Abstract
The purpose of this study is to identify metastasis- associated genes/signaling pathways in basal-like breast tumors. Kaplan–Meier analysis of two public meta-datasets and functional classification was used to identify genes/signaling pathways significantly associated with distant metastasis free survival. Integrated analysis of expression correlation and interaction between mRNAs and miRNAs was used to identify miRNAs that potentially regulate the expression of metastasis-associated genes. The novel metastatic suppressive role of miR-17-5p was examined by in vitro and in vivo experiments. Over 4,000 genes previously linked to breast tumor progression were examined, leading to identification of 61 and 69 genes significantly associated with shorter and longer DMFS intervals of patients with basal-like tumors, respectively. Functional annotation linked most of the pro-metastatic genes to epithelial mesenchymal transition (EMT) process and three intertwining EMT-driving pathways (hypoxia, TGFB and Wnt), whereas most of the anti-metastatic genes to interferon signaling pathway. Members of three miRNA families (i.e., miR-17, miR-200 and miR-96) were identified as potential regulators of the pro-metastatic genes. The novel anti-metastatic function of miR-17-5p was confirmed by in vitro and in vivo experiments. We demonstrated that miR-17-5p inhibition in breast cancer cells enhanced expression of multiple pro-metastatic genes, rendered cells metastatic properties, and accelerated lung metastasis from orthotopic xenografts. In contrast, intratumoral administration of miR-17-5p mimic significantly reduced lung metastasis. These results provide evidence supporting that EMT activation and IFN pathway inactivation are markers of metastatic progression of basal-like tumors, and members of miR-17, miR-200, and miR-96 families play a role in suppressing EMT and metastasis. The metastasis-associated genes identified in this study have potential prognostic values and functional implications, thus, can be exploited as therapeutic targets to prevent metastasis of basal-like breast tumors.
Keywords: Breast cancer, Metastasis, TGFB, Hypoxia, MIR17HG
Introduction
Breast cancer-related death is responsible for about 3 % of female death in the USA and in most cases attributable to distant metastasis [1]. Human breast tumors can be stratified based on transcriptome data into five subtypes, including Luminal A, luminal B, ERBB2-enriched, basal-like, and claudin-low [2, 3]. The molecular subtypes of breast cancer are likely defined by the origin of cells that give rise to tumors, and differ in key genetic alterations, chemotherapy response, metastatic propensity, organ tropism, and overall prognosis [4–9]. Subtype-specific markers have important diagnostic and prognostic values. However, significant molecular heterogeneities and varied patient outcomes have been documented within subtypes of breast cancer, and currently there are no reliable markers to identify patients with high risk of metastasis within subtypes [7, 10, 11]. Therefore, specific prognostic markers for each molecular subtype are needed to improve clinical management of breast cancer patients.
The varied frequency, latency, and pattern of metastasis among different subtype of breast tumors suggest that tumor cells may acquire metastatic competency through distinct molecular mechanisms. Consequently, the likelihood is low to identify genes and pathways that drive metastatic progression independent of tumor subtype. Reported metastatic signatures are usually derived by comparing the expression profiles of metastatic tumors with non-metastatic tumors using heterogeneous patient groups without considering subtype-specific effects [12–16]. Due to multiple confounding factors (e.g., sample size, composition of tumor subtypes, and metastatic latency), these signatures exhibit limited overlap and are more discriminative in defining tumor subtypes than defining metastatic potential independent of subtypes [16]. In this study, we aimed to identify genes associated with distant metastasis survive interval (DMFS) in patients with basal-like tumors, which represent 10–25 % of all breast tumors and exhibit a metastasis rate of ~40 % within 5 years [7, 17].
By using two meta-datasets of primary tumor expression arrays, we identified a panel of genes with both prognostic values and functional implications for distant metastasis of basal-like breast tumors. Integrated analysis of microRNA (miRNA) and mRNA expression suggested that miRNAs of three families (i.e., miR-17, miR-200 and miR-96) play a role in restricting the expression of>60 % of the identified pro-metastatic genes. The anti-metastatic activity of one of the miRNA (miR-17-5p) was confirmed by in vitro and in vivo experiments.
Results
Identification of metastasis-associated genes in basal-like breast tumors
In order to identify genes associated with metastasis in basal-like breast tumors, a set of 4,439 genes was compiled to include: signatures of metastatic tumors [12, 14, 15, 18], breast cancer subtypes [7, 19], cell subpopulations of mammary gland [20], and epithelial-mesenchymal process (EMT) [21]; top 1,000 genes with the highest frequency of genomic alterations [8]; and top 1,000 most variably expressed genes in basal-like breast tumors [8]. The correlation between expression levels of each gene and DMFS intervals of patients with basal-like tumors was first examined by Kaplan–Meier analyses using the online database GOBO (Gene expression-based Outcome for Breast cancer Online), which is based on a meta-dataset containing expression data and metastasis information of 252 basal-like tumors [22]. The molecular subtypes of tumors were defined by the PAM50 classifier [2]. Genes that were found to be significantly associated with DMFS (logrank test p value ≤ 0.05) of patients with basal-like tumors were further examined using Kaplan–Meier plotter, a second program that contains expression data and distant metastasis information of 220 basal-like tumors [23]. These two meta-datasets are composed of overlapping but different expression array data (Supplementary Table 1). This analysis identified 130 genes whose mRNA levels are significantly associated with DMFS intervals of patients with basal-like tumors (logrank test p value ≤ 0.05 in both meta-datasets), among which 61 genes are associated with shorter DMFS interval and 69 genes associated with longer DMFS, designated as pro-metastatic (Table 1) and anti-metastatic genes (Table 2), respectively.
Table 1.
Pro-metastatic genes
| Symbol | Entrez ID | Ch location | Affy ID | P_GOBOl | P_KM | TGFB | Hypoxia | Wnt | EMT | MIR17HG | MIR200 | MIR96 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ACTN1 | 87 | 14q22-24 | 208637_x_at | 0.020 | 0.001 | Y | Y | T | ||||
| AKAP12 | 9590 | 6q24-25 | 210517_s_at | 0.001 | 0.035 | Y | Y | Y | Y | T | T | |
| ANGPTL4 | 51129 | 19p13.3 | 221009_s_at | 0.008 | 0.044 | Y | Y | Y | Y | |||
| BMP8B | 656 | 1p32-35 | 207865_s_at | 0.015 | 0.003 | Y | T | |||||
| CALM1 | 801 | 14q24-31 | 209563_x_at | 0.030 | 0.046 | Y | Y | T | ||||
| CAMK1D | 57118 | 10p13 | 220246_at | 0.031 | 0.052^ | Y | Y | Y | ||||
| CAV1 | 857 | 7q31.1 | 203065_s_at | 0.005 | 0.019 | Y | Y | Y | Y | T | ||
| CAV2 | 858 | 7q31.1 | 213426_s_at | 0.020 | 0.029 | Y | Y | Y | T | |||
| C0L1A2 | 1278 | 7q22.1 | 202310_s_at | 0.018 | 0.023 | Y | Y | Y | Y | T | ||
| COL5A3 | 50509 | 19p13.2 | 218975_at | 0.050 | 0.015 | |||||||
| C0X7A1 | 1346 | 19q13.1 | 204570_at | 0.001 | 0.008 | Y | ||||||
| CRISPLD2 | 83716 | 16q24.1 | 221541_at | 0.001 | 0.011 | Y | Y | Y | ||||
| CRLP1 | 9244 | 19p12 | 206315_at | 0.027 | 0.015 | Y | Y | |||||
| CTDSP2 | 10106 | 12q14.1 | 203445_s_at | 0.018 | 0.004 | Y | Y | T | T | |||
| CYR61 | 3491 | 1p22.3 | 201289_at | 0.001 | 0.044 | Y | Y | Y | Y | |||
| DACT1 | 51339 | 14q23.1 | 219179_at | 0.003 | 0.015 | Y | Y | Y | T | T | T | |
| DKK3 | 27122 | 11p15.2 | 200819 s at | 0.002 | 0.030 | Y | Y | Y | Y | T | ||
| ECM1 | 1893 | 1q21 | 209365_s_at | 0.013 | 0.025 | Y | Y | Y | ||||
| EGR1 | 1958 | 5q31.1 | 201694 s at | 0.035 | 0.003 | Y | Y | Y | T | |||
| EHD2 | 30846 | 19q13.3 | 45297_at | 0.047 | 0.038 | Y | Y | |||||
| ELF3 | 1999 | 1q32.2 | 210827_s_at | 0.042 | 0.001 | Y | Y | Y | ||||
| EMP1 | 2012 | 12p12.3 | 201325_s_at | 0.021 | 0.049 | Y | Y | Y | T | |||
| EMX2 | 2018 | 10q26.1 | 221950_at | 0.009 | 0.018 | Y | Y | T | ||||
| FSTL3 | 10272 | 19p13 | 203592_s_at | 0.026 | 0.012 | Y | Y | Y | ||||
| GBE1 | 2632 | 3p12.3 | 203282_at | 0.043 | 0.047 | Y | ||||||
| GLI1 | 2735 | 12q13.2-13.3 | 206646 at | 0.019 | 0.031 | Y | Y | Y | Y | |||
| HES1 | 3280 | 3q28-29 | 203395_s_at | 0.010 | 0.008 | Y | Y | Y | T | T | ||
| HOXD1 | 3231 | 2q31.1 | 206602_s_at | 0.008 | 0.022 | Y | Y | T | ||||
| HYAL1 | 3373 | 3p21.3-21.2 | 210874_s_at | 0.015 | 0.002 | Y | Y | T | ||||
| INSIG2 | 51141 | 2q14.2 | 209566_at | 0.008 | 0.046 | Y | Y | Y | T | T | T | |
| ITGA5 | 3678 | 12q11-13 | 201389_at | 0.014 | 0.001 | Y | Y | Y | T | |||
| L3MBTL1 | 26013 | 20q13.12 | 216076_at | 0.004 | 0.007 | T | ||||||
| LMNA | 4000 | 1q22 | 214213 × at | 0.029 | 0.007I | Y | Y | Y | ||||
| LPP | 4026 | 3q28 | 202821_s_at | 0.031 | 0.023 | Y | Y | Y | T | T | T | |
| MAPK7 | 5598 | 17p11.2 | 35617 at | 0.033 | 0.015 | Y | Y | T | ||||
| MCAM | 4162 | 11q23.3 | 209086_x_at | 0.024 | 0.000 | Y | Y | Y | T | |||
| MMP10 | 4319 | 11q22.3 | 205680_at | 0.012 | 0.003 | Y | Y | Y | T | |||
| MRC2 | 9902 | 17q23.2 | 37408_at | 0.013 | 0.005 | Y | Y | Y | T | |||
| MTMR9 | 66036 | 8p22-23 | 213278_at | 0.029 | 0.019 | T | T | |||||
| NDEL1 | 81565 | 17p13.1 | 208093 s at | 0.021 | 0.003 | Y | Y | Y | T | |||
| NDUFS4 | 4724 | 5q11.1 | 209303_at | 0.029 | 0.019 | T | ||||||
| N0X4 | 50507 | 11q14.2-21 | 219773 at | 0.013 | 0.035 | Y | Y | Y | T | T | ||
| PMP22 | 5376 | 17p12 | 210139_s_at | 0.000 | 0.020 | Y | Y | Y | T | |||
| PPL | 5493 | 16p13.3 | 203407 at | 0.024 | 0.013 | Y | Y | |||||
| RNASE2 | 6036 | 14q24-31 | 216667_at | 0.043 | 0.017 | |||||||
| S100A10 | 6281 | 1q21 | 200872_at | 0.019 | 0.011 | Y | Y | Y | Y | |||
| SERPINE1 | 5054 | 7q21.3-22 | 202627_s_at | 0.004 | 0.019 | Y | Y | Y | Y | T | ||
| SPOCK1 | 6695 | 5q31 | 202363_at | 0.001 | 0.006 | Y | Y | Y | T | |||
| SPP1 | 6696 | 4q22.1 | 48580 at | 0.003 | 0.018 | Y | Y | Y | Y | |||
| SRPX2 | 27286 | Xq21.33-23 | 205499_at | 0.025 | 0.002 | |||||||
| STMN2 | 11075 | 8q21.13 | 203001 s at | 0.002 | 0.008 | Y | T | |||||
| TAC1 | 6863 | 7q21-22 | 206552_s_at | 0.008 | 0.013 | T | T | |||||
| TAOK1 | 57551 | 17q11.2 | 216310_at | 0.006 | 0.016 | Y | Y | T | T | T | ||
| TGFB1I1 | 7041 | 16p11.2 | 209651_at | 0.000 | 0.034 | Y | Y | Y | Y | T | ||
| THBS3 | 7059 | 1q21 | 209561_at | 0.003 | 0.028 | Y | Y | Y | ||||
| TLN2 | 83660 | 15q15-21 | 212701_at | 0.043 | 0.026 | Y | Y | T | T | |||
| TNC | 3371 | 9q33 | 201645_at | 0.015 | 0.030 | Y | Y | Y | Y | |||
| TPM1 | 7168 | 15q22.1 | 206116_s_at | 0.045 | 0.049 | Y | Y | Y | T | |||
| TRIO | 7204 | 5p15.2 | 208178_x_at | 0.036 | 0.017 | Y | Y | Y | Y | T | T | |
| WDR6 | 11180 | 3p21.31 | 217734_s_at | 0.011 | 0.001 | Y | Y | T | ||||
| WFDC1 | 58189 | 16q24.3 | 219478_at | 0.018 | 0.017 | Y |
P_GOBO and P_KM Log rank test p value of Kaplan–Meier plot analysis using GOBO and KMplotter databases, Y Functionally linked to the indicated pathways, T Putative targets of indicated miRNA families
Table 2.
Anti-metastatic genes
| Symbol | Entrez ID | Affy ID | Ch location | P_GOBOl | P_KM | IFN |
|---|---|---|---|---|---|---|
| ACP2 | 53 | 202767_at | 11 p11-12 | 0.032 | 0.010 | |
| ADAM23 | 8745 | 206046_at | 2q33 | 0.027 | 0.002 | |
| AIM2 | 9447 | 206513_at | 1q22 | 0.003 | 0.003 | Y |
| AP0BEC3G | 60489 | 214995_s_at | 22q13.1-13.2 | 0.011 | 0.000 | Y |
| ARNTL2 | 56938 | 220658_s_at | 12p11.2-12.2 | 0.021 | 0.018 | Y |
| BIRC3 | 330 | 210538_s_at | 11q22 | 0.028 | 0.000 | Y |
| CCL8 | 6355 | 214038_at | 17q11.2 | 0.047 | 0.032 | Y |
| CD74 | 972 | 209619_at | 5q32 | 0.002 | 0.006 | Y |
| CD97 | 976 | 202910_s_at | 19p13 | 0.006 | 0.041 | Y |
| CEACAM21 | 90273 | 214907_at | 19q13.2 | 0.009 | 0.005 | |
| CFLAR | 8837 | 210563_x_at | 2q33-34 | 0.043 | 0.009 | Y |
| CHIT1 | 1118 | 208168_s_at | 1q31-32 | 0.034 | 0.020 | |
| CLEC7A | 64581 | 221698_s_at | 12p13.2 | 0.011 | 0.007 | |
| CLPTM1 | 1209 | 201640_x_at | 19q13.3 | 0.033 | 0.049 | |
| CORO1A | 11151 | 209083_at | 16p11.2 | 0.016 | 0.032 | Y |
| CTSC | 1075 | 201487_at | 11q14.2 | 0.000 | 0.002 | Y |
| CXCL10 | 3627 | 204533_at | 4q21 | 0.018 | 0.001 | Y |
| CXCL11 | 6373 | 211122_s_at | 4q21.2 | 0.022 | 0.011 | Y |
| CXCR6 | 10663 | 206974_at | 3p21 | 0.008 | 0.007 | |
| EMP3 | 2014 | 203729_at | 19q13.3 | 0.014 | 0.040 | |
| ENTPD1 | 953 | 209473_at | 10q24 | 0.003 | 0.003 | Y |
| FASLG | 356 | 210865_at | 1q23 | 0.005 | 0.041 | Y |
| FASN | 2194 | 204780_s_at | 17q25 | 0.038 | 0.002 | Y |
| FUT7 | 2529 | 210506_at | 9q34.3 | 0.026 | 0.022 | |
| GPR18 | 2841 | 210279_at | 13q32 | 0.042 | 0.000 | Y |
| GPR65 | 8477 | 214467_at | 14q31-32.1 | 0.043 | 0.004 | Y |
| GYPC | 2995 | 202947_s_at | 2q14-21 | 0.042 | 0.004 | |
| GZMH | 2999 | 210164_at | 14q11.2 | 0.003 | 0.001 | |
| ICAM3 | 3385 | 204949_at | 19p13.2-13.3 | 0.030 | 0.027 | Y |
| IKZF1 | 10320 | 205038_at | 7p12.2 | 0.006 | 0.029 | |
| IL23A | 51561 | 217326_x_at | 12q13.3 | 0.005 | 0.006 | |
| IL7R | 3575 | 205798_at | 5p13 | 0.014 | 0.008 | Y |
| IRF2 | 3660 | 203275_at | 4q34.1-35.1 | 0.004 | 0.033 | Y |
| IRF8 | 3394 | 204057_at | 16q24.1 | 0.019 | 0.020 | Y |
| IRP9 | 10379 | 203882_at | 14q11.2 | 0.043 | 0.018 | Y |
| NKG7 | 4818 | 213915 at | 19q13.41 | 0.030 | 0.000 | |
| STAT1 | 6772 | M97935 _3_at | 2q32.2 | 0.033 | 0.000 | Y |
| IRF1 | 3659 | 202531_at | 5q31.1 | 0.004 | 0.000 | Y |
| UBE2L6 | 9246 | 201649_at | 11q12 | 0.006 | 0.000 | Y |
| ITGAL | 3683 | 213475_s_at | 16p11.2 | 0.009 | 0.003 | |
| ITK | 3702 | 211339_s_at | 5q31-32 | 0.030 | 0.008 | Y |
| LCK | 3932 | 204891_s_at | 1p34.3 | 0.011 | 0.007 | |
| LPXN | 9404 | 216250_s_at | 11q12.1 | 0.035 | 0.024 | |
| LSP1 | 4046 | 203523_at | 11p15.5 | 0.044 | 0.042 | |
| MECP2 | 4204 | 202616_s_at | Xq28 | 0.023 | 0.018 | |
| MEN1 | 4221 | 202645_s_at | 11q13 | 0.004 | 0.041 | Y |
| MTCP1 | 4515 | 205106_at | Xq28 | 0.027 | 0.003 | |
| NBN | 4683 | 217299_s_at | 8q21 | 0.034 | 0.004 | Y |
| NCKAP1L | 3071 | 209734_at | 12q13.1 | 0.047 | 0.004 | |
| PHTF1 | 10745 | 215285_s_at | 1p13 | 0.002 | 0.012 | |
| PLEK | 5341 | 203470_s_at | 2p13.3 | 0.003 | 0.011 | Y |
| PNOC | 5368 | 205901_at | 8p21 | 0.042 | 0.050 | |
| PRF1 | 5551 | 214617_at | 10q22 | 0.050 | 0.022 | Y |
| PSAP | 5660 | 200866_s_at | 10q21-22 | 0.017 | 0.005 | |
| PSTPIP2 | 9050 | 219938_s_at | 18q12 | 0.032 | 0.024 | Y |
| PTGER4 | 5734 | 204897_at | 5p13.1 | 0.021 | 0.021 | Y |
| PTPRC | 5788 | 212588_at | 1q31-32 | 0.003 | 0.002 | Y |
| RASSF2 | 9770 | 203185_at | 20p13 | 0.026 | 0.015 | Y |
| SELL | 6402 | 204563_at | 1q23-25 | 0.014 | 0.007 | Y |
| SELPLG | 6404 | 209879_at | 12q24 | 0.019 | 0.006 | Y |
| SLAMF8 | 56833 | 219385_at | 1q23.2 | 0.003 | 0.042 | Y |
| SLC7A7 | 9056 | 204588_s_at | 14q11.2 | 0.043 | 0.050 | Y |
| SNX10 | 29887 | 218404_at | 7p15.2 | 0.009 | 0.009 | Y |
| SP110 | 3431 | 209762_x_at | 2q37.1 | 0.006 | 0.034 | Y |
| SP140 | 11262 | 207777_s_at | 2q37.1 | 0.002 | 0.020 | Y |
| TYK2 | 7297 | 205546_s_at | 19p13.2 | 0.035 | 0.047 | Y |
| VCAM1 | 7412 | 203868_s_at | 1p31p32 | 0.048 | 0.019 | Y |
| WHSC1 | 7468 | 209053_s_at | 4p16.3 | 0.002 | 0.016 | |
| WIPF1 | 7456 | 202664_at | 2q31.1 | 0.026 | 0.013 | Y |
P_GOBO and P_KM Log rank test p value of Kaplan–Meier plot analysis using GOBO and KMplotter databases, Y Functionally linked to the indicated pathways, T Putative targets of indicated miRNA families
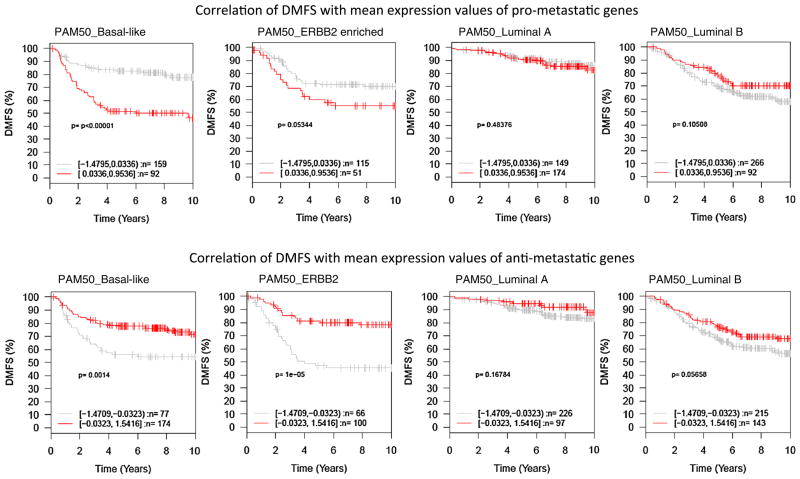
We next examined whether the metastasis-associated genes of basal-like tumors have prognostic values for other subtypes of breast tumors. Log2 expression values of these genes were standardized to have mean 0 and standard deviation 1 cross all tumor samples in the GOBO database. Within each subtype of tumors, patients were split equally into two cohorts, high-expression, and low expression based on mean of the standardized expression values of the pro- or anti-metastatic genes. DMFS intervals of the two patient cohorts for each subtype of tumors were compared by Kaplan–Meier survival plots and logrank p values were calculated. Collectively as genesets, higher expression of the pro-metastatic genes was found to be associated with shorter DMFS interval, whereas higher expression of the anti-metastatic genes associated with longer DMFS interval, of patients with ERBB2-enriched tumors (Fig. 1). However, the expression levels of these metastasis-associated genes were not significantly associated with DMFS interval of patients with luminal tumors (Fig. 1). At individual gene level, 16 genes were found to be coordinately associated with DMFS of patients with basal-like or ERBB2-enriched tumors, including six pro-metastatic genes (ACTN1, COL5A3, ITGA5, NOX4, SPOCK1, and WFDC1) and ten anti-metastatic genes (AIM2, APOBEC3G, CCL8, CTSC, CXCL10, GPR65, ICAM3, LPXN, PLEK, and PTPRC). These observations suggest that metastasis-associated molecular events of basal-like tumors are also involved in metastasis of ERBB2-enriched tumors, but not critical for metastatic progression of luminal tumors.
Fig. 1.
Kaplan–Meier plots of pro-metastatic genes in different subtypes of breast tumors. Collectively as genesets, mean expression values of the pro- and anti-metastatic genes were used to generate Kaplan–Meier plots by using the GOBO program
Signaling pathways driving metastatic progression of basal-like tumors
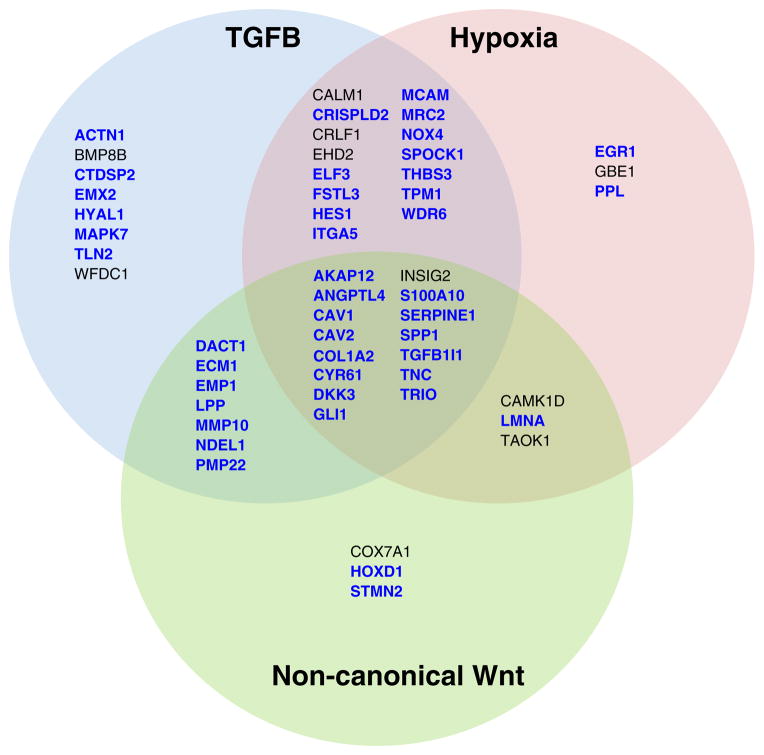
Functional classification uncovered that genes associated with EMT and components of hypoxia, TGFB, and WNT signaling pathways were significantly overrepresented by the pro-metastatic genes (Table 1). A thorough literature search on reported functions of the pro-metastatic genes revealed that 43 genes (~70 %) have been linked to EMT process, and 46, 36, and 27 genes linked to TGFB, hypoxia, and WNT signaling pathways, respectively (Fig. 2). Notably, among the genes linked to WNT pathways are primarily components of the non-canonical Wnt-planar cell polarity pathway (Wnt-PCP, e.g., CAMK1D, DACT1, LMNA, LPP and TAOK1), and genes known to be suppressed by β-catenin (e.g., ANGPTL4, COL1A2, INSIG2, MMP10, NDEL1, SPP1, and TRIO), indicating a role of Wnt-PCP activation, coupled with reduced transcription regulatory activity of β-catenin, in metastatic tumors. The innate immune response genes, especially those induced by interferon (IFN) [24], were found to be significantly overrepresented by the anti-metastatic genes (Table 2), implicating a role of IFN pathway inactivation in metastasis. Together, these findings suggest that EMT activation and IFN signaling pathway inactivation are markers of metastatic progression of basal-like tumors.
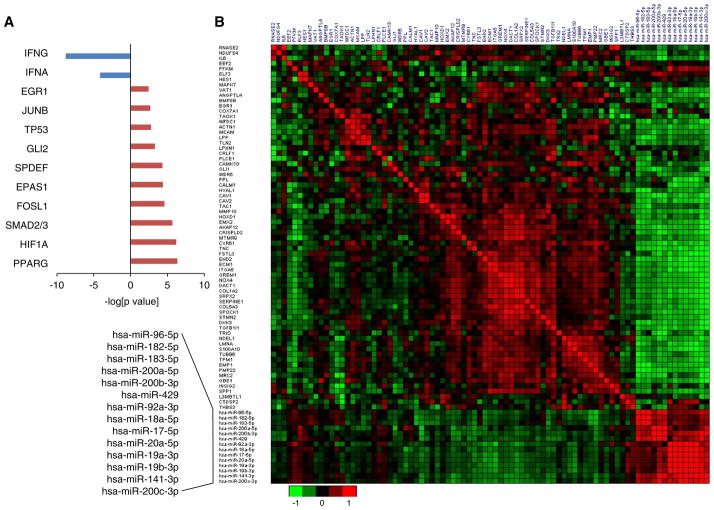
Fig. 2.
Upstream regulators of metastasis-associated genes. a Anti-metastatic genes are predominately regulated by interferon, whereas pro-metastatic genes are regulated by transcription factors involved in TGFB and Hypoxia signaling pathways. The top 10 regulatory transcription factors of pro-metastatic genes are presented. b Inverse correlation of pro-metastatic genes and miRNAs. The heatmap displays the Pearson correlation coefficients of each pairwise combination of mRNA and/or miRNA. Expression Z-scores of mRNAs and miRNAs in basal-like breast tumors (n = 81) were retrieved from the TCGA database and used to calculate Pearson correlation coefficient
Upstream regulators of pro-metastatic genes
To gain insights into the regulation of the metastasis-associated genes, we performed upstream regulator analysis using the ingenuity pathway analysis (IPA) program (Fig. 3a). Among the top 10 transcription factors predicted to regulate the pro-metastatic genes were SMAD2/3 and HIF1A, the master transcription regulators of the TGFB and hypoxia signaling pathways, respectively. The additional eight transcription factors have been shown to play a role in gene regulation in response to hypoxia and/or TGFB [25, 26]. IFNG and IFNA were identified as upstream regulators of anti-metastatic genes. This result reinforces the notion that hypoxic response, TGFB activation, and IFN pathway inactivation promote metastasis of basal-like tumors.
Fig. 3.
Functional annotation links pro-metastatic genes to epithelial– mesenchymal transition (EMT) process and TGFB, hypoxia and noncanonical Wnt signaling pathways. Functional annotation was conducted by using the ingenuity pathway analysis program (IPA) and gene set enrichment analysis (GSEA) based on molecular signatures database (MSigDB). Genes involved in EMT are marked in blue
Since not all TGFB and hypoxia responsive genes are associated with DMFS, activation of transcription factors of TGFB, and hypoxia pathways alone may not be sufficient to establish the expression pattern of metastatic genes. miRNAs have been increasingly recognized as key regulators of metastasis [27]. We previously showed that DROSHA knockdown promoted lung metastasis of basal-like breast cancer cells (MDA-MB-231) in an orthotopic xenograft model, suggesting a role of DROSHA-dependent miRNAs in repressing metastasis [28]. Therefore, we examined whether specific miRNAs play a role in regulating the expression of pro-metastatic genes. We performed integrated analysis of miRNA and mRNA expression to identify inversely correlated mRNA-miRNA pairs in basal-like tumors using the expression data in The Cancer Genome Atlas (TCGA) database [8]. Individual miRNAs were ranked according to the number of prometastatic genes that are inversely correlated with their expression based on Pearson’s correlation analysis. This analysis identified members of three miRNA families as potential regulators of the pro-metastatic genes, including members of miR-17, miR-200 and miR-96 families. Over 60 % of the pro-metastatic genes are predicted targets of the above-mentioned 14 miRNAs (Table 1). The number of putative targets of each of these miRNAs among the prometastatic genes is significantly greater than expected by chance (χ2 p value<0.001), supporting a functional link between these miRNAs and pro-metastatic genes. The Pearson’s correlation efficient of paired miRNA–mRNA is presented in Fig. 2 and Supplementary Table 1. The inverse correlation between the expression of these miRNAs and the pro-metastatic genes was also observed in basal-like tumors included in dataset GSE28884 [29].
Inhibiting miR-17-5p function enhances cell migration, invasion, and anoikis resistance in vitro, and accelerates lung metastasis in vivo
It is known that miRNAs of the miR-200 and miR-96 families inhibit EMT by targeting EMT transcription factors (e.g., SNAI2, ZEB1 and ZEB2) [30, 31]. However, the functions of miR-17 family members in EMT and metastasis have not been well studied. Therefore, we examined whether miR-17-5p functions as metastatic repressor in breast cancer cells. Among the six miRNAs encoded by MIR17HG, miR-17-5p targets the most of the pro-metastatic genes involved in TGFB and hypoxia pathways. Consistently, genes involved in TGFB and hypoxia response were found to be significantly overrepresented by genome-wide targets of miR-17-5p (Supplementary Fig. 1). Thus, we hypothesized that miR-17-5p inhibits metastasis of basal-like tumors through inhibiting EMT activated by TGFB and hypoxia.
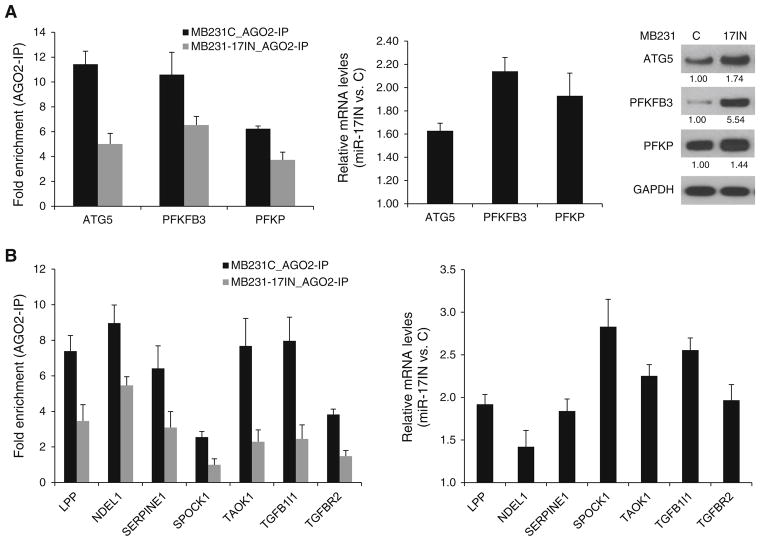
To elucidate, the function of miR-17-5p in basal-like breast cancer cells, a MDA-MB-231 subline (MB231- 17IN) was established by transduction with lentivirus expressing miArrest inhibitor of miR-17-5p (miR-17IN) and mCherry as fluorescent marker. Control cells (MB231- C) were transduced with lentivirus expressing mCherry only. To examine that ectopic expression of miR-17IN can effectively block the function of endogenous miRNA, we measured the interaction of miR-17-5p target mRNAs with RISC (RNA-induced silence complex) by AGO2-immunoprecipitation followed by qPCR analysis. As showed in Fig. 4a, miR-17IN expression reduced RISC binding of three miR-17-5p target mRNAs (ATG5, PFKFB3 and PFKP) that harbor conserved exact matches to positions 2–8 of mature miR-17-5p in their 3′-untranslated regions. The reduced RISC binding was coupled with increased expression levels of these mRNAs and their corresponding proteins (Fig. 4b). We next examined the effect of miR- 17IN on RISC binding and expression levels of several putative targets of miR-17-5p that were identified as pro-metastatic genes. Among the seven genes examined (CAV2, LPP, NDEL1, SERPINE1, SPOCK1, TAOK1 and TGFB1I1), six mRNAs showed reduced RISC binding and elevated expression levels in MB231-17IN cells compared to control MB231-C cells (Fig. 4c). The RISC binding and expression level of TGFBR2, a validated target of miR-17- 5p [32], were examined as positive control.
Fig. 4.
Ectopic expression of miArrest inhibitor of has-miR-17-5p (miR-17IN) in MDA-MB-231 cells (MB231-17IN) resulted in reduced binding to RNA-induced silencing complex (RISC) and elevated expression of target mRNAs. a miR-17IN reduced RISC binding of mRNAs that harbor conserved target sites in their 3′-UTRs with exact match to positions 2–8 of the mature miR-17-5p. RISC binding was examined by AGO2-immunoprecipitation (AGO2-IP) followed by qPCR analysis (upper panel). b miR-17IN increased expression of miR-17-5p target genes at mRNA and protein levels. Expression levels of mRNAs and proteins were examined by qPCR and immunoblotting analysis, respectively. The average fold changes of protein levels from two independent experiments were indicated. c Effect of ectopic expression of miR-17IN on RISC binding and expression of mRNAs that are encoded by pro-metastatic genes and contain miR-17 target sites in their 3′-UTRs. TGFBR2, a validated miR-17 target, is included as a positive control
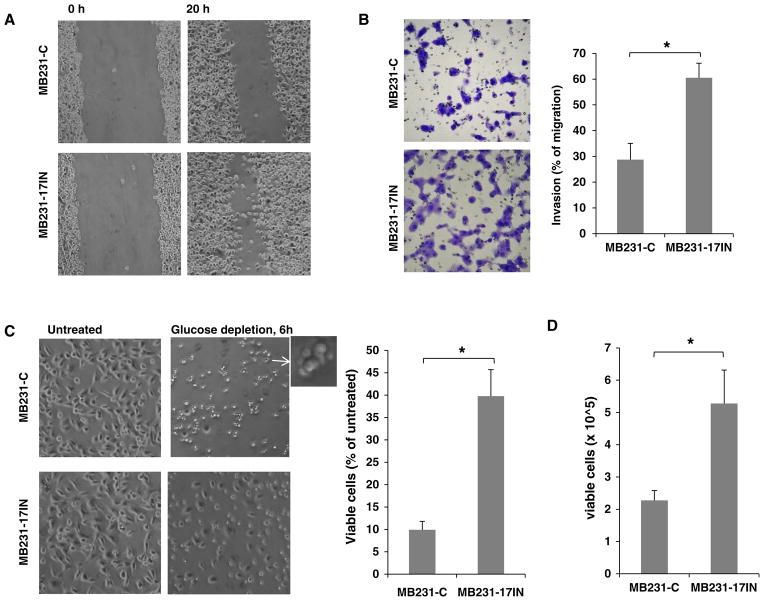
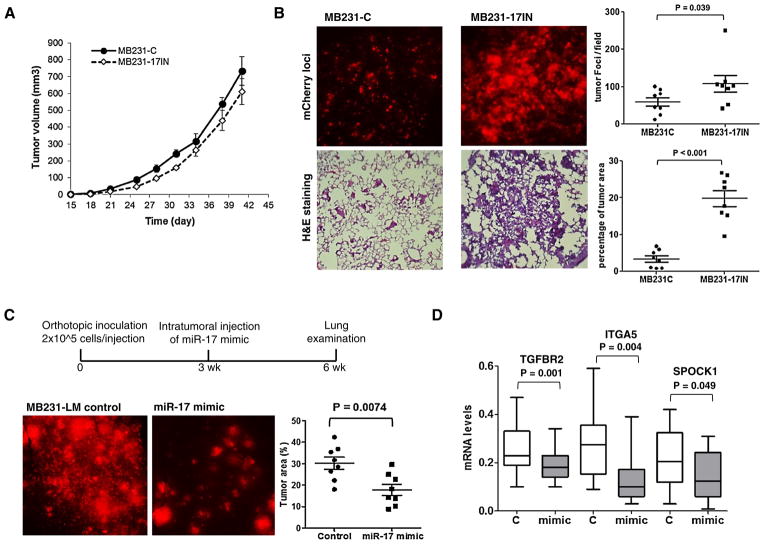
Since most of the pro-metastatic genes targeted by miR- 17 are involved in migration, invasion, and apoptotic resistance [26, 33], we examined the effects of miR-17 inhibition on these cell properties. As shown in Fig. 5, miR-17IN substantially increased cell migration (scratch wound healing assay), invasion (Boyden Chamber transwell invasion assay), and survival (under glucose depletion or anchorage-independent culture). We next examined the in vivo role of miR-17-5p on lung metastasis using an orthotopic xenograft model, in which 4-wk old female NSG (NOD.Cg Prkdcscid Il2rgtm1Wjl/SzJ) mice were inoculated with 5 × 105 cells in the 4th mammary gland fat pads. As shown in Fig. 6, miR-17IN expression had no significant effect on primary tumor growth in mammary gland fat pads, but substantially accelerated spontaneous lung metastasis, increasing tumor burden in lungs by ~6- fold 6 weeks after orthotopic inoculation (percentage of tumor area in lungs: MB231-17IN 19.74 ± 6.12 vs. MB231-C 3.34 ± 2.40; p value<0.001, n = 8). Taken together, these experiments provided evidence supporting a role of miR-17-5p in suppressing metastatic activity of breast tumor cells.
Fig. 5.
Ectopic expression of miArrest inhibitor of has-miR-17-5p (miR-17IN) enhanced metastatic potential of MDA-MB-231 cells. a miR-17IN increased cell mobility as determined by wound scratch healing assay. Cells were imaged at 0 and 20 h after scratch wounds were generated. b miR-17IN enhanced cell invasion as determined by transwell invasion assay with Matrigel-coated Boyden Chambers. c miR-17IN protected cells against apoptosis induced by glucose depletion (6 h). d miR-17IN increased number of viable cell after 4-day suspension culture
Fig. 6.
miR-17-5p suppresses lung metastasis of orthotopic xenografts. a Growth curves of primary tumors generated from cells expressing miR-17IN (MB231-17IN) or control cells (MB231-C). The results were presented as mean ± SE (n = 10). b Metastatic foci of mCherry expressing cells on the dorsal surface of the left lung lobe from mice inoculated with MB231-17IN or control MB231-C cells. The presence of tumor cells in lungs was visualized by H&E staining of formalin-fixed lung section (10 μM). The lung area occupied by metastases foci were quantified using ImageJ program. c Metastatic foci of mCherry expressing cells on the dorsal surface of the left lung lobe from mice inoculated with MB231-LM cells and received intratumoral injection of miR-17-5p mimic or control RNA oligonucleotides. The lung area occupied by metastases foci were quantified using ImageJ program. d Gene expression levels of primary tumors 7-day after treatment with miR-17-5p mimic or control RNA oligonucleotides
Intratumoral delivery of miR-17-5p mimic reduces lung metastasis
Having demonstrated that miR-17-5p inhibition accelerates lung metastasis of MDA-MB-231 cells in vivo, we next examined whether miR-17-5p mimic can be used to block metastasis of a MDA-MB-231 variant (MB231-LM) that was isolated from spontaneous lung metastases of an orthotopic xenograft model. MB231-LM cells were inoculated in both sides at the fourth mammary glands of 4-week old NSG mice to establish orthotopic xenografts. 3 weeks after inoculation, tumors (~150 mm3) received direct injection of miR-17-5p mimic or control oligonucleotides, formulated as neutral lipid-based liposome. Lung metastasis was examined 3 weeks after treatment. As shown in Fig. 6b, intratumoral delivery of miR-17-5p mimic substantially reduced lung metastasis compared to treatment with control oligonucleotides, but showed no significant effect on primary tumor growth. Gene expression analysis of primary tumors at 7 days post-treatment showed that miR-17-5p mimic reduced expression of prometastatic genes ITGA5 and SPOCK1, and a validated miR-17-5p target TGFBR2, in comparison to control tumors (Fig. 6c). These results further support a metastasis-repressive role of miR-17-5p.
Expression of miR-17-5p is inversely correlated with activation of TGFB, hypoxia, and non-canonical WNT pathways in basal-like tumors
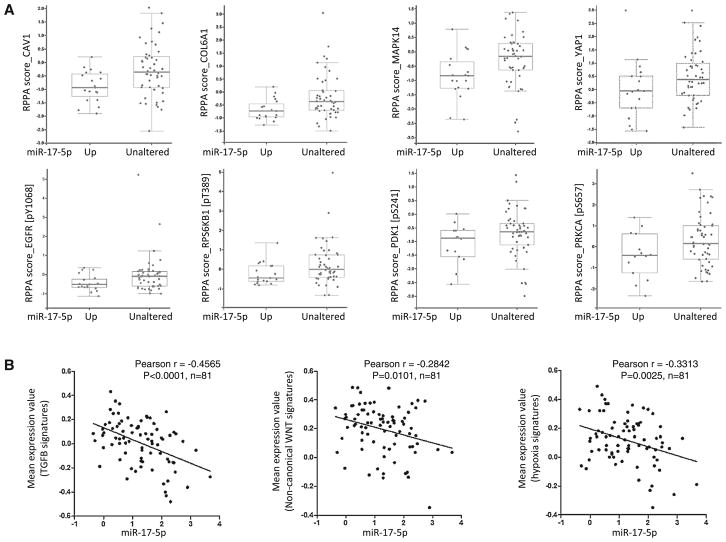
Given the metastasis-suppressing activity of miR-17-5p and its inverse correlation with pro-metastatic genes associated with EMT process, we hypothesized that miR-17-5p plays a role in restraining activation of EMT. Our gene expression data analysis suggests EMT activation in basal-like breast tumors is primarily driven by three intertwining pathways, TGFB, hypoxia, and non-canonical WNT. Therefore, it is likely that miR-17-5p plays a role in repressing the activation of these EMT-driving pathway. To test this, we examined whether miR-17-5p expression level is correlated with activation status of hypoxia, TGFB, and non-canonical WNT activation signaling pathways by examine the protein and gene expression data of previously defined signature genes of these pathways [34]. The TCGA database contains data of 133 proteins/phosphoproteins measured by reverse phase protein array in breast tumors. Correlation analysis within basal-like tumors revealed that miR-17-5p expression is inversely correlated with the abundance of 15 proteins, among which are known targets of TGFB (i.e., CAV1, COL6A1 and MAPK14), hypoxia (i.e., phosphorylated EGFR, ROPS6KB1 and PDK1), or non-canonical WNT pathways (i.e., YAP1 and phosphorylated PRKCA) (Fig. 7a). Next, we examined the correlation between the expression of miR-17-5p and signature genes of TGFB, hypoxia, or non-canonical WNT pathways. The expression data of these signatures were obtained from TCGA database and analyzed to obtain mean expression values for each tumor samples. As shown in Fig. 7b, the expression level of miR-17-5p is inversely correlated with the mean expression values of the signature genes of TGFB, hypoxia, and non-canonical WNT pathways. These correlations are statistically significant, suggesting that TGFB, hypoxia and non-canonical WNT pathways are inactivated in tumors with higher miR-17-5p expression.
Fig. 7.
Correlation between expression of miR-17-5p and activation status of TGFB, hypoxia and non-canonical WNT pathways. a miR- 17-5p expression is inversely correlated with abundance of proteins/ phosphoproteins known to be targeted by TGFB (CAV1, COL6A1 and MAPK14), hypoxia (phosphorylated EGFR, RPS6KB1 and PDK1) or non-canonical WNT pathways (YAP1 and phosphorylated PRKCA). Expression data of proteins/phosphoproteins in basal-like breast tumors, measured by reverse phase protein array, were retrieved from the TCGA database. B. miR-17-5p expression is inversely correlated with mean expression values of signature genes TGFB, hypoxia and non-canonical WNT pathways. Expression data of previously defined signature genes of these pathways in basal-like tumors were retrieved from the TCGA database
Discussion
During metastasis process, tumor cells undergo reversible transitions between epithelial and mesenchymal states [35]. EMT facilitates the execution of early steps in the metastasis cascade by potentiating invasive migration and apoptotic resistance. Most of the pro-metastatic genes identified in our meta-analysis have been linked to the EMT process and three intertwining signaling pathways that are known to activate the EMT program in breast cancer cells, including TGFB, hypoxia, and non-canonical Wnt signaling pathway. Our findings provide evidence supporting a role of EMT driven by hypoxia, TGFB, and non-canonical WNT signaling pathways in metastatic progression of basal-like breast tumors.
Hypoxia is frequently encountered by cells in solid tumors due to insufficient and/or aberrant blood vessel development. Multiple mechanisms have been proposed for hypoxia-induced MET in tumor cells, including activation of latent growth factors and cytokines (e.g., TGFB, EGF, TNFA and TNFSF11) and generation of reactive oxygen species [25, 36]. Release of active TGFBs from tumor stroma is a prominent EMT-promoting event triggered by hypoxia [36]. Non-canonical Wnt-PCP mediated by WNT11 and RYK is known to promote metastasis by coordinating cell polarity, protrusive activity, and directional migration [37–40]. WNT11 was reported to be induced by TGFB [41], providing a direct link between Wnt-PCP activation and TGFB-mediated EMT. Future studies on the regulation of pro-metastatic genes by these signaling pathways will advance our understanding of metastatic progression and provide therapeutic targets.
Most of the metastasis-associated genes exhibit no or low frequency (<10 %) of genomic alterations in basal-like tumors according to the TCGA database. Only five prometastatic genes showed copy number gains in more than 10 % of basal-like tumors, with ECM1, S100A10, THBS3, and LMNA located at 1q21-22 and CAMK1D at 10p13. This observation suggests that expression of metastasis-associated genes is primarily controlled by epigenetic mechanisms, consistent with the transient and reversible nature of metastatic events. Three chromatin modifiers were found to be associated with metastasis, with L3MBTL1 associated with poor DMFS, whereas MECP2 and WHSC1 associated with better prognosis. In addition, six transcription factors (EGR1, ELF3, EMX2, HES1, HOXD1, and GLI1) are associated with poor DMFS, all of which, with the exception of HOXD1, have been linked to TGFB, hypoxia, or Wnt signaling pathway (Table 1). Whether these transcription factors play a role in establishing and maintaining the chromatin structures poised for dynamic transcription regulation during metastasis remain to be examined.
Besides transcription regulators, miRNAs are increasingly recognized as key regulators of metastasis [27]. Integrated analysis of expression correlation and predicted interaction of miRNA–mRNA identified members of three miRNA families (miR-17, miR-200 and miR-96) as repressors of the pro-metastatic genes in basal-like tumors. Unlike the members of the miR-200 and miR-96 families that have been reported to inhibit EMT [30, 31], conflicting results were reported for the role of miRNAs encoded by MIR17HG in metastatic progression of breast cancer [32, 42, 43]. Our experiment results provide evidence supporting a role of miR-17-5p in repressing metastasis of breast cancer.
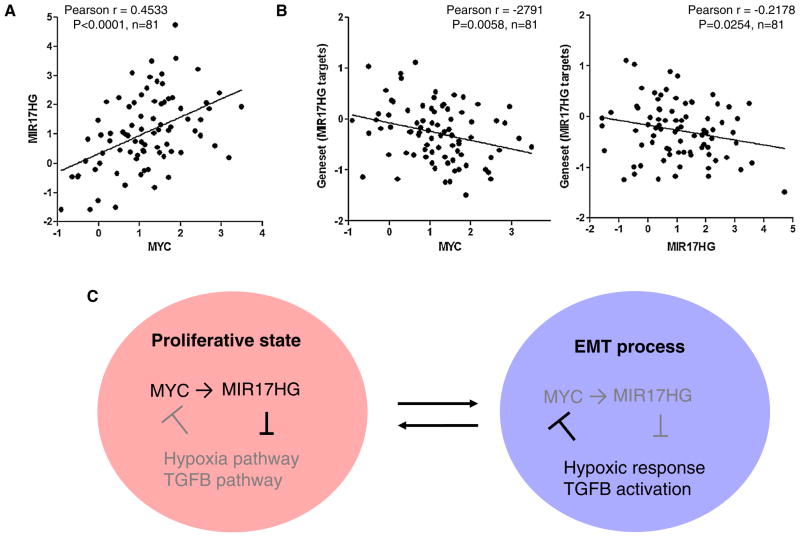
MYC is the predominant transcription activator of MIR17HG [44–46]. Consistently, the expression levels of MIR17HG and MYC were found to be closely correlated in breast tumors (Fig. 8a). Since MYC universally enhances transcription of all active genes in a given cell [47], genes downregulated by MYC are likely targets of transcriptional repressors or miRNAs activated by MYC. Intriguingly, we found that the expression of MYC is inversely correlated with that of the pro-metastatic genes containing target sites of MIR17HG-miRNAs (Fig. 8b). Therefore, we propose that MIR17HG-miRNAs play a role in sustaining MYC-driven proliferative state by targeting genes/pathways involved in EMT, a process that can be activated by a various types of stress through TGFB activation and is intrinsically linked to MYC inactivation and cell growth arrest. This is in line with previous observations that MYC suppresses cell invasive migration and tumor metastasis, whereas TGFB and hypoxia inactivate MYC [48, 49], Taken together, these findings implicate a regulatory network, composed of MYC, MIR17HG, and TGFB pathway, in governing cell phenotypic switch between proliferative and mesenchymal states (Fig. 8c). Downregulation of MIR17HG expression and consequently de-repression of TGFB pathway may be a prerequisite for onset of EMT and metastasis.
Fig. 8.
Correlation between expression of MYC and pro-metastatic genes targeted by MIR17HG-miRNAs. a Positive correlation between expression of MYC and MIR17HG in basal-like breast tumors (TCGA database, n = 81). b Inverse correlation between expression of MYC and pro-metastatic genes potentially targeted by MIR17HG-miRNAs in basal-like breast tumors. Mean expression values of targets of MIR17HG-miRNAs in basal-like tumors (TCGA database, n = 81) were used for the correlation analysis. c A proposed model to depicture a regulatory network composed of MYC, MIR17HG, TGFB, and hypoxia pathways in governing metastatic progression of basal-like breast tumors
Functional analysis of the anti-metastatic genes revealed that downregulation of immune response, especially inactivation of interferon signaling pathway, plays a role in metastatic progression. This observation is consistent with a previous report that higher expression of immune response genes is associated with better prognosis of ESR1/ ERBB2-negative breast tumors [50, 51]. In addition, a recent study showed that inactivation of type 1 IFN signaling pathway in tumor cells promoted, whereas activation of the signaling pathway by IFN treatment suppressed, metastasis of breast tumor xenografts [52]. The suppression of intrinsic IFN signaling pathway of tumor cells may enable metastasis by restricting immunosurveillance during tumor cells circulation and homing in foreign organs. Additional studies are warranted to define the molecular events responsible for IFN pathway inactivation.
In summary, we identified a panel of genes significantly associated with DMFS of patients with basal-like tumors. The pro-metastatic genes are functionally linked to TGFB, hypoxia, and non-canonical WNT signaling pathways and EMT process, whereas the anti-metastatic genes linked to IFN signaling pathway. In addition, members of three miRNA families were identified as potential regulators of the pro-metastatic genes. The novel anti-metastatic function of one of these miRNAs, miR-17-5p, was confirmed by in vitro and in vivo experiments. These metastasis- associated genes have prognostic values and functional implications and can be exploited as potential therapeutic targets for metastasis prevention.
Materials and methods
Cell culture and stable transfection
MDA-MB-231 and MCF7 cells (ATCC, Manassas, VA, USA) were maintained in MEM medium supplemented with 100 units/ml penicillin, 100 μg/ml streptomycin, and 10 % fetal bovine serum. To facilitate tumor imaging, cells were transduced with lentivirus (pEZX-AM03, GeneCopoeia) and selected in medium contained Hygromycin B (200 μg/ml) to stably express mCherry. The mi- Arrest inhibitor of has-miR-17-5p encoded by a lentiviral vector that co-expresses mCherry (HmiR-AN0230-AM03, GeneCopoeia, Rockville, MD, USA) was used to establish sublines defective in miR-17-5p activity.
Primary tumor growth and lung metastasis in orthotopic xenograft model
Tumor cells (5 × 105, 2.5 × 105, and 1 × 106 for MDA-MB- 231, MB231-LM and MCF7, respectively) were surgically inoculated into the 4th inguinal mammary glands of 4-week old female NSG mice (The Jackson Laboratory). To monitor primary tumor growth, mice were inspected twice a week for tumor appearance by manual palpation. Primary tumor sizes were measured by digital calipers and tumor volume was calculated as: Volume = (width2 × - length)/2. To examine metastases in lungs, the dorsal surface of the left lung lobe was imaged using a fluorescent microscope at 10× magnification (NIKON, CFI60), and metastatic foci of mCherry expressing cells were quantified using ImageJ program [53]. The presence of tumor cells in the left lobes was further confirmed by Hematoxylin and Eosin (H&E) staining of formalin-fixed lung sections (10 μM thick) as described previously [28]. All animal studies adhered to protocols approved by the Institutional Animal Care and Use Committee of University of Tennessee Health Science Center.
Intratumoral administration of miR-17-5p mimic
When xenografts of MB231-LM cells reached a volume of ~150 mm3, we injected directly in the tumor mass 1 nmol of miR-17-5p mimic or control RNA oligonucleotides (HMI0264 or HMC0002, respectively; Sigma-Aldrich), formulated as neutral lipid-based liposome by using the MaxSuppressor In Vivo RNA-LANCEr II reagent (Bioo Scientific, Austin, TX, USA). Gene expression-treated tumors and lung metastasis were examined at 7-day and 3-week post-treatment, respectively.
Quantitation of mRNA and miRNA expression using qPCR
Total RNA was prepared using TRIzol (Life Technologies). mRNAs were converted to cDNA by using iScript cDNA Synthesis Kits (BioRad, Hercules, CA, USA). qPCR was performed on the CFX96™ Real-Time PCR Detection System using SYBR Green supermix (BioRad). Expression data of mRNA were normalized to RPL13A using the 2−ΔΔCT method. Primer sequences for mRNAs were obtained from PrimerBank [54].
Migration, invasion, and viability assays
For scratch wound healing assay, confluent cells were put in medium supplemented with 2 % FBS, wound scratches on cell monolayer were generated by using cell combs (Millpore) and imaged at 0 and 20 h. Invasive and cell viability assays were performed as described previously [28].
Statistical analysis
Kaplan–Meier analyses were conducted to identify genes significantly associated with DMFS of patients with basal-like tumors by using two meta-datasets (GOBO and Kaplan– Meier plotter [22, 23]), in which the molecular subtypes of tumors were defined by the PAM50 classifier [2]. The original datasets included in GOBO and Kaplan–Meier plotter that contain expression data and clinical information of basal-like tumors were listed in Supplementary Table 2. Gene function annotation and enrichment analysis were performed by using QIAGEN’s IPA program and molecular signatures database (MSigDB) [34]. miRNA target site mapping was performed using IPA, TargetScan, DIANA-microT- CDS, and miRDB programs [55, 56]. For correlation analysis with genesets, mean expression value of all genes included in a given geneset was used. The expression data of breast tumor samples used for miRNA–mRNA correlation analysis were retrieved from TCGA and GEO (Gene Expression Omnibus, GSE28884). Correlation analysis of expression data and statistical analysis of experiment data were performed using GraphPad Prism 5.
Supplementary Material
Acknowledgments
This work was supported by NIH Grants, CA140346 (to Meiyun Fan).
Footnotes
Electronic supplementary material The online version of this article (doi:10.1007/s10549-014-3040-5) contains supplementary material, which is available to authorized users.
Conflict of interest The authors declare that they have no conflict of interest.
Contributor Information
Meiyun Fan, Email: mfan2@uthsc.edu, Department of Pathology and Laboratory Medicine, 19 South Manassas Street, Memphis, TN 38163, USA, Center for Cancer Research, Memphis, TN 38163, USA.
Aarti Sethuraman, Department of Pathology and Laboratory Medicine, 19 South Manassas Street, Memphis, TN 38163, USA, Center for Cancer Research, Memphis, TN 38163, USA.
Martin Brown, Center for Cancer Research, Memphis, TN 38163, USA.
Wenlin Sun, Department of Pharmacology, University of Tennessee Health Science Center, Memphis, TN 38163, USA.
Lawrence M. Pfeffer, Department of Pathology and Laboratory Medicine, 19 South Manassas Street, Memphis, TN 38163, USA, Center for Cancer Research, Memphis, TN 38163, USA
References
- 1.Society AC. Breast cancer facts & figures 2013–2014 [Google Scholar]
- 2.Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J, Marron JS, Nobel AB, Mardis E, Nielsen TO, Ellis MJ, Perou CM, Bernard PS. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol. 2009;27:1160–1167. doi: 10.1200/JCO.2008.18.1370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Prat A, Parker JS, Karginova O, Fan C, Livasy C, Herschkowitz JI, He X, Perou CM. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 2010;12:R68. doi: 10.1186/bcr2635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Santagata S, Thakkar A, Ergonul A, Wang B, Woo T, Hu R, Harrell JC, McNamara G, Schwede M, Culhane AC, Kindelberger D, Rodig S, Richardson A, Schnitt SJ, Tamimi RM, Ince TA. Taxonomy of breast cancer based on normal cell phenotype predicts outcome. J Clin Investig. 2014;124:859–870. doi: 10.1172/JCI70941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wang ZA, Mitrofanova A, Bergren SK, Abate-Shen C, Cardiff RD, Califano A, Shen MM. Lineage analysis of basal epithelial cells reveals their unexpected plasticity and supports a cell-of-origin model for prostate cancer heterogeneity. Nat Cell Biol. 2013;15:274–283. doi: 10.1038/ncb2697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ince TA, Richardson AL, Bell GW, Saitoh M, Godar S, Karnoub AE, Iglehart JD, Weinberg RA. Transformation of different human breast epithelial cell types leads to distinct tumor phenotypes. Cancer Cell. 2007;12:160–170. doi: 10.1016/j.ccr.2007.06.013. [DOI] [PubMed] [Google Scholar]
- 7.Smid M, Wang Y, Zhang Y, Sieuwerts AM, Yu J, Klijn JG, Foekens JA, Martens JW. Subtypes of breast cancer show preferential site of relapse. Cancer Res. 2008;68:3108–3114. doi: 10.1158/0008-5472.CAN-07-5644. [DOI] [PubMed] [Google Scholar]
- 8.Cancer Genome Atlas N. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70. doi: 10.1038/nature11412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ellis MJ, Perou CM. The genomic landscape of breast cancer as a therapeutic roadmap. Cancer Discov. 2013;3:27–34. doi: 10.1158/2159-8290.CD-12-0462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Metzger-Filho O, Sun Z, Viale G, Price KN, Crivellari D, Snyder RD, Gelber RD, Castiglione-Gertsch M, Coates AS, Goldhirsch A, Cardoso F. Patterns of Recurrence and outcome according to breast cancer subtypes in lymph node-negative disease: results from international breast cancer study group trials VIII and IX. J Clin Oncol. 2013;31:3083–3090. doi: 10.1200/JCO.2012.46.1574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Shah SP, Roth A, Goya R, Oloumi A, Ha G, Zhao Y, Turashvili G, Ding J, Tse K, Haffari G, Bashashati A, Prentice LM, Khattra J, Burleigh A, Yap D, Bernard V, McPherson A, Shumansky K, Crisan A, Giuliany R, Heravi-Moussavi A, Rosner J, Lai D, Birol I, Varhol R, Tam A, Dhalla N, Zeng T, Ma K, Chan SK, Griffith M, Moradian A, Cheng SW, Morin GB, Watson P, Gelmon K, Chia S, Chin SF, Curtis C, Rueda OM, Pharoah PD, Damaraju S, Mackey J, Hoon K, Harkins T, Tadigotla V, Sigaroudinia M, Gascard P, Tlsty T, Costello JF, Meyer IM, Eaves CJ, Wasserman WW, Jones S, Huntsman D, Hirst M, Caldas C, Marra MA, Aparicio S. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature. 2012;486:395–399. doi: 10.1038/nature10933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Van’t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, Linsley PS, Bernards R, Friend SH. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002;415:530–536. doi: 10.1038/415530a. [DOI] [PubMed] [Google Scholar]
- 13.Landemaine T, Jackson A, Bellahcene A, Rucci N, Sin S, Abad BM, Sierra A, Boudinet A, Guinebretiere JM, Ricevuto E, Nogues C, Briffod M, Bieche I, Cherel P, Garcia T, Castronovo V, Teti A, Lidereau R, Driouch K. A six-gene signature predicting breast cancer lung metastasis. Cancer Res. 2008;68:6092–6099. doi: 10.1158/0008-5472.CAN-08-0436. [DOI] [PubMed] [Google Scholar]
- 14.Minn AJ, Gupta GP, Siegel PM, Bos PD, Shu W, Giri DD, Viale A, Olshen AB, Gerald WL, Massague J. Genes that mediate breast cancer metastasis to lung. Nature. 2005;436:518–524. doi: 10.1038/nature03799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bos PD, Zhang XH, Nadal C, Shu W, Gomis RR, Nguyen DX, Minn AJ, van de Vijver MJ, Gerald WL, Foekens JA, Massague J. Genes that mediate breast cancer metastasis to the brain. Nature. 2009;459:1005–1009. doi: 10.1038/nature08021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Culhane AC, Quackenbush J. Confounding effects in “A six-gene signature predicting breast cancer lung metastasis”. Cancer Res. 2009;69:7480–7485. doi: 10.1158/0008-5472.CAN-08-3350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kennecke H, Yerushalmi R, Woods R, Cheang MC, Voduc D, Speers CH, Nielsen TO, Gelmon K. Metastatic behavior of breast cancer subtypes. J Clin Oncol. 2010;28:3271–3277. doi: 10.1200/JCO.2009.25.9820. [DOI] [PubMed] [Google Scholar]
- 18.Wang Y, Klijn JG, Zhang Y, Sieuwerts AM, Look MP, Yang F, Talantov D, Timmermans M, Meijer-van Gelder ME, Yu J, Jatkoe T, Berns EM, Atkins D, Foekens JA. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet. 2005;365:671–679. doi: 10.1016/S0140-6736(05)17947-1. [DOI] [PubMed] [Google Scholar]
- 19.Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Investig. 2011;121:2750–2767. doi: 10.1172/JCI45014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lim E, Vaillant F, Wu D, Forrest NC, Pal B, Hart AH, Asselin-Labat ML, Gyorki DE, Ward T, Partanen A, Feleppa F, Huschtscha LI, Thorne HJ, kConFab, Fox SB, Yan M, French JD, Brown MA, Smyth GK, Visvader JE, Lindeman GJ. Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat Med. 2009;15:907–913. doi: 10.1038/nm.2000. [DOI] [PubMed] [Google Scholar]
- 21.Taube JH, Herschkowitz JI, Komurov K, Zhou AY, Gupta S, Yang J, Hartwell K, Onder TT, Gupta PB, Evans KW, Hollier BG, Ram PT, Lander ES, Rosen JM, Weinberg RA, Mani SA. Core epithelial-to-mesenchymal transition interactome gene-expression signature is associated with claudin-low and metaplastic breast cancer subtypes. Proc Natl Acad Sci USA. 2010;107:15449–15454. doi: 10.1073/pnas.1004900107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ringner M, Fredlund E, Hakkinen J, Borg A, Staaf J. GOBO: gene expression-based outcome for breast cancer online. PLoS ONE. 2011;6:e17911. doi: 10.1371/journal.pone.0017911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, Szallasi Z. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 2010;123:725–731. doi: 10.1007/s10549-009-0674-9. [DOI] [PubMed] [Google Scholar]
- 24.Rusinova I, Forster S, Yu S, Kannan A, Masse M, Cumming H, Chapman R, Hertzog PJ. Interferome v2.0: an updated database of annotated interferon-regulated genes. Nucleic Acids Res. 2013;41:D1040–D1046. doi: 10.1093/nar/gks1215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Majmundar AJ, Wong WJ, Simon MC. Hypoxia-inducible factors and the response to hypoxic stress. Mol Cell. 2010;40:294–309. doi: 10.1016/j.molcel.2010.09.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Massague J. TGFbeta signalling in context. Nat Rev Mol Cell Biol. 2012;13:616–630. doi: 10.1038/nrm3434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pencheva N, Tavazoie SF. Control of metastatic progression by microRNA regulatory networks. Nat Cell Biol. 2013;15:546–554. doi: 10.1038/ncb2769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Fan M, Krutilina R, Sun J, Sethuraman A, Yang CH, Wu ZH, Yue J, Pfeffer LM. Comprehensive analysis of microRNA (miRNA) targets in breast cancer cells. J Biol Chem. 2013;288:27480–27493. doi: 10.1074/jbc.M113.491803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Farazi TA, Horlings HM, Ten Hoeve JJ, Mihailovic A, Halfwerk H, Morozov P, Brown M, Hafner M, Reyal F, van Kouwenhove M, Kreike B, Sie D, Hovestadt V, Wessels LF, van de Vijver MJ, Tuschl T. MicroRNA sequence and expression analysis in breast tumors by deep sequencing. Cancer Res. 2011;71:4443–4453. doi: 10.1158/0008-5472.CAN-11-0608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Park SM, Gaur AB, Lengyel E, Peter ME. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. 2008;22:894–907. doi: 10.1101/gad.1640608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Li XL, Hara T, Choi Y, Subramanian M, Francis P, Bilke S, Walker RL, Pineda M, Zhu Y, Yang Y, Luo J, Wakefield LM, Brabletz T, Park BH, Sharma S, Chowdhury D, Meltzer PS, Lal A. A p21-ZEB1 complex inhibits epithelial-mesenchymal transition through the microRNA 183-96-182 cluster. Mol Cell Biol. 2014;34:533–550. doi: 10.1128/MCB.01043-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Dews M, Fox JL, Hultine S, Sundaram P, Wang W, Liu YY, Furth E, Enders GH, El-Deiry W, Schelter JM, Cleary MA, Thomas-Tikhonenko A. The myc-miR-17~92 axis blunts TGF{beta} signaling and production of multiple TGF{beta}-dependent antiangiogenic factors. Cancer Res. 2010;70:8233–8246. doi: 10.1158/0008-5472.CAN-10-2412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Vanharanta S, Massague J. Hypoxia signaling–license to metastasize. Cancer Discov. 2013;3:1103–1104. doi: 10.1158/2159-8290.CD-13-0481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Liberzon A, Subramanian A, Pinchback R, Thorvaldsdottir H, Tamayo P, Mesirov JP. Molecular signatures database (MsigDB) 3.0. Bioinformatics. 2011;27:1739–1740. doi: 10.1093/bioinformatis/btr260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Nieto MA. Epithelial plasticity: a common theme in embryonic and cancer cells. Science. 2013;342:1234850. doi: 10.1126/science.1234850. [DOI] [PubMed] [Google Scholar]
- 36.Lu X, Kang Y. Hypoxia and hypoxia-inducible factors: master regulators of metastasis. Clin Cancer Res. 2010;16:5928–5935. doi: 10.1158/1078-0432.CCR-10-1360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Luga V, Wrana JL. Tumor–stroma interaction: revealing fibroblast-secreted exosomes as potent regulators of Wnt-planar cell polarity signaling in cancer metastasis. Cancer Res. 2013;73:6843–6847. doi: 10.1158/0008-5472.CAN-13-1791. [DOI] [PubMed] [Google Scholar]
- 38.Macheda ML, Sun WW, Kugathasan K, Hogan BM, Bower NI, Halford MM, Zhang YF, Jacques BE, Lieschke GJ, Dabdoub A, Stacker SA. The Wnt receptor Ryk plays a role in mammalian planar cell polarity signaling. J Biol Chem. 2012;287:29312–29323. doi: 10.1074/jbc.M112.362681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wang Y. Wnt/Planar cell polarity signaling: a new paradigm for cancer therapy. Mol Cancer Ther. 2009;8:2103–2109. doi: 10.1158/1535-7163.MCT-09-0282. [DOI] [PubMed] [Google Scholar]
- 40.Green J, Nusse R, van Amerongen R. The role of ryk and ror receptor tyrosine kinases in wnt signal transduction. Cold Spring Harbor perspectives in biology. 2014;6 doi: 10.1101/cshperspect.a009175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zhang P, Cai Y, Soofi A, Dressler GR. Activation of Wnt11 by transforming growth factor-beta drives mesenchymal gene expression through non-canonical Wnt protein signaling in renal epithelial cells. J Biol Chem. 2012;287:21290–21302. doi: 10.1074/jbc.M112.357202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Fox JL, Dews M, Minn AJ, Thomas-Tikhonenko A. Targeting of TGFbeta signature and its essential component CTGF by miR-18 correlates with improved survival in glioblastoma. RNA. 2013;19:177–190. doi: 10.1261/rna.036467.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Yu Z, Willmarth NE, Zhou J, Katiyar S, Wang M, Liu Y, McCue PA, Quong AA, Lisanti MP, Pestell RG. microRNA 17/20 inhibits cellular invasion and tumor metastasis in breast cancer by heterotypic signaling. Proc Natl Acad Sci USA. 2010;107:8231–8236. doi: 10.1073/pnas.1002080107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Mu P, Han YC, Betel D, Yao E, Squatrito M, Ogrodowski P, de Stanchina E, D’Andrea A, Sander C, Ventura A. Genetic dissection of the miR-17~92 cluster of microRNAs in Myc-induced B-cell lymphomas. Genes Dev. 2009;23:2806–2811. doi: 10.1101/gad.1872909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Becker LE, Lu Z, Chen W, Xiong W, Kong M, Li Y. A systematic screen reveals MicroRNA clusters that significantly regulate four major signaling pathways. PLoS ONE. 2012;7:e48474. doi: 10.1371/journal.pone.0048474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Jin HY, Oda H, Lai M, Skalsky RL, Bethel K, Shepherd J, Kang SG, Liu WH, Sabouri-Ghomi M, Cullen BR, Rajewsky K, Xiao C. MicroRNA-17~92 plays a causative role in lymphoma-genesis by coordinating multiple oncogenic pathways. EMBO J. 2013;32:2377–2391. doi: 10.1038/emboj.2013.178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Nie Z, Hu G, Wei G, Cui K, Yamane A, Resch W, Wang R, Green DR, Tessarollo L, Casellas R, Zhao K, Levens D. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell. 2012;151:68–79. doi: 10.1016/j.cell.2012.08.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Alfano D, Votta G, Schulze A, Downward J, Caputi M, Stoppelli MP, Iaccarino I. Modulation of cellular migration and survival by c-Myc through the downregulation of urokinase (uPA) and uPA receptor. Mol Cell Biol. 2010;30:1838–1851. doi: 10.1128/MCB.01442-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Liu H, Radisky DC, Yang D, Xu R, Radisky ES, Bissell MJ, Bishop JM. MYC suppresses cancer metastasis by direct transcriptional silencing of alphav and beta3 integrin subunits. Nat Cell Biol. 2012;14:567–574. doi: 10.1038/ncb2491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Desmedt C, Haibe-Kains B, Wirapati P, Buyse M, Larsimont D, Bontempi G, Delorenzi M, Piccart M, Sotiriou C. Biological processes associated with breast cancer clinical outcome depend on the molecular subtypes. Clin Cancer Res. 2008;14:5158–5165. doi: 10.1158/1078-0432.CCR-07-4756. [DOI] [PubMed] [Google Scholar]
- 51.Teschendorff AE, Miremadi A, Pinder SE, Ellis IO, Caldas C. An immune response gene expression module identifies a good prognosis subtype in estrogen receptor negative breast cancer. Genome Biol. 2007;8:R157. doi: 10.1186/gb-2007-8-8-r157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Bidwell BN, Slaney CY, Withana NP, Forster S, Cao Y, Loi S, Andrews D, Mikeska T, Mangan NE, Samarajiwa SA, de Weerd NA, Gould J, Argani P, Moller A, Smyth MJ, Anderson RL, Hertzog PJ, Parker BS. Silencing of Irf7 pathways in breast cancer cells promotes bone metastasis through immune escape. Nat Med. 2012;18:1224–1231. doi: 10.1038/nm.2830. [DOI] [PubMed] [Google Scholar]
- 53.Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9:671–675. doi: 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Spandidos A, Wang X, Wang H, Seed B. PrimerBank: a resource of human and mouse PCR primer pairs for gene expression detection and quantification. Nucleic Acids Res. 2010;38:D792–D799. doi: 10.1093/nar/gkp1005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Reczko M, Maragkakis M, Alexiou P, Grosse I, Hatzigeorgiou AG. Functional microRNA targets in protein coding sequences. Bioinformatics. 2012;28:771–776. doi: 10.1093/bioinformatics/bts043. [DOI] [PubMed] [Google Scholar]
- 56.Wang X, El Naqa IM. Prediction of both conserved and nonconserved microRNA targets in animals. Bioinformatics. 2008;24:325–332. doi: 10.1093/bioinformatics/btm595. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.