Abstract
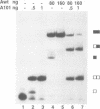
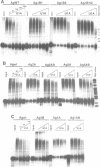
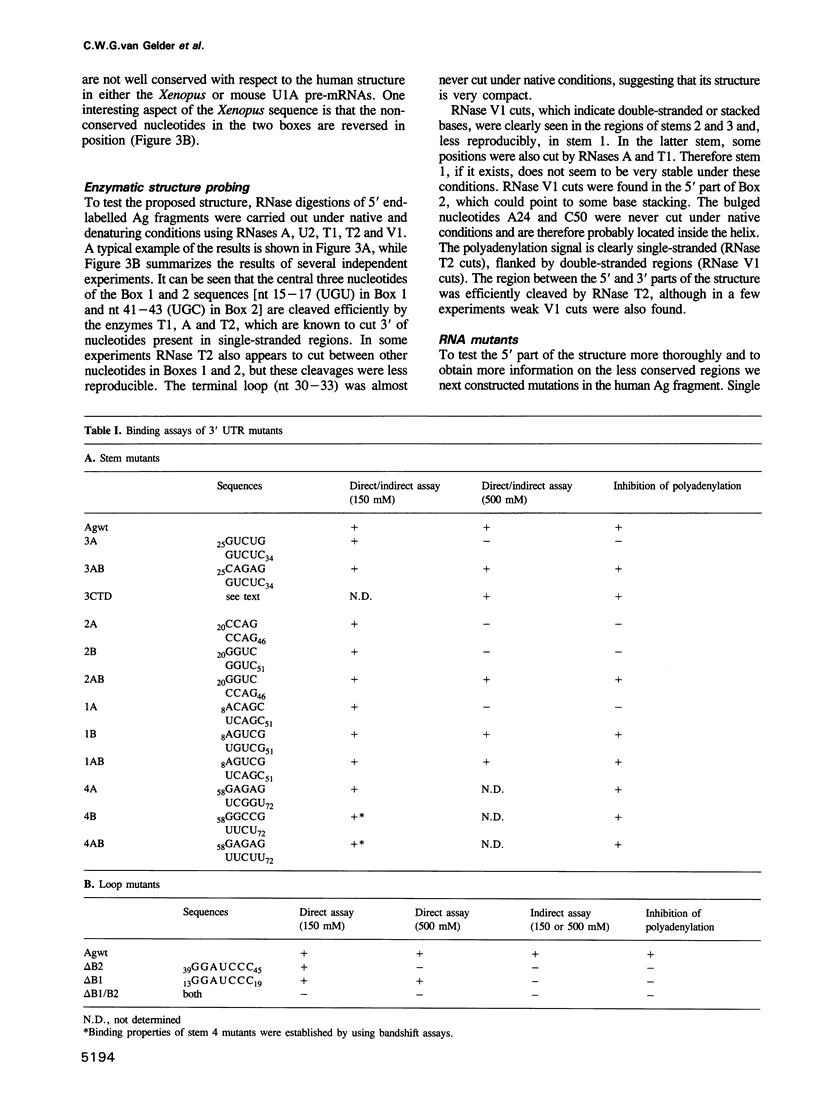
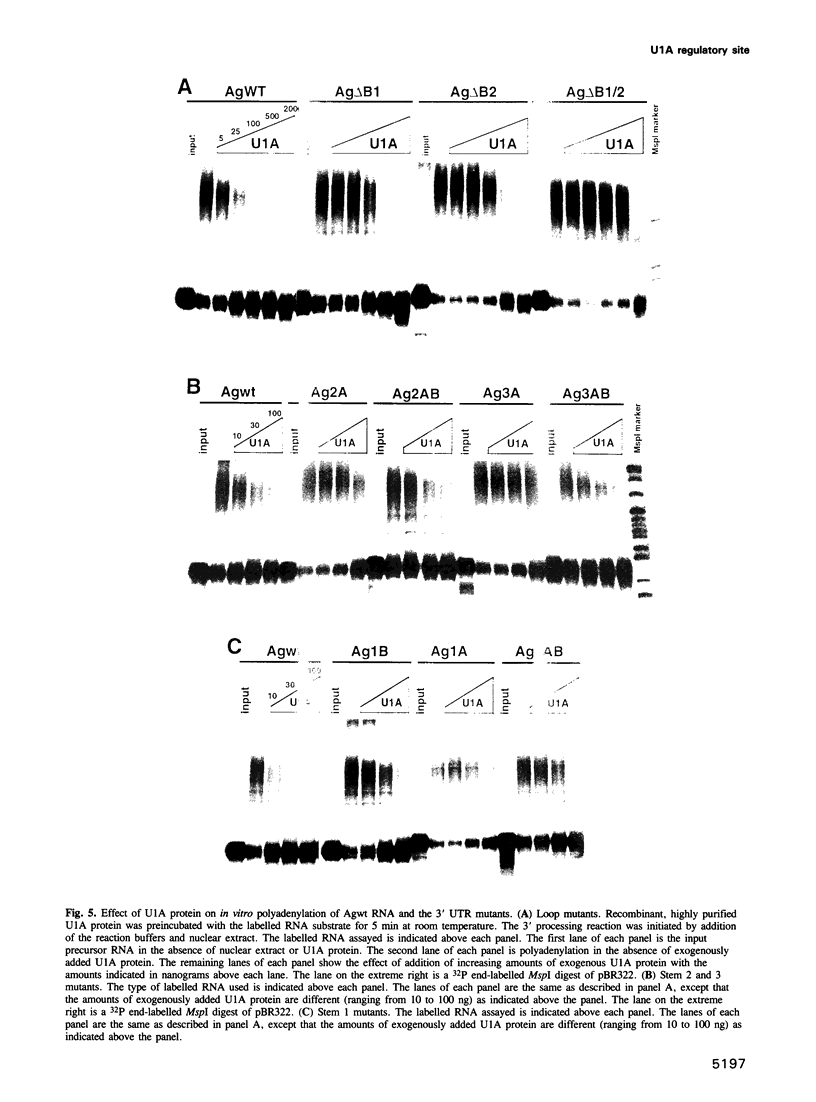
The human U1A protein-U1A pre-mRNA complex and the relationship between its structure and function in inhibition of polyadenylation in vitro were investigated. Two molecules of U1A protein were shown to bind to a conserved region in the 3' untranslated region of U1A pre-mRNA. The secondary structure of this region was determined by a combination of theoretical prediction, phylogenetic sequence alignment, enzymatic structure probing and molecular genetics. The U1A binding sites form (part of) a complex secondary structure which is significantly different from the binding site of U1A protein on U1 snRNA. Studies with mutant pre-mRNAs showed that the integrity of much of this structure is required for both high affinity binding to U1A protein and specific inhibition of polyadenylation in vitro. In particular, binding of a single molecule of U1A protein to U1A pre-mRNA is not sufficient to produce efficient inhibition of polyadenylation.
Full text
PDF
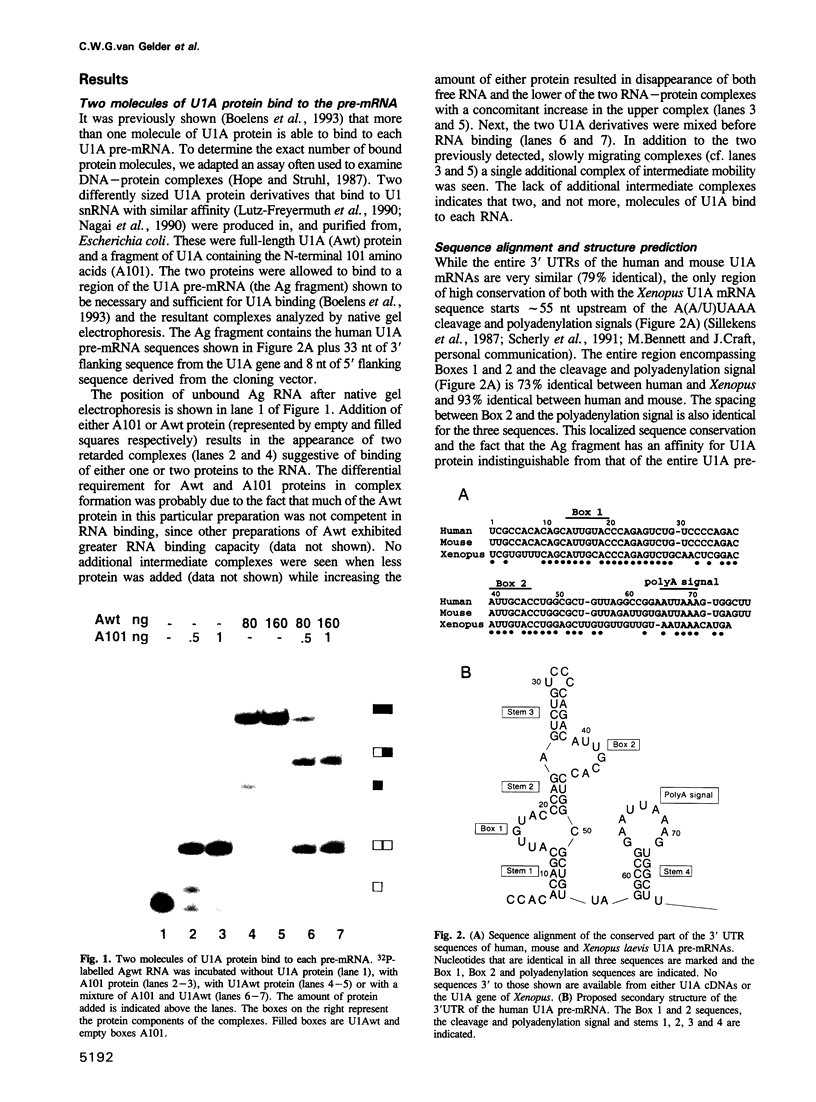
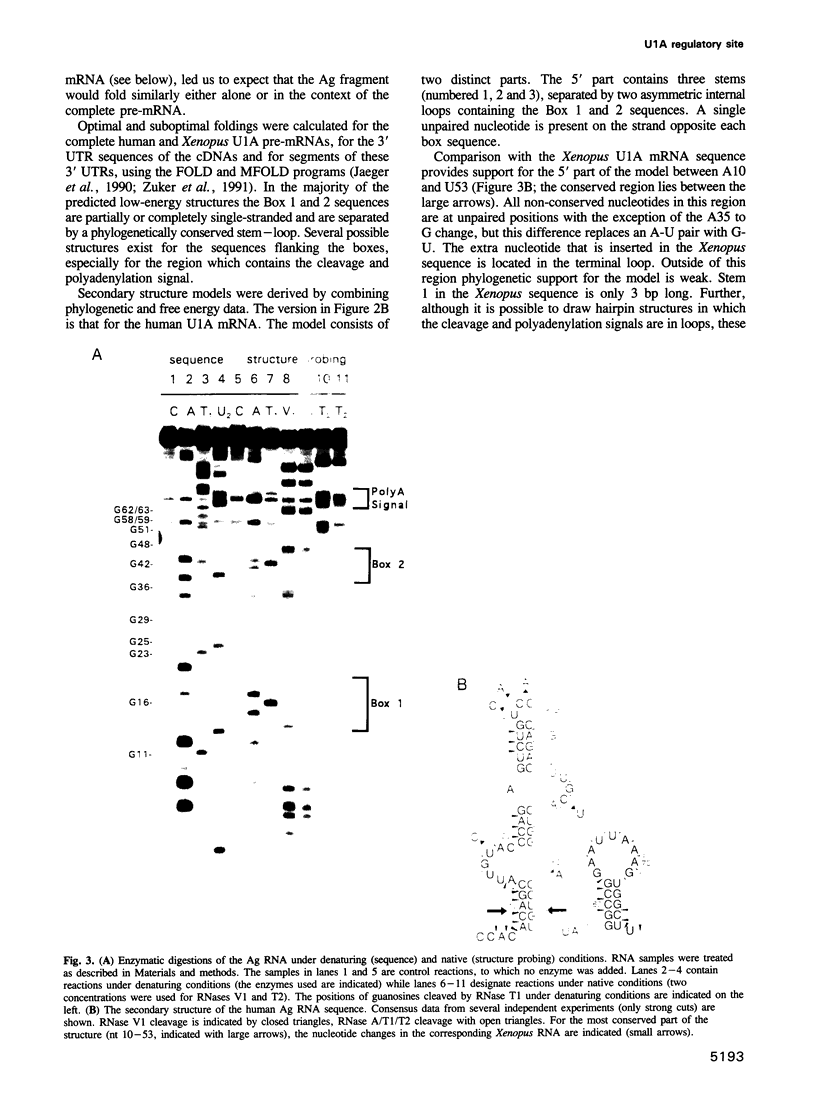





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bach M., Krol A., Lührmann R. Structure-probing of U1 snRNPs gradually depleted of the U1-specific proteins A, C and 70k. Evidence that A interacts differentially with developmentally regulated mouse U1 snRNA variants. Nucleic Acids Res. 1990 Feb 11;18(3):449–457. doi: 10.1093/nar/18.3.449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentley R. C., Keene J. D. Recognition of U1 and U2 small nuclear RNAs can be altered by a 5-amino-acid segment in the U2 small nuclear ribonucleoprotein particle (snRNP) B" protein and through interactions with U2 snRNP-A' protein. Mol Cell Biol. 1991 Apr;11(4):1829–1839. doi: 10.1128/mcb.11.4.1829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boelens W. C., Jansen E. J., van Venrooij W. J., Stripecke R., Mattaj I. W., Gunderson S. I. The human U1 snRNP-specific U1A protein inhibits polyadenylation of its own pre-mRNA. Cell. 1993 Mar 26;72(6):881–892. doi: 10.1016/0092-8674(93)90577-d. [DOI] [PubMed] [Google Scholar]
- Chastain M., Tinoco I., Jr Structural elements in RNA. Prog Nucleic Acid Res Mol Biol. 1991;41:131–177. doi: 10.1016/S0079-6603(08)60008-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehresmann C., Baudin F., Mougel M., Romby P., Ebel J. P., Ehresmann B. Probing the structure of RNAs in solution. Nucleic Acids Res. 1987 Nov 25;15(22):9109–9128. doi: 10.1093/nar/15.22.9109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall K. B., Stump W. T. Interaction of N-terminal domain of U1A protein with an RNA stem/loop. Nucleic Acids Res. 1992 Aug 25;20(16):4283–4290. doi: 10.1093/nar/20.16.4283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamm J., Dathan N. A., Scherly D., Mattaj I. W. Multiple domains of U1 snRNA, including U1 specific protein binding sites, are required for splicing. EMBO J. 1990 Apr;9(4):1237–1244. doi: 10.1002/j.1460-2075.1990.tb08231.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffman D. W., Query C. C., Golden B. L., White S. W., Keene J. D. RNA-binding domain of the A protein component of the U1 small nuclear ribonucleoprotein analyzed by NMR spectroscopy is structurally similar to ribosomal proteins. Proc Natl Acad Sci U S A. 1991 Mar 15;88(6):2495–2499. doi: 10.1073/pnas.88.6.2495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hope I. A., Struhl K. GCN4, a eukaryotic transcriptional activator protein, binds as a dimer to target DNA. EMBO J. 1987 Sep;6(9):2781–2784. doi: 10.1002/j.1460-2075.1987.tb02573.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaeger J. A., Turner D. H., Zuker M. Predicting optimal and suboptimal secondary structure for RNA. Methods Enzymol. 1990;183:281–306. doi: 10.1016/0076-6879(90)83019-6. [DOI] [PubMed] [Google Scholar]
- Jessen T. H., Oubridge C., Teo C. H., Pritchard C., Nagai K. Identification of molecular contacts between the U1 A small nuclear ribonucleoprotein and U1 RNA. EMBO J. 1991 Nov;10(11):3447–3456. doi: 10.1002/j.1460-2075.1991.tb04909.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krol A., Carbon P. A guide for probing native small nuclear RNA and ribonucleoprotein structures. Methods Enzymol. 1989;180:212–227. doi: 10.1016/0076-6879(89)80103-x. [DOI] [PubMed] [Google Scholar]
- Liao X. C., Tang J., Rosbash M. An enhancer screen identifies a gene that encodes the yeast U1 snRNP A protein: implications for snRNP protein function in pre-mRNA splicing. Genes Dev. 1993 Mar;7(3):419–428. doi: 10.1101/gad.7.3.419. [DOI] [PubMed] [Google Scholar]
- Lutz-Freyermuth C., Query C. C., Keene J. D. Quantitative determination that one of two potential RNA-binding domains of the A protein component of the U1 small nuclear ribonucleoprotein complex binds with high affinity to stem-loop II of U1 RNA. Proc Natl Acad Sci U S A. 1990 Aug;87(16):6393–6397. doi: 10.1073/pnas.87.16.6393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lührmann R., Kastner B., Bach M. Structure of spliceosomal snRNPs and their role in pre-mRNA splicing. Biochim Biophys Acta. 1990 Nov 30;1087(3):265–292. doi: 10.1016/0167-4781(90)90001-i. [DOI] [PubMed] [Google Scholar]
- Nagai K., Oubridge C., Jessen T. H., Li J., Evans P. R. Crystal structure of the RNA-binding domain of the U1 small nuclear ribonucleoprotein A. Nature. 1990 Dec 6;348(6301):515–520. doi: 10.1038/348515a0. [DOI] [PubMed] [Google Scholar]
- Nelissen R. L., Sillekens P. T., Beijer R. P., Geurts van Kessel A. H., van Venrooij W. J. Structure, chromosomal localization and evolutionary conservation of the gene encoding human U1 snRNP-specific A protein. Gene. 1991 Jun 30;102(2):189–196. doi: 10.1016/0378-1119(91)90077-o. [DOI] [PubMed] [Google Scholar]
- Scherly D., Boelens W., Dathan N. A., van Venrooij W. J., Mattaj I. W. Major determinants of the specificity of interaction between small nuclear ribonucleoproteins U1A and U2B'' and their cognate RNAs. Nature. 1990 Jun 7;345(6275):502–506. doi: 10.1038/345502a0. [DOI] [PubMed] [Google Scholar]
- Scherly D., Boelens W., van Venrooij W. J., Dathan N. A., Hamm J., Mattaj I. W. Identification of the RNA binding segment of human U1 A protein and definition of its binding site on U1 snRNA. EMBO J. 1989 Dec 20;8(13):4163–4170. doi: 10.1002/j.1460-2075.1989.tb08601.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scherly D., Kambach C., Boelens W., van Venrooij W. J., Mattaj I. W. Conserved amino acid residues within and outside of the N-terminal ribonucleoprotein motif of U1A small nuclear ribonucleoprotein involved in U1 RNA binding. J Mol Biol. 1991 Jun 20;219(4):577–584. doi: 10.1016/0022-2836(91)90651-l. [DOI] [PubMed] [Google Scholar]
- Sillekens P. T., Habets W. J., Beijer R. P., van Venrooij W. J. cDNA cloning of the human U1 snRNA-associated A protein: extensive homology between U1 and U2 snRNP-specific proteins. EMBO J. 1987 Dec 1;6(12):3841–3848. doi: 10.1002/j.1460-2075.1987.tb02721.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsai D. E., Harper D. S., Keene J. D. U1-snRNP-A protein selects a ten nucleotide consensus sequence from a degenerate RNA pool presented in various structural contexts. Nucleic Acids Res. 1991 Sep 25;19(18):4931–4936. doi: 10.1093/nar/19.18.4931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tuerk C., Gauss P., Thermes C., Groebe D. R., Gayle M., Guild N., Stormo G., d'Aubenton-Carafa Y., Uhlenbeck O. C., Tinoco I., Jr CUUCGG hairpins: extraordinarily stable RNA secondary structures associated with various biochemical processes. Proc Natl Acad Sci U S A. 1988 Mar;85(5):1364–1368. doi: 10.1073/pnas.85.5.1364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wahle E., Keller W. The biochemistry of 3'-end cleavage and polyadenylation of messenger RNA precursors. Annu Rev Biochem. 1992;61:419–440. doi: 10.1146/annurev.bi.61.070192.002223. [DOI] [PubMed] [Google Scholar]
- Zuker M., Jaeger J. A., Turner D. H. A comparison of optimal and suboptimal RNA secondary structures predicted by free energy minimization with structures determined by phylogenetic comparison. Nucleic Acids Res. 1991 May 25;19(10):2707–2714. doi: 10.1093/nar/19.10.2707. [DOI] [PMC free article] [PubMed] [Google Scholar]