Abstract
Background
Rapidly driven by the need for developing sustainable sources of nutritionally important fatty acids and the rising concerns about environmental impacts after using fossil oil, oil-plants have received increasing awareness nowadays. As an important oil-rich plant in China, Camellia oleifera has played a vital role in providing nutritional applications, biofuel productions and chemical feedstocks. However, the lack of C. oleifera genome sequences and little genetic information have largely hampered the urgent needs for efficient utilization of the abundant germplasms towards modern breeding efforts of this woody oil-plant.
Results
Here, using the 454 GS-FLX sequencing platform, we generated approximately 600,000 RNA-Seq reads from four tissues of C. oleifera. These reads were trimmed and assembled into 104,842 non-redundant putative transcripts with a total length of ∼38.9 Mb, representing more than 218-fold of all the C. oleifera sequences currently deposited in the GenBank (as of March 2014). Based on the BLAST similarity searches, nearly 42.6% transcripts could be annotated with known genes, conserved domains, or Gene Ontology (GO) terms. Comparisons with the cultivated tea tree, C. sinensis, identified 3,022 pairs of orthologs, of which 211 exhibited the evidence under positive selection. Pathway analysis detected the majority of genes potentially related to lipid metabolism. Evolutionary analysis of omega-6 fatty acid desaturase (FAD2) genes among 20 oil-plants unexpectedly suggests that a parallel evolution may occur between C. oleifera and Olea oleifera. Additionally, more than 2,300 simple sequence repeats (SSRs) and 20,200 single-nucleotide polymorphisms (SNPs) were detected in the C. oleifera transcriptome.
Conclusions
The generated transcriptome represents a considerable increase in the number of sequences deposited in the public databases, providing an unprecedented opportunity to discover all related-genes associated with lipid metabolic pathway in C. oleifera. It will greatly enhance the generation of new varieties of C. oleifera with increased yields and high quality.
Introduction
The tea-oil camellia (Camellia oleifera), a member of the family Theaceae, is well recognized as one of the world's four major woody oil tree together with oil palm, olive and coconut. It is an important and promoted woody oil plant in China, and has been widely utilized in many areas, such as food supplies, inks, lubricants, and cosmetics [1], [2]. The oil content of C. oleifera is up to 50%, usually serving as a high-quality cooking oil. More than 50% cooking oil in southern China, especially in Hunan Province, is tea-oil [3]. Tea oil contains several fatty acids, such as stearic, palmitic, oleic, linoleic and linolenic acids [4], [5]. Approximately ∼90% fatty acids of the C. oleifera are unsaturated fatty acid, comprising ∼83% (almost the highest among all natural oils) oleic acid and ∼7% linoleic acid [3], [5]. Besides, due to the similar oil composition between C. oleifera and olive, the oil of C. oleifera (tea-oil) has also been called “Oriental Olive Oil”. What's more, the tea oil can also reduce serum triglycerides and increase high-density lipoproteins (often regarded as good cholesterol) in humans [6]. Compared with olive and other corn oils, the oil of C. oleifera was also found to have a better stability against oxidation, leading to a suitable nutritional value [7]. Moreover, tea oil is still a good raw material for industrial uses and has been extensively utilized to manufacture soap, margarine, hair-oil, lubricants and rustproof oil. However, in contrast to its vital economic and strategic values, little information and genetic resources are available, which is mainly owing to the absence of genomic and/or transcriptomic resources for this non-model species.
Recent advances in high-throughput next-generation sequencing (NGS) technologies show a great potential for the large-scale production of genomic or transcriptomic data for a non-model species at reasonable costs [8]–[10]. Especially for the 454 pyrosequencing, of which the sequencing reads now approaching the length of traditional Sanger sequences, is ideal for the transcriptome sequencing for species that lacks a sequenced genome [11]. Besides, the latest versions of Newbler assembler from 454 can effectively assembly 454 RNA-Seq reads into putative transcripts, which can be better used for the subsequent gene discovery [12], [13], microarrays design [14] and high throughput SSRs or SNPs identification [15]–[17]. The SSRs and SNPs are often utilized as gene-based genetic markers and widely used for the generation of linkage maps and identification of quantitative trait loci (QTLs).
In this study, we sequenced and de novo assembled the transcriptome of C. oleifera using Roche/454 GS FLX massive parallel pyrosequencing platform for the first time. Our goals were to: 1) characterize the complete transcriptome of C. oleifera, and expand the genetic resources available for Camellia breeding programs; 2) identify the gene-based markers, including SSRs and SNPs, for future genetic analyses; 3) explore the dynamic evolution of orthologous genes between C. oleifera and C. sinensis, and assess the natural selection pressure assigning to genes during the Camellia evolutionary history; 4) discover the genes encoding enzymes that are involved in major metabolic pathways related to lipid biosynthesis and catabolism in C. oleifera, and further analyze the evolutionary process of oil quality determining gene (FAD2) among oil-plants. These datasets and results reported here will provide a public resource and information for future genetic and functional genomic studies in C. oleifera.
Results and Discussion
Transcriptome sequencing and de novo assembly
To comprehensively generate the C. oleifera transcriptome, four cDNA libraries of tender shoots, young leaves, flower buds and flowers were normalized and sequenced using the 454 GS FLX platform. This resulted in 591,033 (∼186 Mb) raw reads with an average length of 315 bp. An overview of the sequencing and data pre-processing is shown in Table 1 . After the removal of adaptors, primer sequences, poly-A tails as well as short and low quality sequences, a total of 554,198 (93.8%, ∼170 Mb) high-quality reads with an average length of 307 bp were retained and used for assembly. Size distribution for these trimmed, size-selected reads is shown in Figure 1a . For assembly, all trimmed and cleaned reads were mutually aligned and assembled using Newbler (version 2.8, Roche, IN, USA), yielding a set of 13,056 contiguous sequences (isotigs) longer than 100 bp, with an N50 of 771 bp. These results showed that more than half of the total assembly length of isotigs was >700 bp ( Table 2 and Figure 1b ). Due to the fact that many genes in the transcriptome are expressed at levels low enough to hinder adequate sampling for 454 sequencing [18], reads that had no apparent overlapping with other reads in the database may contain useful gene information, which could not be obtained from isotigs. Considering this, the remaining 107,369 high-quality reads with length ≥100 bp were also retained and treated as singletons (i.e., reads not assembled into isotigs). Size distribution of these singletons is shown in Figure 1b . To combine the singletons and assembled isotigs and reduce the redundancy among them, CD-HIT [19] (version 4.6) was used to cluster these 120,425 sequences into 104,842 unigenes for further analyses. To our knowledge, so far, the number of sequences that are publicly available for C. oleifera is fewer than 500. Therefore, the transcriptome dataset reported here significantly expands the genetic resources available for C. oleifera in the public database, and can serve as a foundation for future investigation of gene expression, important pathways, molecular genetics and functional genomics in C. oleifera.
Table 1. Summary of the sequencing data of C. oleifera.
| Library | Raw data | Clean data | ||||
| No. of reads | Average length (bp) | Total length (bp) | No. of reads | Average length (bp) | Total length (bp) | |
| Tender shoots | 119,177 | 313.10 | 37,314,251 | 112,055 | 299.57 | 33,567,808 |
| Young leaves | 169,653 | 304.50 | 51,658,715 | 157,874 | 297.34 | 46,942,253 |
| Flower buds | 172,391 | 300.85 | 51,863,705 | 159,944 | 295.85 | 47,320,030 |
| Flowers | 129,812 | 349.95 | 45,427,523 | 124,325 | 342.00 | 42,519,448 |
| Total | 591,033 | 315.15 | 186,264,194 | 554,198 | 307.38 | 170,349,539 |
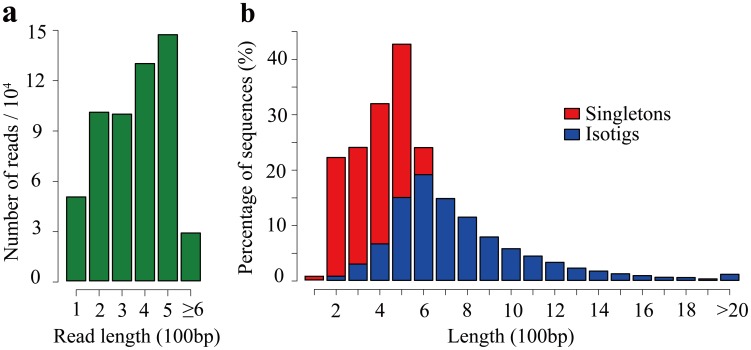
Figure 1. Overview of C. oleifera transcriptome sequencing and assembly.
(a) Length distribution of 454 sequencing reads after filtering and trimming adapters. (b) Length distribution of the singletons and assembled isotigs.
Table 2. Summary of de novo assembly and transcriptome annotation of C. oleifera.
| # Sequences | |
| Assembly | |
| Total number of isotigs | 13,056 |
| Total isotigs length (bp) | 9,442,537 |
| Isotig N50 (bp) | 771 |
| Number of isotigs (length ≥500 bp) | 9,786 |
| Number of isotigs (length ≥1 kb) | 2,095 |
| Number of singletons | 107,369 |
| Total number of unigenes | 104,842 |
| Total unigenes length (bp) | 38,944,469 |
| GC content (%) | 39.1 |
Functional annotation of the C. oleifera transcriptome
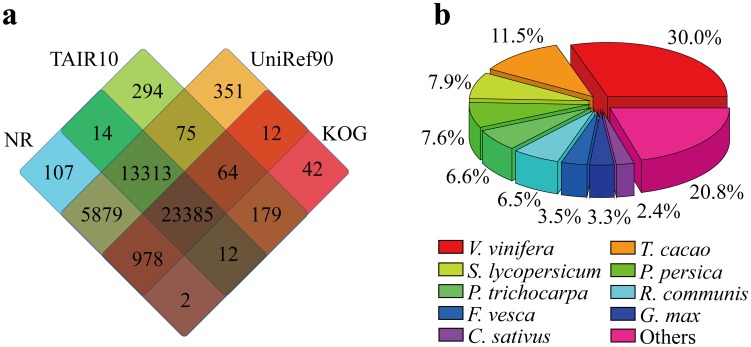
To comprehensively annotate the transcriptome of C. oleifera, several complementary approaches were adopted. First, the 104,842 non-redundant assembled sequences were aligned against four public protein databases, including National Center for Biotechnology Information (NCBI) non-redundant (NR) protein database, Arabidopsis Information Resource database (TAIR, version 10), UniRef90 and Clusters of Orthologous Groups (KOGs) protein database, using BLASTX algorithm [20]. With an E-value threshold of 10−5, a total of 10,739 isotigs (82.3% of total isotigs) and 33,968 singleton sequences (37.0% of total singletons) had the best BLAST matches with known proteins in at least one of the four databases, while 23,385 unigenes (22.3% of total unigenes; 6,428 isotigs plus 16,957 singletons) had significant BLAST matches to proteins in all of the four databases ( Table 2 , Figure 2a and Table S1). The top-hit species distribution of NR BLAST matches is shown in Figure 2b . Approximately 30% of the sequences had significant matches with genes from Vitis vinifera, followed by Theobroma cacao (11.5%), Solanum lycopersicum (7.9%), Prunus persica (7.6%), Populus trichocarpa (6.6%), Ricinus communis (6.5%), Fragaria vesca (3.5%), Glycine max (3.3%), and Cucumis sativus (2.4%).
Figure 2. Characteristics of the homology search of unigenes against four public protein databases.
(a) Venn diagram showing the BLAST searches of C. oleifera transcriptome against the four public protein databases. De novo reconstructed transcript sequences were used to BLAST against public databases including NCBI's NR, TAIR10, UniRef90 and KOG. The number of transcripts that have significant hits (E-value≤10−5) against the four databases is shown in each intersection of the Venn diagram. (b) Species distribution is shown as the percentage of the total homologous sequences (with an E-value≤10−5). We searched the NCBI NR database by using BLASTx and extracted the best hit of each sequence for analysis.
Considering that the conserved domain information within a gene would be useful for interpreting the gene's function, we used the domain-based alignments to further detect the function of the C. oleifera transcriptome. To facilitate the domain annotation, we began by predicting the open reading frame (ORF) for each unigene, and then all the unigenes with detected ORFs were submitted to HMM search against the Pfam database [21] (version 27.0). In total, 18,992 matches were returned and categorized into 4,860 domains/families ( Table 2 and Table S2). The top ten most abundant domains/families are given in Table 3 , which contained “Protein kinase domain”, “Protein tyrosine kinase”, “WD domain, G-beta repeat (WD40)”, “Cytochrome P450”, “RNA recognition motif”, “Reverse transcriptase, RNA-dependent DNA polymerase”, “UDP-glucoronosyl and UDP-glucosyl transferase”, “Mitochondrial carrier protein”, “ABC transporter”, and “PPR repeat family”.
Table 3. The ten most frequently occurred PFAM domains/families in the unigenes of C. oleifera.
| Acc ID | Conserved domain/family | Sequences (n) |
| PF00069 | Protein kinase domain | 440 |
| PF07714 | Protein tyrosine kinase | 220 |
| PF00400 | WD domain, G-beta repeat (WD40) | 178 |
| PF00067 | Cytochrome P450 | 171 |
| PF00076 | RNA recognition motif (RRM_1) | 151 |
| PF07727 | Reverse transcriptase, RNA-dependent DNA polymerase (RVT_2) | 112 |
| PF00201 | UDP-glucoronosyl and UDP-glucosyl transferase | 103 |
| PF00153 | Mitochondrial carrier protein | 86 |
| PF00005 | ABC transporter | 86 |
| PF13041 | PPR repeat family (PPR_2) | 84 |
Besides, we also used the Gene Ontology (GO) classification system to annotate the possible functions of the unigenes. Briefly, using the Blast2GO platform [22]–[24], sequences with a best match from NR database were further assigned with GO terms and Enzyme Commission (EC) numbers. Overall, 27,531 transcripts were assigned to 122,340 GO term annotations with an average number of 4.4 GO terms for each transcript ( Table 2 and Table S3). The distribution of ten most abundant GO terms for biological processes, molecular functions, and cellular components is presented in Figure 3 . Additionally, of the 27,531 sequences annotated with GO terms, 9,993 sequences were assigned with EC numbers and involved in 323 different pathways when the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway mapping based on EC numbers for assignments was carried out ( Table 2 ).
Figure 3. Histogram presentation of the most abundant Gene Ontology (GO) terms assigned to the C. oleifera transcriptome.
The NR based Blast2GO results are summarized into three main GO categories: biological process (BP), cellular component (CC), and molecular function (MF). Only the top ten GO terms for each main function category are shown (blue: BP; red: CC; green: MF). The corresponding GO IDs are presented in parentheses. The x-axis indicates the number of genes assigned to the same GO term. One unigene may be matched to multiple GO terms.
Identification of SSRs, SNPs and Indels
Simple sequence repeats (SSRs) (also called microsatellites), consisting of repeated sequences of 2–6 bp in length, have been widely used in QTL analysis, evaluation of genetic variation and construction of genetic linkage maps [25]–[28]. Although C. oleifera is regarded as an important plant in China, few genetic markers are currently available for this species. The transcriptome data obtained by 454 sequencing provided an excellent source for mining and development of gene-based markers. Here, using a Perl script known as MicroSAtellite (MISA, http://pgrc.ipk-gatersleben.de/misa), we totally detected 2,345 SSRs from 13,056 assembled isotigs (Table S4). Of the 2,345 detected SSRs, di-nucleotide repeats were the most abundant type (1,373; 58.6%), followed by tri-nucleotides (706; 30.1%), hexa-nucleotides (99; 4.2%), penta-nucleotides (93; 4.0%) and tetra-nucleotide (74; 3.2%) ( Figure 4a ). Among the 1,373 di-nucleotide repeats, CT/AG (37.4%) was the most abundant motifs, followed by TC/GA (35.2%) and AT/AT (19.7%). GAA/TTC (10.5%) was the most common motif for tri-nucleotide repeats. Unlike other cereal species [29] like barley, maize, oats, rice, rye, wheat and sorghum, of which tri-nucleotide repeats are the most abundant class of microsatellite, the main SSR type of C. oleifera is di-nucleotide repeat. To the best of our knowledge, although there are much more microsatellites that were developed for the genus Camellia [30]–[32], relatively fewer are publicly available for C. oleifera, suggesting that the SSRs reported here will be particularly useful in the future genetic characterization and germplasm utilization.
Figure 4. Distribution of simple sequence repeats (SSRs), single nucleotide polymorphisms (SNPs) and insertion/deletions (InDels) in C. oleifera isotigs.
(a) Di-, tri-, tetra-, penta- and hexa-nucleotide repeats were analyzed. The x-axis shows the type of the SSRs, whereas y-axis shows total number of SSRs in different classes. (b) Frequencies of different SNPs/InDels. The x-axis indicates the substitution type of SNPs/InDels, while y-axis represents the number of SNPs/InDels for each substitution type.
Besides SSRs analysis, potential SNPs/Indels were also detected. In total, we identified 20,250 high-quality SNPs and 16,906 Indels from the C. oleifera transcriptome (Table S5). Among all the SNPs, transitions (65.6%) were more frequent than tranversions (34.4%) ( Figure 4b ). The number of SNPs detected per transcript was highly variable, ranging from 1 to 106; however, approximately 27.3% of the transcripts contained only one or two SNPs. Furthermore, of the predicted 20,250 SNPs, 15,364 (75.9%) were obtained from isotigs with NR annotation. These SNPs can be considered as priority candidates for marker development and should be very useful for future germplasm utilization and breeding programs for C. oleifera.
Identification of orthologous genes and sequence divergence analysis between C. oleifera and C. sinensis
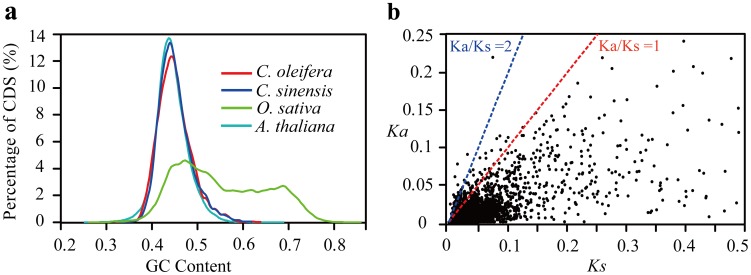
Large-scale transcriptome sequencing of C. sinensis reported previously [33] provides a valuable source data for the comparative analysis of the evolution of Camellia orthologous genes. Using a reciprocal best hit (RBH) method with further relatively strict filters, we totally identified 3,022 putatively orthologous genes between C. sinensis and C. oleifera transcriptomes. Among the 3,022 orthologous genes, the average length was 645.6 bp with an average similarity of 95.74%. The overall GC content of C. oleifera in coding regions was 44.96%, consistent with C. sinensis (∼44.68%) and A. thaliana (∼44.11%) and much lower than O. stativa (∼52.96%) ( Figure 5a ). The 3,022 orthologous genes were also annotated with GO terms, and 1,717 orthologous were found to be involved in molecular functions, 1,571 orthologous were involved in biological processes, and 1,255 orthologous were involved in cellular components.
Figure 5. Characteristics of the 3,022 orthologous genes between C. oleifera and C. sinensis.
(a) Distribution of GC content of CDS sequences among the C. oleifera, C. sinensis, O. sativa and A. thaliana. The CDS sequences of O. sativa (version 7.0) and A. thaliana (version 10) were downloaded from MSU Rice Genome Annotation Project (http://rice.plantbiology.msu.edu/) and TAIR (http://www.arabidopsis.org/), respectively. (b) Distribution of Ka and Ks. The mean Ka/Ks value is 0.39. The red line indicates the threshold of Ka/Ks = 1, whereas the blue line shows the more conservative threshold of Ka/Ks = 2. Analysis was performed using the method by Yang & Nielsen (2000).
To understand the molecular evolution of orthologous genes between C. oleifera and C. sinensis, especially the genes undergoing purifying or positive selections, we calculated the synonymous (Ks) and non-synonymous (Ka) substitution rate for each orthologous gene pair. The mean value of Ka/Ks of these sequence pairs was 0.39. The Ka/Ks ratio (denoted as ω) is widely used to detect selective pressure acting on protein-coding sequences. Ka/Ks>1 indicates a sign of positive (adaptive) selection, Ka/Ks<1 shows a signature of negative (purifying) selection, and Ka/Ks = 1 means the neutral evolution. In our analysis, we found that the majority of sequences pairs (93%; 2,811/3,022) had a Ka/Ks ratio <1, suggesting most of the orthologous genes undergo purifying selection ( Figure 5b ). However, we also identified 211 sequences under accelerated evolution with ratios of Ka/Ks>1 and 38 sequences with ratios of Ka/Ks>2 ( Figure 5b and Table S6). These sequences were found to be mainly involved in “ATP binding (GO: 0005524)”, “integral to membrane (GO: 0016021)” and “oxidation-reduction process (GO: 0055114)”. These fast evolving transcripts may be useful for identifying genes that perhaps strongly underwent positive selection during evolution process and might be responsible for speciation in the Camellia lineage.
Metabolic pathways in C. oleifera
Composed with approximately 82∼84% unsaturated fatty acid [34] and a low content of saturated fat, the oil of C. oleifera has been wildly used in China for cooking oil, lubricants and cosmetics. Considering its high storage of lipids as potential new raw materials for biofuel production, we next focus on the pathways related to lipid metabolism, mainly including fatty acid and triacylglycerols (TAG) metabolic pathways.
Fatty acid biosynthesis and catabolism
The plant fatty acids possess highly edible and industrial values. However, free fatty acids are rarely found in nature but instead are often esterified to a phospholipid (membrane lipids) [35], [36] and glycerolipid (energy-storage) [37]. The de novo biosynthesis of fatty acids, up to a chain length of C16 or C18, from acetyl-CoA mainly involves two enzyme systems: acetyl-CoA carboxylase (ACC) and fatty acid synthase (FAS) complex. In higher plants, both of these two systems are located in the chloroplast and the majority of the component polypeptides are nuclear-encoded. Although the overall fatty acid biosynthesis pathways are well studied in eukaryotes [38], [39], much less are known in the C. oleifera.
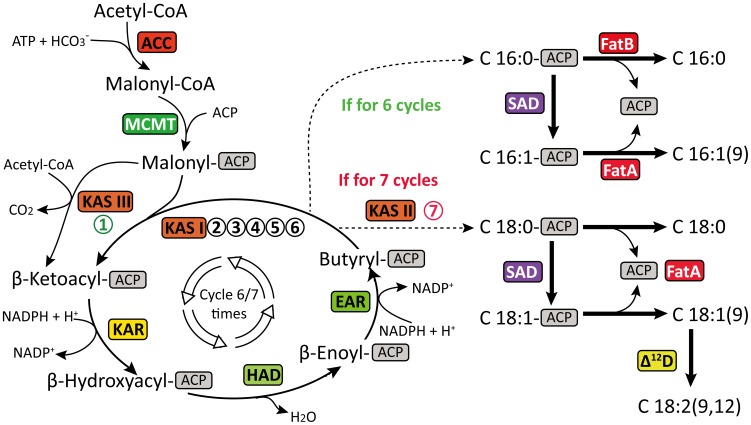
Based on the de novo assembly and functional annotation of the C. oleifera transcriptome, we have successfully identified multiply transcripts encoding the key enzymes involved in the fatty acid biosynthesis and catabolism pathway of C. oleifera ( Table 4 and Table S7). Fatty acid biosynthesis in C. oleifera was derived from the acetyl-CoA, which was initially catalyzed by the acetyl-CoA carboxylase (ACC, EC: 6.4.1.2) to form malonyl-CoA. Next, malonyl-CoA ACP transacylase (MCMT, EC: 2.3.1.39) catalyzed the transfer of acyl carrier protein (ACP) to the malonyl group, producing a malonyl-ACP, the primary substrate for the subsequent elongation. During the elongation process, four reactions were required with the addition of two carbons. First, the beta-ketoacyl-ACP synthase III (KAS III, EC: 2.3.1.180) catalyzed the initial condensation reaction, linked the acetyl group from acetyl-CoA to the malonyl-ACP, and yielded the beta-ketoacyl-ACP containing four carbons. Then, the ketoacyl-ACP was reduced by an NADPH-dependent beta-ketoacyl-ACP reductase (KAR, EC: 1.1.1.100) to generate the beta-hydroxyacyl-ACP. Next, 3-Hydroxyacyl-ACP dehydratase (HAD, EC: 4.2.1.-) removed a molecule of water from beta-hydroxyacyl-ACP to generate an enoyl-ACP, which was finally reduced by NADH to butyryl-ACP in a reaction catalyzed by enoyl-ACP reductase (EAR, EC: 1.3.1.9). The product of the first synthetic cycle, butyryl-ACP, was the substrate for further elongation rounds, each of which used one molecule of malonyl-ACP and released carbon dioxide. Notably, the condensations from C4 to C16 were carried out by beta-ketoactyl-ACP synthase I (KAS I, EC: 2.3.1.41), instead of the beta-ketoactyl-ACP III, although most enzymes used for the further elongation were the same as utilized for generating butyryl-ACP from acetyl-CoA and malonyl-ACP ( Figure 6 ). While the reaction from C16 to C18 was catalyzed by beta-ketoactyl-ACP synthase II (KAS II, EC: 2.3.1.179).
Table 4. Enzymes involved in fatty acid biosynthesis and catabolism identified by the annotation of the C. oleifera transcriptome.
| Enzyme | Symbol | EC Number | Number of unigenes |
| Fatty acid biosynthesis | |||
| Biotin carboxylase | BC | 6.3.4.14 | 1 |
| Biotin carboxyl carrier protein | BCCP | 6.4.1.2 | 8 |
| Acetyl-CoA carboxylase | ACC | 6.4.1.2 | 17 |
| malonyl-CoA-ACP transacylase | MCMT | 2.3.1.39 | 3 |
| Beta-ketoacyl-ACP synthase I | KAS I | 2.3.1.41 | 7 |
| Beta-ketoacyl-ACP synthase II | KAS II | 2.3.1.179 | 3 |
| Beta-ketoacyl-ACP synthase III | KAS III | 2.3.1.180 | 7 |
| Beta-ketoacyl-ACP reductase | KAR | 1.1.1.100 | 8 |
| 3R-hydroxyacyl-ACP dehydrase | HAD | 4.2.1.17; 4.2.1.- | 2 |
| Enoyl-ACP reductase (NADH) | EAR | 1.3.1.9 | 4 |
| Acyl-ACP thioesterase A | FatA | 3.1.2.14 | 4 |
| Acyl-ACP thioesterase B | FatB | 3.1.2.14 | 5 |
| Fatty acid desaturation | |||
| Stearoyl-ACP Desaturase | SAD | 1.14.19.2 | 10 |
| Δ12 (ω6)-Desaturase | Δ12 D | 1.14.19.6 | 3 |
| Fatty acid catabolism | |||
| Long-chain acyl-CoA synthetase | LACS | 6.2.1.3 | 6 |
| Acyl-CoA oxidase | ACX | 1.3.3.6 | 12 |
| Enoyl-CoA hydratase | ECH | 4.2.1.17 | 6 |
| 3s-hydroxyacyl-CoA dehydrogenase | HACDH | 1.1.1.35 | 6 |
| Ketoacyl-CoA Thiolase | KAT | 2.3.1.16 | 12 |
| Acyl-CoA Thioesterase | ACT | 3.1.2.2 | 4 |
| Enoyl-CoA isomerase | Isom | 5.3.3.8 | 4 |
| Dienoyl CoA Isomerase | DCI | 5.3.3.8; 4.2.1.17 | 1 |
| Dienoyl-CoA Reductase | Red | 1.3.1.34 | 9 |
| Trans-2-enoyl-CoA reductase | ECR | 1.3.1.38 | 4 |
| Enoyl-CoA Hydratase 2 | ECH2 | 4.2.1.17 | 5 |
Figure 6. Core reactions of fatty acid biosynthesis reconstructed based on the de novo assembly and annotation of C. oleifera transcriptome.
During fatty acid biosynthesis, two-carbon units are added for each cycle reaction, and the four-step cycle is repeated until the appropriate chain-length is reached. Finally, different types of fatty acids are synthesized. The identified enzymes are shown in boxes and abbreviated as below: ACC, acetyl-CoA carboxylase (EC: 6.4.1.2); MAT, malonyl-CoA ACP transacylase (EC: 2.3.1.39); KAS, beta-ketoacyl-ACP synthase (KAS I, EC: 2.3.1.41; KASII, EC: 2.3.1.179; KAS III, EC: 2.3.1.180); KAR, beta-ketoacyl-ACP reductase (EC: 1.1.1.100); HAD, beta-hydroxyacyl-ACP dehydrase (EC: 4.2.1.-); EAR, enoyl-ACP reductase (EC: 1.3.1.9); AAD, acyl-ACP desaturase (EC: 1.14.19.2); OAH, oleoyl-ACP hydrolase (EC: 3.1.2.14); FatA, Acyl-ACP thioesterase A (EC: 3.1.2.-); Δ12D, Δ12(ω6)-desaturase (EC: 1.4.19.6). The numbers-in-circles indicates the repeat time of the condensation reaction.
For the synthesis of unsaturated fatty acids in plastid, a double bond was introduced to the acyl group esterified to ACP via acyl-ACP desaturase (AAD, EC: 1.14.19.2). The elongation of fatty acids was terminated when the acyl group was removed from the ACP by acyl-ACP thioesterase enzyme, oleoyl-ACP hydrolase (OAH, EC: 3.1.2.14), or when acyl-ACP released the free fatty acid. The final fatty acid composition was determined by the activities of enzymes that used these acyl-ACPs at the termination phase of fatty acid synthesis. We also identified desaturation enzymes Δ12(ω6)-desaturase (Δ12D/FAD2, EC: 1.4.19.6), which desaturated oleic acid (C18:1n-9) to generate linoleic acid (C18:2n-6). The above description showed the pathway responsible for the formation and conversion of fatty acid in C. oleifera and was depicted in Figure 6 . We failed to identify any genes encoding enzymes involving further desaturation and elongation of linoleic acid to form longer chain polyunsaturated fatty acid in our annotated datasets. This is probably related to the certain components of fatty acids in C. oleifera, or the lack of transcriptome sequences from libraries of seed tissues in some case. Besides synthesis, we also identified all the enzymes involving fatty acid catabolism (namely, beta-oxidation pathway) in C. oleifera ( Table 4 and Table S7). The beta-oxidation pathway in C. oleifera involved four enzymes, including acyl-CoA oxidase (ACX, EC: 1.3.3.6), enoyl-CoA hydratase (ECH, EC: 4.2.1.17), 3-hydroxyacyl-CoA dehydrogenase (HACDH, EC: 1.1.1.35), and ketoacyl-CoA thiolase (KAT, EC: 2.3.1.16).
Together, the C. oleifera transcriptome presented here covers most of the enzymes required for the biosynthesis, elongation and metabolism of fatty acids ( Table 4 and Table S7). These identified enzymes contribute to the biochemical and molecular information needed for metabolic engineering of fatty acid synthesis. For example, linoleic acid, or omega-6 fatty acid, is an essential fatty acid for people's health. With the lack of Δ12(ω6)-desaturase (Δ12D, EC: 1.4.19.6), they cannot be synthesize by human body and have to be obtained from the diet. Further genetic engineering with these enzymes will contribute to oil composition, satisfy people's demands for oil and improve our life.
TAG biosynthesis and catabolism
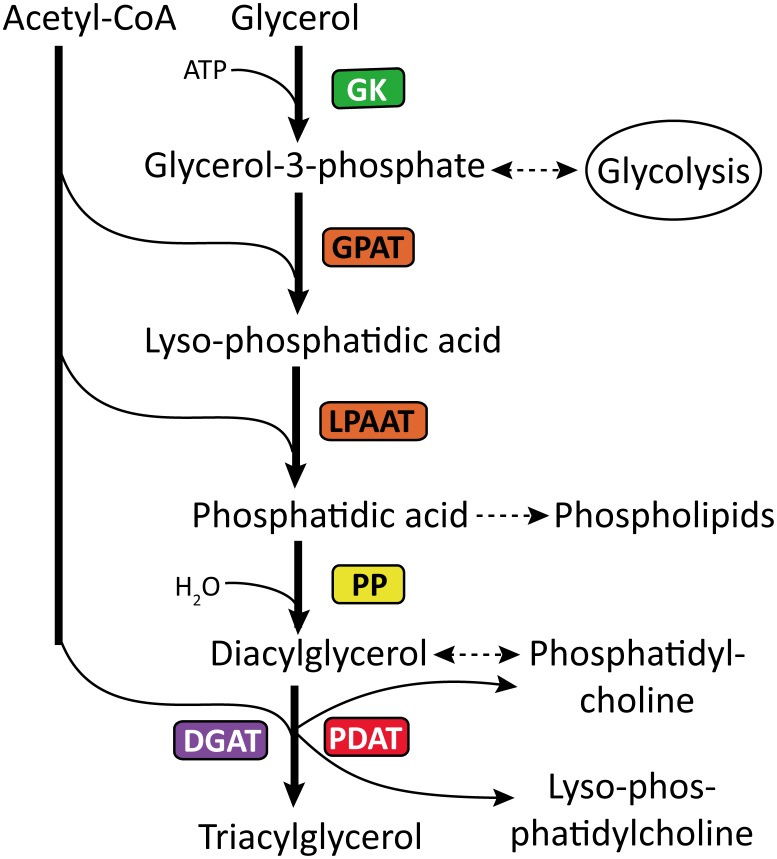
Nearly all eukaryotic organisms and even a few prokaryotes have the ability to synthesize triacylglycerols [40], leaving alone the important oil-plant, C. oleifera. Despite the global process for TAG biosynthesis in C. oleifera has been known, however, merely fewer genes were cloned. Here, according to the functional annotation of C. oleifera transcriptome, transcripts encoding most of the enzymes involving in the TAG biosynthesis pathway were also identified. Table 5 and Table S7 lists all the enzymes obtained above, and Figure 7 shows the TAG pathway reconstructed based on the determined transcripts. The most important route to triacylglycerol biosynthesis is the glycerol-3-phosphate (G-3-P) pathway. Within this pathway, the glycerol-3-phosphate was first produced by either the catabolism of glucose (glycolysis) or the action of the enzyme glycerol kinase (GK, EC: 2.7.1.30) on free glycerol, and subsequently catalyzed by a glycerol-3-phosphate acyltransferase (GPAT, EC: 2.3.1.15) at the position sn-1 to form lyso-phosphatidic acid. The lyso-phosphatidic acid was continually acylated by an acylglycero-3-phosphate acyltransferase (LPAAT, EC: 2.3.1.51) at the position sn-2 to form phosphatidic acid (PA). And then the phosphate group of the PA was removed by an enzyme called phosphatidic acid phosphohydrolase (PP) to generate diacylglycerols (DAG). After the synthesis of DAG, the formation of TAG could occur in two ways ( Figure 7 ). In one pathway, diacylglycerol acyltransferase (DGAT) transferred an acyl group from acetyl-CoA to sn-3 of DAG to form TAG. Another pathway involved a phospholipid∶diacylglycerol acyltransferase (PDAT) that utilized phospholipid as the acyldonor in TAG formation. It is worth mentioning that the last step of TAG pathway was an important determinant of cellular oil content and quality. These genes can be as priority candidates for the further cloning.
Table 5. Enzymes involved in TAG biosynthesis and catabolism identified by the annotation of the C. oleifera transcriptome.
| Enzyme | Symbol | EC Number | Number of unigenes |
| TAG biosynthesis | |||
| Glycerol kinase | GK | 2.7.1.30 | 6 |
| Glycerol-3-phosphate O-acyltransferase | GPAT | 2.3.1.15 | 1 |
| 1-Acyl-sn-glycerol-3-phosphate O-acyltransferase | LPAAT2 | 2.3.1.51 | 5 |
| LPAAT4 | 2.3.1.51 | 3 | |
| LPAAT5 | 2.3.1.51 | 3 | |
| Phosphatidate phosphatase | PP | 3.1.3.4 | 5 |
| Diacylglycerol O-acyltransferase | DGAT1 | 2.3.1.20 | 2 |
| DGAT2 | 2.3.1.20 | 1 | |
| DGAT3 | 2.3.1.20 | 1 | |
| Phospholipid∶diacyglycerol acyltransferase | PDAT1 | 2.3.1.158 | 13 |
| Monoacylglycerol Acyltransferase | MAGAT | 2.3.1.22 | 1 |
| Diacylglycerol Cholinephosphotransferase | DAG-CPT | 2.7.8.2 | 1 |
| Phosphatidylcholine∶diacylglycerol cholinephosphotransferase | PDCT | 2.7.8.* | 1 |
| 1-Acylglycerol-3-Phosphocholine Acyltransferase | LPCAT1 | 2.3.1.23 | 3 |
| Choline Kinase | CK1 | 2.7.1.32 | 4 |
| Choline-Phosphate Cytidylyltransferase | CCT1 | 2.7.7.15 | 4 |
| TAG catabolism | |||
| Triacylglycerol lipase | TAGL | 3.1.1.3 | 18 |
| Monoacylglycerol Lipase (MAGL) | MAGL | 3.1.1.23 | 20 |
Figure 7. Triacylglycerol (TAG) biosynthesis pathway reconstructed based on the de novo assembly and annotation of C. oleifera transcriptome.
Identified enzymes are shown in boxes, including: GK, glycerol kinase (EC: 2.7.1.30); GPAT, glycerol-3-phosphate O-acyltransferase (EC: 2.3.1.15); AGPAT, 1-acyl-sn-glycerol-3-phosphate O-acyltransferase (EC: 2.3.1.51); PP, phosphatidate phosphatase (EC: 3.1.3.4); DGAT, diacylglycerol O-acyltransferase (EC: 2.3.1.20); and PDAT, phopholipid∶ diacyglycerol acyltransferase (EC: 2.3.1.158). The dashed arrows denote reaction(s) in which the enzymes are not shown.
Evolution of the omega-6 fatty acid desaturase 2 (FAD2) genes
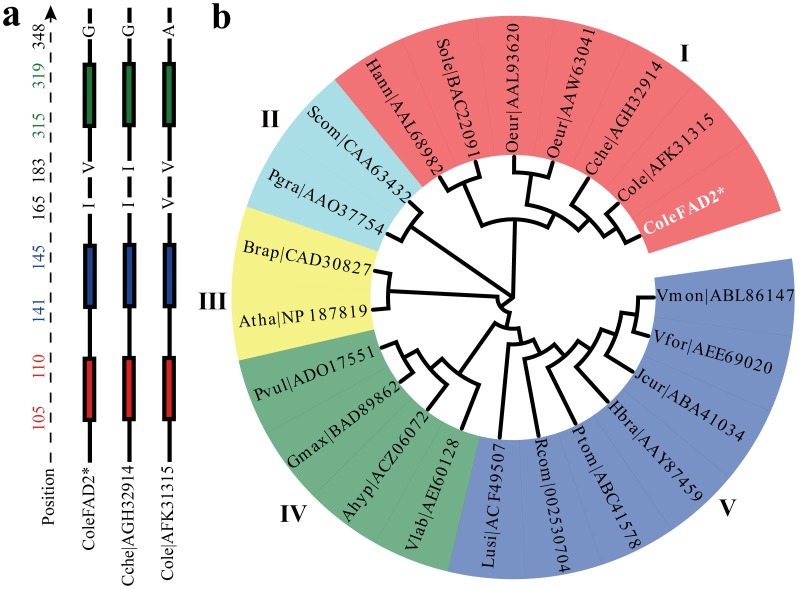
The annotation of the C. oleifera transcriptome also identified a gene of FAD2. The coding sequence (CDS) of this putative FAD2 gene was located in 451–1599 bp of ColeIsotig4522, so-called as ColeFAD2. Besides, two FAD2-like fragments (ColeSingleton23567 and ColeIsotig8486) were also detected in this transcriptome with an average length of 428 bp. Alignment of the deduced polypeptide sequence of the ColeFAD2 together with other two previously cloned FAD2 genes (AFK31315 and AGH32914) displayed extremely high similarity (99.5% and 99.7%, respectively), suggesting that it indeed encodes microsomal oleate desaturase. Despite such high similarities among them, there still existed three inconsistent amino acids ( Figure 8a ). The amino acid Ile (I) at the position 165 of AFK31315 has been converted to a Val (V). In contrast to AGH32914, the amino acid Val (V) at the position 183 has been changed to an Ile (I). And the last inconsistent amino acid appeared in AFK31315 at the position of 348, where the amino acid Gly (G) has been converted to an Ala (A). Similar to the homologous FAD2 genes in other plants, the putative polypeptides of ColeFAD2 also contains three highly conserved histidine-rich motifs (namely, H-boxes; HXXXH, HXXHH, and HXXHH) with conserved spaces between them, which has been shown essential for desaturase activity [41] and responsible for the formation of the diiron-oxygen complex used in biochemical catalysis [42] ( Figure 8a and Figure S1).
Figure 8. Phylogenetic analyses of the FAD2 genes among 20 oil-plants.
(a) The alignment of Cole|AFK31315 (C. oleifera, AFK31315), Cche|AGH32914 (C. chekiangoleosa, AGH32914) and ColeFAD2 (ColeIsotig4522:451–1599) amino acid sequences. The solid black lines indicate conserved amino acids. The filled boxes represent three H-boxes, including HECGH (red box), HRRHH (blue box), and HVAHH (green box). The position (left) is based on FAD2 gene in C. chekiangoleosa (AGH32914). The three inconsistent amino acids were plotted in uppercase letters (black). Multiple sequence alignment was performed using ClustalW [58], [59] package. (b) The amino acid sequences were used for phylogenetic tree analysis. The asterisk indicates the FAD2 gene (ColeFAD2) detected in the assembled C. oleifera transcriptome (ColeIsotig4522:451–1599). I–V represent the five groups of all the 20 oil-plants classified by the sequence similarity. The GenBank accession numbers and the full species names of the genes used here are: Scom|CAA63432 (Solanum commersonii, CAA63432); Atha|NP_187819 (Arabidopsis thaliana, NP_187819); Hann|AAL68982 (Helianthus annuus, AAL68982); Brap|CAD30827 (Brassica rapa, CAD30827); Sole|BAC22091 (Spinacia oleracea, BAC22091); Oeur|AAL93620 (Olea europaea, AAL93620); Pgra|AAO37754 (Punica granatum, AAO37754); Oeur|AAW63041 (Olea europaea, AAW63041); Gmax|BAD89862 (Glycine max, BAD89862); Hbra|AAY87459 (Hevea brasiliensis, AAY87459); Jcur|ABA41034 (Jatropha curcas, ABA41034); Ptom|ABC41578 (Populus tomentosa, ABC41578); Vmon|ABL86147 (Vernicia Montana, ABL86147); Lusi|ACF49507 (Linum usitatissimum, ACF49507); Rcom|002530704 (Ricinus communis, XP_002530704); Ahyp|ACZ06072 (Arachis hypogaea, ACZ06072); Pvul|ADO17551 (Phaseolus vulgaris, ADO17551); Vfor|AEE69020 (Vernicia fordii, AEE69020); Vlab|AEI60128 (Vitis labrusca, AEI60128); Cole|AFK31315 (C. oleifera, AFK31315); Cche|AGH32914 (C. chekiangoleosa, AGH32914).
To investigate phylogenetic relationships of the orthologous FAD2 genes, the deduced amino acid sequence of the identified FAD2 gene (ColeFAD2) was aligned with 21 orthologous FAD2 sequences from 20 oil-plants. Based on the sequence alignments, an unrooted neighbor-joining (NJ) tree was constructed using MEGA software [43] (version 5.05). As illustrated in Figure 8b , these 22 FAD2 genes were mainly classified into five groups. Among these five groups, the three Camellia FAD2 genes and two Olive FAD2 genes, together with each of Spinacia oleracea and Helianthus annuus FAD2 gene, were assigned to Group I, suggesting that they may share analogous functions leading to the similar components of fatty acid, which is regarded as a key factor determining the quality of edible oil. Furthermore, when excluding the two Camellia species from the un-rooted tree, C. oleifera was surprisingly found to be quite close to the Olea europaea but much far from Vernicia montana. This result indicates that parallel evolution may occur between C. oleifera and Olive FAD2 genes, resulting in the creation of analogous structures that have a similar form or function for oil production.
Gene validation and expression analysis
To experimentally confirm that the unigenes obtained from the transcriptome sequencing and assembly were indeed expressed, 17 key unigenes involved in the biosynthesis and metabolism of tea oil were chosen for RT-PCR and qRT-PCR analyses ( Figure 9 ). The detailed information of the selected unigene ID and designed primer pairs in this study were given in Table S8.
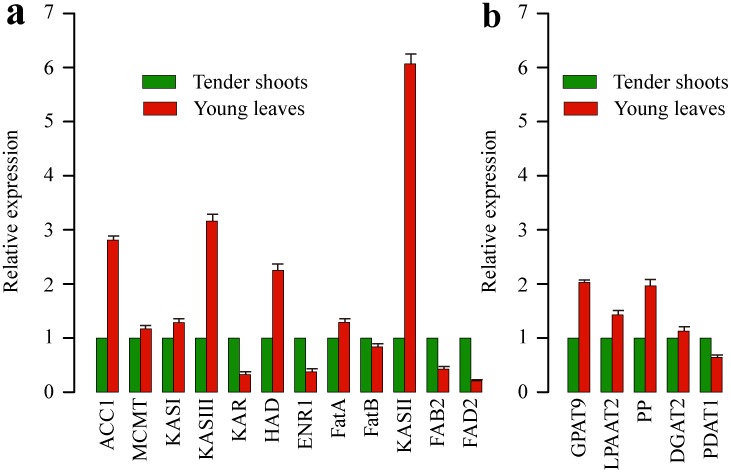
Figure 9. Quantitative RT-PCR validations of the 17 candidate lipid-related genes in the C. oleifera transcriptome.
17 candidate unigenes involved in lipid metabolism including (a) fatty acid and (b) TAG pathways were selected for the quantitative RT-PCR analysis. Standard error of the mean for three biological replicates (nested with three technical replicates) is represented by the error bars. Results represent the mean (± SD) of the three experiments. The translation elongation factor 1-alpha (TEF) gene was chosen as an internal standard.
In the RT-PCR analyses, single bands with the expected sizes were amplified for all 17 unigenes sampled (Table S8), suggesting that the assembled unigenes were reliable and subsequent experiments of gene expression were feasible. In the qRT-PCR analysis, relative expression levels of the selected unigenes were compared between two different tissues. In the fatty acid pathway, unigenes of ACC1, KAS III, HAD and KAS II were expressed much higher in young leaves than in tender shoots ( Figure 9a ), and the highest level of gene expression was observed in KAS II. Expression levels of MCMT, KAS I and FatA in young leaves were slightly higher than those in tender shoots. In comparison, expression levels of KAR, ENR, FatB, FAB2 and FAD2 were low in young leaves but high in tender shoots. For TAG pathway, five selected unigenes had the similar expression pattern except PDAT1 ( Figure 9b ). The overall expression levels of GPAT9, LPAAT2, PP and DGAT2 were higher in young leaves than in tender shoots, while PDAT1 was expressed lower in young leaves but higher in tender shoots. The results of qRT-PCR expression analysis showed that the selected lipid metabolism related unigenes were indeed expressed in the assembled C. oleifera transcriptome and exhibited different expression patterns during the developmental stage of leaves.
Conclusions
Here we sequenced and annotated the transcriptome of C. oleifera. A total of 2,345 SSRs, 20,250 SNPs and 16,906 Indels were detected and can be considered as candidate genetic markers for the future studies. To examine the dynamic evolution of othologous genes between C. sinensis and C. oleifera, we identified 3,022 pairs of orthogous genes, of which 211 were under positive selection with Ks/Ks ratio >1, suggesting that they may probably play a vital role in the evolutionary process of Camellia species. Given the fact that C. oleifera is an important oil-plant in China, thus, in this paper, we mainly focused on the pathways related to lipid metabolism and have successfully identified transcripts associated with fatty acid metabolism, oil accumulation and breakdown in C. oleifera. Moreover, given the economic significance of C. oleifera in the future, it requires further agronomic improvement. To achieve this goal, the genetic and functional genomic resources, especially those genes involved in oil synthesis, accumulation and breakdown as described in this paper, are becoming more important than before. For example, using the genetic engineering technology with genes related to oil synthesis pathway, C. oleifera can be genetically modified to produce transgenic plant with improved oil content and/or composition. Furthermore, due to the importance of gene expression regulation, the entire oil synthesis pathway in C. oleifera can also be engineered in the level of gene expression to increase expression of enzymes responsible for the synthesis of the oleic acid as well as linoleic acids and to decrease expression of enzymes responsible for the breakdown of such compounds. Overall, the C. oleifera transcriptome reported here provides an invaluable new resource for the further genetic engineering and molecular cloning as well as the future functional genomic researches.
Materials and Methods
Plant materials collection, cDNA library construction and 454 sequencing
Four tissues of C. oleifera, including tender shoots, young leaves, flower buds and flowers, were harvested in 2010 from East Park of Kunming Botanic Garden, Yunnan Province, China. All necessary permits were obtained from Wei-bang Sun, who is the director of the Kunming Botanical Garden, the Chinese Academy of Sciences. All samples were flash frozen in liquid nitrogen and stored at −80°C for RNA extraction. Total RNA was extracted by a modified CTAB method. The quality and quantity of total RNA were analyzed using agarose gel electrophoresis and Agilent 2100 Bioanalyzer RNA chip (Agilent Technologies, CA). cDNA library construction and normalization were performed as described previously [44]. All libraries were combined into a single pool and sequenced using the 454 GS-FLX platform (Roche, IN, USA).
Sequence data processing and de novo assembly
The raw reads obtained were first pre-screened to remove adaptors, poly-A tails and contaminants using Seqclean (http://compbio.dfci.harvard.edu/tgi/software/). Low-quality (phred score <20) and short (length <60 bp) reads were trimmed using SolexaQA package [45] (-h 20; -l 60). The trimmed and size-selected reads were then de novo assembled using Newbler software (version 2.8), which performs best for restoring full-length transcripts [44]. Assembled isotigs and singletons were merged and the redundancy among them was removed by CD-HIT (version 4.6) [19] with the similarity threshold of 0.9. The combined and redundancy-removed unigenes were used for the later analysis.
Data deposit
The dataset of 454 sequencing reads were deposited in the NCBI Short Read Archive (SRA) database with the accession number: SRR1472854, SRR1472847, SRR1472843 and SRR1472842. The assembled sequences were available at the NCBI Transcriptome Shotgun Assembly (TSA) database that can be accessed under the accession number GBHI00000000. The version described in this paper is the first version, GBHI01000000.
Unigene functional annotation and pathway assignments
All distinct unigenes (>100 bp) were compared against the NR, Uniref90, TAIR10 and KOG protein databases using the Blastx program with a threshold E-value of 10−5. Gene names were assigned to each unigene according to known genes from the best Blastx hit. The ORF of the unigenes was predicted using ESTscan [46] (version 3.0.3) with a pre-built model for A. thaliana that is distributed with the package, and then searched against PFAM [21] (version 27.0) databases for domain/family annotation using HMMER [47] (version 3.0) with the default parameters. To annotate the unigenes with GO terms, the best Blastx hit from NR database for each transcript was submitted to BLAST2GO platform [22]–[24], and GO terms for each unigene were retrieved based on the relationship between gene names and GO terms. EC number was assigned and parsed based on the BLAST2GO results. To determine metabolic pathways, Kyoto Encyclopedia of Genes and Genomes (KEGG) mapping was used. The sequences with corresponding ECs obtained from Blast2GO were mapped to the KEGG metabolic pathway database. To further enrich the pathway annotation and to identify the BRITE functional hierarchies, assembled sequences were also submitted to the online KEGG Automatic Annotation Server [48] (KAAS; http://www.genome.jp/kegg/kaas/) with bi-directional best hit (BBH) method. The output includes KO assignments and KEGG pathways that are involved with the KO assignments.
Identification of SSRs, SNPs and InDels
Five types of SSRs from di-nucleotides to hexa-nucleotides were identified using the MISA Perl script (http://pgrc.ipk-gatersleben.de/misa/). The minimum repeat unit size for di- nucleotides was set at six, at five for tri- to tetra-nucleotides, and at four for penta- to hexa-nucleotides. Putative SNPs and Indels were detected using ssahaSNP software [49] with the default parameters.
Identification of orthologous genes between C. oleifera and C. sinensis
To identify genes that are putatively orthologous between C. oleifera and C. sinensis, a reciprocal best hit (RBH) method based on the Blastn program was used. Briefly, each sequence from C. oleifera was initially searched against all sequences from C. sinensis using Blastn, and conversely each sequence of C. sinensis was searched against all sequences from C. oleifera. Pairs of sequences that were the best hits each other and sequences longer than 300 bp were regarded as putative orthologs. Due to the limitation of RBH method [50], potential paralogs may not be completely excluded from the ortholog dataset. To remove the possible paralogs, all pairs of sequences were first searched against all plant protein sequences available in GeneBank and only pairs of sequences unambiguously mapped to the same protein with an E-value<1×10−15 were selected as orthologous genes. Similar method and criteria have been used in previous studies [51]–[53].
Estimation of synonymous and non-synonymous substitution rates between orthlogous
To calculate the synonymous (Ks) and non-synonymous (Ka) substitution rate for each orthologous gene pair, each member of a pair of sequences was first searched against all plant protein sequences available in GenBank using Blastx with a significant E-value threshold of 10−15, and the CDS of each orthologous genes was determined based on the best alignment regions. After removing short (<300 bp) and unexpected stop codon(s) containing CDS, the rate of synonymous and non- synonymous substitution was estimated using the maximum likelihood method implemented in codeml program [54] under the F3x4 model [55].
Multiple sequence alignment and phylogenetic analysis of FAD2 genes
Amino acid sequences of the 21 FAD2 genes from 20 oil-plants were downloaded from NCBI GenBank database. The accession numbers and the full species names for these genes were: Scom|CAA63432 (Solanum commersonii, CAA63432); Atha|NP_187819 (Arabidopsis thaliana, NP_187819); Hann|AAL68982 (Helianthus annuus, AAL68982); Brap|CAD30827 (Brassica rapa, CAD30827); Sole|BAC22091 (Spinacia oleracea, BAC22091); Oeur|AAL93620 (Olea europaea, AAL93620); Pgra|AAO37754 (Punica granatum, AAO37754); Oeur|AAW63041 (Olea europaea, AAW63041); Gmax|BAD89862 (Glycine max, BAD89862); Hbra|AAY87459 (Hevea brasiliensis, AAY87459); Jcur|ABA41034 (Jatropha curcas, ABA41034); Ptom|ABC41578 (Populus tomentosa, ABC41578); Vmon|ABL86147 (Vernicia Montana, ABL86147); Lusi|ACF49507 (Linum usitatissimum, ACF49507); Rcom|002530704 (Ricinus communis, XP_002530704); Ahyp|ACZ06072 (Arachis hypogaea, ACZ06072); Pvul|ADO17551 (Phaseolus vulgaris, ADO17551); Vfor|AEE69020 (Vernicia fordii, AEE69020); Vlab|AEI60128 (Vitis labrusca, AEI60128); Cole|AFK31315 (Camellia oleifera, AFK31315); Cche|AGH32914 (Camellia chekiangoleosa, AGH32914). After the sequences were downloaded, the amino acid sequences of the 21 FAD2 genes together with the deduced peptide sequence of ColeFAD2 gene were aligned using MUSCLE [56] (version 3.8.31) with default parameters. Phylogenetic and molecular evolutionary analyses were constructed based on the alignment using MEGA [43] (version 5.05) with the neighbor-joining (NJ) method. Bootstrap values were calculated from 1,000 iterations. Trees were visualized and modified using EvolView [57].
Validation by qRT-PCR
Seventeen unigenes involved in lipid metabolism were selected for the validation using real time qPCR. The gene-specific primer pairs were designed using Primer premier 5.0 software (Premier Biosoft International), and all primer sequences were listed in Table S8. Total RNA was isolated from tender shoots and young leaves of C. oleifera using a modified CTAB method. cDNA was synthesized using the SuperScript VILO cDNA Synthesis Kit (Invitrogen), according to the manufacturer's guidelines. One microgram of RNA was used in each synthesis reaction. Relative mRNA abundance of the selected unigenes was measured using Multicolor Real-Time PCR Detection System (Bio-Rad). The conditions for all reactions were 95°C for 30 s, 40 cycles of 95°C for 15 s, followed by 60°C for 30 s, and 72°C for 20 s. Melting curve analysis was performed by the end of each PCR to confirm the PCR specificity. Three biological replicates of each reaction were performed, and the translation elongation factor 1-alpha (TEF) gene was chosen as an internal standard for normalization. The quantification of qPCR results for each unigene of interest were calculated using the delta-delta Ct (2−ΔΔCt) method. All data were expressed as the mean ± SD after normalization.
Supporting Information
Predicted amino acid sequence of ColeFAD2 (ColeIsotig4522: 451–1599) and alignment with Cole|AFK31315 ( C. oleifera , AFK31315) and Cche|AGH32914 ( C. chekiangoleosa , AGH32914) FAD2 genes. The three point mutations (amino acid: 165, 183 and 348) are show in white background.
(PDF)
Top BLAST hits from NR, UniRef90, TAIR10 and KOG protein databases. BLAST results for all the transcripts with E-value≤10−5 are shown.
(XLS)
Summary of PFAM domains/families in the C. oleifera transcriptome. PFAM search results for all the unigenes with E-value≤10−5 are shown.
(XLS)
Gene Ontology (GO) annotation of the C. oleifera transcripts. Unigenes with the best matches from NR database were further assigned with GO terms using Blast2GO package.
(XLS)
List of SSR motifs detected in the assembled isotigs of C. oleifera . Five types of SSRs were identified from the C. oleifera transcriptome using the MISA Perl script. This list contains the name of unigene, length, SSR number, motif (SSR type), SSR, SSR size, SSR start and SSR end.
(XLS)
Overview of the SNPs detected in the assembled isotigs of C. oleifera . This file contains the detailed information for 20,250 putative SNPs identified from C. oleifera isotigs.
(XLS)
Summary of the 211 orthologous gene pairs ( Ka/Ks ratio >1) between C. oleifera and C. sinensis . The sequence similarity, nonsynonymous substitution rate (Ka), synonymous substitution rate (Ks), omega value (Ka/Ks), and NR annotations are shown.
(XLS)
Detailed information of the lipid metabolism-related genes identified in the C. oleifera transcriptome. The pathway information, family name, family abbreviation, EC number, isoform/gene specific name, isoform/gene specific abbreviation, subcellular location, TAIR10 gene locus, number of unigenes, average unigene length and the corresponding unigene IDs are given.
(XLS)
Primer pairs of the candidate lipid-related unigenes designed for qRT-PCR. Specific primer pairs of seventeen candidate unigenes with potential roles in lipid metabolism designed for real time qRT-PCR using the Primer premier software (version 5.0) are shown.
(XLS)
Acknowledgments
We would like to thank the anonymous reviewers for his/her thorough review and highly appreciate the valuable comments and suggestions, which significantly contributed to improving the quality of the paper.
Data Availability
The authors confirm that all data underlying the findings are fully available without restriction. We have deposited the dataset of 454 sequencing reads in the NCBI Short Read Archive (SRA) database that can be accessed under the accession number: SRR1472842, SRR1472843, SRR1472847 and SRR1472854, respectively. The assembled sequences have been deposited at the NCBI Transcriptome Shotgun Assembly (TSA) database under the accession GBHI00000000. The version described in this paper is the first version, GBHI01000000.
Funding Statement
This work was supported by National Science Foundation of China (U0936603), Key Project of Natural Science Foundation of Yunnan Province (2008CC016), Top Talent Program of Yunnan Province (20080A009), Frontier Grant of Kunming Institute of Botany, CAS (672705232515), the Hundred Talents Program of the Chinese Academy of Sciences (CAS), a grant from the Chinese Academy of Science (KSCX2-YW-N-029), and a start-up grant from Kunming Institute of Botany, CAS to L.-Z. Gao. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Ohlrogge JB (1994) Design of new plant products: engineering of fatty acid metabolism. Plant Physiol 104: 821–826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Yu Y, Ren S, Tan K (1999) Study on climatic regionalization and layer and belt distribution of oiltea camellia quality in China. J Nat Res 14: 123–127. [Google Scholar]
- 3. Shanan H, Ying G (1982) The comprehensive utilization of camellia fruits. Am Camellia Yearbk 37: 104–107. [Google Scholar]
- 4.Stack L, Ruter J (2006) Teaoil Camellia-Eastern “Olive” for the world. In: XXVII International Horticultural Congress-IHC2006: International Symposium on Asian Plants with Unique Horticultural 769: . pp. 43–48. [Google Scholar]
- 5. Xia L, Zhang A, Xiao T (1993) An introduction to the utilization of camellia oil in China. Am Camellia Yearbk 48: 12–15. [Google Scholar]
- 6. Gao J (1993) The importance of camellias as oil plants in China. International Camellia Journal 25: 53–54. [Google Scholar]
- 7. Chen Y-H (2007) Physiochemical properties and bioactivities of tea seed (Camellia oleifera) oil. Clemson University [Google Scholar]
- 8. Hudson ME (2008) Sequencing breakthroughs for genomic ecology and evolutionary biology. Molecular Ecology Resources 8: 3–17. [DOI] [PubMed] [Google Scholar]
- 9. Metzker ML (2009) Sequencing technologies—the next generation. Nature Reviews Genetics 11: 31–46. [DOI] [PubMed] [Google Scholar]
- 10. Shendure J, Porreca GJ, Reppas NB, Lin X, McCutcheon JP, et al. (2005) Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309: 1728–1732. [DOI] [PubMed] [Google Scholar]
- 11. Wall PK, Leebens-Mack J, Chanderbali AS, Barakat A, Wolcott E, et al. (2009) Comparison of next generation sequencing technologies for transcriptome characterization. BMC Genomics 10: 347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Cheung F, Haas BJ, Goldberg SMD, May GD, Xiao YL, et al. (2006) Sequencing Medicago truncatula expressed sequenced tags using 454 Life Sciences technology. BMC Genomics 7: 272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Emrich SJ, Barbazuk WB, Li L, Schnable PS (2007) Gene discovery and annotation using LCM-454 transcriptome sequencing. Genome Research 17: 69–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Bellin D, Ferrarini A, Chimento A, Kaiser O, Levenkova N, et al. (2009) Combining next-generation pyrosequencing with microarray for large scale expression analysis in non-model species. BMC Genomics 10: 555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Barbazuk WB, Emrich SJ, Chen HD, Li L, Schnable PS (2007) SNP discovery via 454 transcriptome sequencing. Plant J 51: 910–918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Blanca J, Canizares J, Roig C, Ziarsolo P, Nuez F, et al. (2011) Transcriptome characterization and high throughput SSRs and SNPs discovery in Cucurbita pepo (Cucurbitaceae). BMC Genomics 12: 104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Trick M, Long Y, Meng JL, Bancroft I (2009) Single nucleotide polymorphism (SNP) discovery in the polyploid Brassica napus using Solexa transcriptome sequencing. Plant Biotechnology Journal 7: 334–346. [DOI] [PubMed] [Google Scholar]
- 18. Hou R, Bao ZM, Wang S, Su HL, Li Y, et al. (2011) Transcriptome Sequencing and De Novo Analysis for Yesso Scallop (Patinopecten yessoensis) Using 454 GS FLX. Plos One 6: e21560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Li W, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22: 1658–1659. [DOI] [PubMed] [Google Scholar]
- 20. Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 3389–3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, et al. (2012) The Pfam protein families database. Nucleic Acids Res 40: D290–301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Conesa A, Gotz S (2008) Blast2GO: A comprehensive suite for functional analysis in plant genomics. Int J Plant Genomics 2008: 619832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, et al. (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21: 3674–3676. [DOI] [PubMed] [Google Scholar]
- 24. Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, et al. (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic acids research 36: 3420–3435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Kong Q, Xiang C, Yu Z (2006) Development of EST-SSRs in Cucumis sativus from sequence database. Molecular Ecology Notes 6: 1234–1236. [Google Scholar]
- 26. Shi J, Dai XG, Chen YN, Chen JH, Shi JS, et al. (2013) Discovery and experimental analysis of microsatellites in an oil woody plant Camellia chekiangoleosa . Plant Systematics and Evolution 299: 1387–1393. [Google Scholar]
- 27. Thiel T, Michalek W, Varshney RK, Graner A (2003) Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor Appl Genet 106: 411–422. [DOI] [PubMed] [Google Scholar]
- 28. Wang XC, Guo L, Shangguan LF, Wang C, Yang G, et al. (2012) Analysis of expressed sequence tags from grapevine flower and fruit and development of simple sequence repeat markers. Mol Biol Rep 39: 6825–6834. [DOI] [PubMed] [Google Scholar]
- 29. Varshney RK, Thiel T, Stein N, Langridge P, Graner A (2002) In silico analysis on frequency and distribution of microsatellites in ESTs of some cereal species. Cellular & Molecular Biology Letters 7: 537–546. [PubMed] [Google Scholar]
- 30. Sharma RK, Bhardwaj P, Negi R, Mohapatra T, Ahuja PS (2009) Identification, characterization and utilization of unigene derived microsatellite markers in tea (Camellia sinensis L.). BMC Plant Biol 9: 53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Wen Q, Xu L, Gu Y, Huang M, Xu L (2012) Development of polymorphic microsatellite markers in Camellia chekiangoleosa (Theaceae) using 454-ESTs. Am J Bot 99: e203–205. [DOI] [PubMed] [Google Scholar]
- 32. Wu HL, Chen D, Li JX, Yu B, Qiao XY, et al. (2013) De Novo Characterization of Leaf Transcriptome Using 454 Sequencing and Development of EST-SSR Markers in Tea (Camellia sinensis). Plant Molecular Biology Reporter 31: 524–538. [Google Scholar]
- 33. Shi CY, Yang H, Wei CL, Yu O, Zhang ZZ, et al. (2011) Deep sequencing of the Camellia sinensis transcriptome revealed candidate genes for major metabolic pathways of tea-specific compounds. BMC Genomics 12: 131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Ma JL, Ye H, Rui YK, Chen GC, Zhang NY (2011) Fatty acid composition of Camellia oleifera oil. Journal Fur Verbraucherschutz Und Lebensmittelsicherheit 6: 9–12. [Google Scholar]
- 35. Drøbak BK, Heras B (2002) Nuclear phosphoinositides could bring FYVE alive. Trends in Plant Science 7: 132–138. [DOI] [PubMed] [Google Scholar]
- 36. Feussner I, Wasternack C (2002) The lipoxygenase pathway. Annu Rev Plant Biol 53: 275–297. [DOI] [PubMed] [Google Scholar]
- 37. Murphy DJ (1990) Storage lipid bodies in plants and other organisms. Prog Lipid Res 29: 299–324. [PubMed] [Google Scholar]
- 38. Costa GG, Cardoso KC, Del Bem LE, Lima AC, Cunha MA, et al. (2010) Transcriptome analysis of the oil-rich seed of the bioenergy crop Jatropha curcas L. BMC Genomics 11: 462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Rismani-Yazdi H, Haznedaroglu BZ, Bibby K, Peccia J (2011) Transcriptome sequencing and annotation of the microalgae Dunaliella tertiolecta: pathway description and gene discovery for production of next-generation biofuels. BMC Genomics 12: 148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Dugail I, Hajduch E (2007) A new look at adipocyte lipid droplets: towards a role in the sensing of triacylglycerol stores? Cellular and Molecular Life Sciences 64: 2452–2458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Shanklin J, Whittle E, Fox BG (1994) Eight histidine residues are catalytically essential in a membrane-associated iron enzyme, stearoyl-CoA desaturase, and are conserved in alkane hydroxylase and xylene monooxygenase. Biochemistry 33: 12787–12794. [DOI] [PubMed] [Google Scholar]
- 42. Broun P, Shanklin J, Whittle E, Somerville C (1998) Catalytic plasticity of fatty acid modification enzymes underlying chemical diversity of plant lipids. Science 282: 1315–1317. [DOI] [PubMed] [Google Scholar]
- 43. Tamura K, Peterson D, Peterson N, Stecher G, Nei M, et al. (2011) MEGA5: Molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Molecular Biology and Evolution 28: 2731–2739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Niu SH, Li ZX, Yuan HW, Chen XY, Li Y, et al. (2013) Transcriptome characterisation of Pinus tabuliformis and evolution of genes in the Pinus phylogeny. BMC Genomics 14: 263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Cox MP, Peterson DA, Biggs PJ (2010) SolexaQA: At-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinformatics 11: 485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Iseli C, Jongeneel CV, Bucher P (1999) ESTScan: a program for detecting, evaluating, and reconstructing potential coding regions in EST sequences. Proc Int Conf Intell Syst Mol Biol 138–147. [PubMed] [Google Scholar]
- 47. Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39: W29–37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M (2007) KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35: W182–185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. van Oeveren J, Janssen A (2009) Mining SNPs from DNA sequence data; computational approaches to SNP discovery and analysis. Methods Mol Biol 578: 73–91. [DOI] [PubMed] [Google Scholar]
- 50. Dalquen DA, Dessimoz C (2013) Bidirectional Best Hits Miss Many Orthologs in Duplication-Rich Clades such as Plants and Animals. Genome Biology and Evolution 5: 1800–1806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Wang JT, Li JT, Zhang XF, Sun XW (2012) Transcriptome analysis reveals the time of the fourth round of genome duplication in common carp (Cyprinus carpio). BMC Genomics 13: 96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Wang XW, Zhao QY, Luan JB, Wang YJ, Yan GH, et al. (2012) Analysis of a native whitefly transcriptome and its sequence divergence with two invasive whitefly species. BMC Genomics 13: 529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Zhang L, Yan HF, Wu W, Yu H, Ge XJ (2013) Comparative transcriptome analysis and marker development of two closely related Primrose species (Primula poissonii and Primula wilsonii). BMC Genomics 14: 329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24: 1586–1591. [DOI] [PubMed] [Google Scholar]
- 55. Goldman N, Yang Z (1994) A codon-based model of nucleotide substitution for protein-coding DNA sequences. Mol Biol Evol 11: 725–736. [DOI] [PubMed] [Google Scholar]
- 56. Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Zhang H, Gao S, Lercher MJ, Hu S, Chen WH (2012) EvolView, an online tool for visualizing, annotating and managing phylogenetic trees. Nucleic Acids Res 40: W569–572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, et al. (2003) Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res 31: 3497–3500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, et al. (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23: 2947–2948. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Predicted amino acid sequence of ColeFAD2 (ColeIsotig4522: 451–1599) and alignment with Cole|AFK31315 ( C. oleifera , AFK31315) and Cche|AGH32914 ( C. chekiangoleosa , AGH32914) FAD2 genes. The three point mutations (amino acid: 165, 183 and 348) are show in white background.
(PDF)
Top BLAST hits from NR, UniRef90, TAIR10 and KOG protein databases. BLAST results for all the transcripts with E-value≤10−5 are shown.
(XLS)
Summary of PFAM domains/families in the C. oleifera transcriptome. PFAM search results for all the unigenes with E-value≤10−5 are shown.
(XLS)
Gene Ontology (GO) annotation of the C. oleifera transcripts. Unigenes with the best matches from NR database were further assigned with GO terms using Blast2GO package.
(XLS)
List of SSR motifs detected in the assembled isotigs of C. oleifera . Five types of SSRs were identified from the C. oleifera transcriptome using the MISA Perl script. This list contains the name of unigene, length, SSR number, motif (SSR type), SSR, SSR size, SSR start and SSR end.
(XLS)
Overview of the SNPs detected in the assembled isotigs of C. oleifera . This file contains the detailed information for 20,250 putative SNPs identified from C. oleifera isotigs.
(XLS)
Summary of the 211 orthologous gene pairs ( Ka/Ks ratio >1) between C. oleifera and C. sinensis . The sequence similarity, nonsynonymous substitution rate (Ka), synonymous substitution rate (Ks), omega value (Ka/Ks), and NR annotations are shown.
(XLS)
Detailed information of the lipid metabolism-related genes identified in the C. oleifera transcriptome. The pathway information, family name, family abbreviation, EC number, isoform/gene specific name, isoform/gene specific abbreviation, subcellular location, TAIR10 gene locus, number of unigenes, average unigene length and the corresponding unigene IDs are given.
(XLS)
Primer pairs of the candidate lipid-related unigenes designed for qRT-PCR. Specific primer pairs of seventeen candidate unigenes with potential roles in lipid metabolism designed for real time qRT-PCR using the Primer premier software (version 5.0) are shown.
(XLS)
Data Availability Statement
The authors confirm that all data underlying the findings are fully available without restriction. We have deposited the dataset of 454 sequencing reads in the NCBI Short Read Archive (SRA) database that can be accessed under the accession number: SRR1472842, SRR1472843, SRR1472847 and SRR1472854, respectively. The assembled sequences have been deposited at the NCBI Transcriptome Shotgun Assembly (TSA) database under the accession GBHI00000000. The version described in this paper is the first version, GBHI01000000.