Abstract
The human genome contains a large number of endogenous retroviruses (HERVs). Their reactivation has frequently been observed in patients with cancer. Considering their role in the carcinogenesis process, we aimed to study the possible relationship between HERVs gene expression and Chronic Lymphocytic Leukemia (CLL). We focused on two viral genes gag and np9, the latter presumably an oncogene. We found that the transcriptional activity of HERV-K np9 gene was greater in CLL patients than in healthy donors. However, gag expression was not significantly increased.
These findings suggest a noteworthy relationship between CLL disease and HERV-K np9 expression.
Keywords: HERVs, np9, CLL
Highlights
-
•
We study the possible relationship between HERVs gene expression and CLL.
-
•
CLL patients express np9 gene at least 5 times higher than in healthy donors.
-
•
The gag gene expression was not significantly increased in CLL samples.
-
•
The expression level of the c-myc was not modified by the over expression of np9.
-
•
This work suggests a possible relationship between CLL and HERV-K np9 expression.
1. Introduction
Human endogenous retroviruses (HERVs) are genetic remnants of ancient retroviral infections of the germ line produced during primate evolution which are now transmitted vertically. To date, approximately 8% of the human genome is composed of such retroviral sequences [1]. Most HERVs are dysfunctional due to numerous mutations and deletions. However, those belonging to HERV-K family, contain sequences which are likely to be transcribed.
Many of these HERVs are transcribed and translated under normal physiological conditions. Nevertheless, reactivation of HERVs has frequently been observed in a variety of human tumors suggesting their potential to contribute to malignant progression [2]. Specifically during hemato-oncological processes, several studies have reported the presence of antibodies against HERV-K, overexpression of HERV genes and also the presence of retroviral particles in primary leukemia cells [2]. In addition, np9, a small regulatory gene encoded by HERV-K, has a role as a potent viral oncogene and as a critical molecular switch of multiple signaling pathways regulating the growth of certain human myeloid and lymphoblastic leukemia cells [3].
CLL is the most common form of leukemia in Western countries and mainly affects elderly individuals. It follows an extremely variable course, with survival ranging from months to decades. Available treatments often induce disease remission, but almost all patients will relapse and there is a consensus that CLL remains incurable. To date, both an unmutated (UM) profile of immunoglobulin (Ig) VH genes as well as the presence of genetic lesions at chromosome 17p13, or at 11q23 constitute poor prognosis indicators [4]. Several studies have begun to shed light on the nature of genetic predisposition of CLL but the basis of this disorder remains unknown [4,5]. Studies that screened for the presence of a virus expressed at the RNA level in human CLL, by using massive sequencing technology, gave no evidence of a putative exogenous viral candidate as a cause for this disease [6].
In order to gain insight into the role of endogenous retrovirus in leukemogenesis, we studied np9 and gag gene expression of HERV-K in Chronic Lymphocytic Leukemia (CLL) patients compared to healthy donors.
The results revealed that 70% of CLL patients express np9 gene at least 5 times higher than healthy donors, and 33% express gag gene twice as high as normal donors. These findings suggest a significant relationship between CLL disease and HERV-K np9 (and possibly gag) expression.
2. Materials and methods
Peripheral blood samples were obtained from 25 patients with a typical diagnosis of B cell-CLL (mean age: 64 years). The control group consisted of 6 healthy volunteers. All patients were followed at the Hospital Maciel from Montevideo and provided an informed consent in accordance with national ethical regulations and the Declaration of Helsinki.
The peripheral blood mononuclear cells (PBMC) were isolated by centrifugation on Ficoll-Hypaque (GE Healthcare). RNA was isolated from 1 to 5×106 cells using the Trizol RNA Isolation Protocol. To minimize genomic DNA contamination each RNA sample was treated with DNA-free™ Kit (Ambion, USA). cDNA synthesis was performed as described previously [7]. Negative controls in which reverse transcriptase was omitted were performed to confirm the exclusion of residual genomic DNA contamination.
For gene expression analyses of np9 and gag genes, we used the KAPA SYBR FAST qPCR Kit Master Mix (2×) Universal (Kapa Biosystems) and a Corbett Rotor-Gene 6000. HERVs-K primer sequences of np9 and gag were previously described [2,8]. The cellular c-myc expression level in CLL patients was also assessed with respect to healthy donors. The results of HERV-K and c-myc expression were normalized to the level of glyceraldehyde 3-phosphate dehydrogenase (GAPDH) transcription using the following primers: forward: 5′- GGTGCTGAGTATGTCGTGGA-3′ and reverse: 5′-ATGCCAGTGAGCTTCCCGTT-3′.
PCR cycling conditions for np9, c-myc and GAPDH amplification were: 3 min at 95 °C and 40 cycles at 95 °C for 3 s, 58 °C for 20 s and 72 °C for 20 s. For HERV-K gag specific amplification, cycling conditions were: 95 °C for 10 min, 40 cycles at 95 °C for 15 s and 60 °C for 1 min. The threshold cycle value (Ct), computed for each of these genes as the average of 2 determinations was used to measure the amount of PCR product. The mean ΔCt was calculated by subtracting the mean Ct of duplicated np9 and gag measurements from the mean Ct of duplicated GAPDH measurements. The ΔΔCt values were calculated with the mean ΔCt of the 6 healthy donors. The expression factor difference was calculated by the formula 2−ΔΔCt. Thus, the calculated relative expression provides data on how much np9 and gag gene expression differs from that found in healthy donors.
3. Results and discussion
In this study, the np9 and gag mRNA expression was established for 25 patients with CLL (Fig. 1). We found that the transcriptional activity of HERV-K np9 gene in CLL patients was higher than in PBMCs from healthy donors. The results revealed that 18 out of 25 CLL patients (70% of CLL patients) express np9 transcripts five-fold higher than in normal PBMCs (Fig. 1A). On the other hand when we analyzed gag gene expression we noted that just 33% of CLL patients expressed elevated gag levels (two-fold) compared to healthy people (Fig. 1B). Both in np9 and gag analysis, a wide variability of expression among patients was observed, with no significant difference between mutated and unmutated subgroups (p=0.65 and 0.09 respectively) (see Fig. 1).
Fig. 1.
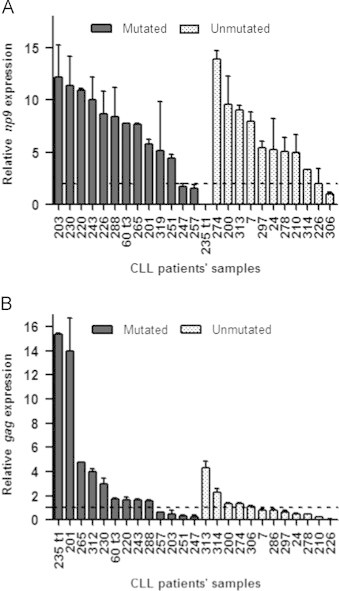
np9 and gag mRNA relative expression levels in CLL patients with mutated and unmutated profile in their immunoglobulin Vh genes. (A) Relative expression of HERV-K np9 gene with respect to healthy donors. The dotted line represents the maximum value of np9 expression from healthy donors as a negative calibrator. (B) Relative expression of HERV-K gag gene with respect to healthy donors. The dotted line represents the maximum value of gag expression from healthy donors as a negative calibrator. The vertical error bars represent the maximum value of each CLL patient׳s sample measured by duplicated.
A previous study regarding HERV-K gag transcriptional activity [2] reported five to eight-fold higher gene expression in 3 B-CLL patients analyzed. However, our study on a large number of B-CLL patients showed that only 2 out of 25 patients exhibited elevated gag expression (5-fold greater than healthy donors) (see Fig. 1B).
The overexpression of HERV-K Np9 protein has been observed in a variety of leukemia cell lines and primary cultures of leukemia cells [3]. It is known that the interaction of Np9 protein abrogates the function of the promyelocytic leukemia zinc finger (PLZF) as a transcriptional repressor of the c-myc proto-oncogene promoter, resulting in the overproduction of this transcription factor and the induction of cell proliferation [9]. Studies of Palacios et al. [10] in CLL report that the proliferative behavior of a small CLL B-cells subpopulation could be related to expression of c-myc oncogene. Unfortunately our studies on c-myc gene expression levels in CLL patients with high and low np9 mRNA expressions showed no significant differences (p=0.19) in the expression of c-myc between these groups, nor with normal PBMCs (Fig. 2). The differences in these results could be explained by the distinct CLL subpopulations analyzed in each work. Other plausible explanation is that np9 mRNA might not be translated or alternatively, that over expression of c-myc through a putative Np9 PLZF pathway may be transient [9].
Fig. 2.
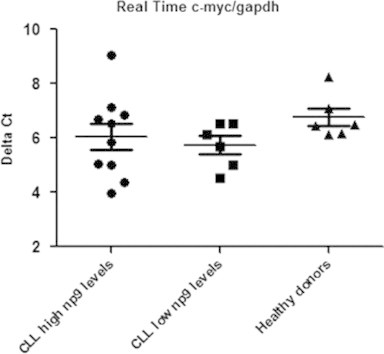
Relative c-myc gene expression normalized to GAPDH expression in CLL patients with high and low np9 expression levels. The horizontal bars represent the median and the error bars the interquartile range. There is no significant differences (p=0.19) in the expression of c-myc gene between the analyzed groups.
To our knowledge, this is the first study of HERV-K np9 mRNA expression in B cell-CLL showing that this molecule is overexpressed in the peripheral blood of CLL patients compared with healthy donors. More studies are needed to evaluate the potential impact of np9 expression on CLL and its role in the regulation of signaling pathways involved in carcinogenesis.
Conflict of interests
The authors declare that they do not have competing interests.
Authors׳ contributions
PM conceived the study, and participated in its design and coordination. SF has made substantial contributions to the design of the study, acquisition of data and analysis. NE and JC have been involved in revising the manuscript critically for important intellectual content. PM and SF wrote the paper. GM, GD and PO made substantial and fundamental contributions to the interpretation and discussion of the results found in this work. AL contributed collecting the samples and selecting the patients. All authors read and approved the final manuscript.
Acknowledgments
This work was supported by Fondo Clemente Estable (Grant FCE_6569) from Agencia Nacional de Investigación e Innovación (ANII) and PEDECIBA, Uruguay.
Contributor Information
Sabrina Fischer, Email: sfischer@cin.edu.uy.
Natalia Echeverría, Email: necheverria@cin.edu.uy.
Gonzalo Moratorio, Email: gonzamoratorio@gmail.com.
Ana Inés Landoni, Email: ailandoni@yahoo.com.
Guillermo Dighiero, Email: dighiero@pasteur.edu.uy.
Juan Cristina, Email: cristina@cin.edu.uy.
Pablo Oppezzo, Email: poppezzo@pasteur.edu.uy.
Pilar Moreno, Email: pmoreno@cin.edu.uy.
References
- 1.Singh S.K. Endogenous retroviruses: suspects in the disease world. Future Microbiol. 2007;2:269–275. doi: 10.2217/17460913.2.3.269. [DOI] [PubMed] [Google Scholar]
- 2.Depil S., Roche C., Dussart P., Prin L. Expression of a human endogenous retrovirus, HERV-K, in the blood cells of leukemia patients. Leukemia. 2002;16:254–259. doi: 10.1038/sj.leu.2402355. [DOI] [PubMed] [Google Scholar]
- 3.Chen T., Meng Z., Gan Y., Wang X., Xu F. The viral oncogene Np9 acts as a critical molecular switch for co-activating beta-catenin, ERK, Akt and Notch1 and promoting the growth of human leukemia stem/progenitor cells. Leukemia. 2013;27:1469–1478. doi: 10.1038/leu.2013.8. [DOI] [PubMed] [Google Scholar]
- 4.Dighiero G., Hamblin T.J. Chronic lymphocytic leukaemia. Lancet. 2008;371:1017–1029. doi: 10.1016/S0140-6736(08)60456-0. [DOI] [PubMed] [Google Scholar]
- 5.Puente X.S., Pinyol M., Quesada V., Conde L., Ordonez G.R. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature. 2011;475:101–105. doi: 10.1038/nature10113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Rego N., Bianchi S, Moreno P., Persson H., Kvist A. Search for an aetiological virus candidate in chronic lymphocytic leukaemia by extensive transcriptome analysis. Br J Haematol. 2012;157:709–717. doi: 10.1111/j.1365-2141.2012.09116.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Oppezzo P., Vasconcelos Y., Settegrana C., Jeannel D., Vuillier F., Legarff-Tavernier M. The LPL/ADAM29 expression ratio is a novel prognosis indicator in chronic lymphocytic leukemia. Blood. 2005;106:650–657. doi: 10.1182/blood-2004-08-3344. [DOI] [PubMed] [Google Scholar]
- 8.Gonzalez-Hernandez M.J., Swanson M.D., Contreras-Galindo R., Cookinham S., King S.R., Noel Jr RJ. Expression of human endogenous retrovirus type K (HML-2) is activated by the Tat protein of HIV-1. J Virol. 2012;86:7790–7805. doi: 10.1128/JVI.07215-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Denne M., Sauter M., Armbruester V., Licht J.D., Roemer K. Physical and functional interactions of human endogenous retrovirus proteins Np9 and rec with the promyelocytic leukemia zinc finger protein. J Virol. 2007;81:5607–5616. doi: 10.1128/JVI.02771-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Palacios F., Moreno P., Morande P., Abreu C., Correa A. High expression of AID and active class switch recombination might account for a more aggressive disease in unmutated CLL patients: link with an activated microenvironment in CLL disease. Blood. 2010;115:4488–4496. doi: 10.1182/blood-2009-12-257758. [DOI] [PubMed] [Google Scholar]


