Abstract
A recent flurry of reports correlates replication timing(RT) with rates of mutation during both evolution and cancer. Specifically, point mutations and copy number losses correlate with late replication, while copy number gains and other rearrangements correlate with early replication. In some cases, plausible mechanisms have been proposed. Point mutation rates may reflect temporal variation in repair mechanisms. Transcription-induced double-strand breaks are expected to occur in transcriptionally active early replicating chromatin. Fusion partners are generally in close proximity, and chromatin in close proximity replicates at similar times. However, temporal enrichment of copy number gains and losses remains an enigma. Moreover, many conclusions are compromised by a lack of matched RT and sequence datasets, filtering out developmental variation in RT, and the use of cell lines to make inferences about germline evolution.
Introduction
Recent evidence has unveiled strong correlations among replication timing (RT), and various forms of genetic mutation in both the germline and cancer [1]. The RT program in mammalian cells is regulated at the level of large (400–800 kb) chromosomal segments (“replication domains”) that replicate at specific times during S phase, with approximately half of these segments replicating at significantly different times in different cell types [2]. For reasons that are still poorly understood there is a near-precise correlation between RT and megabase-scale chromatin interaction spatial compartments defined by genome-wide chromosome-conformation capture (Hi-C), implying that domains in close spatial proximity replicate at similar times [3–5]. Therefore, control of replication invokes mechanisms that are regulated in both space and time. Moreover, early vs late replicating compartments are correlated with active vs inactive transcription and open vs closed chromatin [6–8], which could contribute to varying rates of mutation in the different compartments. In this review, we will summarize recent findings on the correlations of RT to multiple types of mutagenic events in germline and cancer genomes. Potential mechanisms that directly or indirectly couple genome mutation with RT will be discussed.
Replication Timing and Base Substitutions
An initial study spanning 1% of the human genome aligned RT data to either human-primate substitutions or human SNP density, revealing increases of 22% and 53%, respectively, for progressively later replicating DNA [9]. This analysis was followed by genome-wide studies reporting similar results [10,11]. These studies used RT data from highly transformed HeLa cells but, more recently, this concern was addressed by comparing RT profiles from six immortalized lymphoblastoid cell lines derived from father-mother-offspring trios to their matched whole genome sequences. This study found a strikingly more abundant SNP density in late replicating regions, with increases of >2-fold and >6-fold in transition and transversion mutations, respectively [12]. Thus, the removal of population noise in this study may have revealed a more intimate relationship between RT and base substitutions. This correlation is not specific to primates. When polymorphism between 6 Drosophila species was compared to the RT of Kc cells (an embryonic cell line from D. melanogaster), up to 30% increase in divergence was found from early to late replicated regions [13].
There are two important caveats with using somatic cells to draw inferences about evolution. First, it is not possible to distinguish whether mutations arose in the germline (parent to offspring) or during somatic differentiation, since somatic tissue undergoes considerably more cell divisions than germline tissue, which is sequestered early in development. Second, most studies have used replication profiles from somatic cell lines, but the temporal order of replication is significantly different in germline vs. somatic cells [2,6,8,14]. In fact, alignment between divergence of intronic sequences from Mus musculus and Rattus norvegicus reference genomes with RT profiles from mouse embryonic stem cells (ESCs) and induced pluripotent stem cells (iPSCs) revealed only a 10.5% increase from early to late S phase [15], suggesting that mutation rates measured in somatic cells or cell lines may not accurately reflect the germline.
The problem with comparing cell lines to draw conclusions about germline evolution is underscored by studies of various cancer cell types [16–18]. Accumulation of single-nucleotide variation (SNV) in late-replicating/nuclear lamina associated domains (LADs) has been reported during the development of drug resistance in lung cancer [16]. Comparison of SNV in the human germline vs. multiple cancer cell lines revealed a more drastic increase with RT for cancer cells [17,19]. Cancer cells show the same mutational distribution as the germline [17], suggesting that the increased mutational skew may nonetheless arise from similar mechanisms. An important caveat of these cancer cell studies is that, to avoid complications of developmental RT control, they discarded the RT-variant half of the genome from their analyses. Hence, aspects of cancer genome evolution shaped by abnormal developmental control or RT were ignored. Cancer subtypes and even individual cancer clones have their own RT patterns [20], so a complete understanding of the role of replication timing in cancer genome evolution will require studies that compare RT to genome sequences in the same cancer clones.
Altogether, while absolute mutation rates and RT vary by cell type and species, an overall increasing gradient of point mutation rates with later RT has been established in a wide variety of eukaryotic species and probably is conserved among all eukaryotic organisms. This trend is still significant after controlling for covariant features such as recombination hot spots [9], differing tri-nucleotide compositions, G-C content, distance to telomeres, meiotic crossover rate, chromatin compaction [10], and local recombination rates, suggesting (but by no means confirming) a more direct relationship to replication time [13]. Moreover, the somatic vs. germline argument does not apply to single-cell organisms. In Saccharomyces cerevisiae, synonymous SNP and intergenic SNP rates in different strains were found to be elevated >30% in late replicating regions [21]. In fact, late replication was shown to be both necessary and sufficient to cause a high level of point mutations of experimentally manipulated chromosome segments in S. cerevisiae [22].
Proposed Mechanisms for Increasing Point Mutation Rates in Late S Phase
In S. cerevisiae, late S phase templates tend to be repaired with error-prone repair mechanisms [23]. Rev1, in a complex with error-prone Polζ (Rev3/7), increases ~50 fold in late S to G2 phases and deletion of Rev1 significantly lowers mutation rate specifically in late-replicating regions. In contrast, replication forks stalled at lesions during early S phase are accurately repaired by either copying from the correct template or high-fidelity translesion synthesis (Figure 1) [23,24]. Since these proteins are highly conserved [25,26], this mechanism might be common to all eukaryotes. In multi-cellular eukaryotes, early replicating regions are usually gene-rich and transcriptionally active, and lesions in active transcription units tend to be repaired by high fidelity transcription-coupled nucleotide excision repair (Figure 1) [27]. Therefore, different repair mechanisms operating on early vs. late templates could also contribute to the observed trend. Moreover, the substitution rate of replication Timing Transition Regions (TTRs) between replication domains that replicate at significantly different times is higher when those TTRs are interacting more closely with late replicating chromatin [17]. This observation supports the “body-guard” hypothesis (Figure 1) that heterochromatin at the periphery of the nucleus absorbs mutagens to protect interior, more genetically active, chromatin [28]. Other speculative mechanisms have also been proposed. For example, earlier replicating sequences would have more time to be repaired by hypothetical mechanisms that could act after mismatch repair [9]. Also, lower or altered nucleotide pools in late S could trigger mis-incorporation [9], although there is little direct evidence [29].
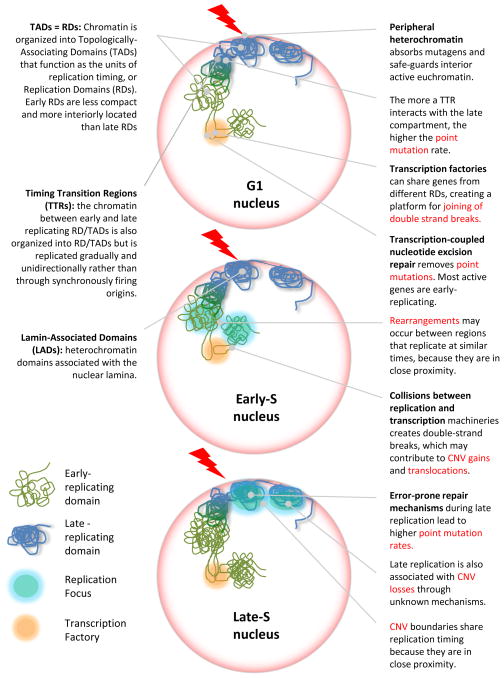
Figure 1. Complex relationship between replication timing and genome variation.
In the G1-nucleus, heterochromatin, localized to the periphery of the nucleus, may form a first defense line for invading mutagens, increasing point mutation rates in late replicating chromatin. Higher levels of transcription in the interior euchromatin could facilitate transcription-coupled nucleotide excision repair (lowering point mutation rates), but could also promote DSBs leading to translocations. In early S phase, collisions between replication and transcription machineries could increase rates of translocations or duplications, which develop either within a domain (observed during reprogramming) or between domains (observed in cancers). In late-S phase, error-prone repair mechanisms cause accumulation of point mutations.
Replication Timing and Copy Number Variation (CNV)
A comparison of CNVs in D. melanogaster and D. simulans with RT from 2 D. melanogaster cell lines found that duplications are enriched in late regions, deletions in early regions [30,31]. No direct measurements of CNV across mammalian species have been reported, but analysis of 331,724 CNVs (in this study called somatic copy-number alterations, SCNAs) in 26 cancer types found the reverse trend to these fly studies [32]. This study demonstrated that the breakpoints for these large (~4.5 Mb) CNV gains are enriched in early replicating regions, whereas those for CNV losses are enriched in late replicating regions, although overall, most CNVs are in constitutively late replicating regions (Figure 1) [32]. Moreover, the boundaries for these large CNVs are of similar RT, and are in close spatial proximity in the nucleus [32]. RT and chromatin proximity together can predict the locations of approximately half of CNVs in the cancer genome [32]. Although these findings was challenged for a lack of statistical significance [33], an analysis using this same dataset successfully modeled CNV distribution using chromosome interaction maps [34]. These cancer studies, similar to the studies of point mutation discussed above, also discarded the RT-variant half of the genome and suffer from the same drawbacks.
A recent study directly examined the relationship between RT-variant domains and CNVs characteristics during human iPSC reprogramming (J. Lu and P. Lerou, submitted). They also found that CNV gains and losses occur preferentially in early and late replicating regions, respectively. Within the RT-variant regions, CNV gains were enriched in domains that replicate earlier in iPSCs as compared to fibroblasts, while losses were mostly in the constitutively late regions, a few of which replicate later in iPSCs than fibroblasts. This study underscores concerns with interpreting studies that remove the RT-variant portion of the genome, since this portion of the genome may have a unique signature of genetic abnormalities.
Proposed Mechanisms for CNV Gains and Losses Linked to Early and Late S Phase
How gains and losses are related to RT is still a matter of speculation. Replicative CNVs could arise from stalled or collapsed forks, which are prone to double-stranded DNA breaks (DSBs) [35,36]. DNA replication is regulated at the level of 400–800kb domains, within which several replicons are simultaneously activated. Since the distribution of origins differs in early and late replicating domains [37,38], the density of simultaneously active forks that could serve as recombination partners may vary. Moreover, regions replicating earlier have twice (or more in cases of re-replication) as many copies as late replicating regions for a significant fraction of the cell cycle, increasing the chance of recombination within vs. between domains [39]. These features of normal replication would tend to favor intra-domain CNVs (~10–300kb), which are more common during evolution and reprogramming ([12,30], J. Lu and P. Lerou, submitted). It seems likely that large inter-domain CNVs (500kb to Mega-base level), which are more commonly seen in cancers [40] arise by different mechanisms than smaller CNVs. Inter-domain CNVs are likely related to the spatial juxtaposition of similar RT domains. One proposed mechanism for large inter-domain CNVs is that domains with similar RT might share replication factories [34], but this is highly unlikely since independent domains have never been observed to share replication factories [41].
Replication-independent CNV mechanisms have also been invoked, including differences in homologous or non-homologous recombination activities [12] and DSB repair mechanisms in different chromatin contexts [42,43]. For example, a recent study has reported that HP1 antagonizes the effect of a histone lysine demethylase, which induces re-replication and thus amplification of specific sites [44], indicating that the HP1-enriched late compartment is suppressed from CNV gains by this mechanism.
Replication Timing, Chromosome Break Points and Translocations
Evolutionary breakpoints between mouse and human are enriched in early replicating regions, and fusion partners generally replicate at similar times in similar cell types from both species [5], consistent with the importance of spatial organization as invoked for CNV. Although these results are subject to the same concerns of using somatic cell RT data to infer evolutionary change, these tendencies are consistent with reports in cancer cells [45,46]. For example, in neuroblastoma cells, >50% of breakpoints mapped to early replicating regions, which is 3.7 times higher than late regions [46]. Additionally, the same trend was found in hematological cancer cells using high resolution RT datasets from a lymphoblastoid cell line [47]. Importantly, in these last two studies, RT data were derived from the same cell lines as translocation data, allowing more direct comparisons. Finally, in a study of acute lymphocytic leukemia, it was found that RT changes associated with translocations can be found in patient samples that lack the translocation [20], suggesting that RT may switch during carcinogenesis upstream of the translocation, placing translocation partners in spatial proximity.
Proposed Mechanisms for Higher Chromosome Break Frequency in Early S Phase
Longstanding cytogenetic evidence [48–56], recently confirmed with chromatin conformation capture [47,57–63], indicates that fusion partners are usually in close spatial proximity before the breaks occur. Since chromatin is organized in the nucleus such that domains of similar RT are in close proximity [4], there is a strong preference for partners to replicate at the same time [47], which is also reported in evolutionary fusion partners between mouse and human [5]. More recently, this concept was tested by experimentally inducing DSBs, revealing that both more common intra-chromosomal fusions and rare translocations were coupled to spatial proximity (Figure 1) [59,62].
The proximity concept explains why fusion partners have similar RT, but the preference for early replication may be due to the promotion of breaks by transcription. Translocations preferentially occur at genic regions, particularly at active transcription start sites [57,59] and translocation signatures in different cancers correlate with tissue-specific transcription profiles [47]. Transcription is a mutagenic process in that it creates topological stress, which leads to DSBs aggravated by topoisomerase inhibition [56]. Moreover, failure of RNA processing during transcription termination leads to genome instability via co-transcriptional R-loop mediated DSB formation [64]. Distal genes regulated by common transcription factors often share transcription factories (Figure 1), placing them in very close proximity under conditions conducive for non-homologous end joining [56,65,66]. In fact, transcriptional induction of TMPRSS2 and ERG (frequent translocation pairs in prostate cancer) results in their movement to a shared transcription factory and dramatically increases their rate of fusion upon exposure to irradiation [67]. Transcription-linked dsDNA breaks may be further exacerbated during replication by head-on collision of transcription and replication machineries (Figure 1) [68]. Since 75% of expressed genes are early replicating [7], transcription-mediated fusions can account for why translocations are enriched in early replicating regions.
Biological Significance of these Complex Correlations
Clearly, one can only speculate as to the significance of mutagenic biases observed in each replication compartment. The “body guard” hypothesis posits that, during somatic development, genes important for general cellular function need to be protected from mutation, while silent genes for alternate developmental lineages are dispensable [28]. From an evolutionary standpoint, late replicating genes could serve as a potential source for gene innovation, with increased point mutations facilitating more rapid evolution of cell-type specific late replicating genes [69]. This may also explain why most housekeeping genes essential for general cell functions exist in the early compartment [69].
Conclusions and Future Directions
It is clear that regions replicated at different times during S phase display distinct patterns of mutation (Figure 1). In yeast, late replication causes enhanced base substitution, possibly due to increased error-prone DNA repair activities in late S phase. A similar correlation is found in higher eukaryotes, but a clear causal relationship has not been established. In the case of CNV and translocations, both result from DSBs but likely involve different mechanisms since they show inverse correlation to RT. There is strong evidence for an involvement of transcription in translocations, which mainly occurs in early replicating regions. The mechanisms linking CNV to RT are more speculative. Both CNV and translocations occur between breakpoints of similar RT, suggesting the importance of sub-nuclear proximity for both events. Correlations between RT and the mutagenic landscape in the germline are consistent with findings in cancer, suggesting common underlying mechanisms. Many existing studies suffer from comparisons between genetic variation and RT in unmatched cell types. Since significant RT changes during embryogenesis [2,6,8,14] and carcinogenesis [20,70] have been well documented, and many high-resolution RT profiles now exist (www.replicationdomain.org and ENCODE), it is critical to analyze the mutations with RT datasets from matching cell types. This would also significantly improve the accuracy in predicting real cancer driver genes [71]. In sum, RT, repair, transcription, and the spatial organization of chromatin confine and refine the genome mutagenic landscapes. Further understanding of the exact molecular mechanism of the RT program and its relationship to genome architecture and transcription should shed light on mechanisms underlying the observed correlation between RT and genome mutational landscape.
Table 1.
Summary of literature
| Type | Study | Species | Year | Source of RT Data | Sequence Data | Hi-C/Lamin alignment |
|---|---|---|---|---|---|---|
| Point mutation[9] | Evolution | Human-Primates | 2009 | 1% of genome, HeLa[72] | Human/chimpanzee reference genomes | None |
| Point mutation[10] | Evolution | Human-Primates | 2010 | HeLa[10] | Human/primate/mouse reference genomes | None |
| Point mutation[15] | Evolution | Mouse | 2010 | mESC and miPSC[73] | Rat/mouse reference genome introns | None |
| Point mutation/CNV[12] | Evolution | Human | 2012 | 6 lymphoblastoid[12] | WGS from father-mother-offspring trios | None |
| Point mutation[19] | Cancer/Evolution | Human | 2012 | 4 cell types; constitutive E&L[2] | SNPs across 6 cancer types, 2 healthy genomes | None |
| Point mutation[17] | Cancer/Evolution | Human | 2013 | 4 cell types; constitutive E&L[2] | Neutral sites across 5 cancer types, 2 healthy genomes | Hi-C |
| Point mutation[16] | Cancer | Human | 2013 | 4 cell types; constitutive E&L[2] | Whole genome/exome of lung cancer cells | Lamin(human fibroblast)[76] |
| Point mutation[13] | Evolution | Fly | 2011 | Kc cells[14] | Synonymous sites in exon cores, putatively unconstrained intronic sites in 6 Drosophila species | None |
| Point mutation[21] | Evolution | Yeast | 2011 | S. cerevisiae[74] | Synonymous intergenic SNPs | None |
| CNV[32] | Cancer | Human | 2011 | 4 cell types; constitutive E&L constitutive E&L[2] | CNVs across 26 cancer types cancer types (Average 4.5Mb ) | Hi-C |
| CNV[31] | Evolution | Fly | 2010 | Kc and C18[14] | Duplications and deletions in 15 natural D. melanogaster populations | None |
| CNV[30] | Evolution | Fly | 2011 | Kc, Bg3 and S2 cells[14,75] | Duplications in 14 natural D. simulans lines | None |
| Break&Fusions[5] | Evolution | Human-mouse | 2010 | Fibroblasts and lymphoblasts[5] | Human and mouse reference genomes | Hi-C |
| Breaks[46] | Cancer | Human | 2005 | neuroblastoma and lymphoblastoid[46] | Breakpoints in neuroblastoma cell lines | None |
| Break&Fusions[47] | Cancer | Human | 2012 | GM06990[73] | Reciprocal chromosomal translocations in leukemia patients (TICdb ) | Hi-C |
Acknowledgments
We apologize to those who could not be cited due to space limitation. We would like to thank J. Lu, M. Thayer, A. Gunjan, K. Hughes, M. Libbrecht and W. Noble for critical reading of the manuscript and helpful discussions. Research in the Gilbert lab is supported by National Institutes of Health grants GM083337, GM085354, and CA161666.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Donley N, Thayer MJ. DNA replication timing, genome stability and cancer: Late and/or delayed DNA replication timing is associated with increased genomic instability. Semin Cancer Biol. 2013 doi: 10.1016/j.semcancer.2013.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Hansen RS, Thomas S, Sandstrom R, Canfield TK, Thurman RE, Weaver M, Dorschner MO, Gartler SM, Stamatoyannopoulos JA. Sequencing newly replicated DNA reveals widespread plasticity in human replication timing. Proc Natl Acad Sci U S A. 2009;107:139–144. doi: 10.1073/pnas.0912402107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Lieberman-Aiden E, van Berkum NL, Williams L, Imakaev M, Ragoczy T, Telling A, Amit I, Lajoie BR, Sabo PJ, Dorschner MO, et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. 2009;326:289–293. doi: 10.1126/science.1181369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ryba T, Hiratani I, Lu J, Itoh M, Kulik M, Zhang J, Schulz TC, Robins AJ, Dalton S, Gilbert DM. Evolutionarily conserved replication timing profiles predict long-range chromatin interactions and distinguish closely related cell types. Genome Res. 2010;20:761–770. doi: 10.1101/gr.099655.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yaffe E, Farkash-Amar S, Polten A, Yakhini Z, Tanay A, Simon I. Comparative analysis of DNA replication timing reveals conserved large-scale chromosomal architecture. PLoS Genet. 2010;6:e1001011. doi: 10.1371/journal.pgen.1001011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hiratani I, Ryba T, Itoh M, Rathjen J, Kulik M, Papp B, Fussner E, Bazett-Jones DP, Plath K, Dalton S, et al. Genome-wide dynamics of replication timing revealed by in vitro models of mouse embryogenesis. Genome Res. 2010;20:155–169. doi: 10.1101/gr.099796.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hiratani I, Takebayashi S, Lu J, Gilbert DM. Replication timing and transcriptional control: beyond cause and effect--part II. Curr Opin Genet Dev. 2009;19:142–149. doi: 10.1016/j.gde.2009.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hiratani I, Ryba T, Itoh M, Yokochi T, Schwaiger M, Chang CW, Lyou Y, Townes TM, Schubeler D, Gilbert DM. Global reorganization of replication domains during embryonic stem cell differentiation. PLoS Biol. 2008;6:e245. doi: 10.1371/journal.pbio.0060245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Stamatoyannopoulos JA, Adzhubei I, Thurman RE, Kryukov GV, Mirkin SM, Sunyaev SR. Human mutation rate associated with DNA replication timing. Nat Genet. 2009;41:393–395. doi: 10.1038/ng.363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chen CL, Rappailles A, Duquenne L, Huvet M, Guilbaud G, Farinelli L, Audit B, d’Aubenton-Carafa Y, Arneodo A, Hyrien O, et al. Impact of replication timing on non-CpG and CpG substitution rates in mammalian genomes. Genome Res. 2010;20:447–457. doi: 10.1101/gr.098947.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cui P, Ding F, Lin Q, Zhang L, Li A, Zhang Z, Hu S, Yu J. Distinct contributions of replication and transcription to mutation rate variation of human genomes. Genomics Proteomics Bioinformatics. 2012;10:4–10. doi: 10.1016/S1672-0229(11)60028-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- **12.Koren A, Polak P, Nemesh J, Michaelson JJ, Sebat J, Sunyaev SR, McCarroll SA. Differential relationship of DNA replication timing to different forms of human mutation and variation. Am J Hum Genet. 2012;91:1033–1040. doi: 10.1016/j.ajhg.2012.10.018. This study finds a stronger correlation between SNP density and late replication by comparing sequences and replication timing profiles in cell lines established from lymphocytes of father-mother-offspring trios. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Weber CC, Pink CJ, Hurst LD. Late-replicating domains have higher divergence and diversity in Drosophila melanogaster. Mol Biol Evol. 2011;29:873–882. doi: 10.1093/molbev/msr265. [DOI] [PubMed] [Google Scholar]
- 14.Schwaiger M, Stadler MB, Bell O, Kohler H, Oakeley EJ, Schubeler D. Chromatin state marks cell-type- and gender-specific replication of the Drosophila genome. Genes Dev. 2009;23:589–601. doi: 10.1101/gad.511809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Pink CJ, Hurst LD. Timing of replication is a determinant of neutral substitution rates but does not explain slow Y chromosome evolution in rodents. Mol Biol Evol. 2010;27:1077–1086. doi: 10.1093/molbev/msp314. [DOI] [PubMed] [Google Scholar]
- 16.Jia P, Jin H, Meador CB, Xia J, Ohashi K, Liu L, Pirazzoli V, Dahlman KB, Politi K, Michor F, et al. Next-generation sequencing of paired tyrosine kinase inhibitor-sensitive and -resistant EGFR mutant lung cancer cell lines identifies spectrum of DNA changes associated with drug resistance. Genome Res. 2013 doi: 10.1101/gr.152322.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- *17.Liu L, De S, Michor F. DNA replication timing and higher-order nuclear organization determine single-nucleotide substitution patterns in cancer genomes. Nat Commun. 2013;4:1502. doi: 10.1038/ncomms2502. This study demonstrates the correlation between base substitution rates and both replication timing and the 3D interactome in cancer. It used replication timing data that represented 50% of the genome that replicates constitutively among 4 cell types. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Watanabe Y, Fujiyama A, Ichiba Y, Hattori M, Yada T, Sakaki Y, Ikemura T. Chromosome-wide assessment of replication timing for human chromosomes 11q and 21q: disease-related genes in timing-switch regions. Hum Mol Genet. 2002;11:13–21. doi: 10.1093/hmg/11.1.13. [DOI] [PubMed] [Google Scholar]
- *19.Woo YH, Li WH. DNA replication timing and selection shape the landscape of nucleotide variation in cancer genomes. Nat Commun. 2012;3:1004. doi: 10.1038/ncomms1982. This study reported that the frequency of somatic SNV in cancer increases with replication time during S phase more drastically than in SNVs from healthy populations. Replication timing data representing 50% of the genome that replicates constitutively between 4 cell types was used. [DOI] [PubMed] [Google Scholar]
- 20.Ryba T, Battaglia D, Chang BH, Shirley JW, Buckley Q, Pope BD, Devidas M, Druker BJ, Gilbert DM. Abnormal developmental control of replication-timing domains in pediatric acute lymphoblastic leukemia. Genome Res. 2012;22:1833–1844. doi: 10.1101/gr.138511.112. This study provides the first comprehensive evaluation of replication timing programs in cancer. Most relevant to this article, RT changes associated with translocations were found in patients that lack the translocation, suggesting that the RT switch may occur upstream of translocation during carcinogenesis, predisposing to the translocation by placing domains in close proximity. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Agier N, Fischer G. The mutational profile of the yeast genome is shaped by replication. Mol Biol Evol. 2011;29:905–913. doi: 10.1093/molbev/msr280. [DOI] [PubMed] [Google Scholar]
- 22.Lang GI, Murray AW. Mutation rates across budding yeast chromosome VI are correlated with replication timing. Genome Biol Evol. 2011;3:799–811. doi: 10.1093/gbe/evr054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Waters LS, Walker GC. The critical mutagenic translesion DNA polymerase Rev1 is highly expressed during G(2)/M phase rather than S phase. Proc Natl Acad Sci U S A. 2006;103:8971–8976. doi: 10.1073/pnas.0510167103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Chang DJ, Cimprich KA. DNA damage tolerance: when it’s OK to make mistakes. Nat Chem Biol. 2009;5:82–90. doi: 10.1038/nchembio.139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Goodman MF, Woodgate R. Translesion DNA Polymerases. Cold Spring Harb Perspect Biol. 2013 doi: 10.1101/cshperspect.a010363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sale JE, Lehmann AR, Woodgate R. Y-family DNA polymerases and their role in tolerance of cellular DNA damage. Nat Rev Mol Cell Biol. 2012;13:141–152. doi: 10.1038/nrm3289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Hanawalt PC, Spivak G. Transcription-coupled DNA repair: two decades of progress and surprises. Nat Rev Mol Cell Biol. 2008;9:958–970. doi: 10.1038/nrm2549. [DOI] [PubMed] [Google Scholar]
- 28.Hsu TC. A possible function of constitutive heterochromatin: the bodyguard hypothesis. Genetics. 1975;79 (Suppl):137–150. [PubMed] [Google Scholar]
- 29.Malinsky J, Koberna K, Stanek D, Masata M, Votruba I, Raska I. The supply of exogenous deoxyribonucleotides accelerates the speed of the replication fork in early S-phase. J Cell Sci. 2001;114:747–750. doi: 10.1242/jcs.114.4.747. [DOI] [PubMed] [Google Scholar]
- 30.Cardoso-Moreira M, Emerson JJ, Clark AG, Long M. Drosophila duplication hotspots are associated with late-replicating regions of the genome. PLoS Genet. 2011;7:e1002340. doi: 10.1371/journal.pgen.1002340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Cardoso-Moreira MM, Long M. Mutational bias shaping fly copy number variation: implications for genome evolution. Trends Genet. 2010;26:243–247. doi: 10.1016/j.tig.2010.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- **32.De S, Michor F. DNA replication timing and long-range DNA interactions predict mutational landscapes of cancer genomes. Nat Biotechnol. 2011;29:1103–1108. doi: 10.1038/nbt.2030. This study unveils the correlation between late replication and large-scale structural changes in cancer cells. They reported that amplicfication boundaries are enriched in early replicating regions, while deletions are enriched in late replicating regions. This study used replication timing data from 50% of the genome that replicates constitutively among 4 cell types. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Bilke S, Gindin Y. Analyzing the association of SCNA boundaries with replication timing. Nat Biotechnol. 2012;30:1043–1045. doi: 10.1038/nbt.2415. author reply 1045–1046. [DOI] [PubMed] [Google Scholar]
- 34.Fudenberg G, Getz G, Meyerson M, Mirny LA. High order chromatin architecture shapes the landscape of chromosomal alterations in cancer. Nat Biotechnol. 2011;29:1109–1113. doi: 10.1038/nbt.2049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Branzei D, Foiani M. Maintaining genome stability at the replication fork. Nat Rev Mol Cell Biol. 2010;11:208–219. doi: 10.1038/nrm2852. [DOI] [PubMed] [Google Scholar]
- 36.Hastings PJ, Lupski JR, Rosenberg SM, Ira G. Mechanisms of change in gene copy number. Nat Rev Genet. 2009;10:551–564. doi: 10.1038/nrg2593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Mesner LD, Valsakumar V, Cieslik M, Pickin R, Hamlin JL, Bekiranov S. Bubble-seq analysis of the human genome reveals distinct chromatin-mediated mechanisms for regulating early- and late-firing origins. Genome Res. 2013 doi: 10.1101/gr.155218.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Guilbaud G, Rappailles A, Baker A, Chen CL, Arneodo A, Goldar A, d’Aubenton-Carafa Y, Thermes C, Audit B, Hyrien O. Evidence for sequential and increasing activation of replication origins along replication timing gradients in the human genome. PLoS Comput Biol. 2011;7:e1002322. doi: 10.1371/journal.pcbi.1002322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Finn KJ, Li JJ. Single-stranded annealing induced by re-initiation of replication origins provides a novel and efficient mechanism for generating copy number expansion via non-allelic homologous recombination. PLoS Genet. 2013;9:e1003192. doi: 10.1371/journal.pgen.1003192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Beroukhim R, Mermel CH, Porter D, Wei G, Raychaudhuri S, Donovan J, Barretina J, Boehm JS, Dobson J, Urashima M, et al. The landscape of somatic copy-number alteration across human cancers. Nature. 2010;463:899–905. doi: 10.1038/nature08822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sporbert A, Gahl A, Ankerhold R, Leonhardt H, Cardoso MC. DNA polymerase clamp shows little turnover at established replication sites but sequential de novo assembly at adjacent origin clusters. Mol Cell. 2002;10:1355–1365. doi: 10.1016/s1097-2765(02)00729-3. [DOI] [PubMed] [Google Scholar]
- 42.Goodarzi AA, Noon AT, Deckbar D, Ziv Y, Shiloh Y, Lobrich M, Jeggo PA. ATM signaling facilitates repair of DNA double-strand breaks associated with heterochromatin. Mol Cell. 2008;31:167–177. doi: 10.1016/j.molcel.2008.05.017. [DOI] [PubMed] [Google Scholar]
- 43.Murga M, Jaco I, Fan Y, Soria R, Martinez-Pastor B, Cuadrado M, Yang SM, Blasco MA, Skoultchi AI, Fernandez-Capetillo O. Global chromatin compaction limits the strength of the DNA damage response. J Cell Biol. 2007;178:1101–1108. doi: 10.1083/jcb.200704140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Black JC, Manning AL, Van Rechem C, Kim J, Ladd B, Cho J, Pineda CM, Murphy N, Daniels DL, Montagna C, et al. KDM4A Lysine Demethylase Induces Site-Specific Copy Gain and Rereplication of Regions Amplified in Tumors. Cell. 2013;154:541–555. doi: 10.1016/j.cell.2013.06.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Fukami M, Tsuchiya T, Vollbach H, Brown KA, Abe S, Ohtsu S, Wabitsch M, Burger H, Simpson ER, Umezawa A, et al. Genomic Basis of Aromatase Excess Syndrome: Recombination- and Replication-Mediated Rearrangements Leading to CYP19A1 Overexpression. J Clin Endocrinol Metab. 2013;98:E2013–2021. doi: 10.1210/jc.2013-2520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Janoueix-Lerosey I, Hupe P, Maciorowski Z, La Rosa P, Schleiermacher G, Pierron G, Liva S, Barillot E, Delattre O. Preferential occurrence of chromosome breakpoints within early replicating regions in neuroblastoma. Cell Cycle. 2005;4:1842–1846. doi: 10.4161/cc.4.12.2257. [DOI] [PubMed] [Google Scholar]
- 47.Shugay M, Ortiz de Mendibil I, Vizmanos JL, Novo FJ. Genomic hallmarks of genes involved in chromosomal translocations in hematological cancer. PLoS Comput Biol. 2012;8:e1002797. doi: 10.1371/journal.pcbi.1002797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Foster HA, Estrada-Girona G, Themis M, Garimberti E, Hill MA, Bridger JM, Anderson RM. Relative proximity of chromosome territories influences chromosome exchange partners in radiation-induced chromosome rearrangements in primary human bronchial epithelial cells. Mutat Res. 2013 doi: 10.1016/j.mrgentox.2013.06.003. [DOI] [PubMed] [Google Scholar]
- 49.Neves H, Ramos C, da Silva MG, Parreira A, Parreira L. The nuclear topography of ABL, BCR, PML, and RARalpha genes: evidence for gene proximity in specific phases of the cell cycle and stages of hematopoietic differentiation. Blood. 1999;93:1197–1207. [PubMed] [Google Scholar]
- 50.Parada LA, McQueen PG, Misteli T. Tissue-specific spatial organization of genomes. Genome Biol. 2004;5:R44. doi: 10.1186/gb-2004-5-7-r44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Parada LA, McQueen PG, Munson PJ, Misteli T. Conservation of relative chromosome positioning in normal and cancer cells. Curr Biol. 2002;12:1692–1697. doi: 10.1016/s0960-9822(02)01166-1. [DOI] [PubMed] [Google Scholar]
- 52.Roix JJ, McQueen PG, Munson PJ, Parada LA, Misteli T. Spatial proximity of translocation-prone gene loci in human lymphomas. Nat Genet. 2003;34:287–291. doi: 10.1038/ng1177. [DOI] [PubMed] [Google Scholar]
- *53.Roukos V, Voss TC, Schmidt CK, Lee S, Wangsa D, Misteli T. Spatial dynamics of chromosome translocations in living cells. Science. 2013;341:660–664. doi: 10.1126/science.1237150. This study addresses the real time dynamics of translocating DSBs using live cell microscopy imaging. It shows that translocations develop preferentially between prepositioned proximal DSBs (Contact-first model), although long-range movement of DSBs does occur infrequently (Breakage-first model) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Branco MR, Pombo A. Intermingling of chromosome territories in interphase suggests role in translocations and transcription-dependent associations. PLoS Biol. 2006;4:e138. doi: 10.1371/journal.pbio.0040138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Nikiforova MN, Stringer JR, Blough R, Medvedovic M, Fagin JA, Nikiforov YE. Proximity of chromosomal loci that participate in radiation-induced rearrangements in human cells. Science. 2000;290:138–141. doi: 10.1126/science.290.5489.138. [DOI] [PubMed] [Google Scholar]
- 56.Cowell IG, Sondka Z, Smith K, Lee KC, Manville CM, Sidorczuk-Lesthuruge M, Rance HA, Padget K, Jackson GH, Adachi N, et al. Model for MLL translocations in therapy-related leukemia involving topoisomerase IIbeta-mediated DNA strand breaks and gene proximity. Proc Natl Acad Sci U S A. 2012;109:8989–8994. doi: 10.1073/pnas.1204406109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- **57.Chiarle R, Zhang Y, Frock RL, Lewis SM, Molinie B, Ho YJ, Myers DR, Choi VW, Compagno M, Malkin DJ, et al. Genome-wide translocation sequencing reveals mechanisms of chromosome breaks and rearrangements in B cells. Cell. 2011;147:107–119. doi: 10.1016/j.cell.2011.07.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Engreitz JM, Agarwala V, Mirny LA. Three-dimensional genome architecture influences partner selection for chromosomal translocations in human disease. PLoS One. 2012;7:e44196. doi: 10.1371/journal.pone.0044196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- **59.Klein IA, Resch W, Jankovic M, Oliveira T, Yamane A, Nakahashi H, Di Virgilio M, Bothmer A, Nussenzweig A, Robbiani DF, et al. Translocation-capture sequencing reveals the extent and nature of chromosomal rearrangements in B lymphocytes. Cell. 2011;147:95–106. doi: 10.1016/j.cell.2011.07.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Kovalchuk AL, Ansarah-Sobrinho C, Hakim O, Resch W, Tolarova H, Dubois W, Yamane A, Takizawa M, Klein I, Hager GL, et al. Mouse model of endemic Burkitt translocations reveals the long-range boundaries of Ig-mediated oncogene deregulation. Proc Natl Acad Sci U S A. 2012;109:10972–10977. doi: 10.1073/pnas.1200106109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- **61.Rocha PP, Micsinai M, Kim JR, Hewitt SL, Souza PP, Trimarchi T, Strino F, Parisi F, Kluger Y, Skok JA. Close proximity to Igh is a contributing factor to AID-mediated translocations. Mol Cell. 2012;47:873–885. doi: 10.1016/j.molcel.2012.06.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- **62.Zhang Y, McCord RP, Ho YJ, Lajoie BR, Hildebrand DG, Simon AC, Becker MS, Alt FW, Dekker J. Spatial organization of the mouse genome and its role in recurrent chromosomal translocations. Cell. 2012;148:908–921. doi: 10.1016/j.cell.2012.02.002. Along with references 57,59 and 61, these four studies addressed principles of spatial constraint on genetic rearrangements by capturing genome-wide translocations with aritificially induced DSBs. These approaches also revealed a strong correlation between translocation and transcription, which mainly occurs within early replicating chromatin. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Lin C, Yang L, Tanasa B, Hutt K, Ju BG, Ohgi K, Zhang J, Rose DW, Fu XD, Glass CK, et al. Nuclear receptor-induced chromosomal proximity and DNA breaks underlie specific translocations in cancer. Cell. 2009;139:1069–1083. doi: 10.1016/j.cell.2009.11.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Santos-Pereira JM, Herrero AB, Garcia-Rubio ML, Marin A, Moreno S, Aguilera A. The Npl3 hnRNP prevents R-loop-mediated transcription-replication conflicts and genome instability. Genes Dev. 2013;27:2445–2458. doi: 10.1101/gad.229880.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Osborne CS, Chakalova L, Mitchell JA, Horton A, Wood AL, Bolland DJ, Corcoran AE, Fraser P. Myc dynamically and preferentially relocates to a transcription factory occupied by Igh. PLoS Biol. 2007;5:e192. doi: 10.1371/journal.pbio.0050192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Dejardin J. How chromatin prevents genomic rearrangements: locus colocalization induced by transcription factor binding. Bioessays. 2011;34:90–93. doi: 10.1002/bies.201100122. [DOI] [PubMed] [Google Scholar]
- 67.Mani RS, Tomlins SA, Callahan K, Ghosh A, Nyati MK, Varambally S, Palanisamy N, Chinnaiyan AM. Induced chromosomal proximity and gene fusions in prostate cancer. Science. 2009;326:1230. doi: 10.1126/science.1178124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Helmrich A, Ballarino M, Nudler E, Tora L. Transcription-replication encounters, consequences and genomic instability. Nat Struct Mol Biol. 2013;20:412–418. doi: 10.1038/nsmb.2543. [DOI] [PubMed] [Google Scholar]
- 69.Juan D, Rico D, Marques-Bonet T, Fernandez-Capetillo O, Valencia A. Late-replicating CNVs as a source of new genes. Biol Open. 2013 doi: 10.1242/bio.20136924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Fritz A, Sinha S, Marella N, Berezney R. Alterations in replication timing of cancer-related genes in malignant human breast cancer cells. J Cell Biochem. 2012;114:1074–1083. doi: 10.1002/jcb.24447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Lawrence MS, Stojanov P, Polak P, Kryukov GV, Cibulskis K, Sivachenko A, Carter SL, Stewart C, Mermel CH, Roberts SA, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. 2013 doi: 10.1038/nature12213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Karnani N, Taylor C, Malhotra A, Dutta A. Pan-S replication patterns and chromosomal domains defined by genome-tiling arrays of ENCODE genomic areas. Genome Res. 2007;17:865–876. doi: 10.1101/gr.5427007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Weddington N, Stuy A, Hiratani I, Ryba T, Yokochi T, Gilbert DM. ReplicationDomain: a visualization tool and comparative database for genome-wide replication timing data. BMC Bioinformatics. 2008;9:530. doi: 10.1186/1471-2105-9-530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Raghuraman MK, Winzeler EA, Collingwood D, Hunt S, Wodicka L, Conway A, Lockhart DJ, Davis RW, Brewer BJ, Fangman WL. Replication dynamics of the yeast genome. Science. 2001;294:115–121. doi: 10.1126/science.294.5540.115. [DOI] [PubMed] [Google Scholar]
- 75.Eaton ML, Prinz JA, MacAlpine HK, Tretyakov G, Kharchenko PV, MacAlpine DM. Chromatin signatures of the Drosophila replication program. Genome Res. 2010;21:164–174. doi: 10.1101/gr.116038.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Guelen L, Pagie L, Brasset E, Meuleman W, Faza MB, Talhout W, Eussen BH, de Klein A, Wessels L, de Laat W, et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature. 2008;453:948–951. doi: 10.1038/nature06947. [DOI] [PubMed] [Google Scholar]