Abstract
Leucine-rich repeat kinase 2 (LRRK2), implicated in familial Parkinson’s disease (PD), was recently identified as a major susceptibility gene for Crohn’s disease (CD) by genome-wide association studies (GWAS). We found that LRRK2 deficiency confers enhanced susceptibility to experimental colitis in mice. Mechanistic studies showed that LRRK2 is a potent negative regulator of NFAT and a component of a previously described RNA-protein complex involving a non-coding RNA repressor of NFAT (NRON). Colitis in LRRK2 deficient mice is exacerbated by enhanced NFAT1 nuclear localization. Moreover, the risk-associated allele Met2397 identified in CD GWAS causes reduced LRRK2 protein expression, which, in light of our unexpected observation that LRRK2 is a negative regulator of NFAT, suggests a pathological mechanism important in human disease.
Leucine-rich repeat kinase 2 (LRRK2), also known as Dardarin, is the most frequently mutated gene in autosomal dominant familial Parkinson’s disease (PD), although the physiological and pathological functions of LRRK2 are not well understood (1, 2). Recently genome-wide association studies (GWAS) have indicated that LRRK2 is also one of the most significant of the 30 genetic susceptibility loci for Crohn’s disease (CD) (3). CD, a common form of inflammatory bowel disease (IBD), is generally thought to develop from a dysregulated immune response to gut luminal biota. There are both genetic and environmental contributions to the development of CD (4). LRRK2 expression is inducible by IFN-γ, which is consistent with the notion that it may play a role in IBD (5). However, the molecular mechanism of this association is not clear.
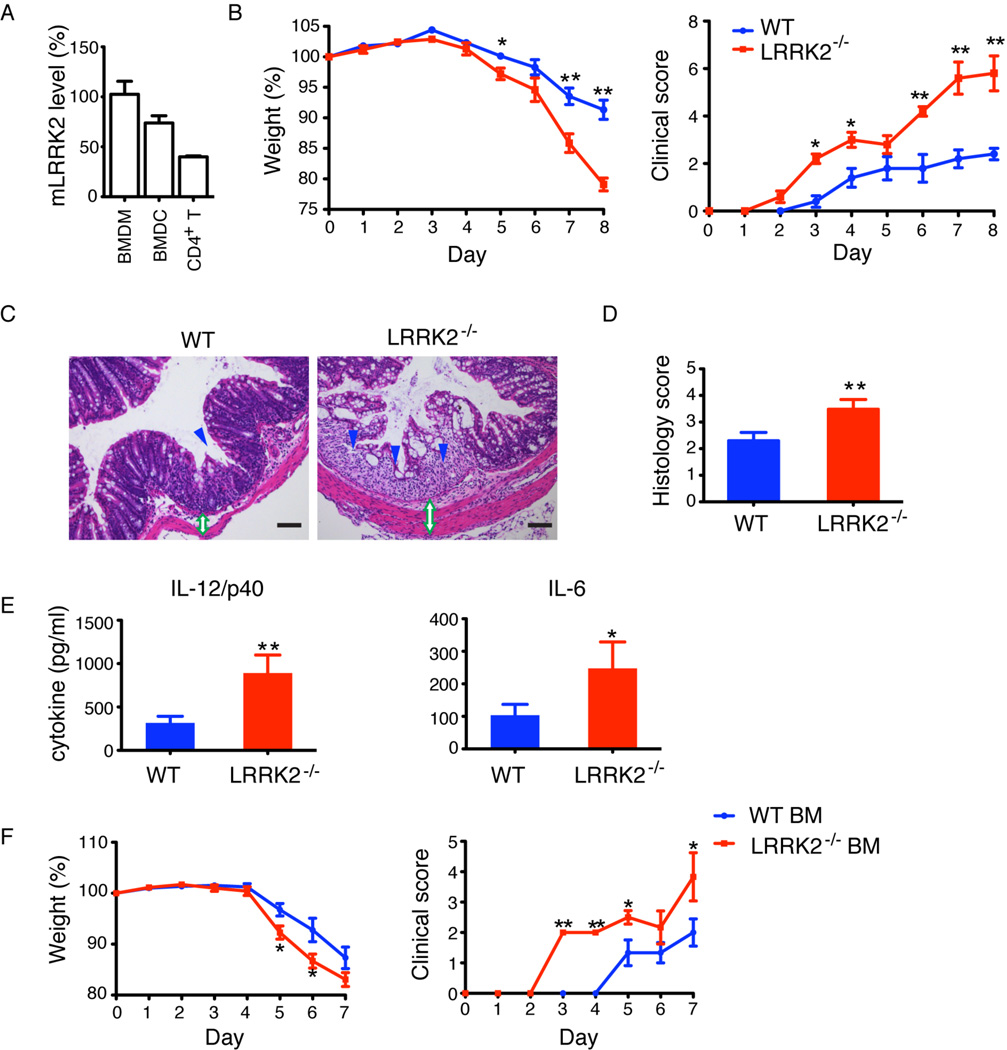
We began our investigation of LRRK2 using a murine model of IBD with LRRK2 deficient (LRRK2−/−) mice. LRRK2−/− mice on a C57BL/6 background do not develop spontaneous IBD and have a normal gastrointestinal tract (Fig. S1). Dextran sodium sulfate (DSS)-induced colitis, a widely used model for IBD that reflects the contribution of the innate immune system in response to acute mucosal damage, depends mainly on the action of the myeloid lineage cells, i.e. macrophages (6). LRRK2 is highly expressed in bone marrow-derived macrophages (BMDM) and dendritic cells (BMDCs) compared to CD4+ T cells (7) (Fig. 1A & Fig. S2), so we examined DSS-induced colitis in LRRK2−/− mice compared to wild type (WT) C57BL/6 mice. These studies revealed that LRRK2−/− mice developed more severe clinical symptoms of IBD including rapid weight loss, severe diarrhea, and bloody stools (Fig. 1B). Similar effects were observed using 5% DSS treatment (Fig. S3). Histological analysis of the colons revealed massive inflammatory infiltrates, thickened walls, and disruption of mucosal structures in LRRK2−/− mice (Fig. 1C, D). The production of the inflammatory cytokines, IL-12 (p40 subunit) and IL-6, was elevated in sera from DSS-treated LRRK2−/− mice (Fig. 1E). Thus, LRRK2 deficiency exacerbates inflammation in DSS treated mice.
Fig. 1.
LRRK2 deficiency exacerbates experimental colitis in mice. (A) Relative levels of LRRK2 mRNA measured by qRT-PCR in bone-marrow derived macrophages (BMDM)(set as 100%), bone-marrow derived dendritic cells (BMDC) and CD4+ T cells. β-actin was used for total mRNA normalization. (B) Mean body weight as a percent of starting weight (left panel) and mean clinical scores (right panel) of WT (blue, n=5) and LRRK2−/− (red, n=5) mice treated with 3% DSS. *P < 0.05. **P < 0.02. (C) Hematoxylin and eosin (H&E) staining of colonic sections. Loss of epithelial crypts (blue arrowhead), transmural inflammation and thickened intestinal wall (green open double arrow) are noted. Scale bar, 200 µm. (D) Mean of histological scores of colon sections. **P < 0.02. (E) Serum cytokines at Day 8. *P < 0.05, **P < 0.02. (F) Mean body weight (left panel) and mean clinical scores (right panel) of WT mice that received either WT or LRRK2−/− bone marrow (BM) (n=6). Data are representative of three independent experiments (A–E) and two independent experiments (F). Error bars represent standard errors of the mean (SEM).
To determine whether the exacerbation of colitis in LRRK2−/− mice was attributable to hematopoietic cells, we reconstituted lethally irradiated WT mice with bone marrow cells either from WT or LRRK2−/− mice and studied DSS-induced colitis in the chimeric mice. We found that mice receiving LRRK2−/− bone marrow cells developed more rapid and more severe colitis, including weight loss, diarrhea, and bloody stools, as compared to those that received WT bone marrow (Fig. 1F). These data suggest that it is LRRK2 deficiency in a cell intrinsic manner in hematopoietic cells that leads to exacerbated colitis, consistent with the observation that LRRK2 is not expressed in intestinal epithelial cells (5). These results prompted us to examine the molecular mechanism by which LRRK2 regulates macrophages which express high levels of this protein.
LRRK2 is a 2527 amino acid protein (286 kD), containing several functional domains, including leucine-rich repeats (LRR), a Ras of complex proteins (Roc) domain, a C-terminal of Roc (COR) domain, a kinase domain, and a WD40 repeat (WD) domain (Fig. 2A). We conducted an in silico search for dLRRK (DRSC15816), the Drosophila ortholog of LRRK2, in the RNAi phenotype database at http://flyrnai.org/ and found that dLRRK scored as a weak hit in a screen for regulators of Nuclear Factor of Activated T cells (NFAT) nuclear translocation, but had no role in other signaling pathways such as Jak-Stat, MAPK, or Wnt-Wingless (8). NFAT was originally identified as a transcription factor in T lymphocytes regulating interleukin-2 (IL-2) transcription in response to Ca2+ influx triggered by antigen recognition (9). Recent studies have found that NFAT regulates innate immune responses in macrophages, dendritic cells, and neutrophils, in addition to having well-recognized roles in T cell cytokine production, neuronal differentiation, stem cell maintenance, and cardiac development (10–16).
Fig. 2.
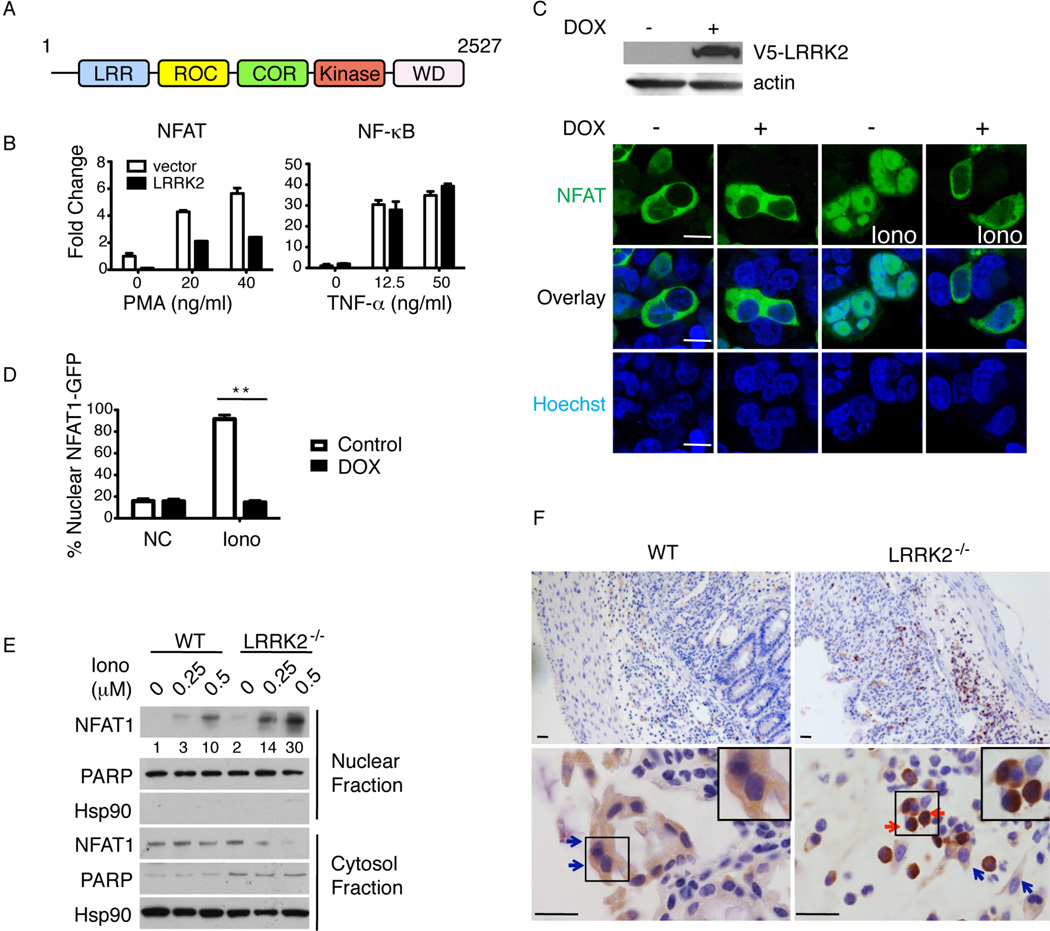
LRRK2 inhibits NFAT1 nuclear function. (A) Schematic of LRRK2 structure. (B) NFAT- and NF-κB-induced luciferase activity in 293T cells transfected with vector or LRRK2 stimulated with 0.5 µM ionomycin and phorbol myristate acetate (PMA) or TNF-α at indicated doses. Data are means of triplicate wells with SEM. (C) Immunoblotting of V5-LRRK2 and actin from 293T cells with a tet-regulated LRRK2 expression construct uninduced (−) or induced with 5 µg ml−1 doxycycline (DOX)(+), top panels. Confocal images of NFAT1-GFP transfected into 293T cells with DOX-inducible V5-LRRK2 before and after 1 µM ionomycin (Iono), Lower panels. Hoechst nuclear staining (blue). Scale bar, 5 µm. (D) Percentage of cells with nuclear NFAT1-GFP among ~100 GFP-positive cells for each condition in (C). Error bars indicate SEM for 10 fields in three experiments. **P < 0.01. (E) Representative immunoblotting of NFAT1 in nuclear vs. cytosolic fractions from BMDM treated with Iono at the indicated doses (µM) for 30’. Numbers below the NFAT1 blot are the quantified relative ratio of NFAT1 in nuclear fractions. (F) Representative images of NFAT1 immunohistochemical staining in colon sections. Cells with cytosolic NFAT1 staining (blue arrows) and nuclear NFAT1 staining (red arrows) are noted. Inset panels, enlarged cells. Scale bar, 25 µm. B–F are representative of three experiments.
The siRNA screen implied that LRRK2 could suppress NFAT activity. Therefore, we first tested the effect of overexpressing LRRK2 on NFAT activation in human embryonic kidney (HEK) 293T cells, which are normally devoid of LRRK2. Using functional luciferase reporter assays, we found that LRRK2 specifically inhibited NFAT- but not NF-κB-dependent transcriptional activity (Fig. 2B). We stably transfected HEK 293T cells, with a DNA construct encoding LRRK2 under the control of a doxycycline (dox)-regulated promoter. As expected, DOX treatment resulted in substantial expression of LRRK2 in these cells (Fig. 2C, top panels). Ectopic expression of an NFAT1-GFP fusion protein allowed us to microscopically observe translocation of NFAT1 from the cytosol to the nucleus in response to ionomycin (iono)-induced Ca2+ flux in these cells (Fig. 2C, bottom panels). We found that overexpressed LRRK2 caused nearly complete retention of NFAT1-GFP in the cytosol despite Iono stimulation (Fig. 2C, D). LRRK2 inhibition of NFAT1-GFP nuclear translocation was also observed in fractionated lysates from 293T cells transiently overexpressing NFAT1-GFP with or without myc-LRRK2 (Fig. S4A). Taken together, these data show that LRRK2 suppresses NFAT1 transcriptional activity by preventing its nuclear translocation. A similar inhibitory effect was observed for other members of the NFAT family, including NFAT2, NFAT3, and NFAT4 (Fig. S4B–D). To evaluate whether endogenous LRRK2 regulates NFAT1, we treated bone marrow derived macrophages (BMDM) from WT and LRRK2−/− mice with different concentrations of Iono, and analyzed by NFAT1 nuclear translocation by fractionation and immunoblotting. We found that LRRK2 deficiency led to enhanced NFAT1 nuclear translocation following Iono treatment (Fig. 2E). These data led us to further investigate whether NFAT1 translocation was altered in the DSS-colitis model. By immunohistochemical staining, we observed marked enhanced NFAT1 nuclear localization in colon sections from LRRK2−/− mice but not WT mice treated with DSS (Fig. 2F). These data indicate that LRRK2 normally helps to retain NFAT1 in the cytoplasm.
We further explored the molecular mechanism of NFAT1 regulation by LRRK2. To examine whether the kinase activity of LRRK2 was required for this activity, we measured NFAT-driven luciferase activity in 293T cells transfected with WT or kinase-dead (KD) LRRK2 (17). Both WT and KD LRRK2 displayed similar inhibitory effects on NFAT activity as assessed using a luciferase reporter (Fig. 3A), suggesting a kinase-independent mechanism. NFAT1 nuclear translocation and activity is thought to be controlled by phosphorylation on a series of serine residues in its N-terminal regulatory domain (9). Different phosphorylation states of NFAT1 can be easily distinguished by their relative mobility shifts on SDS-polyacrylamide gels (18). We treated cells with a “pulse” of Iono, leading to a faster migrating NFAT1 band, and then washed away the Iono to allow rephosphorylation of NFAT1, causing it to migrate more slowly. Overexpression of myc-LRRK2 did not appreciably alter the dephosphorylation and rephosphorylation of ectopically expressed NFAT1-GFP in 293T cells (Fig. 3B). We next investigated whether LRRK2 could block the predominant nuclear localization of a constitutively active form of NFAT1 (caNFAT1) in which the serine residues in the regulatory domain are changed to alanine (19). We transfected 293T cells stably expressing dox-inducible V5-LRRK2 with caNFAT1-GFP and found by confocal microscopy that V5-LRRK2 quite effectively impeded nuclear translocation of caNFAT1-GFP implying that LRRK2 controlled NFAT nuclear translocation directly (Fig. 3C, D). The inhibitory effect on caNFAT1 was also confirmed in cytosolic and nuclear fractions from cells transiently overexpressing myc-LRRK2 (Fig. 3E). Taken together, LRRK2 appears to constrain nuclear translocation of NFAT1 by a dominant negative regulatory mechanism independent of NFAT1 phosphorylation.
Fig. 3.
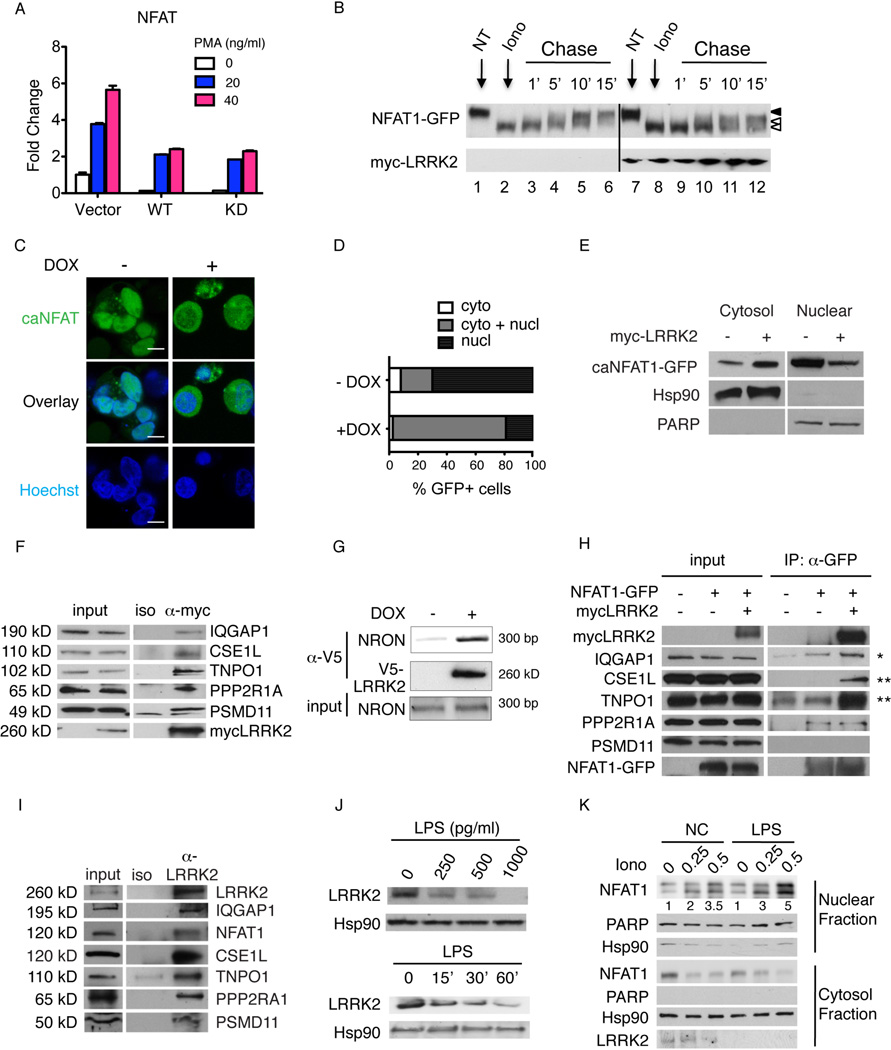
LRRK2 affects NFAT1 cytoplasmic sequestration not phosphorylation. (A) NFAT-induced luciferase activity (mean ± SEM) in 293T cells transfected with vector or LRRK2 WT or kinase dead (KD) variants stimulated with 0.5 µM Iono and indicated doses of PMA. (B) Immunoblotting of NFAT1-GFP from 293T cells transfected with vector control or myc-LRRK2 at the indicated time points after washing out Iono treatment. NT, untreated. Open arrowheads indicate unphosphorylated NFAT1-GFP, and closed arrowhead marks highly phosphophorylated NFAT1-GFP. (C) Confocal images of caNFAT1-GFP transfected into 293T cells with DOX-inducible V5-LRRK2. Hoechst nuclear staining (blue). Scale bar, 5 µm. (D) Percent of cells with predominantly cytoplasmic (cyto), cytoplasmic and nuclear (cyto + nucl), or nuclear (nucl) distribution of caNFAT1-GFP among 100 GFP-positive cells for each condition. (E) Immunoblotting of caNFAT1-GFP in cytosolic vs nuclear fractions from 293T cells transfected with vector control or myc-LRRK2. (F) Immunoprecipitation and immunoblotting of proteins as indicated, using 293T overexpressing myc-LRRK2 as bait. Iso, isotype control Ig for immunoprecipitation. (G) Immunoprecipitation/RT-PCR of NRON using 293T cells with DOX-inducible V5-LRRK2. (H) Immunoprecipitation/immunoblotting for proteins as indicated in 293T cells. *small change; **large change. (I) Immunoprecipitation/immunoblotting, as indicated, using THP1 cells. (J) Immunoblotting of LRRK2 from BMDM treated with lipopolysaccharide (LPS) at the indicated doses for 1 hour (top panels) or at the indicated times (min) (bottom panels) with 1000 pg ml−1 LPS; (K) Immunoblotting of NFAT1 in nuclear vs cytosolic fractions from LPS-primed (1000 pg ml−1 for 1 hour) or untreated (NC) BMDM cells treated with Iono at the indicated doses (µM) for 30 min. A–K represent three experiments.
We reasoned that such LRRK2-mediated inhibition of NFAT1 nuclear localization could be connected to a previously described protein-RNA complex containing a large noncoding RNA, NRON, and 11 proteins involved in sequestering NFAT in the cytoplasm (20). To examine whether LRRK2 was physically associated with the protein-NRON complex, myc-LRRK2 was overexpressed in 293T cells and immunoprecipitated using an anti-myc antibody. Five of the 11 proteins comprising the NRON complex, including IQGAP1, CSE1L, TNPO1, PPP2RA, and PSMD11, coimmunoprecipitated with myc-LRRK2 (Fig. 3F, Fig. S5). The regulatory non-coding RNA, NRON, was also detected in association with LRRK2 by immunoprecipitation followed by RT-PCR using NRON specific primers (Fig. 3G). Moreover, we noted that LRRK2 overexpression enhanced the association of NFAT1 with IQGAP1, CSE1L and TNPO1 upon direct NFAT immunoprecipitation, while the proportion of bound PPP2R1A remained unchanged (Fig. 3H). We observed no coimmunoprecipitation between NFAT1-GFP and PSMD11 with or without myc-LRRK2 (Fig. 3H). We also validated these NRON complex protein interactions occurred with endogenous LRRK2 in human monocyte THP-1 cells (Fig. 3I).
Our data show that LRRK2 deficiency leads to exacerbated colonic inflammatory responses to DSS and that this molecule physically interacts with the NRON complex and retains NFAT1 in cytosol independently of phosphorylation, possibly due to this association. These findings suggested that LRRK2 might act as a dynamic regulator of NFAT1 nuclear localization and that changes in LRRK2 level in response to extrinsic signals might play a regulatory role in immune responses. We were therefore interested in what may regulate LRRK2 protein levels, especially at the posttranslational level. Lipopolysaccharide (LPS) is a common stimulus used to activate macrophages that can not activate NFAT1 by itself (21). We treated BMDM cells with different doses of LPS and observed a substantial reduction of LRRK2 level within one hour of treatment (Fig. 3J). Furthermore, we found that pretreatment of BMDM cells with LPS enhanced NFAT1 nuclear translocation induced by Iono (Fig. 3K). These data suggest that LRRK2 protein levels are actively regulated in BMDMs and that its degradation is important for the level of NFAT1 activation.
We next investigated the functional effect of LRRK2 on NFAT1 inhibition in BMDM cells by examining pro-inflammatory cytokine production following exposure to various stimulators of innate immunity (Fig. 4A, Fig. S6). Zymosan, a cell wall constituent of Candida, commonly found in gut luminal flora, activates NFAT1-dependent cytokine production in macrophages through a Dectin-1-dependent, TLR2-independent mechanism (22). We therefore examined cytokine production by zymosan in either WT or LRRK2−/− BMDM cells. For comparison, we also stimulated BMDM cells with another TLR2 agonist Pam3CSK4 that does not activate NFAT1 in macrophages. We found that LRRK2−/− BMDMs secreted more IL-12 (p40 subunit) and IL-6 in response to zymosan but not to Pam3CSK4, as compared to WT BMDMs. In contrast, cytokine production (IL-12 and IL-6) was similar between WT and LRRK2−/− BMDMs treated with other stimulators (Fig. S5A–B). In addition, LRRK2 deficiency did not affect the production of TNF-α or the inflammasome-dependent release of IL-1β (Fig. 4A). The calcineurin inhibitor FK506 greatly reduced the production of IL-12/p40 and IL-6 in WT and LRRK2−/− BMDMs (Fig. 4B), suggesting that IL-12/p40 and IL-6 production in response to zymosan could be NFAT-dependent. Indeed, immunoblotting of nuclear and cytosolic fractions (Fig. 4C) revealed greater nuclear accumulation of NFAT1 in LRRK2−/− BMDMs following zymosan treatment relative to WT cells. By contrast, LRRK2 deficiency did not significantly alter NF-κB and MAP kinase signaling cascades (Fig. S7).
Fig. 4.
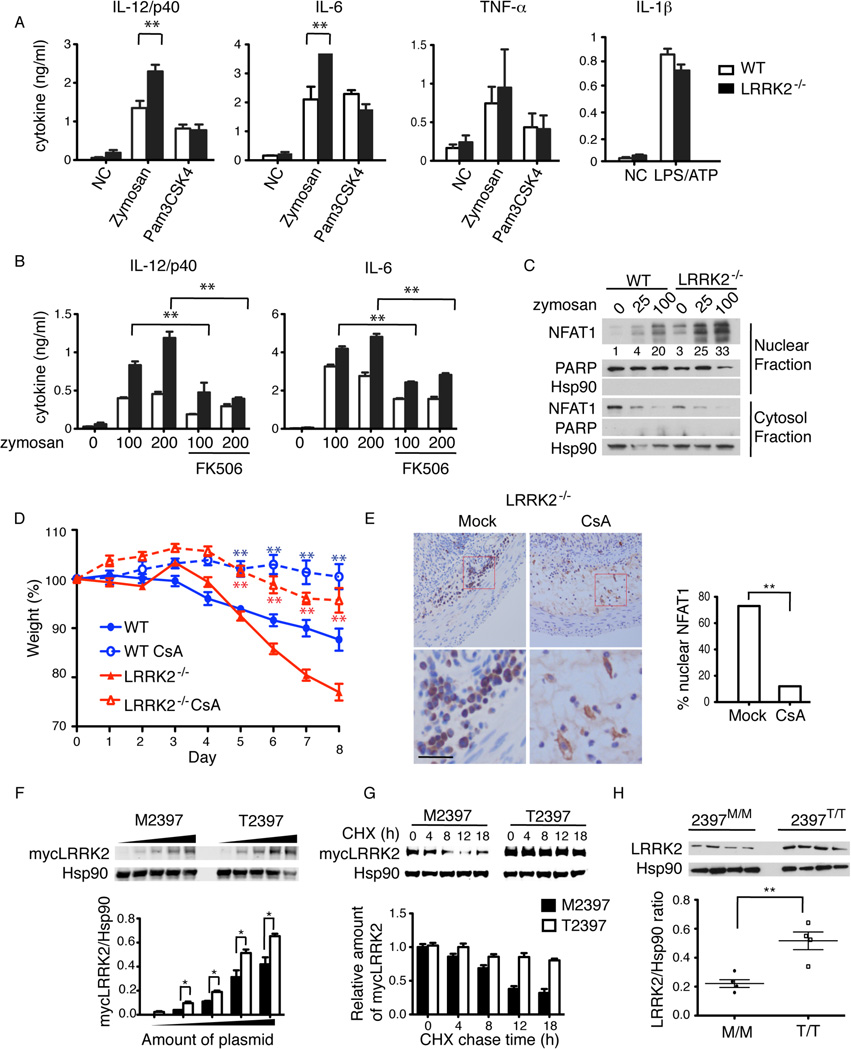
LRRK2 deficiency increases NFAT1 activation in BMDMs and underlies a CD susceptibility polymorphism. (A) IL-12/p40, IL-6 and TNF-α production (mean ± SEM) by WT or LRRK2−/− BMDMs with stimulation of 100 µg ml−1 zymosan or 10 µg ml−1 Pam3CSK4. IL-1β production by BMDMs with 25 ng ml−1 LPS and 5 mM ATP. **P < 0.02. (B) IL-12/p40 and IL-6 production by zymosan (µg ml−1) with or without 2 mM FK506 treatment. (C) NFAT1 immunoblotting in nuclear and cytosol fractions in BMDMs treated with 100 µg ml−1 zymosan for 30 minutes. Numbers below are the quantified relative ratio of NFAT1 in nuclear fractions. (D) Mean body weight (mean ± SEM) of WT and LRRK2−/− receiving DSS with or without 1 mg/mouse/day CsA treatment (n=5). **P < 0.02. P values were calculated between vehicle treated and CsA treated groups. (E) Immunohistochemical NFAT1 staining in colon sections from LRRK2−/− mice receiving vehicle (Mock) or CsA treatment, left panels. Scale bar, 25 µm. Percentage of cells with nuclear NFAT1 among ~100 NFAT1-positive cells for each group, right panel. (F) Immunoblotting of overexpressed M2397 and T2397 forms of mycLRRK2 in 293T cells (top panels). Hsp90 was used as loading control. Ratio of mycLRRK2 and Hsp90 from top panels (bottom panel). *P < 0.05 (G) Cycloheximide (CHX) chase assay of 293T transfected with M2397 or T2397 of mycLRRK2 plasmids treated with 100 µg ml−1 CHX, analyzed by immunoblot with anti-myc antibody to detect mycLRRK2 and anti-Hsp90 to detect Hsp90 as loading control (top panel). Quantification of relative amount of mycLRRK2 (bottom panel). The ratio of mycLRRK2, either M2397 or T2397, to Hsp90 was normalized to their respective starting points at 0 h (set as 1.0). (H) Immunoblotting of LRRK2 in purified peripheral B cells from humans homozygous at 2397, either M/M or T/T (top panels). Scatter plot of quantification of LRRK2 and Hsp90 from top panels (bottom panel). Error bars indicate SEM for each group. **P < 0.01. A–C and H represent three experiments. F and G represent five experiments. D and E represent two experiments.
Based on these observations, we speculated that LRRK2 deficiency accelerated DSS-induced colitis through increased NFAT1 activation in macrophages. We therefore determined whether blocking NFAT1 by inhibiting calcineurin with cyclosporine A (CsA) would alleviate colitis exacerbated by LRRK2 deficiency. CsA has been used to treat DSS-induced colitis in mice (23, 24). However, inhibition of leukocyte recruitment and TGF-β upregulation rather than any effect on NFAT was previously proposed to explain the protective effect of CsA (25, 26). We found that CsA treatment ameliorated weight loss very effectively (Fig. 4D). Consistent with the previous findings, less infiltrating inflammatory cells were observed in colon tissues from CsA-treated mice (Fig. 4E). Importantly, immunohistochemistry revealed that CsA treatment potently inhibited NFAT1 nuclear localization in infiltrating cells in LRRK2−/− mice (Fig. 4E).
Several GWAS identified single-nucleotide polymorphisms (SNPs) at the LRRK2 locus that were associated with CD (3). One of them, rs3761863, leads to a transition mutation 7190 C>T, resulting in a missense amino acid change at position 2397 from Thr to Met, and this amino acid change does not affect the kinase activity (27). We investigated whether this polymorphism would affect LRRK2 protein levels. We found that the high-risk allele M2397 led to a lower overall abundance of LRRK2 protein as compared to the low-risk allele T2397 when each was transiently overexpressed in 293T cells (Fig. 4F). The lower abundance of M2397 correlated with its shorter half-life compared to T2397 (Fig. 4G). To examine whether the polymorphism would affect protein levels of LRRK2 in humans, we compared the levels of LRRK2 in B cells purified from peripheral blood from four 2397M/M homozygous individuals and four 2397T/T homozygous individuals. We found that the constitutive levels of LRRK2 were somewhat variable among individuals however the 2397M/M LRRK2 protein was significantly decreased (Fig. 4H). Therefore, our data suggest that genetically decreased LRRK2 levels in humans may contribute to an increased risk of CD associated with M2397 allele.
In this study, we show that LRRK2 negatively regulates NFAT and that LRRK2 deficiency in mice leads to aberrant activation of macrophages and increased susceptibility to DSS-induced IBD. Biochemical analysis revealed that LRRK2 participates in the NRON complex and likely inhibits NFAT1 activity by promoting its sequestration within the complex and preventing nuclear translocation. LRRK2 removal is important for NFAT nuclear translocation in cells with abundant LRRK2. Our data suggest the concept that NFAT is controlled by two distinct mechanisms – dephosphorylation by calcineurin and release by LRRK2 degradation. These two steps are governed by different stimuli, for instance, Ca2+ influx and LPS, respectively. We propose that LRRK2 regulates NFAT responses to microbial stimuli such as LPS and other TLR ligands, and proper regulation of this response might be very important for IBD. The significantly lower expression level of LRRK2 in T cells compared to macrophages, dendritic cells and B cells may explain why this additional layer of NFAT control previously eluded detection. We suspect that LRRK2 has a dominant influence on NFAT1 in cells highly expressing LRRK2, such as macrophages, dendritic cells, B lymphocytes, and microglia cells. Interestingly, aberrant activation of microglia cells was observed in 20-month-old LRRK2−/− mice but not in WT or LRRK2 transgenic mice (28). Furthermore, our finding that the M2397 LRRK2 protein exhibits decreased stability suggests that constitutively decreased LRRK2 level increases the risk of human CD. LRRK2 is a complex protein with multiple functional domains, and the fact that LRRK2 is implicated in two apparently unrelated diseases, PD and IBD, is puzzling. The PD-related mutation G2019S increases LRRK2 kinase activity, which may affect the cytoskeleton remodeling and microRNA regulation (29, 30). By contrast, the IBD-prone allele M2397 affects LRRK2 accumulation but has no effect on kinase activity suggesting that the pathogenic mechanism in each disease may be distinct. Our study of the role of LRRK2 in IBD provides a successful example of how molecular studies following GWAS can lead to elucidation of the molecular mechanisms of complex diseases such as IBD.
Supplementary Material
Acknowledgements
This work is supported by the Intramural Research Program of the NIH and NIAID. We thank Dr. Mark Cookson and Dr. Nicolas Bidere for reagents; N. Bidere, R. Germain, C. Kanellopoulou, B. Lo, Andrew Snow, and Helen Su for suggestions and comments.
Reference
- 1.Paisan-Ruiz C, et al. Neuron. 2004 Nov 18;44:595. doi: 10.1016/j.neuron.2004.10.023. [DOI] [PubMed] [Google Scholar]
- 2.Zimprich A, et al. Neuron. 2004 Nov 18;44:601. doi: 10.1016/j.neuron.2004.11.005. [DOI] [PubMed] [Google Scholar]
- 3.Barrett JC, et al. Nat Genet. 2008 Aug;40:955. doi: 10.1038/NG.175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Abraham C, Cho JH. N Engl J Med. 2009 Nov 19;361:2066. doi: 10.1056/NEJMra0804647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Gardet A, et al. J Immunol. 2010 Oct 4; [Google Scholar]
- 6.Tlaskalova-Hogenova H, et al. Ann N Y Acad Sci. 2005 Jun;1051:787. doi: 10.1196/annals.1361.122. [DOI] [PubMed] [Google Scholar]
- 7.Maekawa T, Kubo M, Yokoyama I, Ohta E, Obata F. Biochem Biophys Res Commun. 2010 Feb 12;392:431. doi: 10.1016/j.bbrc.2010.01.041. [DOI] [PubMed] [Google Scholar]
- 8.Gwack Y, et al. Nature. 2006 Jun 1;441:646. doi: 10.1038/nature04631. [DOI] [PubMed] [Google Scholar]
- 9.Rao A, Luo C, Hogan PG. Annu Rev Immunol. 1997;15:707. doi: 10.1146/annurev.immunol.15.1.707. [DOI] [PubMed] [Google Scholar]
- 10.Aliprantis AO, Glimcher LH. Adv Exp Med Biol. 2010;658:69. doi: 10.1007/978-1-4419-1050-9_8. [DOI] [PubMed] [Google Scholar]
- 11.Kao SC, et al. Science. 2009 Jan 30;323:651. [Google Scholar]
- 12.Graef IA, et al. Cell. 2003 May 30;113:657. doi: 10.1016/s0092-8674(03)00390-8. [DOI] [PubMed] [Google Scholar]
- 13.Horsley V, Aliprantis AO, Polak L, Glimcher LH, Fuchs E. Cell. 2008 Jan 25;132:299. doi: 10.1016/j.cell.2007.11.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Rohini A, Agrawal N, Koyani CN, Singh R. Pharmacol Res. 2010 Apr;61:269. doi: 10.1016/j.phrs.2009.11.012. [DOI] [PubMed] [Google Scholar]
- 15.Ranger AM, et al. Nature. 1998 Mar 12;392:186. doi: 10.1038/32426. [DOI] [PubMed] [Google Scholar]
- 16.Greenblatt MB, Aliprantis A, Hu B, Glimcher LH. J Exp Med. 2010 May 10;207:923. doi: 10.1084/jem.20092531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Greggio E, et al. J Biol Chem. 2008 Jun 13;283:16906. doi: 10.1074/jbc.M708718200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shaw KT, et al. Proc Natl Acad Sci U S A. 1995 Nov 21;92:11205. doi: 10.1073/pnas.92.24.11205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Monticelli S, Rao A. Eur J Immunol. 2002 Oct;32:2971. doi: 10.1002/1521-4141(2002010)32:10<2971::AID-IMMU2971>3.0.CO;2-G. [DOI] [PubMed] [Google Scholar]
- 20.Willingham AT, et al. Science. 2005 Sep 2;309:1570. doi: 10.1126/science.1115901. [DOI] [PubMed] [Google Scholar]
- 21.Zanoni I, et al. Nature. 2009 Jul 9;460:264. doi: 10.1038/nature08118. [DOI] [PubMed] [Google Scholar]
- 22.Goodridge HS, Simmons RM, Underhill DM. J Immunol. 2007 Mar 1;178:3107. doi: 10.4049/jimmunol.178.5.3107. [DOI] [PubMed] [Google Scholar]
- 23.Murthy SN, et al. Dig Dis Sci. 1993 Sep;38:1722. doi: 10.1007/BF01303184. [DOI] [PubMed] [Google Scholar]
- 24.Melgar S, et al. Int Immunopharmacol. 2008 Jun;8:836. doi: 10.1016/j.intimp.2008.01.036. [DOI] [PubMed] [Google Scholar]
- 25.Soriano-Izquierdo A, et al. Inflamm Bowel Dis. 2004 Nov;10:789. doi: 10.1097/00054725-200411000-00014. [DOI] [PubMed] [Google Scholar]
- 26.Satoh Y, et al. Am J Physiol Gastrointest Liver Physiol. 2009 Sep;297:G514. doi: 10.1152/ajpgi.90608.2008. [DOI] [PubMed] [Google Scholar]
- 27.West AB, et al. Proc Natl Acad Sci U S A. 2005 Nov 15;102:16842. doi: 10.1073/pnas.0507360102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lin X, et al. Neuron. 2009 Dec 24;64:807. [Google Scholar]
- 29.Parisiadou L, Cai H. Commun Integr Biol. 2010 Sep;3:396. doi: 10.4161/cib.3.5.12286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gehrke S, Imai Y, Sokol N, Lu B. Nature. 2010 Jul 29;466:637. doi: 10.1038/nature09191. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.