Abstract
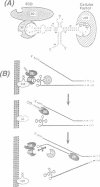
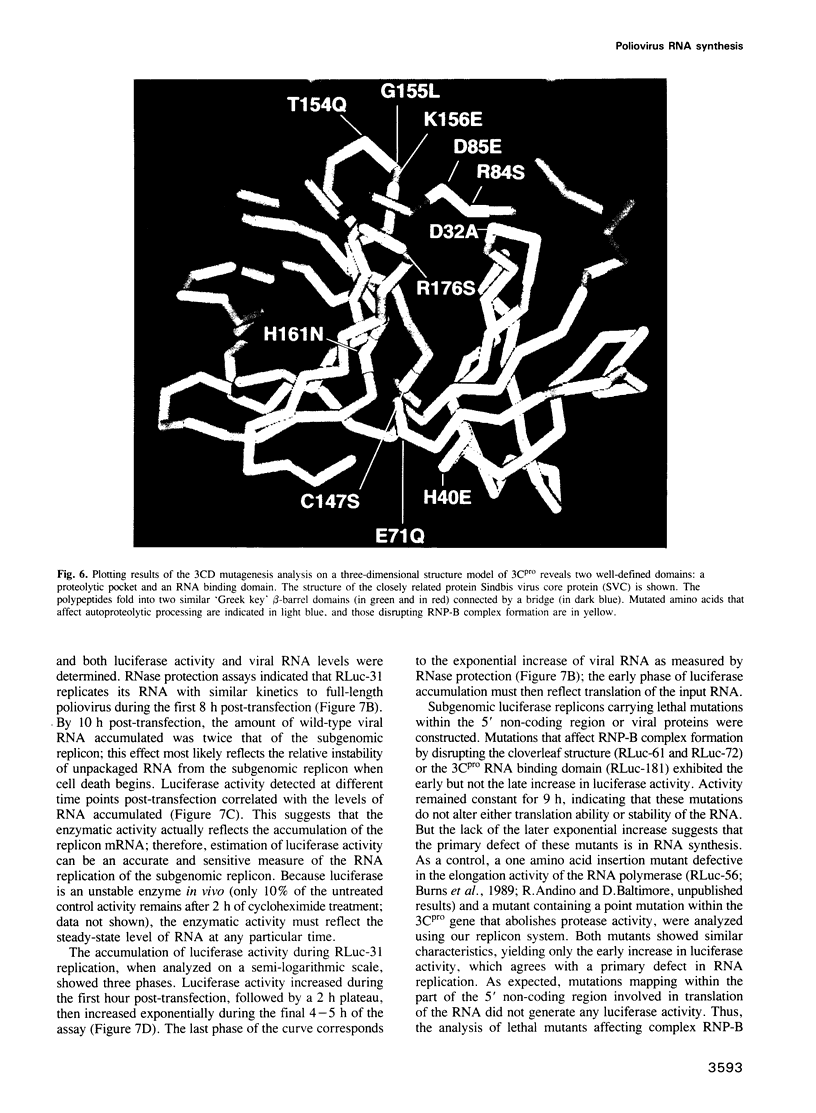
The structure of a ribonucleoprotein complex formed at the 5'-end of poliovirus RNA was investigated. This complex involves the first 90 nucleotides of poliovirus genome which fold into a cloverleaf-like structure and interact with both uncleaved 3CD, the viral protease-polymerase precursor, and a 36 kDa ribosome-associated cellular protein. The cellular protein is required for complex formation and interacts with unpaired bases in one stem-loop of the cloverleaf RNA. Amino acids within the 3C protease which are important for RNA binding were identified by site-directed mutagenesis and the crystal structure of a related protease was used to model the RNA binding domain within the viral 3CD protein. The physiologic importance of the ribonucleic-protein complex is suggested by the finding that mutations that disrupt complex formation abolish RNA replication but do not affect RNA translation or stability. Based on these structural and functional findings we propose a model for the initiation of poliovirus RNA synthesis where an initiation complex consisting of 3CD, a cellular protein, and the 5'-end of the positive strand RNA catalyzes in trans the initiation of synthesis of new positive stranded RNA.
Full text
PDF






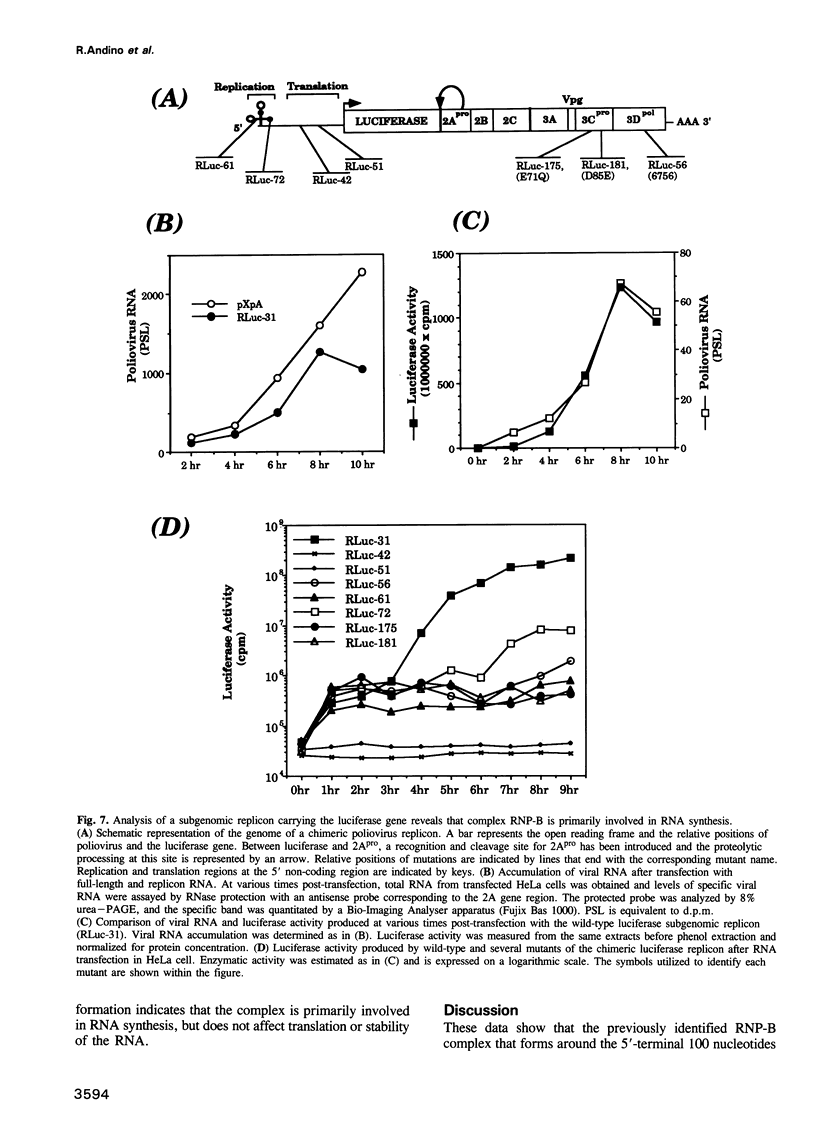




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andino R., Rieckhof G. E., Baltimore D. A functional ribonucleoprotein complex forms around the 5' end of poliovirus RNA. Cell. 1990 Oct 19;63(2):369–380. doi: 10.1016/0092-8674(90)90170-j. [DOI] [PubMed] [Google Scholar]
- Andino R., Rieckhof G. E., Trono D., Baltimore D. Substitutions in the protease (3Cpro) gene of poliovirus can suppress a mutation in the 5' noncoding region. J Virol. 1990 Feb;64(2):607–612. doi: 10.1128/jvi.64.2.607-612.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bazan J. F., Fletterick R. J. Viral cysteine proteases are homologous to the trypsin-like family of serine proteases: structural and functional implications. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7872–7876. doi: 10.1073/pnas.85.21.7872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernstein H. D., Sarnow P., Baltimore D. Genetic complementation among poliovirus mutants derived from an infectious cDNA clone. J Virol. 1986 Dec;60(3):1040–1049. doi: 10.1128/jvi.60.3.1040-1049.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernstein H. D., Sonenberg N., Baltimore D. Poliovirus mutant that does not selectively inhibit host cell protein synthesis. Mol Cell Biol. 1985 Nov;5(11):2913–2923. doi: 10.1128/mcb.5.11.2913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Black D. L., Pinto A. L. U5 small nuclear ribonucleoprotein: RNA structure analysis and ATP-dependent interaction with U4/U6. Mol Cell Biol. 1989 Aug;9(8):3350–3359. doi: 10.1128/mcb.9.8.3350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burns C. C., Lawson M. A., Semler B. L., Ehrenfeld E. Effects of mutations in poliovirus 3Dpol on RNA polymerase activity and on polyprotein cleavage. J Virol. 1989 Nov;63(11):4866–4874. doi: 10.1128/jvi.63.11.4866-4874.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi H. K., Tong L., Minor W., Dumas P., Boege U., Rossmann M. G., Wengler G. Structure of Sindbis virus core protein reveals a chymotrypsin-like serine proteinase and the organization of the virion. Nature. 1991 Nov 7;354(6348):37–43. doi: 10.1038/354037a0. [DOI] [PubMed] [Google Scholar]
- García-Blanco M. A., Jamison S. F., Sharp P. A. Identification and purification of a 62,000-dalton protein that binds specifically to the polypyrimidine tract of introns. Genes Dev. 1989 Dec;3(12A):1874–1886. doi: 10.1101/gad.3.12a.1874. [DOI] [PubMed] [Google Scholar]
- Geigenmüller-Gnirke U., Nitschko H., Schlesinger S. Deletion analysis of the capsid protein of Sindbis virus: identification of the RNA binding region. J Virol. 1993 Mar;67(3):1620–1626. doi: 10.1128/jvi.67.3.1620-1626.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giachetti C., Semler B. L. Role of a viral membrane polypeptide in strand-specific initiation of poliovirus RNA synthesis. J Virol. 1991 May;65(5):2647–2654. doi: 10.1128/jvi.65.5.2647-2654.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorbalenya A. E., Donchenko A. P., Blinov V. M., Koonin E. V. Cysteine proteases of positive strand RNA viruses and chymotrypsin-like serine proteases. A distinct protein superfamily with a common structural fold. FEBS Lett. 1989 Jan 30;243(2):103–114. doi: 10.1016/0014-5793(89)80109-7. [DOI] [PubMed] [Google Scholar]
- Hagino-Yamagishi K., Nomoto A. In vitro construction of poliovirus defective interfering particles. J Virol. 1989 Dec;63(12):5386–5392. doi: 10.1128/jvi.63.12.5386-5392.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harmon S. A., Richards O. C., Summers D. F., Ehrenfeld E. The 5'-terminal nucleotides of hepatitis A virus RNA, but not poliovirus RNA, are required for infectivity. J Virol. 1991 May;65(5):2757–2760. doi: 10.1128/jvi.65.5.2757-2760.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaplan G., Racaniello V. R. Construction and characterization of poliovirus subgenomic replicons. J Virol. 1988 May;62(5):1687–1696. doi: 10.1128/jvi.62.5.1687-1696.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kirkegaard K. Genetic analysis of picornaviruses. Curr Opin Genet Dev. 1992 Feb;2(1):64–70. doi: 10.1016/S0959-437X(05)80324-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kräusslich H. G., Wimmer E. Viral proteinases. Annu Rev Biochem. 1988;57:701–754. doi: 10.1146/annurev.bi.57.070188.003413. [DOI] [PubMed] [Google Scholar]
- Kuhn R. J., Tada H., Ypma-Wong M. F., Semler B. L., Wimmer E. Mutational analysis of the genome-linked protein VPg of poliovirus. J Virol. 1988 Nov;62(11):4207–4215. doi: 10.1128/jvi.62.11.4207-4215.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J. P., Baltimore D. Isolation of poliovirus 2C mutants defective in viral RNA synthesis. J Virol. 1988 Nov;62(11):4016–4021. doi: 10.1128/jvi.62.11.4016-4021.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luthman H., Magnusson G. High efficiency polyoma DNA transfection of chloroquine treated cells. Nucleic Acids Res. 1983 Mar 11;11(5):1295–1308. doi: 10.1093/nar/11.5.1295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pelletier J., Kaplan G., Racaniello V. R., Sonenberg N. Cap-independent translation of poliovirus mRNA is conferred by sequence elements within the 5' noncoding region. Mol Cell Biol. 1988 Mar;8(3):1103–1112. doi: 10.1128/mcb.8.3.1103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Percy N., Barclay W. S., Sullivan M., Almond J. W. A poliovirus replicon containing the chloramphenicol acetyltransferase gene can be used to study the replication and encapsidation of poliovirus RNA. J Virol. 1992 Aug;66(8):5040–5046. doi: 10.1128/jvi.66.8.5040-5046.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosenberg A. H., Lade B. N., Chui D. S., Lin S. W., Dunn J. J., Studier F. W. Vectors for selective expression of cloned DNAs by T7 RNA polymerase. Gene. 1987;56(1):125–135. doi: 10.1016/0378-1119(87)90165-x. [DOI] [PubMed] [Google Scholar]
- Sonenberg N. Regulation of translation by poliovirus. Adv Virus Res. 1987;33:175–204. doi: 10.1016/s0065-3527(08)60318-8. [DOI] [PubMed] [Google Scholar]
- Takegami T., Kuhn R. J., Anderson C. W., Wimmer E. Membrane-dependent uridylylation of the genome-linked protein VPg of poliovirus. Proc Natl Acad Sci U S A. 1983 Dec;80(24):7447–7451. doi: 10.1073/pnas.80.24.7447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toyoda H., Yang C. F., Takeda N., Nomoto A., Wimmer E. Analysis of RNA synthesis of type 1 poliovirus by using an in vitro molecular genetic approach. J Virol. 1987 Sep;61(9):2816–2822. doi: 10.1128/jvi.61.9.2816-2822.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trono D., Andino R., Baltimore D. An RNA sequence of hundreds of nucleotides at the 5' end of poliovirus RNA is involved in allowing viral protein synthesis. J Virol. 1988 Jul;62(7):2291–2299. doi: 10.1128/jvi.62.7.2291-2299.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wimmer E. Genome-linked proteins of viruses. Cell. 1982 Feb;28(2):199–201. doi: 10.1016/0092-8674(82)90335-x. [DOI] [PubMed] [Google Scholar]
- de Wet J. R., Wood K. V., DeLuca M., Helinski D. R., Subramani S. Firefly luciferase gene: structure and expression in mammalian cells. Mol Cell Biol. 1987 Feb;7(2):725–737. doi: 10.1128/mcb.7.2.725. [DOI] [PMC free article] [PubMed] [Google Scholar]