. Visualizing pneumococcal infections in the lungs of live mice using bioluminescent Streptococcus pneumoniae transformed with a novel gram-positive lux transposon. Infect. Immun. 2001;69:3350–3358. doi: 10.1128/IAI.69.5.3350-3358.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Basavanna S, et al. The effects of methionine acquisition and synthesis on Streptococcus pneumoniae growth and virulence. PLoS One. 2013;8 doi: 10.1371/journal.pone.0049638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartel T, et al. Characterization of central carbon metabolism of Streptococcus pneumoniae by isotopologue profiling. J. Biol. Chem. 2012;287:4260–4274. doi: 10.1074/jbc.M111.304311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hammerschmidt S, et al. The host immune regulator factor H interacts via two contact sites with the PspC protein of Streptococcus pneumoniae and mediates adhesion to host epithelial cells. J. Immunol. 2007;178:5848–5858. doi: 10.4049/jimmunol.178.9.5848. [DOI] [PubMed] [Google Scholar]
- Voss S, et al. The choline-binding protein PspC of Streptococcus pneumoniae interacts with the C-terminal heparin-binding domain of vitronectin. J. Biol. Chem. 2013. [DOI] [PMC free article] [PubMed]
- Cartwright K. Pneumococcal disease in western Europe: burden of disease, antibiotic resistance and management. Eur. J. Pediatr. 2002;161:188–195. doi: 10.1007/s00431-001-0907-3. [DOI] [PubMed] [Google Scholar]
- vander Linden M, Al-Lahham A, Nicklas W, Reinert RR. Molecular characterization of pneumococcal isolates from pets and laboratory animals. PLoS One. 2009;4 doi: 10.1371/journal.pone.0008286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brehm , et al. Sequence of the adenine methylase gene of the Streptococcus faecalis plasmid pAM beta 1. Nucleic Acids Res. 1987;15 doi: 10.1093/nar/15.7.3177. [DOI] [PMC free article] [PubMed] [Google Scholar]


