Abstract
Motivation: A holy grail of biological research is a working model of the cell. Current modeling frameworks, especially in the protein–protein interaction domain, are mostly topological in nature, calling for stronger and more expressive network models. One promising alternative is logic-based or Boolean network modeling, which was successfully applied to model signaling regulatory circuits in human. Learning such models requires observing the system under a sufficient number of different conditions. To date, the amount of measured data is the main bottleneck in learning informative Boolean models, underscoring the need for efficient experimental design strategies.
Results: We developed novel design approaches that greedily select an experiment to be performed so as to maximize the difference or the entropy in the results it induces with respect to current best-fit models. Unique to our maximum difference approach is the ability to account for all (possibly exponential number of) Boolean models displaying high fit to the available data. We applied both approaches to simulated and real data from the EFGR and IL1 signaling systems in human. We demonstrate the utility of the developed strategies in substantially improving on a random selection approach. Our design schemes highlight the redundancy in these datasets, leading up to 11-fold savings in the number of experiments to be performed.
Availability and implementation: Source code will be made available upon acceptance of the manuscript.
Contact: roded@post.tau.ac.il
1 INTRODUCTION
Network analysis tools have become over the last decade the method of choice for studying genome-wide data, yielding important insights into gene function, interaction and evolution. Nevertheless, most of these tools, especially in the protein–protein interaction domain, have been limited to pure topological analysis of the pertaining networks, calling for stronger and more expressive network models (Huang and Fraenkel, 2009; Yeger-Lotem et al., 2009; Yosef et al., 2009).
Recently, Boolean network modeling has been successfully attempted at signaling networks, yielding a qualitative functional understanding of signaling pathways and the ability to predict their behavior under different perturbations and environmental cues (Mitsos et al., 2009; Saez-Rodriguez et al., 2009; Sharan and Karp, 2012). However, because of the sparsity of the currently available data, learning such models de novo remains a formidable task, requiring computational strategies to efficiently prioritize experimental conditions that will best reveal the underlying model. We refer the reader to Karlebach and Shamir (2008) for a comprehensive survey of Boolean modeling.
An alternative modeling technique for signaling pathways dynamics based on ordinary differential equations (ODEs) was thoroughly studied (Hughey et al., 2010). These equations offer a mechanistic chemically based view on the change in the level of cellular species as a function of the levels of their interactors. The dependency of such a detailed modeling on the availability of experimental data has triggered two lines of work of algorithmic experimental design (Kreutz and Timmer, 2009): the first addressing the challenge in parameter estimation (Balsa-Canto et al., 2008; Bandara et al., 2009) and the second addressing the model identification problem (Apgar et al., 2008; Harrington et al., 2012; Kremling et al., 2004; Mélykúti et al., 2010). Nevertheless, the application of this formalism to large-scale modeling is limited by the large number of required parameters whose estimation is difficult (Gutenkunst et al., 2007).
In contrast to the relatively rich literature on ODEs, experimental design algorithms for Boolean networks are scarce. Ideker et al. (2000) proposed an experimental design scheme involving two principal entities: a predictor that generates models given an experimental data and a chooser that selects the next experiment to be conducted based on information theoretic principles. These two entities were used in an iterative manner to learn a genetic network from gene expression data. A similar entropy-based criterion was used by Szczurek et al. (2009) to learn regulatory relations downstream to a given signaling pathway. Barrett and Palsson (2006) proposed an experimental design algorithm for learning regulatory networks that maximizes at each step an estimate of the expected information gain. In the context of signaling networks, we have previously sketched a maximum entropy-based experimental design scheme (Sharan and Karp, 2012), but the scheme was not completely defined nor its utility was tested.
Here we propose two comprehensive experimental design strategies. The first realizes the maximum entropy principle to guide the selection of experiments in the context of Boolean networks learning. The second strategy learns de novo experiments that maximize the disagreement between current best-fit models, a criterion that we term maximum difference. For this optimization task, we propose a novel algorithm that considers the entire space of candidate models and possible experiments.
We implement and test these strategies on simulated and real experimental data using two detailed Boolean models for EGFR and IL1 signaling. We show that both strategies can be used to prioritize experiments and discover redundancies among them, considerably outperforming a random-choice scheme. In particular, we find that the maximum difference criterion is superior to all other approaches in all the settings we tested, leading to 5–11-fold savings in the number of experiments to be performed with respect to the available experiment sets.
2 METHODS
2.1 Data retrieval
To evaluate a design scheme, we applied it to prioritize experiments in the context of two signaling systems in human: EGFR signaling, which regulates cellular growth, proliferation, differentiation and motility; and IL1 signaling, which is involved in coordinating the immune response upon bacterial infection and tissue injury. Both systems are well studied, and detailed manual models exist for them. In particular, Samaga et al. (2009) have constructed a comprehensive Boolean model of the EGFR system, which contains 112 molecular species and their associated Boolean functions; Ryll et al. (2011) have created a Boolean model of the IL1 system with 121 molecular species. We retrieved these models from the CellNetAnalyzer repository (http://www.mpi-magdeburg.mpg.de/projects/cna/repository.html).
To learn logical models for these systems, we used data published by the above authors on the activity (phosphorylation) levels of certain proteins under different cellular conditions. Specifically, Samaga et al. measured within the EGFR system the activity levels of 11 proteins under 34 distinct conditions in Hep2G cells. Similarly, Ryll et al. measured within the IL1 system the activity levels of nine proteins under 14 distinct conditions in primary hepatocytes. Following Ryll et al. (2011) and Samaga et al. (2009), we focused our analysis on the measurements at the 30 min time point, representing the early response of each system.
2.2 Experimental design criteria
Previously, an optimal algorithm for learning Boolean models given experimental data was introduced by Sharan and Karp (2012). However, because of the sparsity of experimental data, the learning procedure yields many models, each explaining the data equally well. To overcome this difficulty in model identification, additional experimental data are needed. Here, we studied two strategies to elucidate informative experiments: the first based on a maximum entropy criterion and the other based on a novel criterion, termed maximum difference criterion.
2.2.1 Maximum entropy criterion
In information theory, the entropy statistic is a standard scoring method that quantifies the information encoded in a given random variable, where higher entropy implies a more informative distribution. An experimental design scheme based on the maximum entropy approach has been previously studied in different settings, such as the identification of regulatory interactions (Ideker et al., 2000) and regulatory functions downstream to signaling pathways (Szczurek et al., 2009). We have sketched a maximum entropy-based strategy for experimental design of Boolean network (Sharan and Karp, 2012) but did not implement or demonstrate the utility of this approach.
Here we implement an experimental design strategy based on a maximum entropy approach. At the heart of this strategy is the evaluation of the entropy of a candidate experiment e according to the predicted response of different models. Formally, let re be the response vector of some model to the experimental conditions set in e. Denote by the probability of observing the response re across all the candidate models. Then, the entropy of the experiment is given by
| (1) |
The maximum entropy strategy prioritizes the experiment with the highest entropy from a set of candidate experiments.
Note that computing the entropy requires the calculation of , the distribution of responses over all models that fit the currently available data well. However, enumeration of all such models is intractable. Previous approaches assumed that a set of possible models is given or that the model space can be sampled. In this work, we adopt the latter approach and sample up to a fixed number of best-fit models. Specifically, we solve an integer linear program (ILP) to infer a best-fit model and use the ILP solver to enumerate multiple solutions. The actual number of solutions is varied to study its impact on the performance of our strategy (see below). Other sampling approaches use Monte Carlo simulations; however, these are often computationally intensive and require large running times (Kreutz and Timmer, 2009).
2.2.2 Maximum difference criterion
We propose and implement an intuitive criterion for experimental design strategy based on maximum difference. This criterion is defined as the Hamming distance between two Boolean response vectors. Formally, given an experiment e, let be the response vectors of two models to the experimental conditions defined in e. Then, the difference criterion is defined by
| (2) |
where |·| denotes absolute value. We note that Mélykúti et al. (2010) have previously experimented with an approach based on similar intuition in the context of ODE models.
The maximum difference strategy prioritizes experiments resulting in the highest difference between a pair of models that equally agree with the available experimental data. While considering the difference induced by only two models, this criterion is amenable to efficient computation via an integer linear programming formulation, allowing us to learn a de novo experiment that maximizes this criterion over all optimal models, eliminating the need to enumerate models or to suggest candidate experiments a priori as in the maximum entropy approach.
2.3 Maximum difference learning algorithm
We develop an algorithm to learn an experiment that maximizes the difference criterion over the entire space of optimal models. The input to the algorithm consists of a directed acyclic network over a set of nodes V and a set of experiments E whose outcome is already known.
The algorithm uses the learning algorithm by Sharan and Karp (2012) as a building block and its outline is as follows (see Fig. 1): (i) Duplicate the ILP of Sharan and Karp so that each copy holds a distinct model; these models (M1, M2) are used to evaluate the maximum difference criterion. (ii) Use the term for the objective as defined in the Sharan and Karp formulation and its corresponding optimal value (OPT) to further constrain the copies of the program to describe solutions that optimally agree with the experimental data. (iii) Add to the resulting program new variables and corresponding constraints to represent the experiment to learn () as well as its readouts under each copy (). Finally, (iv) define a new objective to maximize the difference between readouts, as per Equation (2). An optimal solution to this linear program details a maximum difference experiment. Moreover, to maximize the objective, the corresponding variables of the duplicated programs describe two different models.
Fig. 1.
Overview of the experiment learning algorithm. We start from an ILP (Sharan and Karp, 2012) that learns a Boolean network model M whose readouts have OPT disagreements with the experimental data. In our formulation, this program is duplicated so that two models M1 and M2 are learned simultaneously. The models are further constrained so that: they both have at most OPT disagreements with the experimental data and are therefore optimal and that they both simulate an unknown experiment . The objective optimizes the difference between the readouts of the models (, resp.) as per Equation (2)
2.3.1 Implementation details
We first formulate the problem using an ILP and subsequently solve it with a dedicated solver. Generally, an ILP assumes the following form:
| (3) |
| (4) |
| (5) |
Given a directed acyclic graph G with a set of vertices V, each representing a molecular species, Sharan and Karp (2012) learn an optimal model with respect to a given experimental dataset E using an ILP formulation with variables x = (a,t) where ae,v is a binary variable denoting the activity level of species v in an experiment e, and tv represent the Boolean function associated with v. Additionally, their formulation uses constraints to ensure that the activity level of a species v is (i) compatible with the activity levels of its predecessors and its Boolean function; or (ii) determined by the experimental conditions so that , where Ie(v) is the activity level of species v as under the experimental conditions of e.
To generate a program for learning a maximum difference experiment, we first duplicate the variables and constraints in the above program and add the constraint to each copy, where OPT is the value of the objective for an optimal solution to the original program. Thus, the integral variables and represent two models, each of which is optimal with respect to the available experimental data. We also introduce the variables to represent the activity states under a maximum difference experiment . Additionally, we introduce integral variables for every species v indicating whether v is maintained at an inactive state (0), active state (1) or not perturbed (2) in this maximum difference experiment.
We then add the following constraints to the ILP: (i) constraints to maintain the Boolean functions in as in the original program and (ii) constraints to ensure that the activity level of species v matches the perturbation defined by s. Formally where ‘⇒’ indicates an ‘if-then’ operator (see below).
Finally, to maximize the difference between the two models the objective for the ILP is given by ; the expression is modeled using an auxiliary variable dab such that
| (6) |
| (7) |
The complete ILP is as follows (the constraints for Boolean function adherence are omitted for brevity):
| (8) |
| (9) |
| (10) |
| (11) |
| (12) |
| (13) |
A restricted version of the algorithm additionally receives a set of experiments to choose from. In this version, an additional integral variable is introduced to the formulation indicating the chosen experiment. Then, the following constraints are added to the program:
| (14) |
We use the restricted version of the algorithm in the application to the real datasets where the set of experimental conditions was predefined. We also note that, in a similar fashion, additional constraints may be imposed on the algorithm to exclude experiments that are hard or impossible to conduct in a real setting.
Finally, we solve both the restricted and unrestricted versions of the ILP using CPLEX.
2.3.2 If-then operator using ILP
Our construction uses ‘if-then’ clauses to model relationships between constraints. These may be expressed as follows:
| (15) |
or, equivalently, by
| (16) |
To express this operator using ILP, let be a binary variable and C1, C2 be two large constants and consider the following constraints:
| (17) |
| (18) |
when y = 0, the first constraint holds while is, in practice, free to assume any feasible value; similarly, when y = 1, the second constraint must hold, and is not constrained.
In practice, we model ‘if-then’ operators and absolute value terms using the CPLEX built-in facilities.
2.4 Simulation process
For a given signaling system, let m∗ be a Boolean model that is known in advance, and let Ev be a set of experiments for validation. Given an experimental design strategy and some initial subset of experiments E0, we measured the performance of the strategy by the number of additional experiments it used until a model whose predictions perfectly match the validation dataset (Ev) was learned. We call such a model an optimal model. We summarize our results using the third quartile (75th percentile) of the additional experiments distribution, which is robust to outliers in the data.
To generate the simulated datasets, we started with the known model m∗ and a subset of the nodes whose function we wished to learn. We repeatedly simulated experimental data by randomly setting the experimental conditions, i.e. assigning random binary values to a random subset of the nodes and calculating the readouts according to m∗. As mentioned above, the validation set, Ev, was generated independently from the other sets. Additionally, we generated a different set of experiments, Elist, to prioritize out of which the set of initial experiments, E0, was selected. The entire set of experiments in Elist, but not their readouts, was available to design schemes that prioritize experiments (such as the maximum entropy scheme), whereas only E0 was available to methods that infer experiments de novo, namely, to the maximum difference scheme and then random control scheme. To ensure a fair comparison between the different methods Elist was sufficiently large to uncover an optimal model.
To study the different approaches in heterogeneous, yet realistic settings, we varied number of unknown functions in the range 10, 15, 16, 17, 18 and 20 species (the original data contained 16 unknown functions). Similarly, the sets of perturbed and measured species were constrained to a subset of the species of equal size.
For our evaluation, we used initial subsets, E0, ranging in size from 1 to 8 to account for different amounts of prior knowledge pertaining to the system at hand. Initial subsets for which the learned model performed as well as a model that was derived from the entire dataset were omitted. For each size of the initial subset, up to 30 random subsets were repeatedly sampled from Elist, and the third quartile of the number of additional experiments required to construct an optimal model was reported. We repeat these analyses with 10 simulated datasets.
To study the effect of the number of available models on the performance of the maximum entropy design, we fixed the number of unknown species to 16 while increasing the number of available models over 10, 30, 50, 70, 100 and up to 200. To evaluate the stopping criteria, once an experiment was chosen, we enumerated such multiple models for all experimental design strategies and stopped when at least one optimal model was found.
2.5 Significance assessment
We assess the significance of the hypothesis that one strategy requires less experiments relative to another to arrive at an optimal model using a one-sided Wilcoxon paired test as implemented in R.
2.6 Quartiles standard error
We estimate the standard error using Maritz–Jarrett standard error estimation method (Maritz and Jarrett, 1978).
3 RESULTS
3.1 Experimental design schemes for Boolean models
We propose and implement two experimental design schemes for Boolean network models. The first scheme is based on maximum entropy approach, an accepted information theoretic criterion for model selection. Despite its appealing theoretical properties, computing the entropy depends on the challenging task of estimating the distribution of the responses across candidate models and on the availability of a list of candidate experiments.
The second scheme, termed maximum difference, is an intuitive and novel criterion maximizing the disagreement between two candidate models. Using an ILP formulation, we optimally solve this model and uncover a de novo experiment maximizing this criterion. Unique to our approach is its ability to implicitly consider all candidate models and experiments alleviating the need to specify them a priori.
3.2 Evaluation with simulated data
Given a known Boolean model for a signaling system, an experimental design strategy and an initial set of experiments, we measure the performance of the strategy by the number of additional experiments it uses until an optimal model is learned and report the third quartile of the additional attempts in each dataset. Here, an optimal model is one agreeing with the known model on a set of experiments that were not initially available to the experimental design strategy. The simulation procedure (see Section 2) generated experimental conditions uniformly at random to provide an unbiased sample of the experimental space. In contrast, in real published datasets, the choice of experimental conditions is guided and may impact the relative performance of design schemes.
We compared the running times of the maximum difference and maximum entropy strategies when suggesting a single experiment to be conducted (Fig. 2). Expectedly, the time of the maximum entropy approach grew with the number of models being enumerated, whereas the performance of the maximum difference approach was not affected by it, underscoring its advantage in handling complex problems that admit (for a given experimental dataset) many optimal solutions.
Fig. 2.

Runtime comparison. A comparison of the running times of the maximum difference and maximum entropy approaches. Running times are given on two datasets (EGFR and IL1) when computing a single experiment to be conducted as a function of the number of available optimal models
We used the EGFR Boolean model by Samaga et al. and the IL1 model by Ryll et al. to compare the performance of four design strategies: (i) a naive approach choosing experiments at random, (ii) a maximum entropy approach based on a sample of models, (iii) a minimum entropy approach serving as a control and (iv) a maximum difference approach.
First, we sought to study the effect that the number of available models has on the performance of the maximum entropy-based strategy. Therefore, we applied the above evaluation scheme while increasing the number of models.
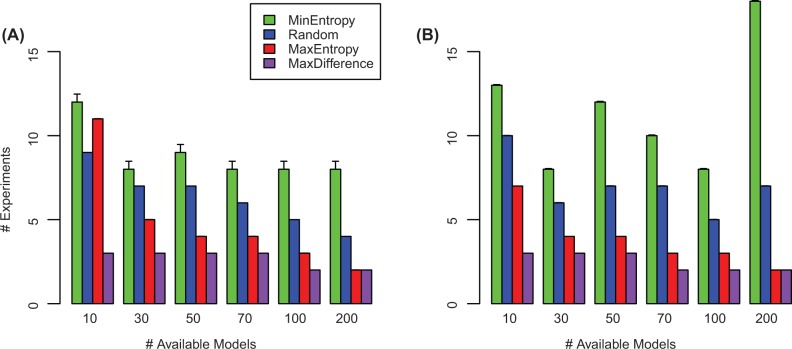
When the simulated data were generated from the EGFR Boolean network model, we found that when only a handful of models were available, the maximum entropy approach performed worse than randomly choosing an experiment, requiring two additional experiments. However, as more models were available the performance relative to the random approach improved. With 200 models available for the evaluation of the entropy, the performance of this approach was comparable with the maximum difference scheme. In our tests, the maximal margin relative to random was obtained when 50 models were available, leading to an improvement of three experiments (see Fig. 3A). The minimum entropy approach performed worse than all other methods; the performance of the method did not vary much when using 30 models or more. Remarkably, the maximum difference approach significantly outperformed all other methods, across the parameter space ( relative to maximum entropy, the next best method, on 70 models). Notably, the advantage in implicitly considering all optimal models was evident as the performance of the method was constant across the parameter range.
Fig. 3.
Sensitivity to the number of available models. The estimation of entropy was dependent on the number of available models. In contrast, the maximum difference learning algorithm optimized over all candidate models. In both panels, the x-axis denotes the number of available models for entropy estimation, and the y-axis denotes the third quartile of the number of experiments required to obtain an optimal model (lower is better). Error bars denote standard error. (A) Simulation with EFGR signaling. (B) simulation with IL1 signaling
We obtained similar results when the simulated datasets were generated using the IL1 signaling model (Fig. 3B). In this case, the maximum entropy approach performed better than the random approach even when only few models were available. Again, only when 200 models where available for the estimation of the entropy criterion the performance of the method was comparable with that of the maximum difference approach. In this dataset, a maximal margin of five experiments relative to random was obtained when 200 models were available. Again, the maximum difference approach outperformed all other methods across the entire parameter space ( relative to maximum entropy on 100 models).
Next, we examined the effect of the size of the learning task (i.e. the number of Boolean functions that need to be learned) on the performance of the different methods. To this end, we applied our evaluation scheme while fixing the number of models and varying the number of unknown functions in the range of 10–20, guided by the 16 unknown functions in the EGFR model.
When simulating the datasets through the EGFR model, the performance of the maximum entropy approach was closer to the random scheme than to the maximum difference design scheme (Fig. 4A). Still, maximum entropy designs were consistently better than random ( for 18 unknown functions) with a margin of two experiments. Additionally, with higher uncertainty, the number of additional experiments grew. For 10 unknown functions the maximum entropy strategy also performed similar to random. However, in this case, the space of possible models was greatly reduced to a point where even the random selection required four experiments for convergence, thus, room for improvement was limited to begin with.
Fig. 4.
Sensitivity to the number of unknown functions. Increasing the number of unknown functions led to increment in the number of experiments that are required to uncover the underlying model in all but the maximum difference strategy, which retained an almost constant performance. In both panels, the x-axis denotes the number of unknown functions, and the y-axis denotes the third quartile of the number of experiments required to obtain an optimal model (lower is better). Error bars denote standard error. (A) Simulation with EGFR signaling. (B) Simulation with IL1 signaling
Notably, the maximum difference approach significantly outperformed all other methods while increasing the marginal gap as the number of functions increased ( relative to maximum entropy for 15 unknown functions). For example, the reduction in required experiments relative to maximum entropy, which was the next best-performing method increased from one experiment for 15 unknown functions to five experiments for 18 unknown functions.
A similar behavior was observed when simulating the datasets through the IL1 model. Again, the maximum entropy approach performed significantly better than the random design scheme ( for 20 unknown functions). Still the maximum difference approach was superior to the maximum entropy scheme in all but a single scenario where 17 functions were predicted. Notably, the maximum difference approach required as little as half of the experiments required by the maximum entropy approach, which was the next best method, to converge when running the simulation with 10, 16, 18 and 20 missing functions. Furthermore, the performance of the maximum difference approach was nearly constant regardless of the size of the learning task.
3.3 Application to real data
To examine the utility of the different approaches in a more realistic setting, we applied them to the available datasets of phosphorylation measurements under different experimental conditions for the EGFR (34 experiments) and IL1 (14 experiments) systems. We ran the different design strategies with increasing subsets of the data as starting points and measured the number of experiments needed to obtain a model that was as good as the one learned from all the available experiments. The results are depicted in Figure 5.
Fig. 5.
Performance evaluation on real data. The x-axis denotes the number of initial experiments, and the y-axis denotes the third quartile of the number of additional experiments required to reconstruct a model fitting the data as well as a model obtained from the all the available experimental data. (A) Results on the EGFR system. (B) Results on the IL1 system
In this setting, we could not apply the maximum difference and random strategies in a straightforward manner, as we cannot simulate de novo experiments. Instead, we applied restricted versions of these strategies, allowing them to choose only experiments that were included in the available data. A key difference between the maximum difference and maximum entropy strategies in this setting is that the former implicitly considered all possible models where the later required a sample of models to evaluate the maximum entropy criterion.
In both datasets, the maximum difference approach performed best, regardless of the number of initial experiments. On the EGFR dataset, maximum difference performed best with a significant advantage over the maximum entropy approach (; 1.75 average difference between third quartiles). Both methods significantly outperformed the random selection with margins of experiments for maximum entropy and experiments for the maximum difference design.
Similar results were obtained for the IL1 system, where data for only 14 experimental conditions were available. Even with this small amount of data the reduction in the number of additional experiments that were required by the maximum difference strategy was statistically significant. Specifically, the maximum difference approach required on average 1.25 less experiments than both the maximum entropy approach and the random approach . Interestingly, the maximum entropy design performed comparably with the random approach in this dataset.
The analyses of the real datasets revealed that the maximum difference approach required less than three experiments to learn an optimal model, achieving a 5–11-fold improvement over the respective sets of available experiments.
4 DISCUSSION
In this article, we studied two approaches for experimental design in Boolean networks. Our main contribution is in the development of an algorithm to optimally solve the maximum difference design criterion while exploring the space of all optimal models under all possible experiments. In addition, we implemented a method based on the well-studied criterion of maximum entropy and demonstrated its utility over a random selection of experiment as well as its limitations under varying conditions. Our evaluation of these schemes indicated that under many conditions, especially in the face of scarce data and increasing complexity of the underlying system, our novel maximum difference approach outperforms the maximum entropy scheme.
Our findings suggest that current studies might suffer from redundant experimentation with respect to the available models of the systems at hand. Furthermore, results on simulated data suggest that by adopting an experimental design scheme, much of the redundancy may be eliminated. Thus, our approach should be beneficial for the study of systems whose underlying model is sufficiently detailed and may be formalized as a Boolean network.
On the methodological side, the maximum difference approach is limited to considering the differences between pairs of models, calling for a generalized approach that considers multiple models. Additionally, in our current sampling procedure, we rely on the ILP solver to retrieve a diverse family of models. Maximum entropy and similar approaches should benefit from the development of other strategies that better sample the model space.
Acknowledgement
The authors thank Dana Silverbush for her valuable comments.
Funding: Edmond J. Safra Center for Bioinformatics at Tel Aviv University (to N.A.) and the I-CORE Program of the Planning and Budgeting Committee and The Israel Science Foundation (757/12 to R.S.).
Conflicts of Interest: none declared.
REFERENCES
- Apgar JF, et al. Stimulus design for model selection and validation in cell signaling. PLoS Comput. Biol. 2008;4:e30. doi: 10.1371/journal.pcbi.0040030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balsa-Canto E, et al. Computational procedures for optimal experimental design in biological systems. IET Syst. Biol. 2008;2:163–172. doi: 10.1049/iet-syb:20070069. [DOI] [PubMed] [Google Scholar]
- Bandara S, et al. Optimal experimental design for parameter estimation of a cell signaling model. PLoS Comput. Biol. 2009;5:e1000558. doi: 10.1371/journal.pcbi.1000558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrett CL, Palsson BO. Iterative reconstruction of transcriptional regulatory networks: an algorithmic approach. PLoS Comput. Biol. 2006;2:e52. doi: 10.1371/journal.pcbi.0020052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gutenkunst RN, et al. Universally sloppy parameter sensitivities in systems biology models. PLoS Comput. Biol. 2007;3:1871–1878. doi: 10.1371/journal.pcbi.0030189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrington HA, et al. Parameter-free model discrimination criterion based on steady-state coplanarity. Proc. Natl Acad. Sci. USA. 2012;109:15746–15751. doi: 10.1073/pnas.1117073109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang S-S, Fraenkel E. Integrating proteomic, transcriptional, and interactome data reveals hidden components of signaling and regulatory networks. Sci. Signal. 2009;2:ra40. doi: 10.1126/scisignal.2000350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughey JJ, et al. Computational modeling of mammalian signaling networks. Wiley Interdiscip. Rev. Syst. Biol. Med. 2010;2:194–209. doi: 10.1002/wsbm.52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ideker TE, et al. Discovery of regulatory interactions through perturbation: inference and experimental design. Pac. Symp. Biocomput. 2000;5:305–316. doi: 10.1142/9789814447331_0029. [DOI] [PubMed] [Google Scholar]
- Karlebach G, Shamir R. Modelling and analysis of gene regulatory networks. Nat. Rev. Mol. Cell Biol. 2008;9:770–780. doi: 10.1038/nrm2503. [DOI] [PubMed] [Google Scholar]
- Kremling A, et al. A benchmark for methods in reverse engineering and model discrimination: problem formulation and solutions. Genome Res. 2004;14:1773–1785. doi: 10.1101/gr.1226004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kreutz C, Timmer J. Systems biology: experimental design. FEBS J. 2009;276:923–942. doi: 10.1111/j.1742-4658.2008.06843.x. [DOI] [PubMed] [Google Scholar]
- Maritz JS, Jarrett RG. A note on estimating the variance of the sample median. J. Am. Stat. Assoc. 1978;73:194–196. [Google Scholar]
- Mélykúti B, et al. Discriminating between rival biochemical network models: three approaches to optimal experiment design. BMC Syst. Biol. 2010;4:38. doi: 10.1186/1752-0509-4-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitsos A, et al. Identifying drug effects via pathway alterations using an integer linear programming optimization formulation on phosphoproteomic data. PLoS Comput. Biol. 2009;5:e1000591. doi: 10.1371/journal.pcbi.1000591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ryll A, et al. Large-scale network models of IL-1 and IL-6 signalling and their hepatocellular specification. Mol. Biosyst. 2011;7:3253–3270. doi: 10.1039/c1mb05261f. [DOI] [PubMed] [Google Scholar]
- Saez-Rodriguez J, et al. Discrete logic modelling as a means to link protein signalling networks with functional analysis of mammalian signal transduction. Mol. Syst. Biol. 2009;5:331. doi: 10.1038/msb.2009.87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samaga R, et al. The logic of EGFR/ErbB signaling: theoretical properties and analysis of high-throughput data. PLoS Comput. Biol. 2009;5:e1000438. doi: 10.1371/journal.pcbi.1000438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharan R, Karp RM. Proceedings of the 16th Annual international conference on Research in Computational Molecular Biology. 2012. Reconstructing boolean models of signaling. RECOMB’12. pp. 261–271. Springer-Verlag, Berlin, Heidelberg. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szczurek E, et al. Elucidating regulatory mechanisms downstream of a signaling pathway using informative experiments. Mol. Syst. Biol. 2009;5:287. doi: 10.1038/msb.2009.45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yeger-Lotem E, et al. Bridging high-throughput genetic and transcriptional data reveals cellular responses to alpha-synuclein toxicity. Nat. Genet. 2009;41:316–323. doi: 10.1038/ng.337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yosef N, et al. Toward accurate reconstruction of functional protein networks. Mol. Syst. Biol. 2009;5:248. doi: 10.1038/msb.2009.3. [DOI] [PMC free article] [PubMed] [Google Scholar]