Abstract
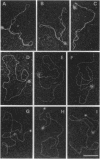
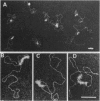
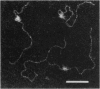
Electron microscopic visualization indicates that the transcription activator NRI (NTRC) binds with exceptional selectivity and efficiency to a sequence-induced superhelical (spiral) segment inserted upstream of the glnA promoter, accounting for its observed ability to substitute for the natural glnA enhancer. The cooperative binding of NRI to the spiral insert leads to protein oligomerization which, at higher concentration, promotes selective coating of the entire superhelical segment with protein. Localization of NRI at apical loops is observed with negatively supercoiled plasmid DNA. With a linear plasmid, bending of DNA is observed. We confirm that NRI is a DNA-bending protein, consistent with its high affinity for spiral DNA. These results prove that spiral DNA without any homology to the NRI-binding sequence site can substitute for the glnA enhancer by promoting cooperative activator binding to DNA and facilitating protein oligomerization. Similar mechanisms might apply to other prokaryotic and eukaryotic activator proteins that share the ability to bend DNA and act efficiently as multimers.
Full text
PDF
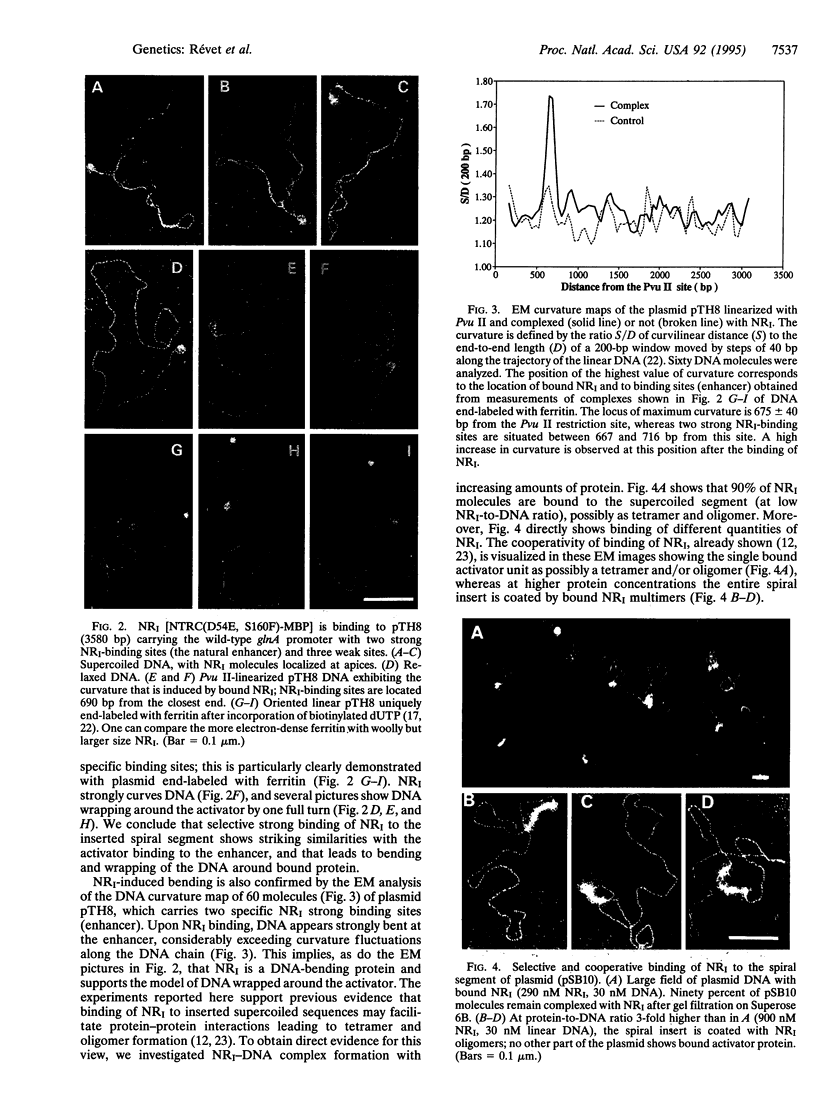


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Banerji J., Rusconi S., Schaffner W. Expression of a beta-globin gene is enhanced by remote SV40 DNA sequences. Cell. 1981 Dec;27(2 Pt 1):299–308. doi: 10.1016/0092-8674(81)90413-x. [DOI] [PubMed] [Google Scholar]
- Brahms G., Brahms S., Magasanik B. A sequence-induced superhelical DNA segment serves as transcriptional enhancer. J Mol Biol. 1995 Feb 10;246(1):35–42. doi: 10.1016/s0022-2836(95)80067-0. [DOI] [PubMed] [Google Scholar]
- Collado-Vides J., Magasanik B., Gralla J. D. Control site location and transcriptional regulation in Escherichia coli. Microbiol Rev. 1991 Sep;55(3):371–394. doi: 10.1128/mr.55.3.371-394.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crothers D. M. Upsetting the balance of forces in DNA. Science. 1994 Dec 16;266(5192):1819–1820. doi: 10.1126/science.7997876. [DOI] [PubMed] [Google Scholar]
- Dubochet J., Bednar J., Furrer P., Stasiak A. Z., Stasiak A., Bolshoy A. A. Determination of the DNA helical repeat by cryo-electron microscopy. Nat Struct Biol. 1994 Jun;1(6):361–363. doi: 10.1038/nsb0694-361. [DOI] [PubMed] [Google Scholar]
- Herendeen D. R., Kassavetis G. A., Barry J., Alberts B. M., Geiduschek E. P. Enhancement of bacteriophage T4 late transcription by components of the T4 DNA replication apparatus. Science. 1989 Sep 1;245(4921):952–958. doi: 10.1126/science.2672335. [DOI] [PubMed] [Google Scholar]
- Hunt T. P., Magasanik B. Transcription of glnA by purified Escherichia coli components: core RNA polymerase and the products of glnF, glnG, and glnL. Proc Natl Acad Sci U S A. 1985 Dec;82(24):8453–8457. doi: 10.1073/pnas.82.24.8453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klose K. E., North A. K., Stedman K. M., Kustu S. The major dimerization determinants of the nitrogen regulatory protein NTRC from enteric bacteria lie in its carboxy-terminal domain. J Mol Biol. 1994 Aug 12;241(2):233–245. doi: 10.1006/jmbi.1994.1492. [DOI] [PubMed] [Google Scholar]
- Klose K. E., Weiss D. S., Kustu S. Glutamate at the site of phosphorylation of nitrogen-regulatory protein NTRC mimics aspartyl-phosphate and activates the protein. J Mol Biol. 1993 Jul 5;232(1):67–78. doi: 10.1006/jmbi.1993.1370. [DOI] [PubMed] [Google Scholar]
- Kustu S., North A. K., Weiss D. S. Prokaryotic transcriptional enhancers and enhancer-binding proteins. Trends Biochem Sci. 1991 Nov;16(11):397–402. doi: 10.1016/0968-0004(91)90163-p. [DOI] [PubMed] [Google Scholar]
- Laundon C. H., Griffith J. D. Curved helix segments can uniquely orient the topology of supertwisted DNA. Cell. 1988 Feb 26;52(4):545–549. doi: 10.1016/0092-8674(88)90467-9. [DOI] [PubMed] [Google Scholar]
- Magasanik B. Reversible phosphorylation of an enhancer binding protein regulates the transcription of bacterial nitrogen utilization genes. Trends Biochem Sci. 1988 Dec;13(12):475–479. doi: 10.1016/0968-0004(88)90234-4. [DOI] [PubMed] [Google Scholar]
- Moreau P., Hen R., Wasylyk B., Everett R., Gaub M. P., Chambon P. The SV40 72 base repair repeat has a striking effect on gene expression both in SV40 and other chimeric recombinants. Nucleic Acids Res. 1981 Nov 25;9(22):6047–6068. doi: 10.1093/nar/9.22.6047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muzard G., Théveny B., Révet B. Electron microscopy mapping of pBR322 DNA curvature. Comparison with theoretical models. EMBO J. 1990 Apr;9(4):1289–1298. doi: 10.1002/j.1460-2075.1990.tb08238.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller M. M., Gerster T., Schaffner W. Enhancer sequences and the regulation of gene transcription. Eur J Biochem. 1988 Oct 1;176(3):485–495. doi: 10.1111/j.1432-1033.1988.tb14306.x. [DOI] [PubMed] [Google Scholar]
- Ninfa A. J., Reitzer L. J., Magasanik B. Initiation of transcription at the bacterial glnAp2 promoter by purified E. coli components is facilitated by enhancers. Cell. 1987 Sep 25;50(7):1039–1046. doi: 10.1016/0092-8674(87)90170-x. [DOI] [PubMed] [Google Scholar]
- Porter S. C., North A. K., Wedel A. B., Kustu S. Oligomerization of NTRC at the glnA enhancer is required for transcriptional activation. Genes Dev. 1993 Nov;7(11):2258–2273. doi: 10.1101/gad.7.11.2258. [DOI] [PubMed] [Google Scholar]
- Ptashne M. Gene regulation by proteins acting nearby and at a distance. Nature. 1986 Aug 21;322(6081):697–701. doi: 10.1038/322697a0. [DOI] [PubMed] [Google Scholar]
- Reitzer L. J., Magasanik B. Transcription of glnA in E. coli is stimulated by activator bound to sites far from the promoter. Cell. 1986 Jun 20;45(6):785–792. doi: 10.1016/0092-8674(86)90553-2. [DOI] [PubMed] [Google Scholar]
- Révet B. M., Sena E. P., Zarling D. A. Homologous DNA targeting with RecA protein-coated short DNA probes and electron microscope mapping on linear duplex molecules. J Mol Biol. 1993 Aug 5;232(3):779–791. doi: 10.1006/jmbi.1993.1431. [DOI] [PubMed] [Google Scholar]
- Su W., Porter S., Kustu S., Echols H. DNA-looping and enhancer activity: association between DNA-bound NtrC activator and RNA polymerase at the bacterial glnA promoter. Proc Natl Acad Sci U S A. 1990 Jul;87(14):5504–5508. doi: 10.1073/pnas.87.14.5504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theveny B., Revet B. DNA orientation using specific avidin-ferritin biotin end labelling. Nucleic Acids Res. 1987 Feb 11;15(3):947–958. doi: 10.1093/nar/15.3.947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss V., Claverie-Martin F., Magasanik B. Phosphorylation of nitrogen regulator I of Escherichia coli induces strong cooperative binding to DNA essential for activation of transcription. Proc Natl Acad Sci U S A. 1992 Jun 1;89(11):5088–5092. doi: 10.1073/pnas.89.11.5088. [DOI] [PMC free article] [PubMed] [Google Scholar]