Abstract
Bacillus thuringiensis represents one of the six species of "Bacillus cereus group" in the genus Bacillus within the family Bacillaceae. Strain Sbt003 was isolated from soil and identified as B. thuringiensis. It harbors at least seven plasmids and produces three shapes of parasporal crystals including oval, bipyramidal and rice. SDS-PAGE analysis of spore-crystal suspension of this strain reveals six major protein bands, which implies the presence of multiple parasporal crystal genes. Bioassay of this strain reveals that it shows specific activity against nematodes and human cancer cells. In this study, we report the whole genomic shotgun sequences of Sbt003. The high-quality draft of the genome is 6,175,670 bp long (including chromosome and plasmids) with 6,372 protein-coding and 80 RNA genes.
Keywords: The Next-Generation sequencing, parasporal crystal protein, Bacillus thuringiensis
Introduction
Bacillus thuringiensis, B. cereus, B. anthracis and other three species constitute the "Bacillus cereus group", a nontaxonomic term, within the genus Bacillus and family Bacillaceae [1]. These species were classified as separate species mainly based on their distinct phenotypes, although extensive genomic studies on strains of these species using different techniques have suggested that they form a single species [2-5]. Strain Sbt003 belongs to the species B. thuringiensis. The type strain of the species produces one or more parasporal crystal proteins showing specific activity against certain larvae from various orders of insects [6]. The specific role and the abundant number of genes encoding of insecticidal crystal proteins of this species have attracted much attention from both academic and industrial researchers. Dozens of B. thuringiensis strains have been sequenced, and dozens more are on their way. In this study, we present a summary classification and a set of features for B. thuringiensis Sbt003, together with the description of the genomic sequencing and annotation.
Classification and features
B. thuringiensis strain Sbt003 harbors at least 7 plasmids and produces three different shapes of parasporal crystals including oval, bipyramidal and rice (Figure 1A, Figure 1B and Table 1). SDS-PAGE analysis of spore-crystal suspension of this strain reveals six major protein bands of 168.8, 148.5, 133.5, 117.2, 107.9 and 103.1 kDa, which implies the presence of multiple parasporal crystal genes (Figure 1C).
Figure 1.
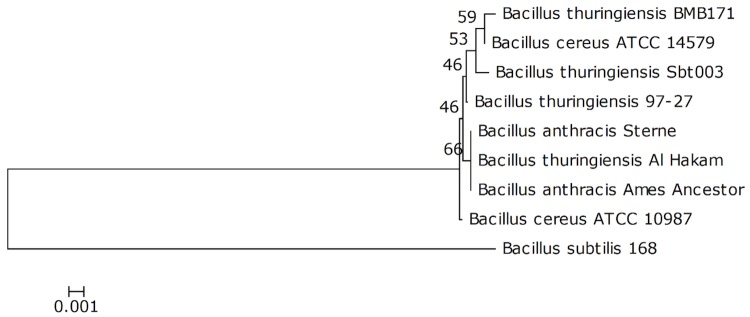
General characteristics of Bacillus thuringiensis Sbt003.(A) Agarose gel electrophoresis of total DNA of Sbt003. Lane M, molecular mass standard, Lambda DNA/HindIII; Lane 1, Sbt003. (B) Phase contrast micrograph of Sbt003 sporulated culture. (C) SDS-PAGE analysis of crystal proteins of Sbt003. Lane M, molecular mass standard; Lane 1, Sbt003.
Table 1. Classification and general features of B. thuringiensis Sbt003 according to the MIGS recommendations [7].
| MIGS ID | Property | Term | Evidence codea |
|---|---|---|---|
| Domain Bacteria | TAS [8] | ||
| Phylum Firmicutes | TAS [9-11] | ||
| Class Bacilli | TAS [12,13] | ||
| Current classification | Order Bacillales | TAS [14,15] | |
| Family Bacillaceae | TAS [14,16] | ||
| Genus Bacillus | TAS [14,17,18] | ||
| Species Bacillus thuringiensis | TAS [14,19] | ||
| Type strain HD73 | |||
| Gram stain | Gram-positive | NAS | |
| Cell shape | Rod-shaped | IDA | |
| Motility | Motile | NAS | |
| Sporulation | Spore-forming | IDA | |
| Temperature range | Room temperature | NAS | |
| Optimum temperature | 28°C | IDS | |
| Carbon source | Organic carbon source | NAS | |
| Energy source | Organic carbon source | NAS | |
| MIGS-6 | Habitat | Soil | IDA |
| MIGS-6.3 | Salinity | Salt tolerant | NAS |
| MIGS-22 | Oxygen | Aerobic | NAS |
| MIGS-14 | Pathogenicity | Avirulent | NAS |
| MIGS-4 | Geographic location | Hubei, China | IDA |
| MIGS-4.1 | Latitude | 29-31N | |
| MIGS-4.2 | Longitude | 111-114E | |
| MIGS-4.3 | Depth | 5-10cm | |
| MIGS-4.4 | Altitude | About 35m | |
| MIGS-5 | Sample collection time | 2000 | IDA |
a) Evidence codes - IDA: Inferred from Direct Assay; TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample, but based on a generally accepted property for the species, or anecdotal evidence). These evidence codes are from the Gene Ontology project [20].
A representative genomic 16S rDNA sequence of strain Sbt003 was searched against GenBank database using BLAST [21]. Sequences showing more than 97% identity to the 16S rDNA of Sbt003 were selected for phylogentic analysis, and a 16S rDNA sequence from B. subtilis subsp. subtilis str. 168 was used as the outgroup. Nine sequences were aligned with ClustalW algorithm. The tree was reconstructed using neighbor joining with the Kimura 2-parameter substitution model. The phylogenetic tree was assessed by bootstrapping 1,000 times, and the consensus tree is shown in Figure 2.
Figure 2.
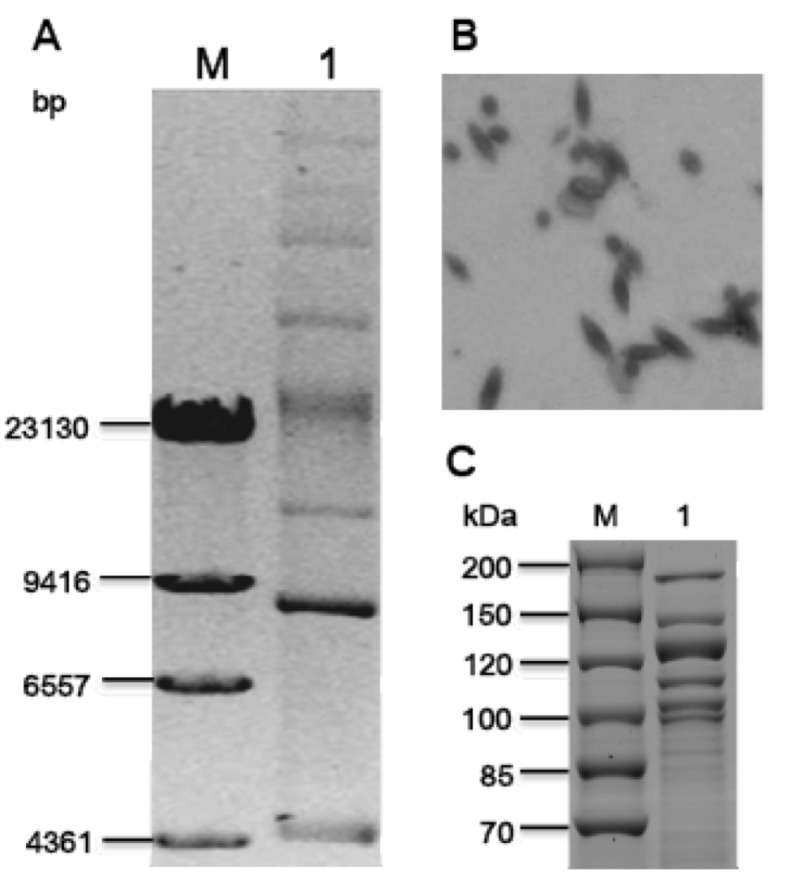
Neighbor-joining phylogenetic tree generated using MEGA 4 based on 16S rRNA sequences. The strains and their corresponding GenBank accession numbers (and, when applicable, draft sequence coordinates) for 16S rDNA sequences are: A, B. thuringiensis serovar konkukian str. 97-27 (AE017355.1): 9337-10763; B, B. thuringiensis BMB171 (CP001903): 9217-10643; C, B. subtilis subsp. subtilis str. 168 (NC_000964): 9839-11263; D, B. cereus ATCC 10987 (NC_003909): 9335-10761; E, B. anthracis str. 'Ames Ancestor' (NC_007530): 9335-10761; F, B. anthracis str. Sterne (NC_005945): 9336-10762; G, B. thuringiensis str. Al Hakam (NC_008600): 9336-10762; H, B. cereus ATCC 14579 (NC_004722): 28956-30382.
Genome sequencing and annotation
Genome project history
This organism was selected for sequencing due to its specific activity against nematodes and human cancer cells. The complete high quality draft genome sequence is deposited in GenBank. The Beijing Genomics Institute (BGI) performed the sequencing and NCBI staff used the Prokaryotic Genome Automatic Annotation Pipeline (PGAAP) to complete the annotation. A summary of the project is given in Table 2.
Table 2. Genome sequencing project information.
| MIGS ID | Property | Term |
|---|---|---|
| MIGS-31 | Finishing quality | High Quality Draft |
| MIGD-28 | Libraries used | Two genomic libraries, one Illumina paired-end library (500 bp inserted size); one Illumina mate-pair library (2 kb inserted size) |
| MIGS-29 | Sequencing platform | Illumina Hiseq 2000 |
| MIGS-31.2 | Sequencing coverage | 488 × |
| MIGS-30 | Assemblers | SOAPdenovo 1.05 version |
| MIGS-32 | Gene calling method | Glimmer and GeneMark |
| GenBank Data of Release | Pending | |
| NCBI project ID | 175950 | |
| Project relevance | Biotechnological |
Growth conditions and DNA isolation
B. thuringiensis Sbt003 was grown in 50 mL Luria broth for 6 hours at 28°C. DNA was isolated by incubating the cells with lysozyme (10 mg/mL) in 2 mL TE (50 mM Tris base, 10 mM EDTA, 20% sucrose, pH8.0) at 4°C for 6 hours. 4 mL 2% SDS was added and the mixture was incubated at 55°C for 30 min; 2 mL 5M NaCl were added, and the mixture was incubated at 4°C for 10 min. DNA was purified by organic extraction and ethanol precipitation.
Genome sequencing and assembly
The genome of B. thuringiensis Sbt003 was sequenced using Illumina Hiseq 2,000 platform (with a combination of a 100-bp paired-end reads sequencing from a 500-bp genomic library and a 90-bp mate-paired reads sequencing from a 2-kb genomic library). Reads with average quality scores below Q30 or having more than 3 unidentified nucleotides were eliminated. Using SOAPdenovo 1.05 version, 22,295,588 paired-end reads (achieving ~325 fold coverage [2.01 Gb]) and 11,166,312 mate-paired reads (achieving ~ 163 fold coverage [1.00 Gb]) were assembled de novo [22]. The assembly is considered a high-guality draft and consists of 104 contigs arranged in 61 scaffolds with a total size of 6,175,670 bp. According to bioinformatic analysis, we identified two large plasmids belonging to ori44-type and repB-type plasmids, respectively. The former plasmid has two ori44-type replicons. We propose it represents a fusion of two plasmids and its estimated size is about 200 kb. The latter plasmid has an expected size of at least 90 kb, according to the sequence of contig0027, which is typical of repB-type plasmids (80 ~ 90 kb). In addition, we identified five other plasmids from the plasmid pattern (see Figure 1A). The expected sizes of the smaller three are 13 kb, 8kb and 4kb, respectively, while the sizes of the larger two can't be deduced either from the plasmid pattern or by bioinformatic analysis.
Genome annotation
Genome annotation was completed using the Prokaryotic Genomes Automatic Annotation Pipeline (PGAAP). Briefly, protein-coding genes were predicted using a combination of GeneMark and Glimmer [23-25]. Ribosomal RNAs were predicted by sequence similarity searching using BLAST against an RNA sequence database and/or using Infernal and Rfam models [26,27]. Transfer RNAs were predicted using tRNAscan-SE [28]. In order to detect missing genes, a complete six-frame translation of the nucleotide sequence was done and predicted proteins (generated above) were masked. All predictions were then searched using BLAST against all proteins from complete microbial genomes. Annotation was based on comparison to protein clusters and on the BLAST results. Conserved domain Database and Cluster of Orthologous Group information were then added to the annotation.
Genome Properties
The high-quality draft assembly of the genome consists of 104 contigs in 61 scaffolds, with an overall 35.21% G+C content. Of the 6,452 genes predicted, 6,372 were protein-coding genes, and 80 RNAs were also identified. The majority of the protein-coding genes (66.67%) were assigned a putative function while the remaining ones were annotated as hypothetical proteins (Table 3). The distribution of genes into COGs functional categories is presented in Table 4.
Table 3. Genome Statistics.
| Attribute | Value | % of total |
|---|---|---|
| Genome size (bp) | 6,175,670 | 100.00 |
| DNA coding region (bp) | 4,818,828 | 78.03 |
| DNA G+C content (bp) | 2,174,469 | 35.21 |
| Number of scaffolds | 61 | - |
| Extrachromosomal elements | > 300 kb | > 4.86 |
| Total genes | 6,452 | 100.00 |
| tRNA genes | 70 | 1.08 |
| rRNA genes | 10 | 0.16 |
| rRNA operons | 0** | - |
| Protein-coding genes | 6,372 | 98.76 |
| Pseudo gene (Partial genes) | 0 (49) | 0 (0.76%) |
| Genes with function prediction (proteins) | 4248 | 66.67% |
| Genes assigned to COGs | 4,334 | 68.02% |
| Genes with signal peptides | 437 | 6.86 |
| CRISPR repeats | 0 | 0 |
**none of the rRNA operons appears to be complete due to unresolved assembly problems.
Table 4. Number of genes associated with the general COG functional categories.
| Code | Value | % age | Description |
|---|---|---|---|
| J | 224 | 4.404 | Translation, ribosomal structure and biogenesis |
| A | 0 | 0.0 | RNA processing and modification |
| K | 485 | 9.536 | Transcription |
| L | 374 | 7.354 | Replication, recombination and repair |
| B | 1 | 0.020 | Chromatin structure and dynamics |
| D | 48 | 0.944 | Cell cycle control, cell division, chromosome partitioning |
| Y | 0 | 0 | Nuclear structure |
| V | 143 | 2.812 | Defense mechanisms |
| T | 225 | 4.424 | Signal transduction mechanisms |
| M | 254 | 4.994 | Cell wall/membrane/envelope biogenesis |
| N | 59 | 1.160 | Cell motility |
| Z | 1 | 0.020 | Cytoskeleton |
| W | 1 | 0.020 | Extracellular structures |
| U | 65 | 1.278 | Intracellular trafficking, secretion, and vesicular transport |
| O | 122 | 2.399 | Posttranslational modification, protein turnover, chaperones |
| C | 215 | 4.227 | Energy production and conversion |
| G | 310 | 6.095 | Carbohydrate transport and metabolism |
| E | 480 | 9.438 | Amino acid transport and metabolism |
| F | 109 | 2.143 | Nucleotide transport and metabolism |
| H | 156 | 3.067 | Coenzyme transport and metabolism |
| I | 140 | 2.753 | Lipid transport and metabolism |
| P | 309 | 6.076 | Inorganic ion transport and metabolism |
| Q | 124 | 2.438 | Secondary metabolites biosynthesis, transport and catabolism |
| R | 783 | 15.395 | General function prediction only |
| S | 458 | 9.005 | Function unknown |
| 2038 | 31.98 | Not in COGs |
The whole genomic sequence and the coding sequence of Sbt003 were analyzed by BtToxin_scanner [29], and eight potential crystal protein sequences were identified. Among these, four were considered to be full-length (locus tags: C797_02099, C797_12066, C797_12568 and C797_27783) while the others were considered to be truncated (Locus tags: C797_02094, C797_12046, C797_12061, C797_18417).
Acknowledgements
This work was supported by grants from the National High Technology Research and Development Program (863) of China (2011AA10A203), China 948 Program of Ministry of Agriculture (2011-G25), the National Basic Research Program (973) of China (2009CB118902), the National Natural Science Foundation of China (31170047 and 31171901), and the Genetically Modified Organisms Breeding Major Projects of China (2009ZX08009-032B).
Reference
- 1.Vilas-Bôas GT, Peruca AP, Arantes OM. Biology and taxonomy of Bacillus cereus, Bacillus anthracis, and Bacillus thuringiensis. Can J Microbiol 2007; 53:673-687 10.1139/W07-029 [DOI] [PubMed] [Google Scholar]
- 2.Helgason E, Caugant DA, Lecadet MM, Chen Y, Mahillon J, Lovgren A, Hegna I, Kvaloy K, Kolsto AB. Genetic diversity of Bacillus cereus/B. thuringiensis isolates from natural sources. Curr Microbiol 1998; 37:80-87 10.1007/s002849900343 [DOI] [PubMed] [Google Scholar]
- 3.Helgason E, Okstad OA, Caugant DA, Johansen HA, Fouet A, Mock M, Hegna I, Kolsto AB. Bacillus anthracis, Bacillus cereus, and Bacillus thuringiensis--one species on the basis of genetic evidence. Appl Environ Microbiol 2000; 66:2627-2630 10.1128/AEM.66.6.2627-2630.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ticknor LO, Kolsto AB, Hill KK, Keim P, Laker MT, Tonks M, Jackson PJ. Fluorescent amplified fragment length polymorphism analysis of Norwegian Bacillus cereus and Bacillus thuringiensis soil isolates. Appl Environ Microbiol 2001; 67:4863-4873 10.1128/AEM.67.10.4863-4873.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Helgason E, Tourasse NJ, Meisal R, Caugant DA, Kolsto AB. Multilocus sequence typing scheme for bacteria of the Bacillus cereus group. Appl Environ Microbiol 2004; 70:191-201 10.1128/AEM.70.1.191-201.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Schnepf E, Crickmore N, Van Rie J, Lereclus D, Baum J, Feitelson J, Zeigler DR, Dean DH. Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol Mol Biol Rev 1998; 62:775-806 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen M, Angiuoli SV, et al. Towards a richer description of our complete collection of genomes and metagenomes "Minimum Information about a Genome Sequence " (MIGS) specification. Nat Biotechnol 2008; 26:541-547 10.1038/nbt1360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 1990; 87:4576-4579 10.1073/pnas.87.12.4576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gibbons NE, Murray RGE. Proposals Concerning the Higher Taxa of Bacteria. Int J Syst Bacteriol 1978; 28:1-6 10.1099/00207713-28-1-1 [DOI] [Google Scholar]
- 10.Garrity GM, Holt JG. The Road Map to the Manual. In: Garrity GM, Boone DR, Castenholz RW (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 1, Springer, New York, 2001, p. 119-169. [Google Scholar]
- 11.Murray RGE. The Higher Taxa, or, a Place for Everything...? In: Holt JG (ed), Bergey's Manual of Systematic Bacteriology, First Edition, Volume 1, The Williams and Wilkins Co., Baltimore, 1984, p. 31-34. [Google Scholar]
- 12.List of new names and new combinations previously effectively, but not validly, published. List no. 132. Int J Syst Evol Microbiol 2010; 60:469-472 10.1099/ijs.0.022855-0 [DOI] [PubMed] [Google Scholar]
- 13.Ludwig W, Schleifer KH, Whitman WB. Class I. Bacilli class nov. In: De Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 3, Springer-Verlag, New York, 2009, p. 19-20. [Google Scholar]
- 14.Skerman VBD, McGowan V, Sneath PHA. Approved Lists of Bacterial Names. Int J Syst Bacteriol 1980; 30:225-420 10.1099/00207713-30-1-225 [DOI] [PubMed] [Google Scholar]
- 15.Prévot AR. In: Hauderoy P, Ehringer G, Guillot G, Magrou. J., Prévot AR, Rosset D, Urbain A (eds), Dictionnaire des Bactéries Pathogènes, Second Edition, Masson et Cie, Paris, 1953, p. 1-692. [Google Scholar]
- 16.Fischer A. Untersuchungen über bakterien. Jahrbücher für Wissenschaftliche Botanik 1895; 27:1-163 [Google Scholar]
- 17.Cohn F. Untersuchungen über Bakterien. Beitr Biol Pflanz 1872; 1:127-224 [Google Scholar]
- 18.Gibson T, Gordon RE. Genus I. Bacillus Cohn 1872, 174; Nom. gen. cons. Nomencl. Comm. Intern. Soc. Microbiol. 1937, 28; Opin. A. Jud. Comm. 1955, 39. In: Buchanan RE, Gibbons NE (eds), Bergey's Manual of Determinative Bacteriology, Eighth Edition, The Williams and Wilkins Co., Baltimore, 1974, p. 529-550. [Google Scholar]
- 19.Berliner E. Über die Schlaffsucht der Mehlmottenraupe (Ephestia kuhniella Zell) und ihren Erreger Bacillus thuringiensis n. sp. Zeitschrift für angewandte Entomologie Berlin 1915; 2:29-56.
- 20.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25:25-29 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol 1990; 215:403-410 [DOI] [PubMed] [Google Scholar]
- 22.Li R, Zhu H, Ruan J, Qian W, Fang X, Shi Z, Li Y, Li S, Shan G, Kristiansen K, et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res 2010; 20:265-272 10.1101/gr.097261.109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Besemer J, Lomsadze A, Borodovsky M. GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res 2001; 29:2607-2618 10.1093/nar/29.12.2607 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Delcher AL, Harmon D, Kasif S, White O, Salzberg SL. Improved microbial gene identification with GLIMMER. Nucleic Acids Res 1999; 27:4636-4641 10.1093/nar/27.23.4636 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lukashin AV, Borodovsky M. GeneMark.hmm: new solutions for gene finding. Nucleic Acids Res 1998; 26:1107-1115 10.1093/nar/26.4.1107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Griffiths-Jones S, Bateman A, Marshall M, Khanna A, Eddy SR. Rfam: an RNA family database. Nucleic Acids Res 2003; 31:439-441 10.1093/nar/gkg006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Eddy SR. A memory-efficient dynamic programming algorithm for optimal alignment of a sequence to an RNA secondary structure. BMC Bioinformatics 2002; 3:18 10.1186/1471-2105-3-18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 1997; 25:955-964 10.1093/nar/25.5.0955 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ye W, Zhu L, Liu Y, Crickmore N, Peng D, Ruan L, Sun M. Mining new crystal protein genes from Bacillus thuringiensis on the basis of mixed plasmid-enriched genome sequencing and a computational pipeline. Appl Environ Microbiol 2012; 78:4795-4801 10.1128/AEM.00340-12 [DOI] [PMC free article] [PubMed] [Google Scholar]


